Introduction
Autophagy associated with drug resistance is a major
obstacle in cancer chemotherapy (1,2). It has
been reported that autophagy-deficient tumors are typically more
sensitive to chemotherapeutic agents than are their
autophagy-proficient counterparts (3). In the previous study, it was discovered
that human drug-resistant K562/ADM leukemia cells preferentially
emerged with higher autophagy activity compared with the parental,
sensitive K562 cells following starvation treatment (4). However, autophagy is considered to have
a dual role in tumor cell resistance, since previous research has
shown conflicting roles for autophagy with regard to drug
resistance in different cancer cell lines; specifically, autophagy
either promotes or suppresses resistance via various mechanisms
(5–7).
Resveratrol (trans-3,4′,5-trihydroxystilbene;
Res), a natural polyphenolic compound, has been reported to have
numerous beneficial effects, including anti-oxidant and
anti-inflammatory activities, and an inhibitory effect on tumor
growth (8,9). Yan et al (10) reported that Res not only induced
leukemia cell apoptosis and erythroid differentiation, but also
autophagy. A previous report has indicated that Res induces
autophagy in different cancer cell lines via either a pro-survival
or pro-death mechanism (11).
Additionally, a number of studies have demonstrated that Res
increases the sensitivity of malignancies, including melanoma,
prostate and non-small lung cancer, to chemotherapy (12,13).
Although Res is known to induce apoptosis and autophagy in
different types of tumor cells, few studies have explored the
effects of Res-induced autophagy and its association with apoptosis
in drug-resistant leukemia cells.
The molecular regulators interconnecting between
autophagy and apoptosis, including B-cell lymphoma 2 (Bcl-2),
Bcl-2-associated X protein (Bax) and beclin 1, have been suggested
to act as switching points that are critical for the outcome of
tumor cells, and lysosomes have been reported to initiate the cell
death pathway in autophagic cells (14–16). It
has been demonstrated that Res activates a lysosome-dependent
cytotoxic pathway that results in caspase-dependent cell death in
colorectal cancer cells (17). Other
results have shown that lysosomal cathepsin L mediates Res-induced
autophagy and apoptotic cell death in cervical cancer cells
(18). However, few studies have
investigated the interrelation of autophagy, apoptosis and drug
resistance in leukemia. The molecular pathway by which autophagy
leads to apoptosis in drug-resistant conditions was explored in the
present study. Furthermore, the present study also examined the
fate of K562/ADM cells during sustained Res exposure. To the best
of our knowledge, the present findings are the first to show that
autophagy-dependent apoptosis is mediated by lysosomal cathepsin D
(Cath D).
Materials and methods
Reagents
Resveratrol (Res), MTT, dimethyl sulfoxide (DSMO),
3-methyladenine (3-MA), monodansylcadaverine (MDC),
microtubule-associated protein 1A/1B-light chain 3 (LC3) antibody
and RPMI-1640 were purchased from Sigma-Aldrich (Merck KGaA,
Darmstadt, Germany). Fetal bovine serum (FBS) was obtained from
Sijiqing Biological Engineering Materials Co., Ltd. (Hangzhou,
China). Antibodies against beclin 1, Bcl-2, p62, cleaved caspase-3
and Bax were obtained from Cell Signaling Technology, Inc.
(Danvers, MA, USA).
Cell culture
K562/ADM ADM-resistant cell line was kindly provided
by Dr Zhang Jiwang from Hematological laboratory, Ruijin Hospital
Affiliated to the Shanghai Jiao Tong University School of Medicine
(Shanghai, China). Cells were grown in RPMI-1640 medium
supplemented with 10% (v/v) FBS, 2 mM glutamine, 100 U/ml
penicillin and 100 U/ml streptomycin at 37°C in a humidified
atmosphere containing 5% CO2. K562/ADM cells were
stimulated with 5 mg/l Adriamycin (ADM; Shenzhen Main Luck
Pharmaceuticals, Inc., Shenzhen, China) every 45 days to maintain
increased drug resistance and then experiments were performed after
2 weeks of culturing without ADM. Cells in the exponential phase
were used in the experiments.
Cell viability assay
Cells were incubated with various concentrations of
Res and were collected for determination of the half-maximal
inhibitory concentrations (IC50) of Res using an MTT
assay. Briefly, K562/ADM cells were seeded at a density of
1×105 cells/ml in 96-well plates. The cells were treated
with 0, 20, 40, 80, 120 and 160 µmol/l of Res for 24, 48 or 72 h at
37°C. The controls were treated with 0 µmol/l of Res. A total of 10
µl MTT (5 mg/ml) was added to each well, followed by incubation for
4 h at 37°C. 100 µl DMSO was added to each of the wells, and
incubation for 20 min at 37°C to dissolve the formazan crystals.
The absorbance at 490 nm was measured using a PowerWave X plate
reader (Bio-Tek Instruments, Inc., Winooski, VT, USA). Moreover, an
autophagic inhibitor 3-MA (5 mmol/l) was added to the cells 3 h
prior to Res treatment to confirm the role of autophagy in
Res-induced cell death.
Transmission electron microscopy
K562/ADM cells (treated with Res at 0, 40 or 80
µmol/l at 37°C for 48 h) were fixed with 2.5% glutaraldehyde and 2%
paraformaldehyde in 0.1 mol/l phosphate buffer for 90 min at room
temperature, followed by post-fixation with 1% osmium tetroxide for
30 min (10). Subsequently, the
cells were gradually dehydrated in 10% graded series of 50–100%
ethanol and propylene oxide, and embedded in Epon 812 resin.
Finally, the blocks were cut into 70-nm-thick ultrathin sections
with a microtome, which were then stained with uranyl acetate for
30 min and lead citrate for 15 min at 37°C for observation. The
ultrastructure of the cells was examined using a JEMI1230
transmission electron microscope (JEOL, Ltd., Tokyo, Japan). The
controls were treated with 0 µmol/l of Res.
MDC staining
K562/ADM cells (treated with Res at 0, 40 or 80
µmol/l at 37°C for 24 h) were stained with 0.05 mmol/l MDC at 37°C
for 30 min. The cells were washed three times with PBS and
autophagic vacuoles were observed with a fluorescence inverted,
phase contrast microscope (Olympus Corporation, Tokyo, Japan).
Flow cytometric analysis
K562/ADM cells were pre-incubated with or without
3-MA for 3 h and treated with Res at 0, 40, or 80 µmol/l for 48 h
prior to detection of cell apoptosis using an Annexin V-FITC
Apoptosis Detection Kit (BD Biosciences, San Jose, CA, USA). A
total of 1×106 cells were collected and suspended in 100
ml binding buffer containing 0.25 mg of Annexin V-FITC. After 15
min of incubation in the dark at room temperature, cells were
washed by PBS and suspended in the binding buffer with 5 mg of PI.
Then, the stained cells were analyzed using a flow cytometer (EPICS
XL; Beckman Coulter, Inc., Miami, FL, USA) and analyzed using
Windows Multiple Document Interface for Flow Cytometry software
(version 2.8; The Scripps Research Institute, La Jolla, CA,
USA).
Western blotting
At the end of the treatment with Res at 0, 40, or 80
µmol/l for 24, 48 and 72 h, the cells were washed twice with PBS
and lysed in ice-cold RIPA lysis buffer (Beijing Solarbio Science
& Technology Co. Ltd., Beijing, China). Total protein
concentration was determined using a BCA protein quantitative kit
(cat no., P0010s; Beyotime Institute of Biotechnology, Haimen,
China). Protein extracts (40 µg/lane) were separated by 10–12%
SDS-PAGE and electrophoretically transferred onto polyacrylamide
difluoride membranes (EMD Millipore, Billerica, MA, USA). The
membranes were blocked in 5% non-fat milk for 1 h at room
temperature, followed by overnight incubation with primary
antibodies at 4°C. The primary antibodies used were as follows:
Beclin 1 (cat. no. 3738), LC3 (cat. no. ABC929; EMD Millipore),
Bcl-2 (cat. no. 2872), cleaved caspase-3 (cat. no. 9661), Bax (cat.
no. 2774); p62 (cat. no. 5114), Cath D (cat. no. sc-10725; Santa
Cruz Biotechnology, Inc., Dallas, TX, USA) (all 1:1,000), P-gp
(1:500; cat. no. BA1351-2; Boster Biological Technology, Wuhan,
China) and β-actin (1:700; cat. no. sc-47778; Santa Cruz
Biotechnology, Inc.). Subsequently, the membranes were washed three
times for 5 min with PBS in Tween 20 and incubated again with
horseradish peroxidase-conjugated goat anti-rabbit (cat. no.
ZB-2301) or goat anti-mouse (cat. no. ZB-2305) antibodies (both
1:10,000; OriGene Technologies, Inc., Beijing, China) at 37°C for 1
h. Protein bands were visualized using enhanced chemiluminescence
reagents (EMD Millipore). Western blots were scanned using an
Infrared Imaging System (LI-COR Biosciences, Lincoln, NE, USA) and
the bands were quantified using Image J software (version 1.45S;
National Institutes of Health; Bethesda, MD, USA). β-actin was used
as a reference protein.
Statistical analysis
Statistical analysis was performed using SPSS 16.0
software (SPSS, Inc., Chicago, IL, USA). All the experiments were
repeated at least three times. Data are presented as the mean ±
standard deviation. One-way analysis of variance followed by
Tukey's least significant difference post-hoc test was used to
analyze statistical differences among groups under different
conditions. P<0.05 was considered to indicate a statistically
significant difference.
Results
Res induces autophagy in K562/ADM
cells
To determine whether Res induces autophagy in
K562/ADM cells, autophagic bodies were investigated using
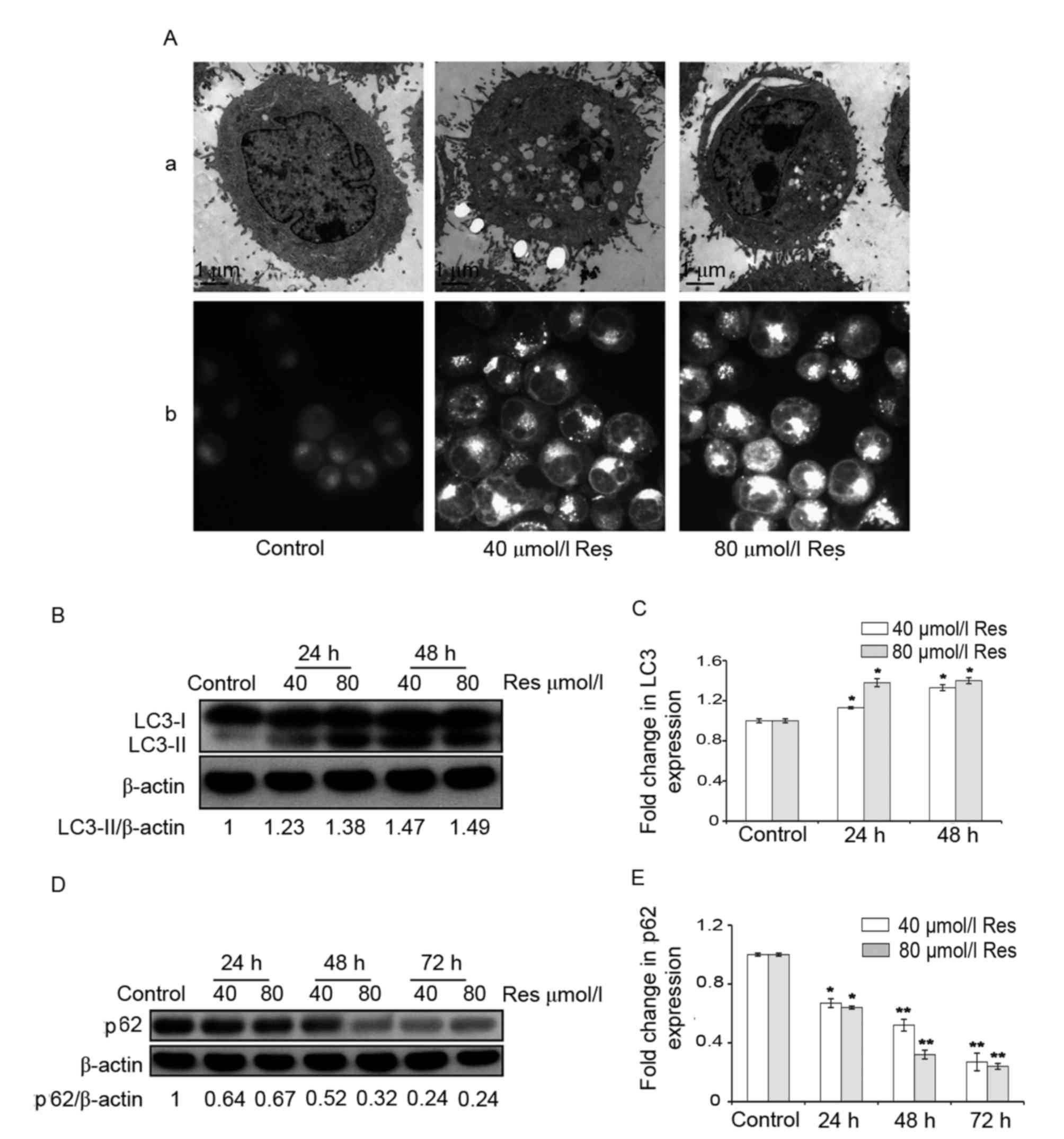
transmission electron microscopy. Compared with the control group,
a large number of autophagic vacuoles appeared in the cytoplasm
when K562/ADM cells were treated with 40 or 80 µmol/l Res (Fig. 1A-a). Furthermore, the number of
MDC-positive fluorescent points and the amount of engulfed
cytoplasmic material in Res-treated cells was markedly greater
compared with that in the control group, as observed under a
fluorescence microscope (Fig.
1A-b).
The expression of autophagic LC3 protein was
determined using western blotting. There are two forms of LC3:
LC3-I and LC3-II (3,19). During the process of autophagy, the
cytoplasmic form, LC3-I, is converted into LC3-II, which is the
form that is bound to the autophagosome membrane (3,19). The
results revealed that, following treatment with 40 or 80 µmol/l
Res, the relative ratio of LC3-II/β-actin increased in K562/ADM
cells compared with the control (Fig.
1B). This difference was indicated to be statistically
significant (P<0.05; Fig.
1C).
In addition to LC3, the expression level of p62
protein, which is another autophagy-specific biomarker was also
explored (19,20). The p62 protein acts as a signaling
hub through its involvement in the formation of autophagosomes and
is degraded during autophagy (19,20). The
results indicated that p62 protein expression levels were decreased
by Res in a time-dependent manner, suggesting an increase in
autophagic flux (Fig. 1D).
Furthermore, the difference between the Res-treated cells and the
control was statistically significant (P<0.05; Fig. 1E).
Inhibition of autophagy attenuates the
Res-induced apoptosis of K562/ADM cells
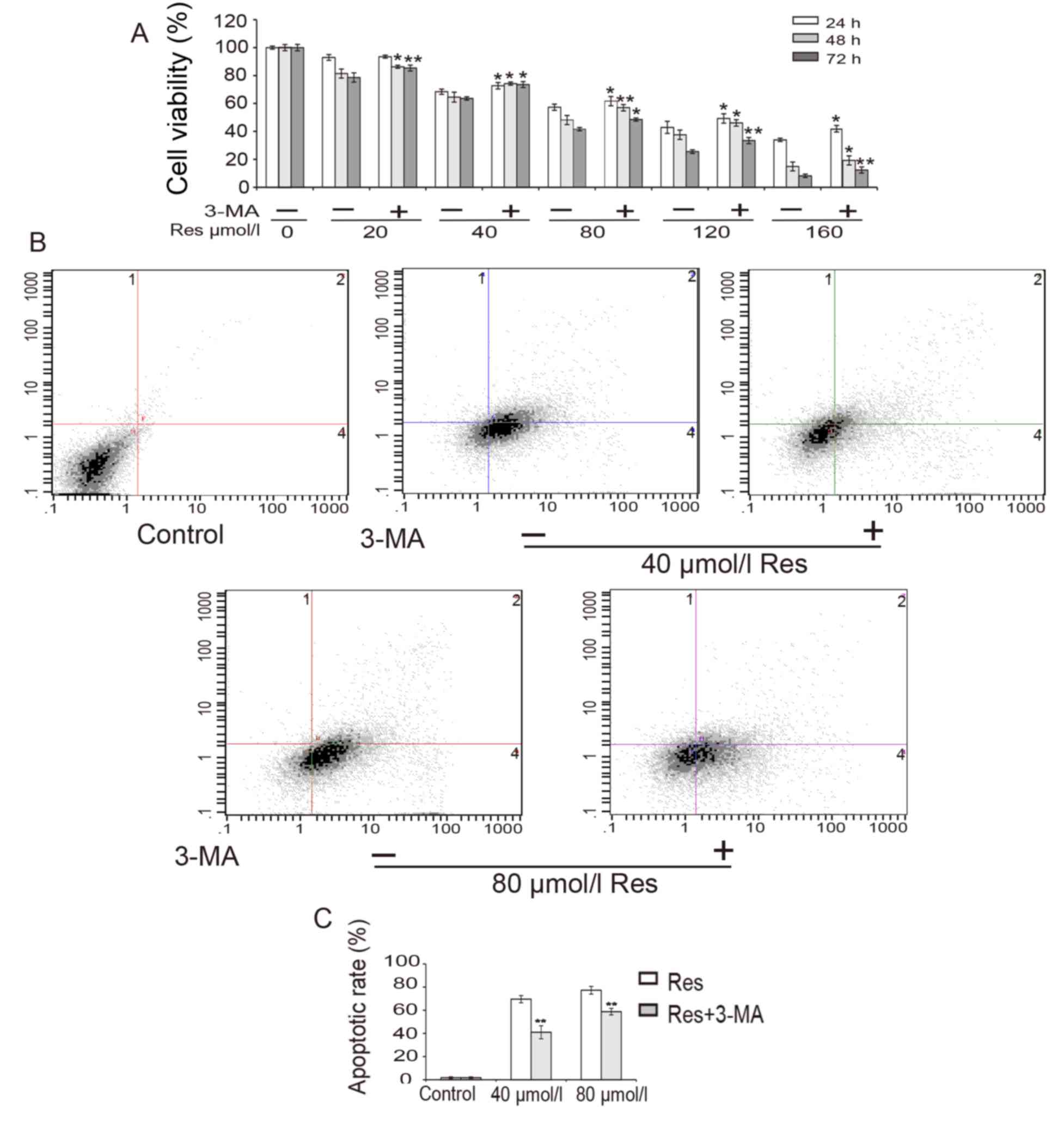
The effect of Res on the viability of K562/ADM cells
was investigated. K562/ADM cells were treated with different
concentrations of Res (0–160 µmol/l) for 24, 48 or 72 h, and cell
viability was assessed by MTT assay. As shown in Fig. 2A, cell viability decreased in a time-
and dose-dependent manner. The IC50 of Res in K562/ADM
cells at 48 h was calculated to be 57.7 µmol/l. These data indicate
that Res reduced the survival of K562/ADM cells. In addition, the
results revealed that cell viability was restored when cells were
treated with 5 mmol/l 3-MA for 3 h prior to treatment with Res
(Fig. 2A).
To determine the association between cell death and
apoptosis, a flow cytometry experiment was conducted (Fig. 2B and C). The results revealed that
the proportion of apoptotic cells increased following treatment
with Res in a dose-dependent manner, which suggested that Res
induced apoptosis in K562/ADM cells. However, following
pre-treatment with 3-MA, the proportion of apoptotic cells was
significantly decreased from 69.6 to 41.0% for 40 µmol/l Res
treatment and from 77.3 to 58.8% for 80 µmol/l Res treatment
(P<0.01; Fig. 2C).
These data demonstrated that Res inhibited cell
viability and that Res-induced apoptosis was reduced by 3-MA in
K562/ADM cells. These results suggest that the Res-induced
apoptosis of K562/ADM cells was autophagy-dependent.
Protein expression levels of Bax/Bcl-2
and beclin 1 increase during the Res-induced autophagy and
apoptosis of K562/ADM cells
To further explore the mechanism by which Res
induces apoptosis, the expression levels of cleaved caspase-3,
Bcl-2, Bax, beclin 1 and Cath D proteins were determined using
western blotting (Fig. 3). Following
treatment with 40 or 80 µmol/l Res for 24 and 48 h, the expression
levels of cleaved caspase-3 increased in a dose-dependent manner in
K562/ADM cells (Fig. 3A). In
addition, differences in the protein expression levels of cleaved
caspase-3 between the control and Res-treated cells were
statistically significant (P<0.05; Fig. 3B). The data suggested that Res
significantly induced apoptosis through the activation of
caspase-3.
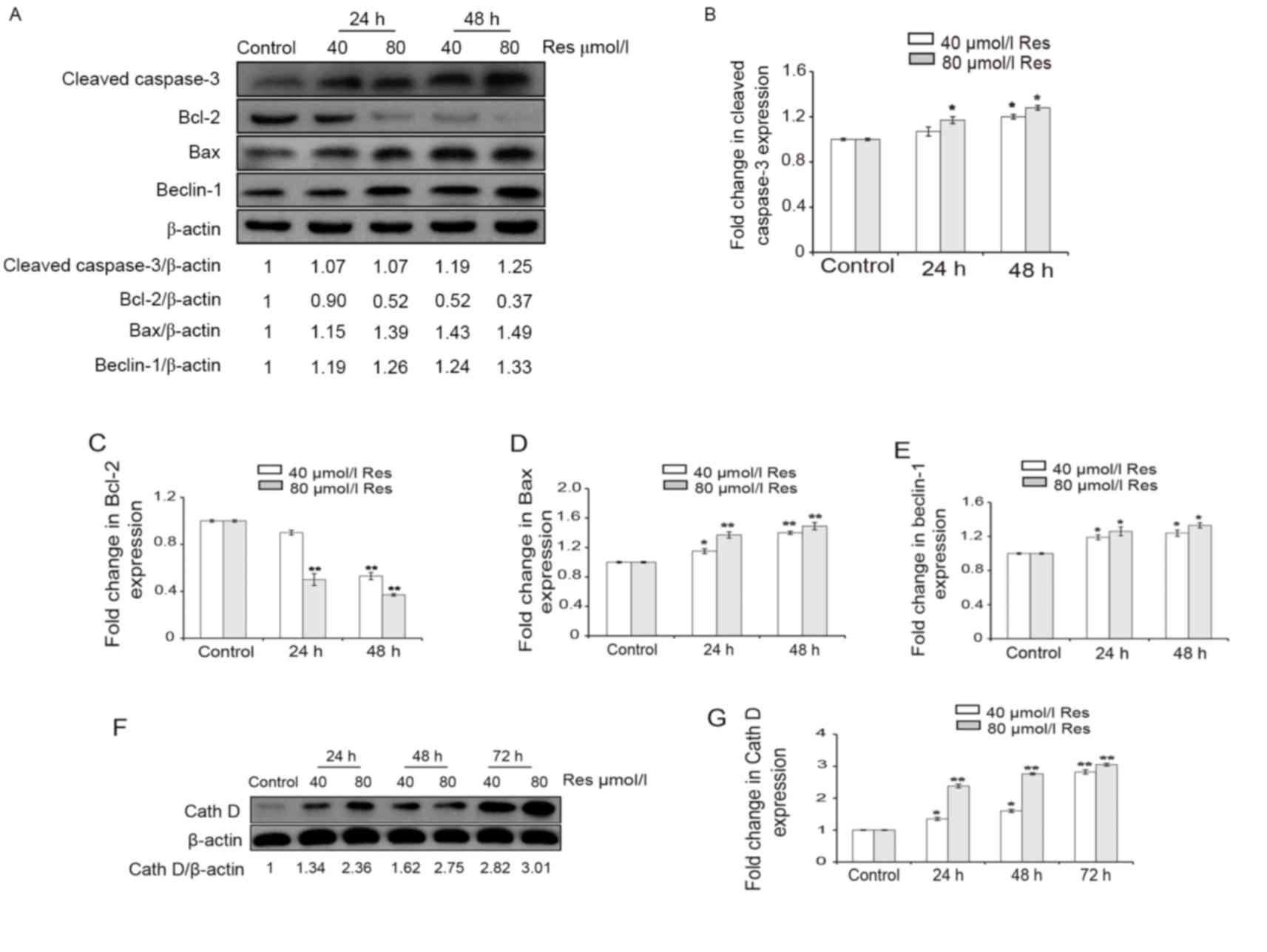 | Figure 3.Protein expression levels of cleaved
caspase-3, Bcl-2, Bax, beclin 1 and Cath D were detected by western
blotting. Cells were treated with 40 and 80 µmol/l Res for 24, 48,
or 72 h. β-actin served as an internal control. (A) Blot images of
cleaved caspase-3, Bcl-2, Bax and beclin 1 protein expression.
Quantification of (B) cleaved caspase-3, (C) Bcl-2, (D) Bax and (E)
beclin 1 protein expression levels. (F) Blot image of Cath D
protein expression. (G) Quantification of Cath D protein expression
levels. Data are represented as the mean ± standard deviation of
triplicate experiments. The control group was treated with 0 µmol/l
Res. *P<0.05 and **P<0.01 vs. control group. Res,
resveratrol; Bcl-2, B-cell lymphoma 2; Bax, Bcl-2 associated X
protein; Cath D, cathepsin D. |
The Bcl-2 family is a crucial integrator of cellular
survival and death signals (14,16).
Bcl-2 is an anti-apoptotic protein that has the ability to inhibit
autophagy and apoptosis through interactions with beclin 1 and
Bax/Bak, respectively, which are mediated through their
Bcl-2-homology-3-binding pockets (14,16). The
western blotting results indicated that the expression levels of
Bcl-2 were decreased by Res (Fig. 3A and
C); this was accompanied by increased expression levels of Bax
protein in a dose-dependent manner (Fig.
3A and D). Statistically significant differences were detected
in the protein expression levels of Bax and Bcl-2 in Res-treated
cells compared with the control (P<0.05; Fig. 3C and D). Therefore, the increased
expression levels of Bax and decreased expression levels of Bcl-2
may be important in the apoptotic process in K562/ADM cells
following treatment with Res.
Beclin 1, another hallmark of autophagosomes, is a
well-known protein with respect to the formation of autophagic
vesicles (1,5). Western blotting results in the present
study demonstrated that the expression level of beclin 1 protein
was increased by 1.19- to 1.33-fold following treatment with Res,
further corroborating the hypothesis that autophagy is induced by
Res treatment (Fig. 3A and E). In
addition, the differences in beclin 1 protein expression levels
between the Res-treated cells and the control were statistically
significant (P<0.05; Fig. 3E).
These results suggest that complex interactions of the
anti-apoptotic Bcl-2 and pro-autophagic beclin 1 proteins
co-regulate the ultimate fate of Res-induced K562/ADM cells with
respect to apoptosis.
Cath D is activated during res-induced
autophagy and apoptosis in K562/ADM cells
Lysosomal proteases, also known as cathepsins,
participate in the activation of the intrinsic programmed cell
death pathway (17,19). Cath D, a major intracellular aspartic
protease, is typically involved in apoptosis through its release
from lysosomes (17,19). For this reason, whether Cath D is a
possible mediator of Res-induced cytotoxicity was investigated in
the present study. The western blotting results indicated that Cath
D expression levels were increased by res in a time- and
concentration-dependent manner, with an increase of ~3-fold in the
80 µmol/l Res-treated group at 72 h compared with the control group
(Fig. 3F and G). Furthermore, the
differences in Cath D protein expression levels between the
Res-treated cells and the control were statistically significant
(P<0.05; Fig. 3G). This suggests
that lysosomal Cath D may be involved in the process of Res-induced
autophagy and the subsequent progression to apoptotic death in
K562/ADM cells treated with Res.
Discussion
Res has been shown to induce cell death through
various mechanisms in different cancer cell lines (2,11,12,19).
Various antitumor effects of Res have been described previously,
including the inhibition of DNA polymerase, protein kinase C,
cyclooxygenase-2 or interleukin-6 activities, and the suppression
of transcription factors such as nuclear factor-κB (4,11,12,21,22).
The present study revealed that Res increased autophagosome
formation, decreased cell viability and induced apoptotic death in
drug-resistant K562/ADM cells. It was hypothesize that the
mediation of these effects by Res may involve the permeation of Res
through the lysosomal membrane, resulting in the release of Cath D
from lysosomes into the cytosol. In addition, the released Cath D
may trigger caspase-dependent cell death through interaction with
Bcl-2 and beclin 1 and the activation of Bax.
Yan et al (10) reported that Res inhibited the
proliferation of K562 leukemia cells and induced apoptosis and
autophagy, and demonstrated the contribution of Res-induced
autophagy to erythroid differentiation in K562 cells. However, the
present results indicated that autophagy preceded apoptosis in
K562/ADM cells following Res treatment. To investigate the role of
autophagy in Res-induced cell death, 3-MA, an autophagic inhibitor
(4,15), was used to block autophagy and
resulted in the marked attenuation of Res-induced apoptosis. These
data indicated that Res-induced apoptosis may be
autophagy-dependent in K562/ADM cells.
In the present study, autophagic responses increased
significantly following Res treatment for 24 h in K562/ADM cells.
As observed under a transmission electron microscope, a large
number of autophagic bodies appeared in K562/ADM cells following
Res treatment. This finding was further supported by evaluation of
the expression levels of LC3 using western blotting. LC3 is
considered to be a specific marker of autophagic activity (3,19). The
LC3-II/β-actin ratio was significantly increased in K562/ADM cells
treated with 40 or 80 µmol/l Res for 24 or 48 h. Previous studies
have indicated that the LC3-II turnover assay is suitable for
monitoring autophagic flux in the presence of a lysosomal inhibitor
(23,24). To distinguish whether autophagosome
accumulation was due to enhanced autophagic activity or reduced
autophagosome degradation, the protein expression levels of p62
were determined in the present study. p62, an autophagy-related
protein, is involved in the formation of autophagosomes and
degraded through autophagy (19,20). The
present results showed that the protein expression level of p62 was
significantly decreased in Res-treated cells compared with the
control. These data therefore indicated that Res induced a high
level of LC3-II protein by the induction of autophagic activity
rather than by blocking autophagosome degradation in K562/ADM
cells.
Beclin 1 is required for the initiation stage in the
formation of the autophagosome (1,5). In the
present study, an increase in beclin1 protein expression levels was
observed in Res-treated cells compared with the control, suggesting
an enhanced induction of autophagy. Beclin 1 was originally
identified as a Bcl-2-interacting protein. Previous studies have
demonstrated that beclin 1 and Bcl-2 interact in a multiprotein
complex on the membrane of the endoplasmic reticulum; this
observation in turn suggests a mechanism by which these proteins
assist in mediating the intricate relationship between apoptosis
and autophagy (1,16). It is known that beclin 1 generates a
pro-apoptotic protein fragment and also interacts with the
anti-apoptotic Bcl-2 family protein to inhibit autophagy and
initiate mitochondrial apoptosis (22). Prabhu et al (25) reported that the silencing of key
regulators of autophagy, such as beclin 1 and autophagy protein
(ATG)5, significantly enhanced Res-induced caspase activation. In
the present study, western blotting results revealed that Res
significantly increased the expression levels of Bax and decreased
those of Bcl-2. The interaction of beclin 1 with Bcl-2 in the
cytosolic compartment may increase efficient Bax translocation to
mitochondria, leading to permeabilization of the mitochondrial
membrane, cytochrome c release and activation of a caspase
cascade, and finally, initiation of caspase-dependent cell death
(2,10,11).
Lysosomes contain numerous hydrolases, which degrade
intracellular and extracellular delivered materials, and have been
implicated in the regulation of cell death (17,19).
Various stimuli may induce lysosomal membrane permeabilization,
thereby releasing a class of hydrolytic enzymes, the cathepsins,
from the lysosomal compartment into the cytosol, which contributes
to apoptosis (17,18). Cath D has been suggested to be a key
regulator of apoptosis via its catalytic activity, and there is
also evidence to support the hypothesis that cathepsins act in
concert with caspases in apoptotic cell death (17). However, little is known regarding the
mechanism by which Cath D mediates the cytotoxic agent-induced
autophagic death of resistant leukemia cells. In the present study,
a significantly higher protein expression level of Cath D was
observed during the autophagy and apoptosis of Res-induced K562/ADM
cells, and this indicated that Cath D may have a key role in
Res-induced autophagy-dependent apoptotic death in these cells.
Trincheri et al (17)
considered Bax to be the probable target of Cath D in Res-mediated
cytotoxicity. Furthermore, Yelamanchili et al (26) observed that the upregulation of Cath
D correlated with cellular damage leading to typical
caspase-dependent activation of apoptosis. The present study data
revealed that cleaved caspase-3 and Cath D protein expression
levels were significantly increased following Res treatment and,
therefore, may be involved in mediating the apoptotic effect of Res
in K562/ADM cells. Specifically, the death pathway may be initiated
by the promotion of lysosomal membrane permeabilization and release
of lysosomal Cath D to the cytosol to trigger the activation of Bax
and the selective mitochondrial release of apoptosis-inducing
factors, eventually resulting in activation of the caspase
proteolytic cascade and apoptosis (17,19,24).
Autophagy has a cytoprotective function under stress
conditions, such as during cytotoxic drug administration, and may
result in increased drug resistance (1). It has been reported that autophagy
increases the resistance of colorectal cancer stem-like cells to
photodynamic therapy-induced apoptosis (1,27,28).
Similarly, cisplatin-activated autophagy, involving the activation
of ATG7, has been indicated to increase liver cancer cell
resistance to cisplatin (29). Eum
et al (1) suggested that the
inhibition of autophagy may resensitize resistant tumor cells to
anticancer therapy. Previous observations by our group highlighted
the elevation of autophagic activity in drug-resistant K562/ADM
cells (15). These findings suggest
that cancer cells may activate the autophagic process to fight
against cytotoxin-induced cell injury by recycling damaged
organelles and proteins, resulting in drug resistance. However,
under sustained exposure to cytotoxic agents, autophagic machinery
could be recruited to trigger both caspase-dependent and
-independent lethal pathways (17,19). In
addition, P-glycoprotein serves a significant role in causing
multidrug resistance (30). The
expression of P-glycoprotein was also examined, but its expression
did not change, indicating that it did not participate in the
autophagy cell death induced by resveratrol (data not shown).
Therefore, it was hypothesize that ongoing exposure to Res drives
the autophagic K562/ADM cells to a higher degree of autophagy and
activates Bax, a downstream effector of Cath D (17,19,26). The
activation of Bax promotes lysosomal membrane permeabilization with
release of Cath D from lysosomes (17,18). In
turn, Cath D contributes to apoptosis through the clearance of
dying cells, activation of Bax and subsequent amplification of
apoptotic signaling.
In summary, the present study revealed that Res
promoted autophagy in leukemia-drug-resistant K562/ADM cells and
that this enhanced autophagy was involved in the Res-induced
apoptosis of K562/ADM cells, with apoptosis occurring via a series
of events that potentially involves the interaction between Bcl-2
and beclin 1, the activation of Bax and the release of lysosomal
Cath D. Res cytotoxicity was associated with the up-regulation of
Cath D expression, suggesting that Cath D may be the key to
triggering Res-induced cell death (17). Based on the present data, it was
hypothesize that K562/ADM cells increase their autophagic activity
to counteract cytotoxin-induced cellular injury and sustain drug
resistance under physiological conditions. However, with ongoing
exposure to Res or other anticarcinogens, the drug-resistant cells
are forced to maintain high levels of autophagy for a long period
of time, resulting in injury to the cells due to the activation of
a Bcl-2/beclin1-Cath D/Bax loop. This loop may promote a series of
changes: Lysosome permeabilization and the release of Cath D,
Bax-mediated mitochondrial permeabilization with cytochrome
c release, activation of the caspase cascade and apoptosis.
Therefore, manipulating the balance of autophagy and apoptosis via
activation of the Cath D-Bax loop may have great therapeutic
potential for overcoming drug resistance in leukemia.
Acknowledgements
The present study was supported by the Fundamental
Research Funds for the Central Universities (grant no.
lzujbky-2014-143) and National Natural Science Foundation of China
(grant no. 81541025).
References
|
1
|
Eum KH and Lee M: Targeting the autophagy
pathway using ectopic expression of Beclin1 in combination with
rapamycin in drug-resistant v-Ha-ras-transformed NIH 3T3 cells. Mol
Cells. 31:231–238. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Lin CJ, Lee CC, Shih YL, Lin TY, Wang SH,
Lin YF and Shih CM: Resveratrol enhances the therapeutic effect of
temozolomide against malignant glioma in vitro and in vivo by
inhibiting autophagy. Free Radic Biol Med. 52:377–379. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Burada F, Nicoli ER, Ciurea ME, Uscatu DC,
Ioana M and Gheonea DI: Autophagy in colorectal cancer: An
important switch from physiology to pathology. World J Gastrointest
Oncol. 7:271–284. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Chen J, Wei HL, Chen J and Xie B:
Antitumor effect of arsenic trioxide in human K562 and K562 ADM
cells by autophagy. Toxicol Mech Methods. 22:512–519. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Shuhua W, Chenbo S, Yangyang L, Xiangqian
G, Shuang H, Tangyue L and Dong T: Autophagy-related genes raptor,
rictor and beclin 1 expression and relationship with multidrug
resistance in colorectal carcinoma. Hum Pathol. 46:1752–1759. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Panda PK, Mukhopadhyay S, Das DN, Sinha N,
Naik PP and Bhutia SK: Mechanism of autophagic regulation in
carcinogenesis and cancer therapeutics. Semin Cell Dev Biol.
39:43–55. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Galluzzi L, Pietrocola F, Bravo-San Pedro
JM, Amaravadi RK, Baehrecke EH, Cecconi F, Codogno P, Debnath J,
Gewirtz DA, Karantza V, et al: Autophagy in malignant
transformation and cancer progression. EMBO J. 34:856–880. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhao XY, Li GY, Liu Y, Chai LM, Chen JX,
Zhang Y, Du ZM, Lu YJ and Yang BF: Resveratrol protects against
arsenic trioxide-induced cardiotoxicity in vitro and in vivo. Br J
Pharmacol. 154:105–113. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Barjot C, Tournaire M, Castagnino C, Vigor
C, Vercauteren J and Rossi JF: Evaluation of antitumor effects of
two vine stalk oligomers of resveratrol on a panel of lymphoid and
myeloid cell lines: Comparison with resveratrol. Life Sci.
81:1565–1574. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yan HW, Hu WX, Zhang JY, Wang Y, Xia K,
Peng MY and Liu J: Resveratrol induces human K562 cell apoptosis,
erythroid differentiation and autophagy. Tumour Biol. 35:5381–5388.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Naponelli V, Modernelli A, Bettuzzi S and
Rizzi F: Roles of autophagy induced by natural compounds in
prostate cancer. Biomed Res Int. 2015:1218262015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Fang Y, DeMarco VG and Nicholl MB:
Resveratrol enhances radiation sensitivity in prostate cancer by
inhibiting cell proliferation and promoting cell senescence and
apoptosis. Cancer Sci. 103:1090–1098. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Fang Y, Bradley MJ, Cook KM, Herrick EJ
and Nicholl MB: A potential role for resveratrol as a radiation
sensitizer for melanoma treatment. J Surg Res. 183:645–653. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Pattingre S, Tassa A, Qu X, Garuti R,
Liang XH, Mizushima N, Packer M, Schneider MD and Levine B: Bcl-2
antiapoptotic proteins inhibits Beclin1-dependent autophagy. Cell.
122:927–939. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chen J, Chen J, Xie B and Wei HL: Acquired
multidrug resistance in human K562/ADM cells is associated with
enhanced autophagy. Toxicol Mech Methods. 23:678–683. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yang J and Yao S: JNK-Bcl-xL-Bax/Bak
pathway mediates the crosstalk between matrine-induced autophagy
and apoptosis via interplay with Beclin1. Int J Mol Sci.
16:25744–25758. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Trincheri NF, Nicotra G, Follo C, Castino
R and Isidoro C: Resveratrol induces cell in colorectal cancer
cells by a novel pathway involving lysosomal cathepsin D.
Carcinogenesis. 28:922–931. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hsu KF, Wu CL, Huang SC, Wu CM, Hsiao JR,
Yo YT, Chen YH, Shiau AL and Chou CY: Cathepsin L mediates
resveratrol-induced autophagy and apoptotic cell death in cervical
cancer cells. Autophagy. 5:451–460. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Jang BG, Choi BY, Kim JK, Kim MJ, Sohn M
and Suh SW: Impairment of autophagic flux promotes glucose
reperfusion-induced neuro2A cell death after glucose deprivation.
PloS One. 8:e764662013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhang J, Ma K, Qi T, Zhang Q, Li G and
Chiu JF: P62 regulates resveratrol-mediated Fas/Cav-1 complex
formation and transition from autophagy to apoptosis. Oncotarget.
6:789–801. 2015.PubMed/NCBI
|
|
21
|
Zhang X, Xu W, Su J, Chu M, Jin H, Li G,
Tan C, Wang X and Wang C: The prosurvival role of autophagy in
resveratrol-induced cytotoxicity in GH3 cells. Int J Mol Med.
33:987–993. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Bhardwaj A, Sehthi G, Vadhan-Raj S,
Bueso-Ramos C, Takada Y, Gaur U, Nair AS, Shishodia S and Aggarwal
BB: Resveratrol inhibits proliferation, induces apoptosis and
overcomes chemoresistance through down-regulation of STAT3 and
nuclear factor-kappaB-regulated antiapoptotic and cell survival
gene products in human multiple myeloma cell. Blood. 109:2293–2302.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Liu YN, Wang YX, Liu XF, Jiang LP, Yang G,
Sun XC, Geng CY, Li QJ, Chen M and Yao XF: Citreoviridin induces
ROS-dependent autophagic cell death in human liver HepG2 cells.
Toxicon. 95:30–37. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Chen QY, Shi JG, Yao QH, Jiao DM, Wang YY,
Hu HZ, Wu YQ, Song J, Yan J and Wu LJ: Lysosomal member
permeabilization is involved in curcumin-induced apoptosis of A549
lung carcinoma cells. Mol Cell Biochem. 359:389–398. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Prabhu V, Srivastava P, Yadav N, Amadori
M, Schneider A, Seshadri A, Pitarresi J, Scott R, Zhang H,
Koochekpour S, et al: Resveratrol depletes mitochondrial DNA and
inhibition of autophagy enhances resveratrol-induced caspase
activation. Mitochondrion. 13:493–499. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yelamanchili SV, Chaudhuri AD, Flynn CT
and Fox HS: Upregulation of cathepsin D in the caudate nucleus of
primates with experimental parkinsonism. Mol Neurodegener.
6:522011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wei MF, Chen MW, Chen KC, Lou PJ, Lin SY,
Hung SC, Hsiao M, Yao CJ and Shieh MJ: Autophagy promotes
resistance to photodynamic therapy-induced apoptosis selectively in
colorectal cancer stem-like cells. Autophagy. 10:1179–1192. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Kim DH, Lee NY, Sung WJ, Baek JH, Kim JG,
Sohn SK, Suh JS, Lee KS and Lee KB: Multidrug resistance as a
potential prognostic indicator in acute myeloid leukemia with
normal karyotype. Acta Haematol. 114:78–83. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Xu N, Zhang J, Shen C, Xia L, Xue F and
Xia Q: Cisplatin-induced down regulation of mir-199a-5p increases
drug resistance by activating autophagy in HCC cell. Biochem
Biophys Res Cimmun. 423:826–831. 2012. View Article : Google Scholar
|
|
30
|
Xue C, Wang C, Liu Q, Meng Q, Sun H, Huo
X, Ma X, Liu Z, Ma X, Peng J and Liu K: Targeting P-glycoprotein
expression and cancer cell energy metabolism: Combination of
metformin and 2-deoxyglucose reverses the multidrug resistance of
K562/Dox cells to doxorubicin. Tumor Biol. 37:8587–8597. 2016.
View Article : Google Scholar
|