Introduction
Transmembrane protein 174 (TMEM174) is a type III
transmembrane protein that lacks a clear signal peptide. The N and
C terminals are located inside of the cell. TMEM174 was originally
identified from a large gene pool by high-throughput cell screening
technology. This technique is used for the isolation of functional
genes and for the analysis into the mechanisms of gene function
(1). Dependent on its transmembrane
helices, TMME174 overexpression is able to promote the
transcriptional activity of activator protein 1 (AP-1), which is
partly mediated by the ERK pathway (2). Our previous study demonstrated high
TMEM174 expression in the kidney and also revealed its potential
involvement with renal cancer based on its capacity to stimulate
cell proliferation (2). The kidney
is vital for the maintenance of the salt and water balance within
the body, in order to keep a stable internal environment, i.e., for
homeostasis. However, the expression of TMEM174 in renal cancer and
normal renal tissues remains to be elucidated. In the present
study, RNA in situ hybridization was used to detect the
TMEM174 gene expression in various malignant renal cancer and
normal renal tissues. The aim of this study was to provide a
theoretical basis for the molecular mechanisms of the development
of kidney cancer.
Materials and methods
Tissue microarray
Tissue microarrays were purchased from Shaanxi Chao
Ying Biotechnology, Ltd., Co. (Xi’an, China). Information on each
specimen, consisting of patient age, gender, organization,
pathological diagnosis, clinical grade, tumor-node-metastasis
classification, clinical stage, specimen type and result
information, was available. TNM staging, clinical staging and
pathological grading were determined based on the American Joint
Committee on Cancer manual (3).
Specimens were used from a total of 208 cases, including 178 cases
of renal cancer and nephritis, 20 adjacent tissues and 10 normal
tissues.
Preparation of digoxigenin-labeled probes
for RNA in situ hybridization
The sense and anti-sense probes that matched the
TMEM174 core responding sequence were as follows: Anti-sense,
5′-GAGCATTGTGTTATTATATCAG*AATA GCCTCTAGCGAGGGAGAGAGTATATT-3′DIG and
sense, 5′-ATATACTCTCTCCCTCGCTAGAGGC*TATTCTGATA
TAATAACACAATGCTCA-3′DIG. The asterisks indicate that the 3′
terminals were labeled with digoxigenin. All probes were
synthesized by Shanghai Sangon Biological Engineering Technology
and Services Co., Ltd. (Shanghai, China).
RNA in situ hybridization
Hybridization procedures were performed in this
study with the RNA&ISH kit (KD2084, Roche Diagnostics,
Indianapolis, IN, USA), based on the manufacturer’s instructions,
with certain modifications. The glassware was washed, rinsed in
distilled deionized water and autoclaved prior to use. Gloves were
worn when the glassware and slides were handled to prevent RNase
contamination of the tissue. The hybridization conditions were as
follows: Probe concentration, 10 ng/μl; antibody titer, 1:400;
washing temperature, room temperarture; dyeing temperature, 37°C;
and dyeing time, 2 h. Deparaffinized sections were mounted on
Denhardt-coated glass slides (D2532; Sigma Aldrich, St. Louis, MO,
USA) and treated with pepsin (0.25 mg/ml in DEPC
H2O-HCl; Sigma Aldrich) for 30 min in a 37°C water bath.
The treated sections were then processed for in situ
hybridization at 42–47°C for 24 h. The hybridization mixture
contained the labeled oligonucleotide probe, 50% formamide, 10
mmol/l Tris-HCl, 1 mmol/l vanadyl-ribonucleoside complex (94740;
Sigma Aldrich), 1 mmol/l CTAB (855820; pH 7.0; Sigma Aldrich), 0.15
mol/l Nacl, 1 mmol/l EDTA (pH 7.0), 1X Denhardt’s mixture and 10%
dextran sulfate. Subsequent to hybridization, the slides were
washed three times, for 30 min each time, in 0.1 mol/l
Tris-buffered saline (TBS) at room temperature. The slides were
then treated with TBS [100 mmol/l Tris (pH 7.5) and 150 mmol/l
NaCl] containing a 1% blocking reagent (Roche Diagnostics,
Shanghai, China) and 0.03% Triton X-100 for 30 min at room
temperature, and incubated for 30 min with antidioxigenin alkaline
phosphatase-conjugated antibodies (Roche Diagnostics) diluted at
1:4,000 in TBS containing 0.03% Triton X-100 and a 1% blocking
reagent. Subsequent to being washed three times, for 15 min each
time in TBS and 0.05% Tween 20, the slides were rinsed in a
diammonimum phosphate (DAP)-buffer [100 mmol/l Tris (pH 9.5) 100
mmol/l NaCl, 50 mmol/l MgCl2] and hybridization signals
were subsequently visualized using nitroblue tetrazolium and
5-bromo-4-chloro-3-indolyl phosphate as substrates [DAP-buffer (100
mmol/l Tris, pH 9.5, 100 mmol/l NaCl and 50 mmol/l
MgCl2) in 10% PVA (341584; Sigma Aldrich)].
Results
Association between TMEM174 expression
and renal pathological cell types
TMEM174 gene expression in the various malignant
renal cancer and normal renal tissues was detected by RNA in
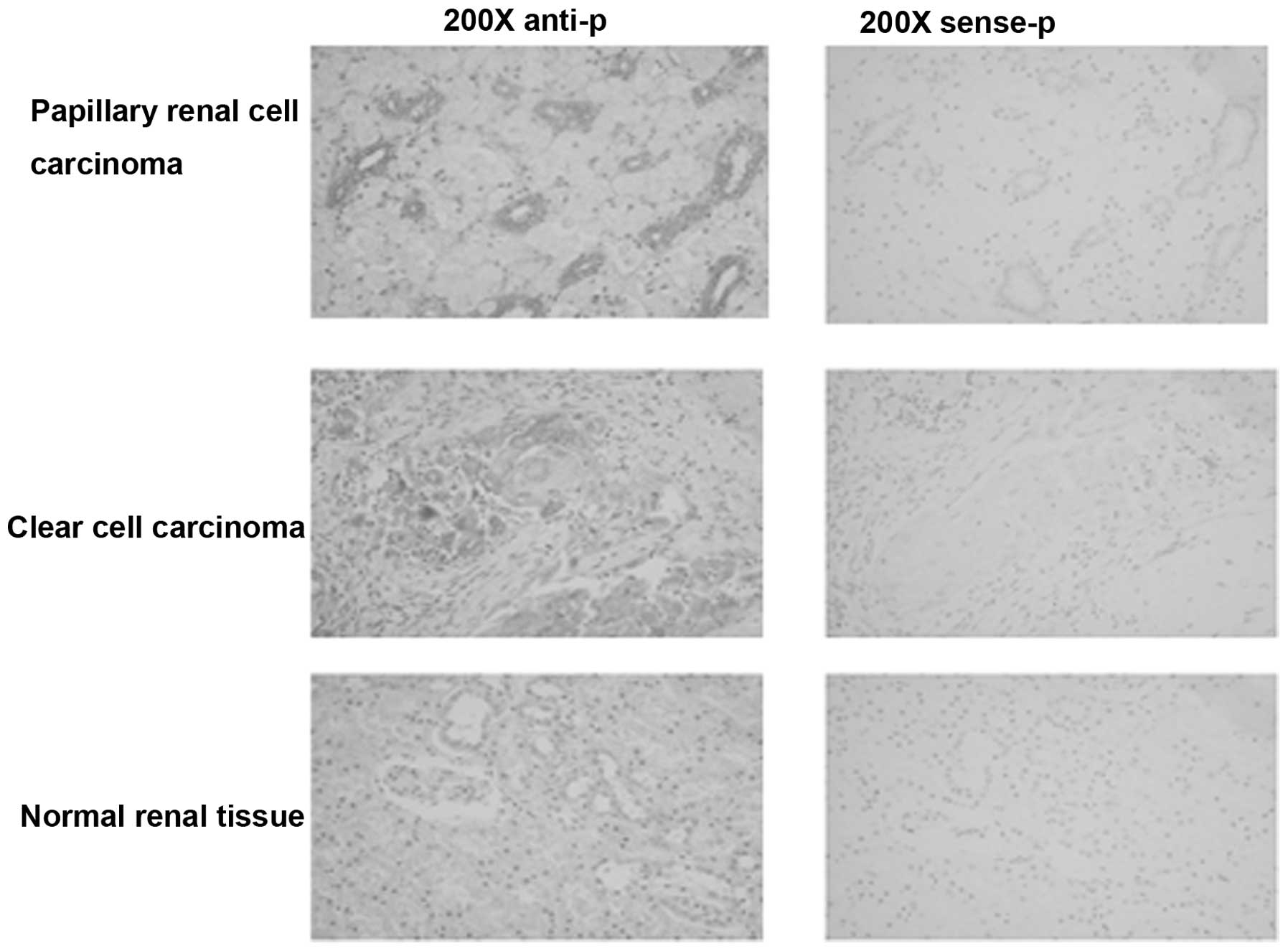
situ hybridization. As shown in Table I and Fig. 1, TMEM174 exhibits differential
expression in renal tissues. TMEM174 had a high positive rate of
expression in squamous cell carcinoma with necrosis, papillary
renal cell carcinoma and transitional cell carcinoma, and a low
positive rate of expression in clear cell carcinoma, interstitial
nephritis, undifferentiated carcinoma, retroperitoneal metastatic
clear cell carcinoma, adrenal gland metastatic clear cell
carcinoma, pelvic cavity metastatic chromophobe carcinoma, severe
atypical hyperplasia of transitional epithelium and hyperplasia.
Extremely weak expression was observe in collecting duct carcinoma,
Wilms’ tumor, chronic pyelonephritis, acute pyelonephritis, cancer
adjacent normal renal tissue and normal renal tissue.
 | Table IAssociation between TMEM174 expression
and renal pathological cell type. |
Table I
Association between TMEM174 expression
and renal pathological cell type.
| Renal tumors | Number of tumors | TMEM174 mRNA-positive
rumors (0–1+/1+/2+/3+), n | TMEM174 mRNA-negative
tumors, n |
|---|
| Clear cell
carcinoma | 59 | 24 (9/12/3/0) | 35 |
| Collecting duct
carcinoma | 4 | 0 (0/0/0/0) | 4 |
| Squamous cell
carcinoma with necrosis | 10 | 9 (0/5/3/1) | 1 |
| Wilms’ tumor | 20 | 4 (4/0/0/0) | 16 |
| Papillary renal cell
carcinoma | 20 | 20 (3/8/9/0) | 0 |
| Transitional cell
carcinoma | 35 | 32 (5/16/11/0) | 3 |
| Undifferentiated
carcinoma | 4 | 4 (2/2/0/0) | 0 |
| Retroperitoneal
metastatic clear cell carcinoma | 4 | 4 (0/4/0/0) | 0 |
| Adrenal gland
metastatic clear cell carcinoma | 4 | 4 (1/3/0/0) | 0 |
| Pelvic cavity
metastatic chromophobe carcinoma | 2 | 2 (0/2/0/0) | 0 |
| Hyperplasia (sparse
renal tubule tissue) | 2 | 2 (0/2/0/0) | 0 |
| Severe atypical
hyperplasia of transitional epithelium | 2 | 2 (0/2/0/0) | 0 |
| Chronic
pyelonephritis | 2 | 2 (2/0/0/0) | 0 |
| Acute
pyelonephritis | 2 | 1 (1/0/0/0) | 1 |
| Interstitial
nephritis | 6 | 4 (4/0/0/0) | 2 |
| Cancer adjacent
normal renal tissue | 20 | 10 (10/0/0/0) | 10 |
| Normal renal
tissue | 10 | 4 (4/0/0/0) | 6 |
Association between TMEM174 expression
and renal pathological clinical stage
The association between TMEM174 expression and renal
pathological clinical stage was analyzed. As shown in Table II, the THEM174 expression rate was
58% in phase I tissue samples, 50% in phase II tissue samples, 71%
in phase III tissue samples and 50% in phase IV tissue samples.
 | Table IIAssociation between TMEM174 expression
and renal pathological clinical stage. |
Table II
Association between TMEM174 expression
and renal pathological clinical stage.
| Clinical stage | n | TMEM174 positive
rate, % |
|---|
| I | 108 | 58 |
| II | 26 | 50 |
| III | 14 | 71 |
| IV | 4 | 50 |
Discussion
Each year in the United States, nearly 55,000
individuals are diagnosed with kidney cancer. In order to be
effectively treated, an early diagnosis and effective surgical
therapy are required. In more advanced-stage cases with involvement
of the renal vein and lymph nodes or invasion through the renal
cortex, surgical therapy frequently fails (4–8).
Kidney cancer cell progression is a coordinated process that
comprises cell cycle dysregulation and a specific gene expression
program to determine tissue identity (9). The TMEM174 gene is highly expressed in
the kidney tissues.
The AP-1 transcription factor is a heterodimeric
protein formed from proteins of the c-Fos, ATF, c-Jun and JDP
families. In response to a range of stimuli, including cytokines,
stress, growth factors and viral and bacterial infections, AP-1
regulates gene expression (10).
The cellular transcription factor cAMP-response element binding
protein (CREB) binds to specific DNA sequences referred to as cAMP
response elements (CREs), thereby regulating downstream gene
transcription (11). TMEM174
overexpression has been shown to enhance the transcriptional
activity of AP-1 and promote cell proliferation (2). In addition, our recent studies
demonstrated that the CREB and AP-1 transcription factors are
involved in the transcriptional regulation of the TMME174 gene
(12).
When antibodies are not available, RNA in
situ hybridization is a useful method that allows the
determination of the transcriptional expression pattern of a gene
(13). With this technique, the
expression of multiple RNA species may be assayed using distinct
RNA-labeled probes, or the RNA and protein localization within
larval tissues may be examined. In the present study, RNA in
situ hybridization analysis was used to detect the expression
of TMEM174, and the results showed that its expression is varies
among differing renal tissues. These results indicate that TMEM174
may have a significant role in the development of renal cancer.
Acknowledgements
This study was supported by grants from the National
Natural Science Foundation of China (nos. 81072093, 30671092 and
81302323) and the Natural Science Foundation of Hebei Province
(nos. C2009001260, C2014209140 and C2013209024).
References
|
1
|
Wan D, Gong Y, Qin W, et al: Large-scale
cDNA transfection screening for genes related to cancer development
and progression. Proc Natl Acad Sci USA. 101:15724–15729. 2004.
|
|
2
|
Wang P, Sun B, Hao D, Zhang X, Shi T and
Ma D: Human TMEM174 that is highly expressed in kidney tissue
activates AP-1 and promotes cell proliferation. Biochem Biophys Res
Commun. 394:993–999. 2010.
|
|
3
|
Greene FL, Page DL, Fleming ID, et al:
AJCC Cancer Staging Manual. 6th edition. Springer-Verlag; New York:
2002
|
|
4
|
Eto M and Naito S: Molecular targeting
therapy for renal cell carcinoma. Int J Clin Oncol. 11:209–213.
2006.
|
|
5
|
Costa LJ and Drabkin HA: Renal cell
carcinoma: new developments in molecular biology and potential for
targeted therapies. Oncologist. 12:1404–1415. 2007.
|
|
6
|
Stillebroer AB, Oosterwijk E, Oyen WJ,
Mulders PF and Boerman OC: Radiolabeled antibodies in renal cell
carcinoma. Cancer Imaging. 7:179–188. 2007.
|
|
7
|
Mellado B and Gascón P: Molecular biology
of renal cell carcinoma. Clin Transl Oncol. 8:706–710. 2006.
|
|
8
|
Weight CJ, Kaouk JH, Hegarty NJ, et al:
Correlation of radiographic imaging and histopathology following
cryoablation and radio frequency ablation for renal tumors. J Urol.
179:1277–1283. 2008.
|
|
9
|
Chen X, Ruan A, Wang X, et al: miR-129-3p,
as a diagnostic and prognostic biomarker for renal cell carcinoma,
attenuates cell migration and invasion via downregulating multiple
metastasis-related genes. J Cancer Res Clin Oncol. May 7–2014.(Epub
ahead of print).
|
|
10
|
Hess J, Angel P and Schorpp-Kistner M:
AP-1 subunits: quarrel and harmony among siblings. J Cell Sci.
117:5965–5973. 2004.
|
|
11
|
Bourtchuladze R, Frenguelli B, Blendy J,
et al: Deficient long-term memory in mice with a targeted mutation
of the cAMP-responsive element-binding protein. Cell. 79:59–68.
1994.
|
|
12
|
Hu F, Meng Y, Gou L and Zhang X: Analysis
of promoters and CREB/AP-1 binding sites of the human TMEM174 gene.
Exp Ther Med. 6:1290–1294. 2013.
|
|
13
|
Jensen E: Technical review: In situ
hybridization. Anat Rec (Hoboken). 9–May;2014.(Epub ahead of
print). DOI: 10.1002/ar.22944
|