Introduction
Cutaneous melanoma or malignant melanoma (MM) is a
highly malignant skin tumor that develops from melanocytes
(1–3).
MM was considered uncommon previously, but its incidence has
increased remarkably over the past 30 years (4). Surgery to remove the tumor and
chemotherapy are the routine treatments for early-stage melanoma
(5). However, due to the high degree
of genetic instability and mutations, and less sensitivity to
classical chemotherapeutic treatments, there is high rate of cancer
spreading to other parts of the body in patients with MM (1,4,6). Therefore, the present study explored the
possible mechanism of MM pathogenesis and its signaling
pathways.
Chemokines are cell cytokines induced by
inflammation (7). Chemokines and
their signaling receptors have been shown to regulate cellular
migration, cell invasion, motility, cell growth, cellular
interactions with the extracellular matrix and tumor development
(8,9).
C-X-C chemokine receptor type 7 (CXCR7) is a protein that interacts
with C-X-C motif chemokine ligand 12 (CXCL12), also known as
stromal cell-derived factor 1 (SDF-1), and CXCL11 in signal
transduction (10). CXCR7 expression
is associated with increased pathological inflammation and tumor
development (10). The CXCR7-CXCL12
complex serves an important role in tumor invasion and
organ-specific metastasis (10).
However, the roles of CXCR7 in skin cancer invasion, tumorigenesis
and tumor development are still not well understood.
Squamous cell carcinoma (SCC), basal cell carcinoma
(BCC) and MM are three common skin tumors (3,11,12). The present study explored the effects
of CXCR7 on melanoma progression and development in the tumor
tissue of human patients with MM, cultured cell lines and
transplanted tumor mouse models.
Transwell chamber assays have been used for cell
migration and homing assays (13,14). The
porous polycarbonate membrane at the bottom of the upper chamber
has certain permeability. However, inactivated cells cannot pass
the membrane freely. Activated migrating cells possess the ability
to pass through the polycarbonate membrane by secreting hydrolytic
enzymes to dissolve the porous basement membrane and bind to the
matrigel membrane in the bottom chamber (15). The present study used this Transwell
system to test the CXCR7 effects on melanoma cell migration and
invasion.
Materials and methods
Patients and specimen collection
Human skin tumors from all selected patients were
collected from January 2008 to December 2014 at the Institute of
Dermatology, Chinese Academy of Medical Sciences (Nanjing, China)
and the Affiliated Hospital of Hebei University (Hebei, China). The
Chinese Academy of Medical Sciences (Nanjing, China) and the
Affiliated Hospital of Hebei University (Hebei, China)
Institutional Review Boards approved the present study and the
protocol. Consent for surgery was obtained from all patients upon
informing them of the nature of the disease, its treatment options
and its possible outcomes. Classification into squamous SCC, BCC or
MM was determined by histopathology through immunohistochemistry
staining. Tumor tissues from 30 cases of skin SCC were harvested.
The clinical profiles of patients with SCC included 16 males and 14
females, aged 34–82 years old (median age, 61.20 years). The tumor
progression time ranged from 1 month to 25 years (median, 2.78
years). Among these SCC tumor cases, 12 were located on the head
and face, 1 on the back, 6 on the hands, 9 on the limbs and 2 on
the genital area. Among these SCC cases, 18 (60%) were located on
exposed parts of the body (i.e., head, face and hands) and 12 (40%)
on non-exposed parts. These patients included 12 cases of
high-differentiated SCC (SCC-H) and 18 cases of low-differentiated
SCC (SCC-L). In addition, 25 cases of skin BCC were collected,
comprising 10 males and 15 females, aged 33–80 years old (mean age,
62.12 years). The tumor progression time for BCC ranged from 2
months to 11 years (median, 32.78 months). There were 17 (68%)
cases on exposed parts of the body and 8 (32%) on non-exposed
parts. A total of 30 cases of invasive skin MM samples were
collected. Among these, 18 were males and 12 females, aged 35–75
years old (median age, 61.26 years), and the tumor progression time
ranged from 3 to 19 months (median, 12.10 months).
Immunohistochemistry staining
CXCR7 protein was detected by immunohistochemistry.
The tumor tissues were paraffin embedded and sliced into ~4-µm
thick sections. Sections were deparaffinized with xylene and
rehydrated in decreasing concentrations of ethanol. Sections were
then heated in a microwave for 20 min in 10 mM sodium citrate
buffer for antigen retrieval. Sections were next incubated with a
CXCR7 antibody (cat no., PA3-069, Thermo Fisher Scientific, Inc.,
Waltham, MA, USA; dilution, 1:2,000) at 4°C overnight. Following
three washes in PBS, a peroxidase-conjugated secondary antibody
(cat. no., P447, Dako; Agilent Technologies, Inc., Santa Clara, CA,
USA; dilution, 1:5,000) followed by Pierce DAB Substrate kit (cat.
no., 34002, Invitrogen; Thermo Fisher Scientific, Inc.) were
applied for 30 and 3 min, respectively, at room temperature. Next,
sections were dehydrated in decreasing concentrations of ethanol
and cover slipped. Negative control sections were incubated with an
anti-mouse immunoglobulin G (IgG) antibody (cat. no., sc-2025;
Santa Cruz Biotechnology, Inc., Dallas, TX, USA; dilution,
1:5,000). The stained sections were examined with a light
microscope (IX51; Olympus Corporation, Tokyo, Japan) and
photographed. MaxVision software (version 3.0; MaxPac 8200/8000 ML,
Max Vision LLC, Madison, AL, USA) was used to measure CXCR7
staining intensity. The staining intensity criteria of positive
cells was as follows: 0, no staining; 1, light yellow staining; 2,
brown staining; and 3, dark brown staining. The scoring was
performed by two independent pathologists based on the
aforementioned color category. The percentages of positively
stained cells were also considered in the evaluations. Thus, a
score of 0 was assigned to samples with no positive cells; samples
with <10% of positive cells received a score of 1; those with
10–50% of positive cells received a score of 2; those with 51–80%
of positive cells received a score of 3; and those with >80% of
positive cells received a score of 4. Finally, these two values
(staining intensity and percentage of positive cells) were
multiplied to classify the specimens into CXCR7 low (final score,
0–6) or high (final score, 7–12) expression. Hematoxylin and eosin
(H&E) staining was performed to monitor tissue morphology.
Cell culture and transfection
The melanoma cell lines A375 and M14 were purchased
from the American Type Culture Collection (Manassas, VA, USA)
(16,17). The cells were grown in Dulbecco's
modified Eagle's medium (DMEM) containing 10% fetal bovine serum
(FBS) (both from Gibco; Thermo Fisher Scientific, Inc., Waltham,
MA, USA) in an incubator at 37°C and 5% CO2.
For transfection, cells were seeded in 24-well
plates. To assess the effects of CXCR7 on the biological function
of M14 cells (M14 was selected for its higher level of CXCR7
compared with A375 cell), CXCR7 small hairpin RNA (shRNA) or empty
vectors (Genscript Biotech Corporation, Nanjing, China) were
transfected into M14 cells using FuGENE HD Transfection Reagent
(Promega Corporation, Madison, WI, USA). M14 cells were selected
for their higher level of CXCR7 expression compared with A375
cells. The antibiotic G418 (1,000 µg/ml) (Invitrogen; Thermo Fisher
Scientific, Inc.) was used for 2 weeks for cell selection upon
shRNA transfection. CXCR7 shRNA- or vector shRNA-transfected M14
cells were subcloned and selected under a light microscope (IX51;
Olympus Corporation, Tokyo, Japan). The selected CXCR7 shRNA- and
vector shRNA-transfected M14 cell clones were cultured in 500 µg/ml
G418 for further experiments.
CXCR7 shRNA design and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
CXCR7 short interfering (si)RNA sequence was
obtained from Thermo Fisher Scientific, Inc. (Silencer®
Select siRNAs). CXCR7 shRNA was constructed based on the siRNA
sequence by Genscript Biotech Corporation (Nanjing, China). Three
pairs of shRNA were designed, and the pair with the best shRNA
efficiency was selected. Western blotting was performed to confirm
CXCR7 protein knocked down. Total RNA from cells was isolated using
TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.). Total
RNA quality and concentration were measured by ultraviolet
spectrophotometry. The absorbance (A) 260/A280 range was set at
1.9–2.1 to assure good RNA quality. RNA (1 µg/sample) was used for
RT to synthesize complementary DNA using random primers provided in
the High-Capacity cDNA Reverse Transcription kit (Applied
Biosystems; Thermo Fisher Scientific, Inc.). qPCR was performed to
determine CXCR7 and β-actin messenger RNA (mRNA) expression. CXCR7
primers [CXCR-forward (F), 5′-CCCTGCATCCATTCTCTCTT-3′ and
CXCR7-reverse (R), 5′-CCTGTTGCAAAACTGTCAGC-3′] and β-actin primers
(β-actin-F, 5′-GAAGGATTCCTATGTGGGCGAC-3′ and β-actin-R,
5′-AGCCTGGATAGCAACGTACATGG-3′) were used for qPCR using a PCR kit
(GoTaq® Real-Time qPCR and RT-qPCR Systems for Dye-Based
Detection, Promega Corporation). The reactions were performed under
the following conditions: 95°C for 3 min, followed by 45 cycles at
94°C for 20 sec, 60°C for 30 sec and 72°C for 20 sec. CXCR7 mRNA
levels obtained from qPCR were normalized to β-actin mRNA levels
yielding arbitrary units (18).
Western blotting
Upon harvesting, M14 or A375 cells were washed with
PBS, and whole cell protein samples were extracted with
radioimmunoprecipitation assay buffer [0.5% Nonidet P-40, 10 mM
Tris (pH 7.4), 150 mM NaCl and 1% SDS] containing a Protease
Inhibitor Cocktail (Sigma-Aldrich; Merck KGaA, Darmstadt, Germany).
Proteins (70 µg/well) were separated by 10% SDS-PAGE and
transferred to polyvinylidene difluoride membranes. The primary
antibodies used for western blotting were CXCR7 antibody () (cat.
no., PA3-069, Thermo Fisher Scientific, Inc., dilution, 1:2,000)
and mouse anti β-actin antibody (cat. no., sc-8432; Santa Cruz,
Biotechnology, Inc., Dallas, TX, USA; dilution, 1:5,000). Each
membrane was incubated with one primary antibody (in 5% BSA) at 4°C
in a cold room overnight. The secondary antibody ECL horseradish
peroxidase-conjugated sheep anti-mouse IgG (cat. no., NA931;
dilution, 1:5,000), or ECL HRP-conjugated donkey anti-rabbit IgG
(cat. no., NA934V; dilution, 1:5,000) (GE Healthcare Life Sciences,
Chalfont, UK) was incubated with the membrane at room temperature
(25°C) for 1 h. Amersham ECL Prime Western Blotting Detection
Reagent (GE Healthcare Life Sciences) was used to detect the
chemiluminescent signals in an X-ray film.
Transwell chamber experimental
settings
The Transwell chamber was purchased from Corning
Incorporated (Corning, NY, USA), and included a polycarbonate
membrane coated with matrigel that acted as an artificial basement
membrane as previous described (15).
The 24-well Transwell chamber was first incubated with 500 µl DMEM
without FBS for 2 h. The medium was then replaced with 500 µl cells
in 10% FBS-DMEM (0.6×106 cells/ml). These cells included
non-transfected, empty vector shRNA-transfected and CXCR7
shRNA-transfected M14 cells (M14 cell was selected for its higher
CXCR7 level compared to A375 cells) (300,000 cells/well), which
were seeded in the upper chamber. Recombinant Human CXCR12 (cat.
no., 350-NS/CF, R&D Systems Inc., Minneapolis, MN, USA) (10
ng/ml) was added to all wells. The cells were incubated in the
Transwell chamber for 24 h. The bottom chamber of the Transwell
membrane was then stained with Cell Counting kit-8 (Dojindo
Molecular Technologies, Inc., Kumamoto, Japan) for 20 min. The
cells were next washed with PBS to remove the excess dye and air
dried for 1 h. The number of cells that passed through the membrane
was then counted under a light microscope (IX51; Olympus
Corporation, Tokyo, Japan). All experiments were repeated three
times.
Animal studies
All animal experiments were approved by the Chinese
Academy of Medical Sciences and Affiliated Hospital of Hebei
University Animal Policy and Welfare Committee, and complied with
the National Institutes of Health guidelines (Guide for the Care
and Use of Laboratory Animals) (https://www.ncbi.nlm.nih.gov/books/NBK54050/). Wild
type BALB/c nude mice were obtained from Shanghai Laboratory Animal
Center (Shanghai, China). There were 18 mice (5 weeks old; 9
female, 9 male; average weight 18–19 g) used in the experiment. The
mice were housed at temperatures ~23°C with 40–60% humidity and 12
h light/12 h dark cycle. Food and water were accessible at
libitum. The effects of CXCR7 on the tumor progression of skin
melanoma were tested in transplanted mice. Untreated, empty vector
shRNA-transfected and CXCR7 shRNA-transfected M14 cells were
harvested. Each cell suspension (1×106 cells/100 µl) was
subcutaneously inoculated into the mice right armpit. Tumor size
was measured from day 11 following tumor transplantation. Tumor
volume (TV) was calculated by the following formula:
TV=1/2xaxb2, where a and b are the tumor length and
width, respectively. Mice were sacrificed by carbon dioxide at day
21 post-tumor transplantation. The 20 l plastid chamber with a lid
was used for mice euthanasia. Compressed CO2 gas in
cylinders was used for source of CO2 with gas inflow to
the chamber regulated at a speed ~30–50% of the chamber or cage
volume/min to provide a final concentration of 50% for ~1 min.
After the gas was stopped, the animal was then observed until all
muscle activity and signs of life have been absent for ≥30 sec. The
tumors were harvested, weighted and recorded. The maximum weight
was ~1.70 g.
Statistical analysis
SPSS statistical software (version 13.0; SPSS, Inc.,
Chicago, IL, USA) was used for data analysis. Data are expressed as
the mean ± standard deviation of ≥3 sample replicates, unless
stated otherwise. *P<0.05, **P<0.01, or ***P<0.001 was
considered to indicate a statistically significant difference.
Results
CXCR7 expression in three types of
skin cancer tissue
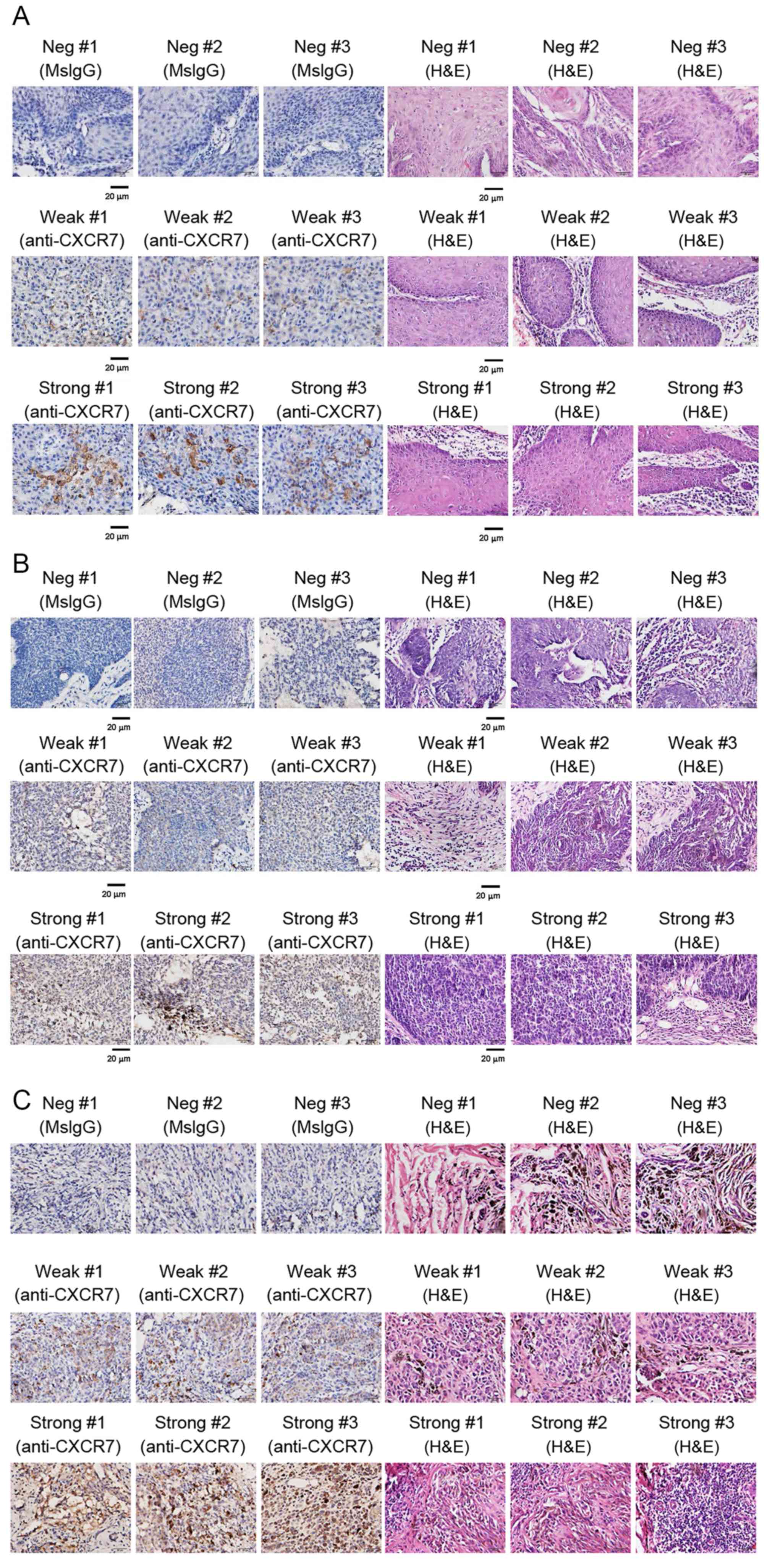
Immunohistochemistry staining was performed to
assess the expression of CXCR7 in skin cancer tissues. H&E
staining was also performed to monitor cancer tissue morphology
(Fig. 1A-C). CXCR7 protein-positive
samples were stained in brown color. CXCR7 protein was located in
the cytoplasm and cell membrane (Fig.
1A-C). CXCR7 protein expression levels were compared between
SCC, BCC and MM, i.e., the three types of skin cancer tissues
evaluated in the present study (Table
I). It was observed that MM had the highest (80%, 24 out of 30
cases) percentage of CXCR7 protein expression, followed by SCC-L
with 38.9%. SCC-H and BCC had 8.3 and 8.0% of high CXCR7 protein
expression levels, respectively (Tables
I and II). These findings
indicated that CXCR7 is involved in tumor progression.
 | Table I.Patient's CXCR7 protein levels in SCC,
MCC and MM cases, according to their pathology details and
anti-CXCR7 staining score. |
Table I.
Patient's CXCR7 protein levels in SCC,
MCC and MM cases, according to their pathology details and
anti-CXCR7 staining score.
| Pathology number | Anti-CXCR7 score |
|---|
| SCC |
|
|
14792 | 1 |
|
14615 | 1 |
|
14552 | 2 |
|
14502 | 8 |
|
14365 | 2 |
|
14325 | 3 |
|
14298 | 3 |
|
14147 | 8 |
|
14143 | 6 |
|
14005 | 4 |
|
13945 | 4 |
|
13911 | 0 |
|
13905 | 0 |
|
13874 | 4 |
|
13866 | 3 |
|
13759 | 6 |
|
12114 | 9 |
|
12203 | 0 |
|
12102 | 8 |
|
12009 | 8 |
|
11658 | 4 |
|
11411 | 4 |
|
11402 | 8 |
|
11252 | 0 |
|
10892 | 2 |
|
10598 | 3 |
|
10454 | 4 |
|
10336 | 8 |
|
10215 | 3 |
|
10112 | 8 |
| BCC |
|
|
14955 | 3 |
|
14912 | 0 |
|
14905 | 0 |
|
14478 | 4 |
|
14117 | 8 |
|
13819 | 6 |
|
13856 | 0 |
|
13848 | 1 |
|
13722 | 1 |
|
13692 | 2 |
|
13189 | 2 |
|
12751 | 4 |
|
12502 | 3 |
|
12452 | 0 |
|
12441 | 3 |
|
12438 | 2 |
|
12098 | 2 |
|
11791 | 3 |
|
11302 | 4 |
|
11217 | 4 |
|
11009 | 4 |
|
10953 | 8 |
|
10255 | 0 |
|
10233 | 1 |
|
10115 | 3 |
| MM |
|
|
14576 | 8 |
|
14291 | 8 |
|
14002 | 8 |
|
13879 | 3 |
|
13762 | 8 |
|
13412 | 9 |
|
13255 | 9 |
|
12908 | 9 |
|
12745 | 9 |
|
12702 | 8 |
|
12537 | 6 |
|
12524 | 8 |
|
12515 | 8 |
|
12355 | 8 |
|
12118 | 8 |
|
12035 | 8 |
|
11387 | 8 |
|
10652 | 9 |
|
10557 | 4 |
|
10102 | 9 |
|
09711 | 9 |
|
09658 | 9 |
|
09356 | 2 |
|
09110 | 8 |
|
09005 | 9 |
|
08562 | 4 |
|
08312 | 4 |
|
08254 | 8 |
 | Table II.Summary of CXCR7 protein levels in
SCC, MCC and MM patients. |
Table II.
Summary of CXCR7 protein levels in
SCC, MCC and MM patients.
|
|
| CXCR7 protein
expression levelsa |
|---|
| Group | Cases, n | Low, n | High, n | High, % |
|---|
| SCC-T | 30 | 22 | 8 | 26.7 |
| SCC-H | 12 | 11 | 1 |
8.3 |
| SCC-L | 18 | 11 | 7 | 38.9 |
| BCC | 25 | 23 | 2 |
8.0 |
| MM | 30 | 6 | 24 | 80.0 |
CXCR7 expression in melanoma cell
lines M14 and A375
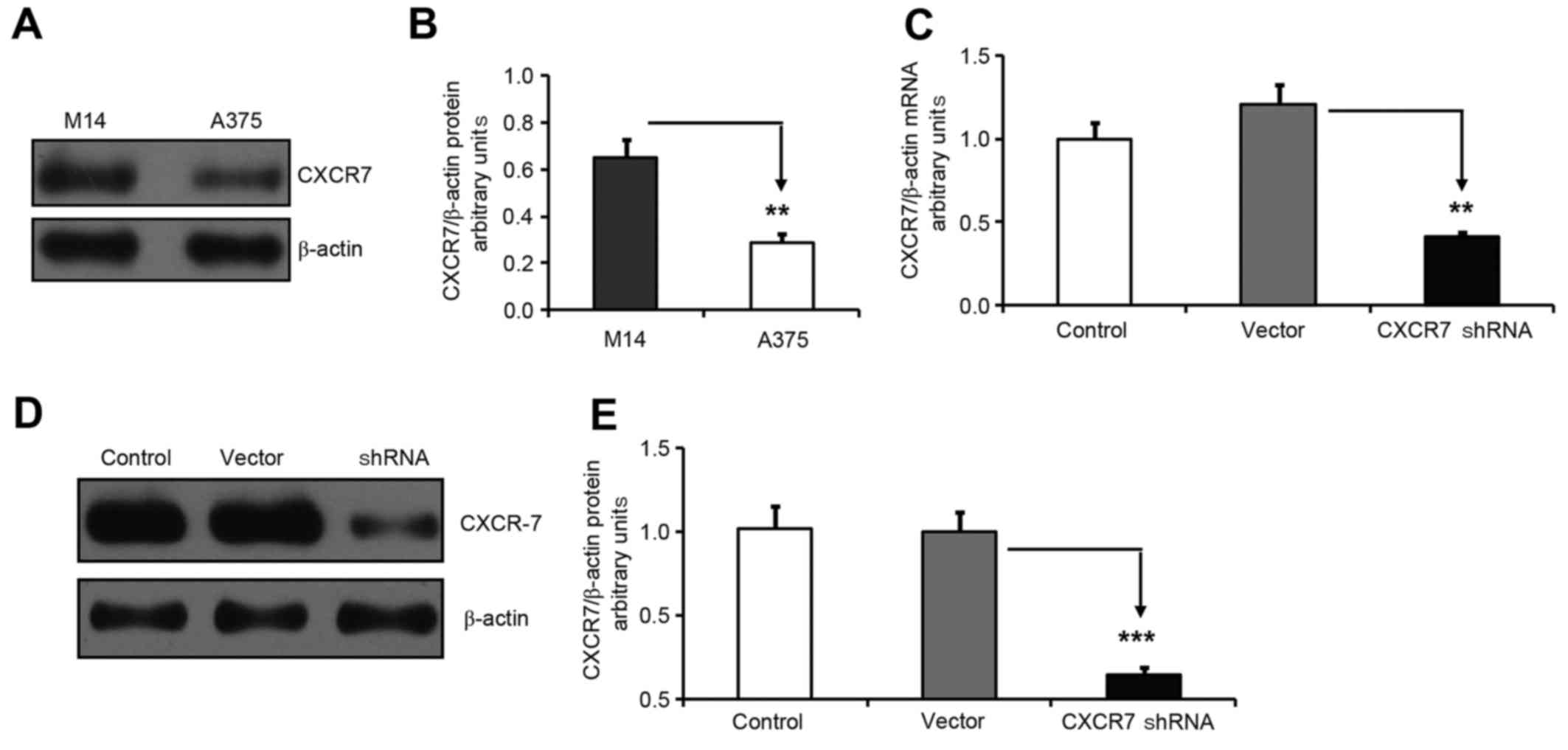
Western blotting was performed to determine the
CXCR7 protein expression levels in melanoma M14 and A375 cells.
β-actin protein expression was used as a control. The CXCR7/β-actin
protein ratio indicated that M14 cells had much higher CXCR7
protein levels compared with those of A375 cells (Fig. 2A and B). Therefore, M14 cells were
selected for our experiments.
CXCR7 shRNA significantly reduces
CXCR7 mRNA and protein expression in M14 cells
shRNA was used to knock down CXCR7 expression in M14
cells. G418 was used for clone selection following shRNA
transfection. Green fluorescent protein observation by fluorescence
microscopy indicated that 95% of cells were transfected with the
CXCR7 shRNA or empty vector. qPCR data revealed that CXCR7 shRNA
significantly inhibited CXCR7 mRNA expression in M14 cells
(Fig. 2C). Western blotting for CXCR7
and β-actin further confirmed the knocked down of CXCR7 protein
expression in M14 cells (Fig. 2D and
E).
CXCL12 cytokine induces M14 cell
invasion
Cells that are able to pass through a polycarbonate
membrane in matrigel Transwell chamber assays are used to study
cell migration and invasion (15).
CXCL12 has been demonstrated to stimulate cell biological functions
(10,19,20). In
the present study, CXCL12 effects on M14 cell invasive activity
were tested in a matrigel Transwell chamber assay. The M14 cells
that passed through the Transwell chamber and bound to the matrigel
were counted. It was observed that both 10 and 100 ng/ml CXCL12
significantly increased the number of M14 cells that passed through
the Transwell chamber, thus confirming that CXCL12 induced M14 cell
invasive ability (Fig. 3A and B).
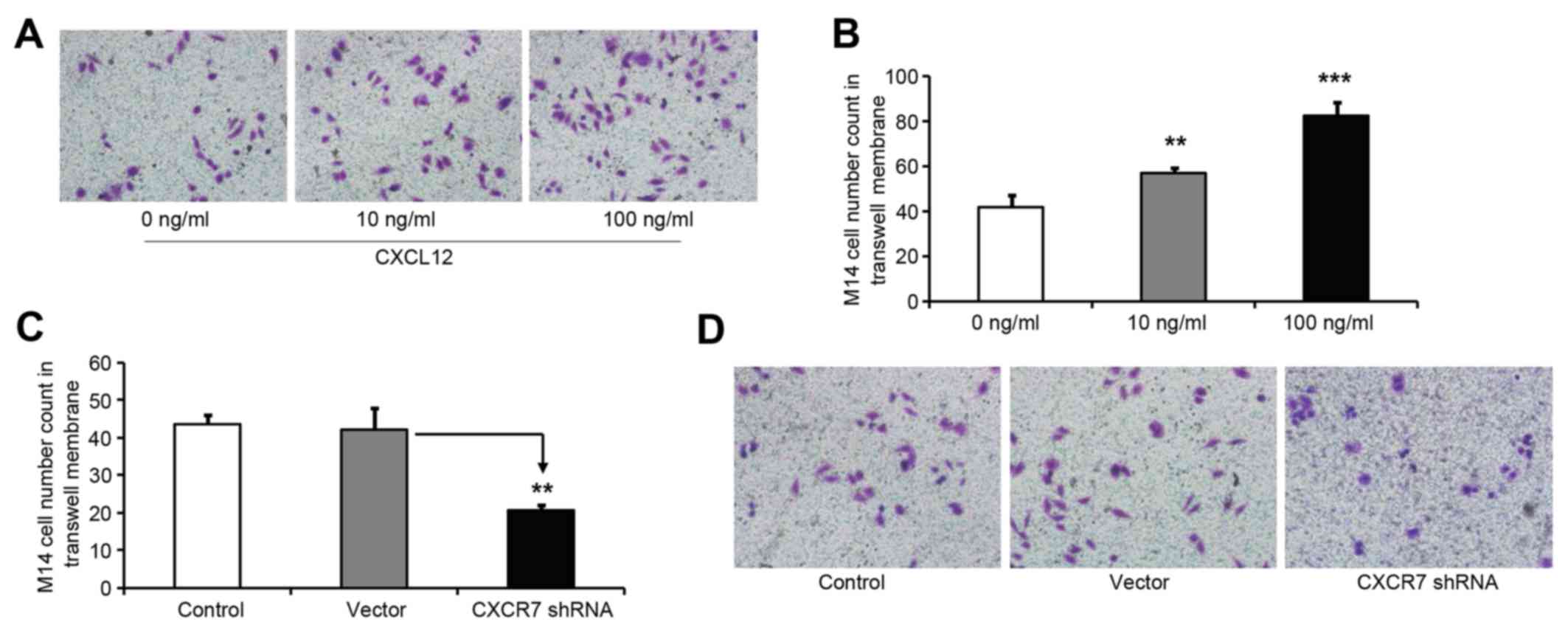 | Figure 3.CXCL12 cytokine induces M14 cell
invasive activity, and CXCR7 knocked down inhibits CXCR12-induced
M14 cell invasive activity. M14 cells (10,000 cells/2 ml) were
seeded in each Transwell chamber with 0, 10 or 100 ng/ml CXCL12 for
24 h. The cells that passed through the Transwell chamber and bound
to the matrigel were recorded and counted. (A) Images of M14 cells
bound to the matrigel membrane (magnification, ×400 (B) CXCL12
increased M14 cell invasive activity. The M14 cell count in the
Transwell matrigel membrane was 41.67±5.51 (0 ng/ml CXCL12),
56.67±2.52 (10 ng/ml CXCL12) and 82.23±6.11 (100 ng/ml CXCL12).
P-values were obtained for the comparisons between the indicated
treatments and 0 ng/ml CXCR12 (**P<0.01; ***P<0.001). (C)
CXCR7 shRNA significantly reduced M14 cell counts in in the
matrigel membrane. The M14 cell count in the Transwell matrigel
membrane was 43.63±2.15 (control), 42.00±6.02 (vector) and
20.62±1.50 (CXCR7 shRNA). **P<0.01 was obtained for the
comparison between vector and CXCR7 shRNA. There were not
significantly different cell counts between the control and vector
groups. These results indicated that CXCR7 downregulation inhibited
M14 cell migration and invasion. (D) Images of M14 cells in the
matrigel membrane (magnification, ×400). Untreated M14 cells
(control), empty vector shRNA-transfected M14 cells (vector) and
CXCR7 shRNA-transfected M14 cells (10,000 cells/2 ml/well) were
plated in Transwell chamber wells with 10 ng/ml CXCR12 for 24 h.
The cells that passed through the membrane and bound to the
matrigel were recorded and counted. A lower cell number was
observed in CXCR7 shRNA-transfected M14 cells compared with that in
control and vector-transfected M14 cells. CXCR7, C-X-C chemokine
receptor type 7; shRNA, small hairpin RNA; CXCL12, C-X-C motif
chemokine ligand 12. |
CXCR7 knockdown reduces CXCL12-induced
invasive activity in M14 cells
The present study used 10 ng/ml CXCL12 to assess the
CXCR7 biological function in M14 cells. The M14 cell invasive
ability was monitored in a matrigel Transwell chamber assay with
CXCR7 shRNA-transfected, empty vector shRNA-transfected and
non-transfected M14 cells. It was observed that CXCR7 knockdown
significantly reduced the invasive activity of M14 cells compared
with that of shRNA vector-transfected and control cells. By
contrast, there was not a significant difference in the number of
M14 cells between shRNA vector-transfected and control cells
(Fig. 3C and D). These data indicated
that CXCR7 may regulate M14 cell migration and invasion.
CXCR7 shRNA inhibits melanoma tumor
development in mice
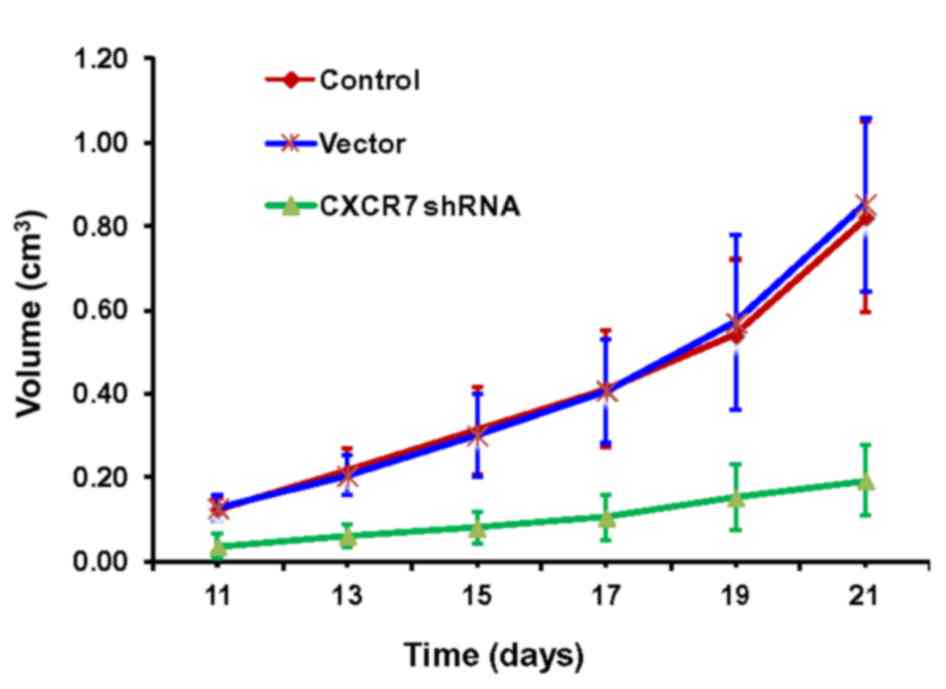
The effects of CXCR7 on transplanted skin melanoma
tumor progression were tested in mice. The transplanted tumors were
monitored daily, and the sizes were measured from day 11 to day 21
post-transplantation. It was observed that the tumor size grew
continually in the control and vector groups. By contrast, the
tumor size in the mice transfected with CXCR7 shRNA did not grow as
much as that of the control or shRNA vector-transfected mice
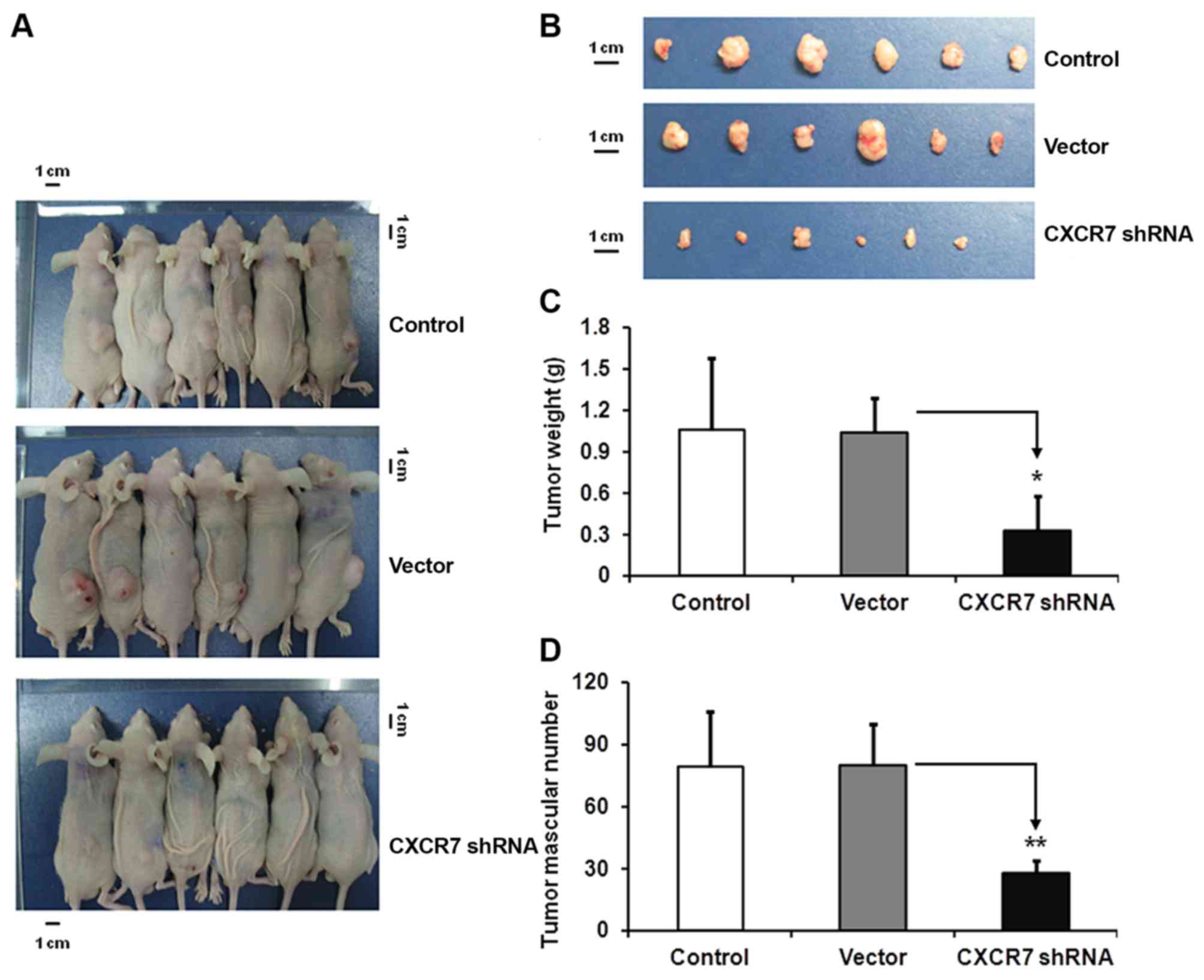
(Fig. 4). Tumors were harvested at
day 21 post-implantation. Smaller tumor sizes were observed in
CXCR7 shRNA mice compared with those in control and shRNA vector
mice (Fig. 5A and B). It has been
reported that vasculature serves an important role in skin cancer
inflammation (21), therefore the
tumor weight and tumor vascular number were also compared. The
average tumor weight was significantly lower in the CXCR7 shRNA
group compared with that in the control and vector shRNA groups
(*P<0.05; Fig. 5C). In addition,
the tumor vascular number was also significantly lower in CXCR7
shRNA mice compared with that in vector shRNA and control mice
(**P<0.01; Fig. 5D). These
findings further demonstrated the effects of CXCR7 on tumor
development in a mouse model.
Discussion
In the present study, a higher percentage (80%) of
positive CXCR7 protein staining was observed in MM tumors compared
with that in SSC and BCC tumors from patients, indicating CXCR7
involvement in melanoma cancer progression and development. It was
observed that M14 melanoma cells expressed high levels of CXCR7.
CXCR7 shRNA significantly knocked down CXCR7 mRNA and protein
expression in M14 cells, which subsequently inhibited M14 cell
biological functions, including cell migration, invasion and
metastasis. There are several reports indicating that CXCR7
expression was significantly higher in tumor tissue than in
adjacent normal tissue (10,22). The higher rate of cancer spreading to
other parts of the body was associated with higher CXCR7 levels in
cancer (23), further supporting the
CXCR7-mediated induction of skin melanoma invasion and
metastasis.
Several studies have shown that melanoma cell lines
co-express high levels of chemokine factor receptors and ligands
(24). Chemokines and tumor cell
surface receptors are involved in post-melanoma angiogenesis, tumor
cell growth, invasion and metastasis of malignant processes
(25). Furthermore, high levels of
cell surface CXCR4 expression were associated with melanoma tumor
surface ulceration, tumor invasion and increased mortality
(26). CXCR3 expression levels were
associated with tumor cell infiltration depth in primary invasive
melanoma (27). High expression of
CXCR3 and its ligands CXCL9 and CXCL10 led to actin polymerization
and invasion in B16F10 murine melanoma models (26,27). The
expression of the chemokine receptors CCR7, CCR10, CXCR3 and CXCR4
was associated with melanoma lymph node metastasis in mouse models
(19,23,27,28). CXCR7
interaction with the CXCL12 receptor has been demonstrated to
increase the CXCL12-induced normal epidermal cell migration
(29). Furthermore, it was observed
that the chemokine ligand for CXCR7 was CXCL12 (also known as
SDF-1), which also binds CXCR4 (30).
CXCR7 has been demonstrated to heterodimerize with CXCR4 and
regulate CXCL12-mediated G protein signaling (28). These findings supported CXCR7 roles on
melanoma biological behavior. CXCR7 is involved in multiple
functions, including tumor growth, survival and metastasis, in
several human tumors such as breast cancer, pancreatic cancer,
papillary thyroid carcinoma and prostate cancer (19,20,31). High
CXCR7 expression may promote angiogenesis, enhance tumor cell
invasion, and promote tumor invasion and metastasis (10,22,28,29).
Hypoxia-inducible factor-1α has been demonstrated to increase CXCR7
expression and activate epidermal growth factor receptor signal
transduction, and subsequently promote the growth of cancer cells
in prostate cancer (32). It has been
demonstrated that overexpression of CXCR7 in malignant human
myeloid cells resulted in nuclear factor-κB activation, followed by
mitogen-activated protein kinase p42/44 and AKT phosphorylation,
and subsequently increased cancer cell migration (22).
Therefore, a model can be proposed in which CXCR7
interacts with CXCL12 to activate the chemokine receptor signaling
pathway, and to increase melanoma cell migration and invasion. It
can be speculated that CXCR7 is a potential therapeutic target for
cutaneous melanoma treatments. Further characterization of
CXCR7-induced chemokine receptor signaling pathways is
warranted.
References
|
1
|
Balch CM, Soong SJ, Gershenwald JE,
Thompson JF, Reintgen DS, Cascinelli N, Urist M, McMasters KM, Ross
MI, Kirkwood JM, et al: Prognostic factors analysis of 17,600
melanoma patients: Validation of the American Joint Committee on
Cancer melanoma staging system. J Clin Oncol. 19:3622–3634. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Balch CM, Soong SJ, Smith T, Ross MI,
Urist MM, Karakousis CP, Temple WJ, Mihm MC, Barnhill RL, Jewell
WR, et al: Long-term results of a prospective surgical trial
comparing 2 cm vs. 4 cm excision margins for 740 patients with 1–4
mm melanomas. Ann Surg Oncol. 8:101–108. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Slominski A, Wortsman J, Carlson AJ,
Matsuoka LY, Balch CM and Mihm MC: Malignant melanoma. Arch Pathol
Lab Med. 125:1295–1306. 2001.PubMed/NCBI
|
|
4
|
Balch CM, Buzaid AC, Soong SJ, Atkins MB,
Cascinelli N, Coit DG, Fleming ID, Gershenwald JE, Houghton A Jr,
Kirkwood JM, et al: Final version of the American Joint Committee
on Cancer staging system for cutaneous melanoma. J Clin Oncol.
19:3635–3648. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
DeSantis CE, Lin CC, Mariotto AB, Siegel
RL, Stein KD, Kramer JL, Alteri R, Robbins AS and Jemal A: Cancer
treatment and survivorship statistics, 2014. CA Cancer J Clin.
64:252–271. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Morton DL, Wen DR, Wong JH, Economou JS,
Cagle LA, Storm FK, Foshag LJ and Cochran AJ: Technical details of
intraoperative lymphatic mapping for early stage melanoma. Arch
Surg. 127:392–399. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Baggiolini M and Loetscher P: Chemokines
in inflammation and immunity. Immunol Today. 21:418–420. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Balkwill F: Cancer and the chemokine
network. Nat Rev Cancer. 4:540–550. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Rossi D and Zlotnik A: The biology of
chemokines and their receptors. Annu Rev Immunol. 18:217–242. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sánchez-Martin L, Sánchez-Mateos P and
Cabañas C: CXCR7 impact on CXCL12 biology and disease. Trends Mol
Med. 19:12–22. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Beutner KR, Geisse JK, Helman D, Fox TL,
Ginkel A and Owens ML: Therapeutic response of basal cell carcinoma
to the immune response modifier imiquimod 5% cream. J Am Acad
Dermatol. 41:1002–1007. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Rowe DE, Carroll RJ and Day CL Jr:
Prognostic factors for local recurrence, metastasis, and survival
rates in squamous cell carcinoma of the skin, ear, and lip.
Implications for treatment modality selection. J Am Acad Dermatol.
26:976–990. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Shaw LM: Tumor cell invasion assays.
Methods Mol Biol. 294:97–105. 2005.PubMed/NCBI
|
|
14
|
Yang FC, Atkinson SJ, Gu Y, Borneo JB,
Roberts AW, Zheng Y, Pennington J and Williams DA: Rac and Cdc42
GTPases control hematopoietic stem cell shape, adhesion, migration,
and mobilization. Proc Natl Acad Sci USA. 98:5614–5618. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tsareva SA, Moriggl R, Corvinus FM,
Wiederanders B, Schütz A, Kovacic B and Friedrich K: Signal
transducer and activator of transcription 3 activation promotes
invasive growth of colon carcinomas through matrix
metalloproteinase induction. Neoplasia. 9:279–291. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Giard DJ, Aaronson SA, Todaro GJ, Arnstein
P, Kersey JH, Dosik H and Parks WP: In vitro cultivation of human
tumors: Establishment of cell lines derived from a series of solid
tumors. J Natl Cancer Inst. 51:1417–1423. 1973. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sulit HL, Golub SH, Irie RF, Gupta RK,
Grooms GA and Morton DL: Human tumor cells grown in fetal calf
serum and human serum: Influences on the tests for lymphocyte
cytotoxicity, serum blocking and serum arming effects. Int J
Cancer. 17:461–468. 1976. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Larionov A, Krause A and Miller W: A
standard curve based method for relative real time PCR data
processing. BMC Bioinformatics. 6:622005. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sun X, Cheng G, Hao M, Zheng J, Zhou X,
Zhang J, Taichman RS, Pienta KJ and Wang J: CXCL12/CXCR4/CXCR7
chemokine axis and cancer progression. Cancer Metastasis Rev.
29:709–722. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang J, Shiozawa Y, Wang J, Wang Y, Jung
Y, Pienta KJ, Mehra R, Loberg R and Taichman RS: The role of
CXCR7/RDC1 as a chemokine receptor for CXCL12/SDF-1 in prostate
cancer. J Biol Chem. 283:4283–4294. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Huggenberger R and Detmar M: The cutaneous
vascular system in chronic skin inflammation. J Investig Dermatol
Symp Proc. 15:24–32. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Tarnowski M, Liu R, Wysoczynski M,
Ratajczak J, Kucia M and Ratajczak MZ: CXCR7: A new SDF-1-binding
receptor in contrast to normal CD34(+) progenitors is functional
and is expressed at higher level in human malignant hematopoietic
cells. Eur J Haematol. 85:472–483. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
D'Alterio C, Consales C, Polimeno M,
Franco R, Cindolo L, Portella L, Cioffi M, Calemma R, Marra L,
Claudio L, et al: Concomitant CXCR4 and CXCR7 expression predicts
poor prognosis in renal cancer. Curr Cancer Drug Targets.
10:772–781. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Pinto S, Martinez-Romero A, O'Connor JE,
Gil-Benso R, San-Miguel T, Terrádez L, Monteagudo C and Callaghan
RC: Intracellular coexpression of CXC- and CC-chemokine receptors
and their ligands in human melanoma cell lines and dynamic
variations after xenotransplantation. BMC Cancer. 14:1182014.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Richmond A, Yang J and Su Y: The good and
the bad of chemokines/chemokine receptors in melanoma. Pigment Cell
Melanoma Res. 22:175–186. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Longo-Imedio MI, Longo N, Treviño I,
Lázaro P and Sánchez-Mateos P: Clinical significance of CXCR3 and
CXCR4 expression in primary melanoma. Int J Cancer. 117:861–865.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Monteagudo C, Martin JM, Jorda E and
Llombart-Bosch A: CXCR3 chemokine receptor immunoreactivity in
primary cutaneous malignant melanoma: Correlation with
clinicopathological prognostic factors. J Clin Pathol. 60:596–599.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Levoye A, Balabanian K, Baleux F,
Bachelerie F and Lagane B: CXCR7 heterodimerizes with CXCR4 and
regulates CXCL12-mediated G protein signaling. Blood.
113:6085–6093. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lee E, Han J, Kim K, Choi H, Cho EG and
Lee TR: CXCR7 mediates SDF1-induced melanocyte migration. Pigment
Cell Melanoma Res. 26:58–66. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Sierro F, Biben C, Martinez-Muñoz L,
Mellado M, Ransohoff RM, Li M, Woehl B, Leung H, Groom J, Batten M,
et al: Disrupted cardiac development but normal hematopoiesis in
mice deficient in the second CXCL12/SDF-1 receptor, CXCR7. Proc
Natl Acad Sci USA. 104:14759–14764. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Salazar N, Muñoz D, Kallifatidis G, Singh
RK, Jordà M and Lokeshwar BL: The chemokine receptor CXCR7
interacts with EGFR to promote breast cancer cell proliferation.
Mol Cancer. 13:1982014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Singh RK and Lokeshwar BL: The
IL-8-regulated chemokine receptor CXCR7 stimulates EGFR signaling
to promote prostate cancer growth. Cancer Res. 71:3268–3277. 2011.
View Article : Google Scholar : PubMed/NCBI
|