Introduction
Human breast cancer is the most frequently diagnosed
cancer and the leading cause of cancer-related death among females.
It was shown in the GLOBOCAN 2008 estimates that breast cancer
accounted for 23% of total cancer cases and 14% of cancer
mortalities (1). Due to the
progression and metastasis of this tumor, the prognosis of patients
with advanced stage breast cancer remains poor despite treatment
with various therapy types. Better understanding of the molecular
mechanisms underlying the progression of breast cancer is required
for prevention and treatment.
Use of small interfering RNA (siRNA) to silence a
target gene is a simple and effective method, particularly when
specific siRNA is employed (2). By
introducing double-stranded RNA homologous to a particular message
and causing its sequence-specific degradation, RNA interference
(RNAi) technique provides a rapid method with which to deplete mRNA
as a post-transcriptional regulation.
Jumonji domain containing 2A (JMJD2A) was identified
and characterized in 2004 (3). It
is widely expressed in human tissues and cell lines. Moreover, the
expression level of JMJD2A mRNA was found to be high in several
cell types including prostate cancer, U2OS osteosarcoma, HT1376
bladder carcinoma and human T-cell lymphotropic virus 1-infected
cell lines (4,5). JMJD2A belongs to the
cancer-associated gene family of JMJD2 proteins (3), which, containing a JmjC domain, are
lysine trimethyl-specific histone demethylases capable of
catalyzing the demethylation of trimethylated H3K9 (H3K9me3) and
H3K36 (H3K36me3) (6–8). However, the existing literature
rarely pays close attention to the function of JMJD2A in breast
cancer. It remains unclear whether the inhibition caused by
knockdown of JMJD2A is a special case or a common phenomenon.
Effects of knockdown of JMJD2A on other human breast cancer cell
lines are unknown.
This study transfected chemically synthesized
JMJD2A-specific siRNA into human breast cancer cell line MCF-7. The
transcription level of JMJD2A mRNA, expression level of JMJD2A
protein and the biological characteristics of MCF-7 cells, such as
cell proliferation, migration and invasion ability, were
investigated.
Materials and methods
JMJD2A siRNA synthesis
JMJD2A siRNA was chemically synthesized by Qiagen
Technology Co., Ltd. (Shanghai, China). The sense sequence of the
synthesized siRNA duplexes was: 5′-GAGUUAUCAACUCAAGAUA-3′, and its
antisense sequence was: 5′-UAUCUUGAGUUGAUAACUC-3′. siRNA was
diluted to 20 μmol/l with RNase-free water.
Cell transfection
The MCF-7 human breast cancer cell line, which was
preserved in our laboratory in logarithmic growth phase, was seeded
into 6-well plates, at a density of 5×105 cells per well
and cultured in regular growth medium 24 h prior to transfection.
Regular growth medium was replaced by serum-free Opti-MEM (Gibco,
Invitrogen, Carlsbad, CA, USA) 8 h later. Transfection compounds
were prepared in three groups as follows: siRNA group (100 μl of
Opti-MEM, 6 μl of HiPerFect transfection reagent and 5 μl of JMJD2A
siRNA), negative control group (100 μl of Opti-MEM, 6 μl of
HiPerFect transfection reagent and 5 μl of negative control siRNA)
and blank control group (100 μl of Opti-MEM). HiPerFect
transfection reagent and negative control siRNA were purchased from
Qiagen Technology Co. Ltd (Shanghai, China). Transfection compounds
were placed at room temperature for 10 min and then dropped into
6-well plates, with 2200 μl of bulk volume per well. Both Opti-MEM
and transfection compounds were replaced with complete medium 24 h
after transfection. FAM-siRNA was transfected to measure the
transfection efficiency simultaneously.
Quantitative real-time PCR
Total RNA of the three groups was extracted with the
RNAiso reagent kit (Takara, Dalian, China) 48 h after transfection.
cDNA was generated by reverse transcription of 2 μg of total RNA
using random primers and PrimeScript RT Master Mix (Perfect Real
Time) (Takara) in a total reaction volume of 40 μl. The sequences
of forward and the reverse oligonucleotide primers, specific to
JMJD2A and housekeeping genes, were designed using Primer5
software. Primers (5′-CCAGAACCAACCAGGAGC-3′ and 5′-TTCACT
GCGCGAGACCAT-3′ for JMJD2A; 5′-TGGCACCCAGCA CAATGAA-3′ and
5′-CTAAGTCATAGTCCGCCTAGAA GCA-3′ for β-actin) were synthesized by
Shanghai Daweike Biotechnology Co. Ltd. (Shanghai, China).
Real-time quantitative PCR was performed
in an ABI PRISM 7500 real-time system
A 10-fold dilution of each cDNA was amplified in a
20-μl volume, using the SYBR Premix Ex Taq™ (Perfect Real Time)
(Takara), at a final concentration of 0.2 μM for each primer. PCR
cycle conditions were set as follows: 95°C for 30 sec, and 40
cycles of 95°C for 5 sec and 60°C for 34 sec. The amplification
specificity was evaluated with melting curve analysis. Threshold
cycle Ct, which correlates inversely with the target mRNA levels,
was calculated using the second derivative maximum algorithm
provided by the iCycler software. For JMJD2A, the mRNA levels were
normalized to β-actin mRNA levels (9).
Western blot analysis
Cells in different groups were homogenized 72 h
after transfection in western blot analysis buffer containing 10 mM
Tris-HCl (pH 7.4), 150 mM NaCl, 1% (v/v) Triton X-100, 1% sodium
deoxycholate, 0.1% SDS, 5 mM EDTA, 1 mM PMSF, 0.28 kU/l aprotinin,
50 mg/l leupeptin, 1 mM benzamidine and 7 mg/l pepstain A. The
homogenate was then centrifuged at 12,000 rpm for 10 min at 4°C and
the supernatant was retained and preserved at −80°C for later use.
Protein concentration was determined using a BCA kit (Pierce;
Thermo Fisher Scientific, Rockford, IL, USA). Protein (20 μg) from
each group was subject to electrophoresis on a 10% SDS-PAGE gel
using a constant current. Proteins were transferred to
nitrocellulose membranes on a semidry electrotransferring unit and
incubated with monoclonal rabbit anti-human JMJD2A antibody (Cell
Signaling Technology, Danvers, MA, USA; 1:1000) in Tris-buffered
saline containing 0.1% Tween-20 (TBST) and 5% nonfat dry milk
overnight at 4°C.
After overnight incubation with the primary
antibodies, membranes were washed and incubated with HRP-labeled
goat anti-rabbit secondary antibody (Santa Cruz Biotechnology Inc.,
Santa Cruz, CA, USA) in TBST for 2 h. Immunoreactivity was detected
with enhanced chemiluminescent autoradiography (ECL kit, Amersham).
The membranes were reprobed with β-actin (Cell Signaling
Technology; 1:1000) after stripping. The signal intensity of
primary antibody binding was quantitatively analyzed with Sigma
Scan Pro 5 and was normalized to a loading control, β-actin
(10).
Flow cytometric anlysis (FCM)
Cells in different groups were collected with
trypsinization 72 h after transfection and washed twice with PBS.
Cells were fixed in 70% ethanol for 1 h at room temperature. After
centrifugation, the cell pellet was resuspended in PBS (pH 7.4),
containing 100 μl RNase A (1 mg/ml) and 400 μl propidium iodide (50
μg/ml). The cells were incubated for 30 min at room temperature,
and the DNA content was determined by flow cytometry using a
FACScan flow cytometer at 488 nm and the data were analyzed using
Lightcycle software. The experiment was performed three times in
triplicate (11). Proliferation
indices (PI) were calculated as follows: PI = (S + G2/M)/(G0/G1 + S
+ G2/M) x 100%.
WST-8 assay
MCF-7 cells were seeded into 96-well plates at a
density of 5×103 cells per well and cultured in regular
growth medium 24 h before transfection. Regular growth medium was
replaced by serum-free Opti-MEM 8 h later. These cells were grouped
similarly to cell transfection. The bulk volume of the transfection
compounds was 100 μl per well. Opti-MEM and transfection compounds
were replaced by complete medium 24 h after transfection. After
incubation for 72 h, MCF-7 cells were incubated for an additional 2
h with 20 μl of WST-8 dye (Beyotime Institute of Biotechnology,
Haimen, China). Absorbance of the three groups was measured with a
microplate reader (Model 550, Bio-Rad, Hercules, CA, USA) at a
wavelength of 450 nm (A450). All experiments were carried out eight
times (12).
In vitro cell migration and invasion
assay
The cells in different groups were treated with
trypsin and re-suspended as single-cell solutions 24 h after
transfection. A total of 2×105 cells in 0.5 ml of
serum-free RPMI 1640 medium were seeded on an 8-μm-pore
polycarbonate membrane Boyden chamber insert in a
Transwell® apparatus (Costar, Cambridge, MA, USA),
either coated with (invasion) or without (migration) Matrigel (BD
Biosciences, San Jose, CA, USA). RPMI-1640 (600 μl) containing 20%
FBS was added to the lower chamber. After incubation for 72
(invasion) or 36 h (migration) at 37°C in a 5% CO2
incubator, the cells on the top surface of the insert were removed
with a cotton swab. The cells that migrated to the bottom surface
of the insert were fixed in 100% methanol for 2 min, stained in
Giemsa for 2 min, rinsed in PBS and then subjected to microscopic
inspection (x200). Values for invasion and migration were obtained
by counting the number of stained cells in five fields per membrane
and data represented the average of three independent experiments
(13).
Statistical analysis
The data were presented as the means ± standard
error (SE) for MCF-7 cells in each group. Statistical analysis was
carried out by one-way ANOVA followed by Dunnett’s or Student’s
t-test (two means comparison). Statistical analysis was carried out
using the related programs in SPSS 12.0. Differences were
considered to indicate statistical significance at P<0.05.
Results
JMJD2A siRNA synthesis
The sequence of chemically synthesized JMJD2A siRNA
was consistent with the requirements, and the purity reached 98%.
This met the experimental requirements.
Observation of cell transfection
results
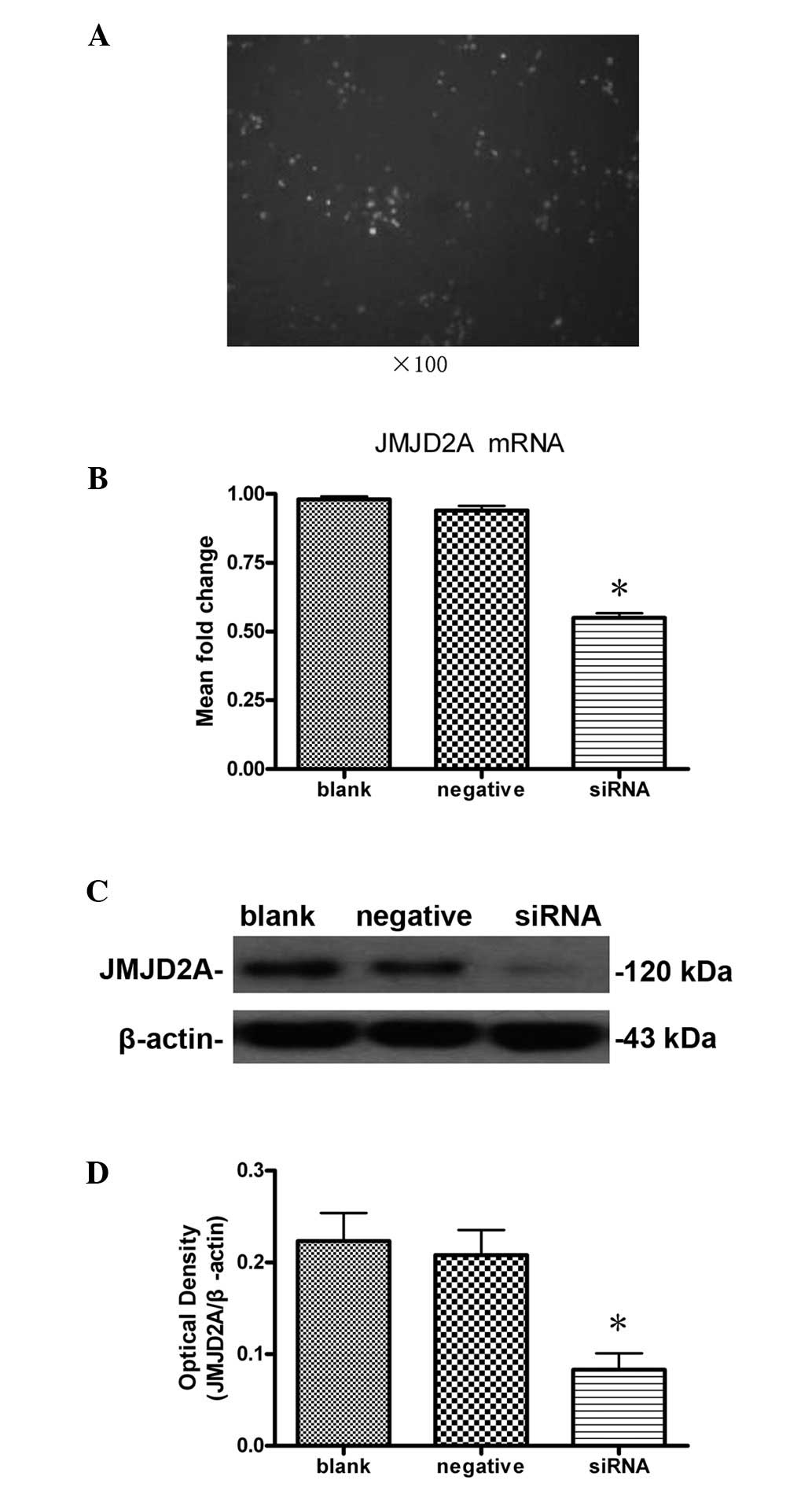
MCF-7 cells transfected with FAM-siRNA were
subjected to fluorescence microscopy 8 h after transfection. Cells
that showed green fluorescence were considered to be successfully
transfected. As shown in Fig. 1A,
cell transfection was successful and HiPerFect transfection reagent
was effective.
Downregulation of JMJD2A mRNA levels by
transfection with JMJD2A-specific siRNA
According to the results of quantitative real-time
PCR (Fig. 1B), no significant
difference (P>0.05) was detected in the levels of JMJD2A mRNA
between the blank (0.98±0.02) and negative (0.94±0.03) control
groups. The mRNA expression in the siRNA group (0.55±0.03) was
significantly lower than that of the blank (P<0.05) and negative
(P<0.05) control groups. These data indicate that JMJD2A mRNA
levels in MCF-7 cells decreased significantly following
transfection with JMJD2A siRNA. Transfection with JMJD2A-specific
siRNA may result in JMJD2A mRNA degradation to silence the JMJD2A
gene.
Inhibition of JMJD2A protein expression
in MCF-7 cells by transfection with JMJD2A-specific siRNA
Western blot analysis showed that the level of
JMJD2A protein expression in the siRNA group (0.083±0.031) were
significantly lower than those in the blank (0.223±0.053) and
negative (0.208±0.047) control groups (P<0.05; Fig. 1C and D), whereas the difference
between the blank and negative control groups was not significant
(P>0.05; Fig. 1C and D). These
data indicated that JMJD2A-specific siRNA silencing significantly
reduced the levels of JMJD2A mRNA and protein expression in MCF-7
cells.
Changes in cell cycle and inhibition of
proliferation in MCF-7 cells due to knockdown of JMJD2A
Cell cycle analysis by FCM revealed that JMJD2A
siRNA was capable of inducing changes in the cell cycle of MCF-7
cells. The mean value of the experiments was shown in Fig. 2A and B. There were no significant
differences (P>0.05) in the percentages of cells at each phase
between the blank and negative control groups. Compared with the
blank (44.24±1.86%) and negative (46.37±1.29%) control groups there
was a significant difference (P<0.05) in the percentage of cells
in the G0/G1 phase in the siRNA group (53.80±1.80%). Similarly,
there was a significant difference (P<0.05) in the percentage of
cells in the S phase in the siRNA group (36.55±1.52%) compared to
the blank (47.06±1.26%) and negative (44.72±1.86%) control groups.
However, there was no significant difference (P>0.05) in the
percentage of cells in the G2/M phase in the siRNA group
(9.64±1.26%), compared with blank (8.70±0.63%) and negative
(8.58±1.35%) control group. Knockdown of JMJD2A was capable of
increasing the percentage of cells in the G0/G1 phase and
decreasing the percentage of cells in the S phase. The results
indicate that the treatment was capable of arresting cells at the
G1/S checkpoint and delaying the cell cycle into the S phase.
Furthermore, proliferation indices (PI) of each group were
calculated. We found that there was a significant difference
(P<0.05) in the PI of the siRNA group (46.19±1.80%) versus those
of the blank (55.76±1.86%) and negative (53.63±1.29%) control
groups. Our results revealed changes in the cell cycle after
transfection, indicating that cell proliferation was inhibited by
transfection.
Additionally, the WST-8 assay was performed to test
the effects of transfection with JMJD2A siRNA on the proliferation
of MCF-7 cells in three different groups. As shown in Fig. 2C, there was no significant
difference (P>0.05) in the average absorbance between the blank
(1.73±0.02) and negative (1.69±0.03) control groups. The average
absorbance in the siRNA group (1.45±0.05) was significantly lower
than these values in the blank (P<0.05) and negative control
groups (P<0.05). Absorbance is representative of the cell
proliferation in WST-8 assay. The WST-8 assay results were
consistent with the FCM results. These data indicated that
transfection with JMJD2A siRNA significantly reduced the
proliferation of MCF-7 cells.
Suppression of MCF-7 cell migration and
invasion in vitro by knockdown of JMJD2A
As shown in Fig. 3,
cell migration was significantly decreased in the siRNA group
compared with the blank (P<0.05) and negative (P<0.05)
control groups. Cells in the siRNA group showed significantly
decreased invasiveness compared with the blank (P<0.05) and
negative control groups (Fig. 4;
P<0.05). These results demonstrated that transfection with
JMJD2A siRNA reduced the migration and invasion of MCF-7 cells.
Discussion
In this study, the results of the quantitative
real-time PCR and Western blot analysis revealed that knockdown of
JMJD2A in human breast cancer cell line MCF-7 was successful.
Furthermore, knockdown of JMJD2A caused inhibition of
proliferation, migration and invasion of MCF-7 cells. Human breast
cancer is the leading cause of cancer-related death among females
due to its powerful invasive ability and early metastatic
potential. Understanding the pathological mechanism of breast
cancer and identification of treatment target sites is a priority.
This study provides a new perspective in understanding the
molecular mechanisms underlying the progression of breast cancer,
as well as identifies a potential therapeutic target in breast
cancer.
Recent studies indicate that not only gene
dysfunction but also histone modifications are involved in breast
tumorigenesis (14). Coincidently,
JMJD2A is a lysine trimethyl-specific histone demethylase with the
capability of catalyzing the demethylation of H3K9me3 and H3K36me3
(6–8). H3K9 modifications are reported in
numerous biological phenomena including germ cell development, X
chromosome inactivation and DNA damage repair and apoptosis, and
deregulated histone methylation is associated with tumorigenesis
(15–17). Local chromatin architecture is
strongly influenced by post-translational histone modifications
such as methylation. Various histone modifications and histone
variants combine to the proposition of the regulatory histone code.
The histone code at least partly determines the transcriptional
potential for a specific gene or a genomic region (18–20).
H3K9me3 is predominant in coding regions of certain active genes
and is enriched in heterochromatin, with a generally repressive
nature (21–24). The intragenic permissive chromatin
regions are flanked by H3K9me3 and the maintenance of the
intragenic chromatin boundary appears to function as a checkpoint
in elongation (25). The balance
between methylated and demethylated histones may be broken by high
expression of endogenous JMJD2A, which catalyzes demethylation of
H3K9me3 excessively. Suv39H1, which is an H3K9 histone
methyltransferase, functions as a tumor suppressor by maintaining
H3K9 methylation levels (26,27).
Based on the literature, we hypothesize that JMJD2A may take part
in breast tumorigenesis through demethylation of H3K9me3 acting as
transcriptional activators.
The inhibition due to knockdown of JMJD2A appears to
support our hypothesis. However, the mechanism of JMJD2A in breast
cancer is elusive. Here, we discuss two possible pathways involved
according to the present literature.
One possible pathway is that JMJD2A may be involved
in the estrogen signaling pathway. A recent study focused on JMJD2
family proteins, particularly JMJD2B, which is considered to have a
similar function to that of JMJD2A in breast cancer. The study
demonstrated that JMJD2B comprises a key component of the estrogen
signaling pathway and the establishment of local epigenetic state
and chromatin structure required for proper induction of
ER-responsive genes (28). JMJD2B,
which interacts with ERα and components of the SWI/SNF-B chromatin
remodeling complex, was recruited to ERα target sites, demethylated
H3K9me3 and facilitated transcription of ER-responsive oncogenes
including MYB, MYC and CCND1, and knockdown of JMJD2B severely
impaired estrogen-induced cell proliferation and, consequently, the
tumor formation capacity of breast cancer cells. Our results are
consistent with this. JMJD2A may have the same mechanism as
JMJD2B.
The other possible candidate is the pRB-E2F complex
pathway. Depletion of JMJD2A caused only a marginal defect in ER
target gene induction in contrast to JMJD2B (28), which indicates another pathway in
which JMJD2A may be involved. JMJD2A has molecular characterization
in binding both retinoblastoma protein (pRB) and histone
deacetylases (HDACs) (4).
Associating with pRB, JMJD2A may recruit HDACs to the pRB-E2F
complex, change the chromatin structure at the E2F-responsive
promoters and induce repression transcription from E2F-dependent
promoters (29,30). E2F1, 4 and their complexes with
HDAC play an important role in down-regulating the expression of
the maternally imprinted tumor suppressor gene ARHI in breast
cancer cells (31). Expression of
ARHI is markedly downregulated in breast cancer, and reactivation
of ARHI expression in breast cancer cells is associated with
decreased H3K9me3, which is demethylated by JMJD2A (32). These studies support the
possibility of the pRB-E2F complex pathway.
Participating in either the estrogen signaling
pathway or pRB-E2F complex pathway or both, JMJD2A may play a
diverse role in human breast cancer. Though the exact mechanism
remains unclear, inhibition effects from knockdown of JMJD2A
indicate that JMJD2A participates in human breast cancer and may be
a potential therapeutic target.
To date, there are few studies focusing on JMJD2A in
human breast cancer. Our research verified the inhibition effects.
JMJD2A may be a potential therapeutic target in human breast
cancer. However, the present results were based on studies in a
single cell line in vitro. They have merely exposed the ‘tip
of the iceberg’ of the role of JMJD2A in human breast cancer.
Further studies to elucidate the pleiotropic functions of JMJD2A
and its contribution to human breast cancer in vitro and
in vivo are required.
Acknowledgements
The work was supported by the National
Science Foundation of China (nos. 81172897 and 81072512).
References
|
1
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar
|
|
2
|
Sen GL and Blau HM: A brief history of
RNAi: the silence of the genes. FASEB J. 20:1293–1299. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Katoh M and Katoh M: Identification and
characterization of JMJD2 family genes in silico. Int J
Oncol. 24:1623–1628. 2004.PubMed/NCBI
|
|
4
|
Gray SG, Iglesias AH, Lizcano F,
Villanueva R, Camelo S, Jingu H, Teh BT, Koibuchi N, Chin WW,
Kokkotou E and Dangond F: Functional characterization of JMJD2A, a
histone deacetylase- and retinoblastoma-binding protein. J Biol
Chem. 280:28507–28518. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Shin S and Janknecht R: Activation of
androgen receptor by histone demethylases JMJD2A and JMJD2D.
Biochem Biophys Res Commun. 359:742–746. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Trojer P and Reinberg D: Histone lysine
demethylases and their impact on epigenetics. Cell. 125:213–217.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Whetstine JR, Nottke A, Lan F, Huarte M,
Smolikov S, Chen Z, Spooner E, Li E, Zhang G, Colaiacovo M and Shi
Y: Reversal of histone lysine trimethylation by the JMJD2 family of
histone demethylases. Cell. 125:467–481. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Nottke A, Colaiácovo MP and Shi Y:
Developmental roles of the histone lysine demethylases.
Development. 136:879–889. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhang XD, Wang Y, Wang Y, Zhang X, Han R,
Wu JC, Liang ZQ, Gu ZL, Han F, Fukunaga K and Qin ZH: p53 mediates
mitochondria dysfunction-triggered autophagy activation and cell
death in rat striatum. Autophagy. 5:339–350. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Luo CL, Li BX, Li QQ, Chen XP, Sun YX, Bao
HJ, Dai DK, Shen YW, Xu HF, Ni H, et al: Autophagy is involved in
traumatic brain injury-induced cell death and contributes to
functional outcome deficits in mice. Neuroscience. 184:54–63. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Dai HY, Liu L, Qin SK, He XM and Li SY:
Lobaplatin suppresses proliferation and induces apoptosis in the
human colorectal carcinoma cell Line LOVO in vitro. Biomed
Pharmacother. 65:137–141. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Tang Y, Huang B, Sun L, Peng X, Chen X and
Zou X: Ginkgolide B promotes proliferation and functional
activities of bone marrow-derived endothelial progenitor cells:
involvement of Akt/eNOS and MAPK/p38 signaling pathways. Eur Cell
Mater. 21:459–69. 2011.
|
|
13
|
Li L, Zhang C, Li X, Lu S and Zhou Y: The
candidate tumor suppressor gene ECRG4 inhibits cancer cells
migration and invasion in esophageal carcinoma. J Exp Clin Cancer
Res. 29:1332010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Jovanovic J, Rønneberg JA, Tost J and
Kristensen V: The epigenetics of breast cancer. Mol Oncol.
4:242–254. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Martin C and Zhang Y: The diverse
functions of histone lysine methylation. Nat Rev Mol Cell Biol.
6:838–849. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Müller-Tidow C, Klein HU, Hascher A, Isken
F, Tickenbrock L, Thoennissen N, Agrawal-Singh S, Tschanter P,
Disselhoff C, Wang Y, Becker A, Thiede C, Ehninger G, zur Stadt U,
Koschmieder S, Seidl M, Müller FU, Schmitz W, Schlenke P,
McClelland M, Berdel WE, Dugas M and Serve H; Study Alliance
Leukemia: Profiling of histone H3 lysine 9 trimethylation levels
predicts transcription factor activity and survival in acute
myeloid leukemia. Blood. 116:3564–3571. 2010.
|
|
17
|
Cloos PA, Christensen J, Agger K and Helin
K: Erasing the methyl mark: histone demethylases at the center of
cellular differentiation and disease. Genes Dev. 22:1115–1140.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Schübeler D, MacAlpine DM, Scalzo D,
Wirbelauer C, Kooperberg C, van Leeuwen F, Gottschling DE, O’Neill
LP, Turner BM, Delrow J, Bell SP and Groudine M: The histone
modification pattern of active genes revealed through genome-wide
chromatin analysis of higher eukaryote. Genes Dev. 18:1263–1271.
2004.PubMed/NCBI
|
|
19
|
Shilatifard A: Chromatin modifications by
methylation and ubiquitination: implications in the regulation of
gene expression. Annu Rev Biochem. 75:243–269. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Xu D, Bai J, Duan Q, Costa M and Dai W:
Covalent modifications of histones during mitosis and meiosis. Cell
Cycle. 8:3688–3694. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Mikkelsen TS, Ku M, Jaffe DB, Issac B,
Lieberman E, Giannoukos G, Alvarez P, Brockman W, Kim TK, Koche RP,
et al: Genome-wide maps of chromatin state in pluripotent and
lineage-committed cells. Nature. 448:553–560. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Barski A, Cuddapah S, Cui K, Roh TY,
Schones DE, Wang Z, Wei G, Chepelev I and Zhao K: High-resolution
profiling of histone methylations in the human genome. Cell.
129:823–837. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Brinkman AB, Roelofsen T, Pennings SW,
Martens JH, Jenuwein T and Stunnenberg HG: Histone modification
patterns associated with the human X chromosome. EMBO Rep.
7:628–634. 2006.PubMed/NCBI
|
|
24
|
Vakoc CR, Mandat SA, Olenchock BA and
Blobel GA: Histone H3 lysine 9 methylation and HP1gamma are
associated with transcription elongation through mammalian
chromatin. Mol Cell. 19:381–391. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Gomes NP and Espinosa JM: Gene-specific
repression of the p53 target gene PUMA via intragenic CTCF-Cohesin
binding. Genes Dev. 24:1022–1034. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Peters AH, O’Carroll D, Scherthan H,
Mechtler K, Sauer S, Schöfer C, Weipoltshammer K, Pagani M, Lachner
M, Kohlmaier A, et al: Loss of the Suv39h histone
methyltransferases impairs mammalian heterochromatin and genome
stability. Cell. 107:323–337. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Braig M, Lee S, Loddenkemper C, Rudolph C,
Peters AH, Schlegelberger B, Stein H, Dörken B, Jenuwein T and
Schmitt CA: Oncogene-induced senescence as an initial barrier in
lymphoma development. Nature. 436:660–665. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Kawazu M, Saso K, Tong KI, McQuire T, Goto
K, Son DO, Wakeham A, Miyagishi M, Mak TW and Okada H: Histone
demethylase JMJD2B functions as a co-factor of estrogen receptor in
breast cancer proliferation and mammary gland development. PLoS
One. 6:e178302011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Takaki T, Fukasawa K, Suzuki-Takahashi I
and Hirai H: Cdk-mediated phosphorylation of pRB regulates HDAC
binding in vitro. Biochem Biophys Res Commun. 316:252–255.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Lai A, Kennedy BK, Barbie DA, Bertos NR,
Yang XJ, Theberge MC, Tsai SC, Seto E, Zhang Y, Kuzmichev A, et al:
RBP1 recruits the mSIN3-histone deacetylase complex to the pocket
of retinoblastoma tumor suppressor family proteins found in limited
discrete regions of the nucleus at growth arrest. Mol Cell Biol.
21:2918–2932. 2001. View Article : Google Scholar
|
|
31
|
Lu Z, Luo RZ, Peng H, Huang M, Nishmoto A,
Hunt KK, Helin K, Liao WS and Yu Y: E2F-HDAC complexes negatively
regulate the tumor suppressor gene ARHI in breast cancer. Oncogene.
25:230–239. 2006.PubMed/NCBI
|
|
32
|
Yu Y, Xu F, Peng H, Fang X, Zhao S, Li Y,
Cuevas B, Kuo WL, Gray JW, Siciliano M, Mills GB and Bast RC Jr:
NOEY2 (ARHI), an imprinted putative tumor suppressor gene in
ovarian and breast carcinomas. Proc Natl Acad Sci USA. 96:214–219.
1999. View Article : Google Scholar : PubMed/NCBI
|