Introduction
Glioblastoma (GBM) is a solid invasive tumor
originating from the supporting cells of the central nervous system
(1,2), accounting for 40 to 50% of intracranial
tumors. According to its growth pattern, it is divided as diffuse
glioma and localized glioma. The former is common in adults and has
a higher degree of malignancy (3–6). Most
patients with gliomas have moderate advanced stages when they seek
medical advice. Surgical treatment is the main routine treatment
(7,8), but the effect is not ideal. Studies
show that the median survival time is only 12–15 months, and the 2-
and 5-year survival rates are 20 and 5%, respectively (9,10).
Therefore, early diagnosis and treatment, play an important role in
improving the survival rate of glioma patients. CAPG is a member of
the gelysin superfamily. Studies have shown that the expression of
CAPG in tumor cells is significantly higher than that in normal
tissues (11). However, there are
few related studies on the expression in glioma. The Cancer Genome
Atlas (TCGA) is a joint genome of the National Cancer and Oncology
Institute (NCI) and the National Human Genome Research Institute
(NHGRI) that uses large-scale genome sequencing to map all human
cancers. The variogram was drawn and has been systematically
analyzed, aiming to find all the minor mutations in carcinogenic
and anti-oncogenic genes. This study analyzed the TCGA gene
expression profiling chip and RNAseq data in combination with tumor
tissues of patients with clinical glioma, explored the clinical
expression characteristics of CAPG in human glioma, and provided
new ideas and methods for early diagnosis and treatment of
glioma.
Materials and methods
Data analysis method based on TCGA
Selection of data
The gene expression profiling chip RNA-seq level 3
data and pathological data of GBM in TCGA database were downloaded
and sorted, which contained 529 disease samples and 10 normal
samples, which were directly used to analyze differential genes. In
this study, 515 samples of low grade gliomas (LGG) and 152 samples
of high grade gliomas (HGG) were selected.
The study was approved by the Ethics Committee of
the First Affiliated Hospital of Xinjiang Medical University
(Urumqi, China). Patients who participated in this research had
complete clinical data. The signed informed consents were obtained
from the patients or the guardians.
Obtaining differential genes
Our data was normalized and t-test was done using
the Affy and Limma packages in the R language, and then the data
was filtered according to P<0.05 and |FC|≥2. The performance of
CardGene is shown in Table I.
 | Table I.Performance of CardGene. |
Table I.
Performance of CardGene.
| Gene | FC | P-value | FDR | Total samples | Normal samples | Cancer samples |
|---|
| CAPG | 5.953 | 7.920E-15 | 1.104E-13 | 558 | 10 | 548 |
HCS screening
The number of target genes was reduced by the
following criteria: i) the genes reported in relevant SCI papers
were removed; ii) removing multiple transmembrane protein genes;
iii) removing undefined gene annotation (e.g., with an open reading
frame in the description); iv) removing the number of PubMed
articles more than 60; and v) removing filtered gene from existing
experimental data in key gene databases. The relevant data sources
are highly reliable gene disease databases. And the final list of
genes was randomly condensed to determine which list of genes to be
analyzed (Table II).
 | Table II.CardGene genetic information. |
Table II.
CardGene genetic information.
| Gene ID | Gene name | Number of
transcripts | Number of Pubmed
documents | Novoseek disease
relationships for the gene | MalaCard disease
relationships for the gene |
|---|
| 822 | CAPG | 6 | 54 | 0 | 0 |
Clinical experimental methods
Organization of acquisition
From 2016 to 2017, 25 cases of glioma tissue
specimens diagnosed in the Neurosurgery Department of the First
Affiliated Hospital of Xinjiang Medical University were collected
as the experimental group, aged from 21 to 58 years (mean
43.12±10.12), 15 males and 10 females. None of the patients
received anti-tumor therapy such as chemotherapy, radiotherapy or
biological therapy before operation, and each case was confirmed by
complete clinical and pathological data. Fifteen normal brain
tissue specimens were collected from traumatic decompression
surgery in the same period as the control group. All specimens were
collected within half an hour after isolation and stored in a
refrigerator at −80°C.
Experimental methods
Total RNA was extracted from the glioma tissue using
the TRIzol reagent (Invitrogen Life Technologies, Inc.; Thermo
Fisher Scientific, Inc., Waltham, MA, USA). RT-PCR was used to
evaluate the expression of CAPG in the cell lines. The cDNA was
obtained by reverse transcription kit and stored at −20°C for later
use. The total RNAs were extracted by RNA isolation kit (GE
Healthcare, Munich, Germany), and the cDNAs were prepared by using
transcriptor first strand cDNA synthesis kit (Roche Diagnostics
GmbH, Mannheim, Germany), carried out on the ABI 7500, SYBR Green
system (Bio-Rad Laboratories, Inc., Hercules, CA, USA). The
conditions of qPCR were as follows: One cycle at 50°C for 2 min and
95°C for 10 min and followed by 40 cycles of denaturation at 95°C
for 15 sec and annealing extension at 55°C for 1 min. The primers
used included the following: CAPG F: CGAACACTCAGGTGGAGATT and R:
TCCAGTCCTTGAAAAATTGC; GAPDH F: TGCACC ACCAACTGCTTAGC and R:
GGCATGGACTGTGGTCAT GAG. After amplification, the melting curve was
analyzed to determine the specificity of the PCR products, and the
threshold cycle values of CAPG and GAPDH were obtained. The
relative quantity of CAPG was calculated by 2−ΔΔCq as
the reference (12).
Immunohistochemistry
Τissues were harvested and fixed in 10% formalin at
room temperature for one week, then embedded in paraffin, from
which 4 μm-thick sections were made. The slides were
incubated with 3% hydrogen peroxide to quench endogenous peroxidase
activities. After heat induced antigen retrieval, the specimens
were blocked with 5% normal goat serum (ImmunoReagents Inc.,
Raleigh, NC, USA) for 1 h at room temperature, and incubated with
antibodies against CAPG (cat. no. ab155688; dil, 1:50) at 4°C
overnight, and donkey anti-goat IgG-HRP secondary antibody (cat.
no. ab205723; dil, 1:200) both from Abcam (Cambridge, UK). Slides
underwent color development with 3–3′-diaminobenzidine (DAB) and
were counterstained with hematoxylin. Photomicrographs were taken
with a light Olympus microscope BX51 (Olympus Corporation, Tokyo,
Japan). The staining intensity was scored on a scale of 0 to 3 as
negative, weak, medium and strong, respectively. The stained area,
which was calculated as the percentage of positively stained cells
relative to the total cells, was scored on a scale of 0 to 4: 0
(0%), 1 (1–25%), 2 (26–50%), 3 (51–75%) and 4 (76–100%). The
overall score was calculated by multiplying the intensity score and
the staining area score. Samples were categorized into four grades:
An overall score equal to 0 was graded as ‘−’; an overall score
equal to 1, 2, 3 or 4 was graded as ‘+’; an overall score equal to
5, 6, 7 or 8 was graded as ‘++’; an overall score equal to 9, 10,
11 or 12 was graded as ‘+++’. The stained tissue sections were
analyzed by two pathologists without any knowledge regarding the
patient clinical information. Based on the immunohistochemical
grades, the patients were divided into two groups: the high
expression group, which included patients with grades ‘++’ or
‘+++’, and the low expression group, including patients graded as
‘-’ or ‘+’.
Data collection
The data of age, sex, ethnicity, surgical approach,
tissue type of tumor, metastasis condition, pathological staging
and follow-up for one year of survival were collected.
Statistical analysis
Data was entered in excel, and analyzed by R
language. Kolmogorov-Smirnov test was used to test the normality of
relative quantity of CAPG mRNA. The measurement data were analyzed
by t-test and variance analysis, and the counting data were tested
by Chi-square test. Spearman's correlation was used for correlation
analysis, and log-rank test was used to test the statistical
significance of survival analysis. Kaplan-Meier was used in order
to generate the curves. The test level was 0.05, P<0.05 was
considered to indicate a statistically significant difference.
Results
Results of TCGA data analysis method
Results of CardGene analysis
Previous studies have yielded differential results
for normal and human glioma CAPG genes expression. The vertical
coordinates shown below were the corresponding signal values of
genes in the expression microarray, and the transverse axis was
from normal and cancer tissues, respectively. Fig. 1, clearly shows the difference of gene
expression distribution between the two, the expression of CAPG in
cancer tissue was higher than that in normal tissue
(P<0.05).
Analysis of significant differences in
gene expression levels at various clinical data levels
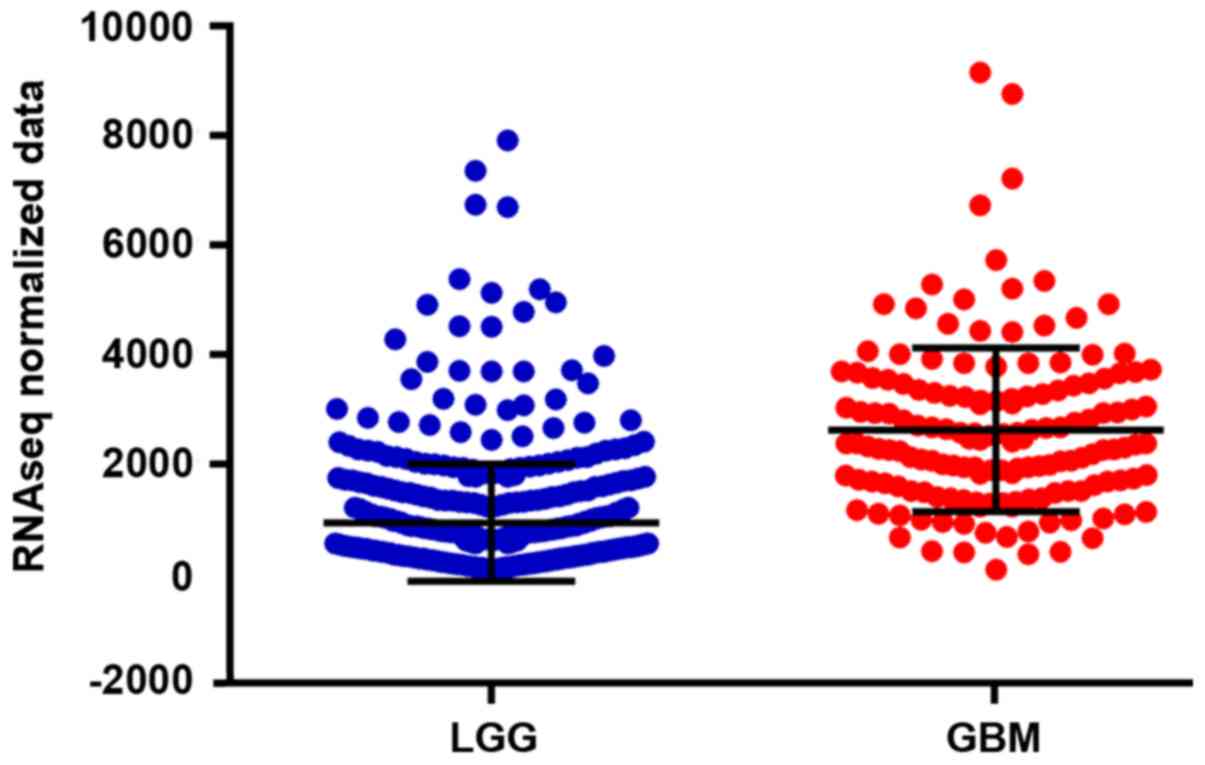
The results of this study showed that the expression
of CAPG was significantly different in patients with different age,
sex and pathological stage, suggesting that the expression of CAPG
might be related to the above-mentioned clinical pathological
stage. The results of statistical analysis are shown in Table III, and the high expression rate of
CAPG aged >46 years, male and pathological stages of HGG were
higher than that aged <46, female and pathological stages of
LGG. Pathological information is shown in Fig. 2.
 | Table III.Analysis of significant differences in
gene expression levels at various clinical data levels. |
Table III.
Analysis of significant differences in
gene expression levels at various clinical data levels.
|
| Expression levels of
objective gene |
|
|---|
|
|
|
|
|---|
| Variables | Low | High | P-value |
|---|
| Age |
|
| <0.001 |
| ≤46 | 224 (65.88) | 116 (34.12) |
|
|
>46 | 110 (33.64) | 217 (66.36) |
|
| Sex |
|
| 0.037 |
| Male | 179 (46.61) | 205 (53.39) |
|
|
Female | 155 (54.77) | 128 (45.23) |
|
| Pathological
stage |
|
| <0.001 |
| LGG | 324 (62.91) | 191 (37.09) |
|
| HGG | 10 (6.58) | 142 (93.42) |
|
Correlation between gene expression
level and clinical data
The results of correlation analysis showed that the
expression level of CAPG was significantly correlated with age, sex
and pathological stage of the patients, and was positively
correlated with age and pathological stage, and negatively
correlated with sex (P<0.001; Table
IV).
 | Table IV.Correlation between gene expression
level and clinical data. |
Table IV.
Correlation between gene expression
level and clinical data.
| Variables | Expression level | Age | Sex | Pathological
stage |
|---|
| Expression level |
| r | 1.000 | 0.322 | −0.081 | 0.473 |
|
P-value | – | <0.001 | 0.037 | <0.001 |
| Age |
| r | 0.322 | 1.000 | −0.011 | 0.397 |
|
P-value | <0.001 | – | 0.785 | <0.001 |
| Sex |
| r | −0.081 | −0.011 | 1.000 | −0.083 |
|
P-value | 0.037 | 0.785 | – | 0.032 |
| Pathological
stage |
| r | 0.473 | 0.397 | −0.083 | 1.000 |
|
P-value | <0.001 | <0.001 | 0.032 | – |
Univariate survival analysis
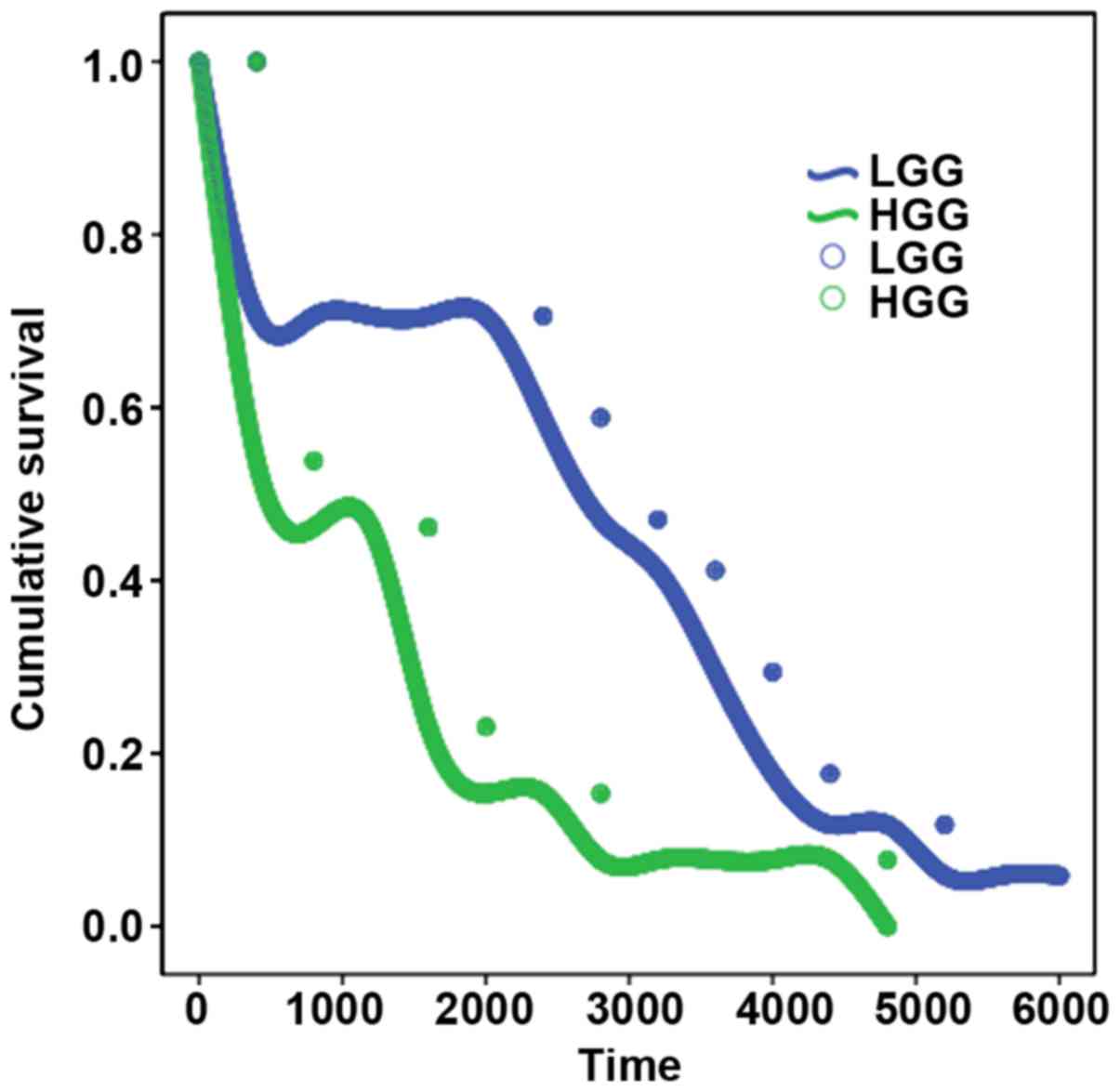
The results of survival analysis showed that the
expression level of CAPG had a significant effect on the survival
time of patients, and the prognosis of patients with higher
expression level of CAPG was poor. The difference of log-rank test
was significant (P<0.05; Table V
and Fig. 3).
 | Table V.Level of gene expression and survival
test of clinical data. |
Table V.
Level of gene expression and survival
test of clinical data.
| Variables | Total no. | Events | P-value |
|---|
| Expression |
|
HGG | 332 | 76 | <0.001 |
|
LGG | 332 | 169 |
|
|
Total | 664 | 245 |
|
Clinical experimental results
Normality test
Kolmogorov-Smirnov test was used to test the
normality of relative quantity of CAPG mRNA, Z=3.211, P<0.001,
which is a skewed distribution.
Expression of CAPG in glioma
tissues
The results of IHC staining showed that the
expression of CAPG in LGG and HGG tissues was higher than that in
normal tissues (Fig. 4A), the
difference was statistically significant (P<0.05; Fig 4B).
Characteristics of CAPG mRNA
expression in glioma
The results showed that the expression of CAPG mRNA
in glioma was different among different sex, lymph node metastasis
and pathological tissue types. The expression of CAPG in male,
lymphatic metastasis and poorly differentiated tumors was higher
than that in female, non-lymphatic metastasis and highly
differentiated tumors, and the difference was statistically
significant (P<0.005; Table
VI).
 | Table VI.Expression of CAPG mRNA in
clinic. |
Table VI.
Expression of CAPG mRNA in
clinic.
| Variables | No. | Relative expression
of CAPG mRNA | P-value |
|---|
| Age |
|
<50 | 14 | 0.30 (0.17,
0.45) | 0.143 |
|
>50 | 11 | 0.32 (0.15,
0.47) |
|
| Sex |
|
Male | 15 | 0.33 (0.15,
0.41) | 0.004 |
|
Female | 10 | 0.29 (0.11,
0.57) |
|
| Ethnicity |
|
Han | 19 | 0.30 (0.15,
0.58) | 0.209 |
|
Minorities | 6 | 0.32 (0.13,
0.51) |
|
| Lymphatic
metastasis |
|
Have | 12 | 0.35 (0.20,
0.50) | <0.001 |
| No | 13 | 0.25 (0.13,
0.62) |
|
| Pathological
types |
|
HGG | 10 | 0.33 (0.21,
0.51) | 0.013 |
|
LGG | 15 | 0.27 (0.15,
0.39) |
|
|
Normal | 15 | 0.13 (0.09,
0.30) |
|
Analysis of CAPG mRNA and survival
time
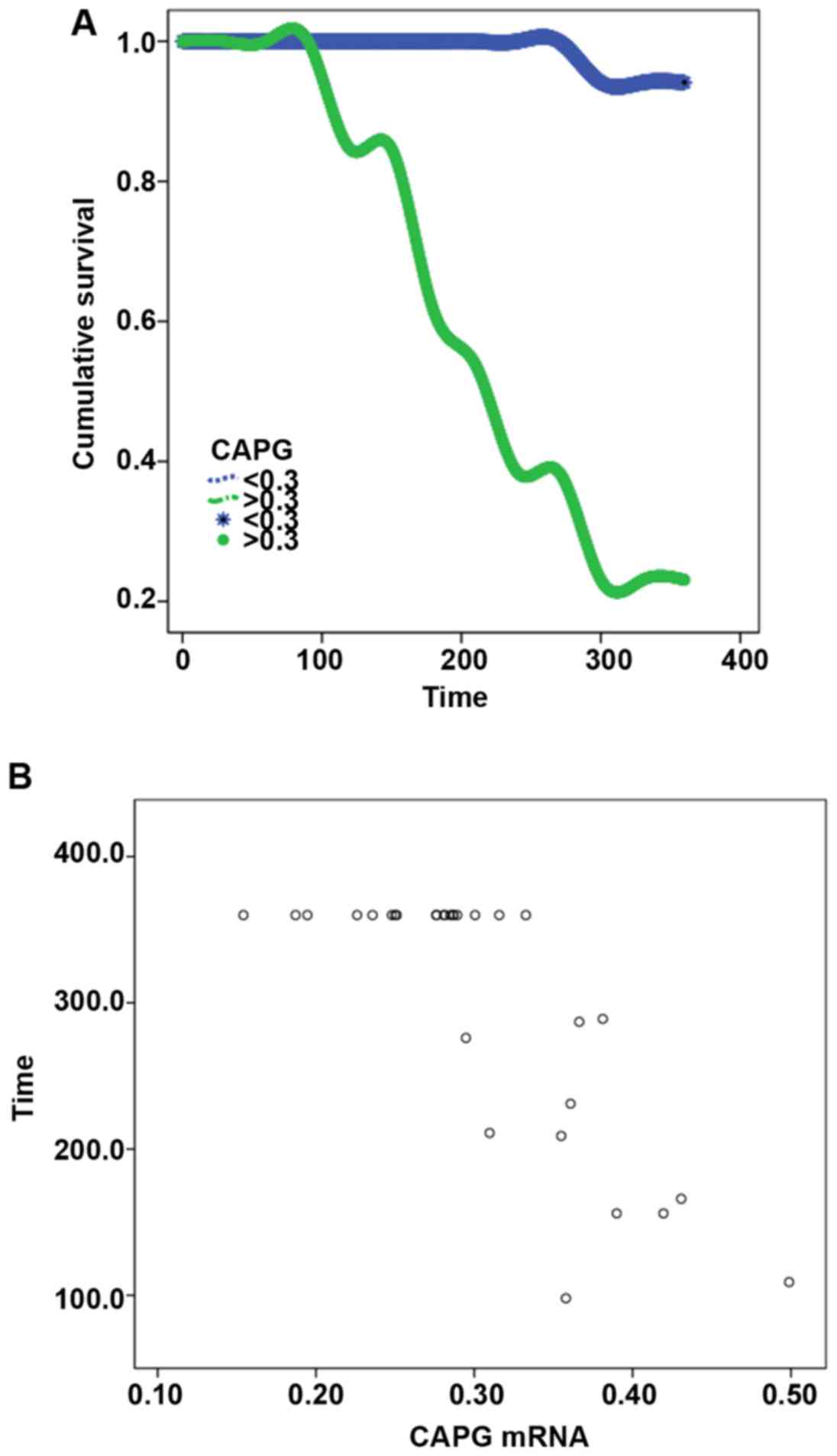
Among the patients who were followed up for one
year, 11 died and 14 survived. The expression level of CAPG had a
significant effect on the survival time of patients. Patients with
higher expression levels of CAPG had shorter survival time, and
there was a significant difference in log-rank test (P<0.001).
Pearson's correlation analysis of survival time and CAPG expression
showed a negative correlation between survival time and CAPG
(r=−0.792, P<0.001; Table VII
and Fig. 5).
 | Table VII.The result of survival test. |
Table VII.
The result of survival test.
| Variables | Total no. | Event | Log-rank | P-value |
|---|
| Expression |
|
<0.3 | 12 | 1 | 15.3321 | <0.001 |
|
>0.3 | 13 | 10 |
|
|
|
Total | 664 | 245 |
|
|
Multi-factor analysis
The statistically significant index was used in
Table II as the independent
variable, and the expression of CAPG mRNA was logarithmically
transformed (normalized) and then used as a dependent variable for
multiple linear regression analysis, αin=0.05, αout=0.01. Stepwise
regression analysis showed that lymphatic metastasis and
histopathological type were independent factors affecting the
expression of CAPG. The expression of CAPG in tumor tissues with
lymphatic metastasis was higher than that in non-lymphatic
metastatic tissues, and the lower the differentiation degree, the
higher the expression of CAPG in tumor tissues with lymphatic
metastasis. The difference was statistically significant, as shown
in Table VIII.
 | Table VIII.Analysis of multiple linear
regression. |
Table VIII.
Analysis of multiple linear
regression.
| Index | B | S.E | Beta | P-value |
|---|
| Variable | 48.321 | 10.098 |
| <0.001 |
| Sex | 10.991 | 9.032 | 0.435 | 0.213 |
| Lymphatic
metastasis | −8.123 | 1.276 | 0.389 | <0.001 |
| Tissue type | 11.587 | 3.121 | 0.320 | 0.005 |
Discussion
Glioma is a common malignant tumor in the central
nervous system, the morbidity is approximately 80% in the alignant
tumors of brain. In intracranial malignant tumors, the GBM, which
has the highest degree of malignancy, is one of the main causes of
death. In recent years, the important role of cytoskeleton protein
in tumor prognosis has been paid increasing attention, and
cytoskeleton recombination is an important factor in invasion and
metastasis of tumor cells (13,14).
CAPG is an important actin-binding protein, which was first found
in human alveolar macrophages, and it is Ca2+ dependent
actin-binding protein which is composed of 348 amino acids. It
regulates the length of actin by capping and cutting microfilaments
to promote the cross-linking of actin and the reorganization of the
cytoskeleton (15). Therefore, CAPG
might be a new potential factor to promote tumorigenesis and as a
new target of anti-cancer drugs.
Various biological public databases have been
applied with the advent of the big data era, such as the Gene
Expression Omnibus (GEO) gene expression database, the Survey
Epidemiology and End Results (SEER) database and the gene Atlas of
TCGA database (16–18). TCGA is led by the NCI, and is a major
project to conquer cancer in USA. It systematically provides
sequencing and chip data of cancer genomics, provides a large
genomic and clinical data for researchers in cancer basic science
and gene transfer chemistry, and provides a large data basis for
mining meaningful genomes and discovering biological mechanisms
affecting tumor occurrence, development, differentiation and
metastasis.
Our study aimed to analyze the data of gene
expression profile-chip and pathology which were selected from 529
disease samples and 10 normal GBM in TCGA database. The results
showed that the expression of CBM CAPG gene was higher than that in
normal tissues. A further analysis of the clinical data showed that
the expression of CAPG in males with the high expression rate of
CAPG aged >46 years and pathological stages of HGG were higher
than that in females aged ≤46, with pathological stages of LGG. The
results of correlation analysis also showed that CAPG expression
level was positively correlated with age and pathological stage,
and negatively correlated with sex. Survival analysis showed that
the expression level of CAPG had a significant impact on the
survival time, and patients with higher expression level of CAPG
had poor prognosis.
Above are the results of TCGA data analysis.
However, there would be various kinds of gene expression due to
different countries, regions and races. In order to further verify
the clinical features of CAPG expression in GMB tumor tissues, we
randomly selected 25 tumor tissues of glioma patients from
neurosurgery department in the First Affiliated Hospital of XJMU of
China as the experimental group, and 15 normal brain tissues which
underwent traumatic decompression surgery as the control group.
RT-qPCR showed that the expression of CAPG mRNA in glioma tissues
was higher than that in normal tissues, and the expression level in
males with lymphatic metastasis and low differentiation was higher
than that in females, without lymphatic metastasis and high
differentiation. The results of survival analysis after one year's
follow-up showed that the expression level had a significant impact
on the survival time. The survival time of patients with higher
expression level was short, and Pearson correlation analysis showed
that there was negative correlation between the survival time and
CAPG.
The results above showed that CAPG was highly
expressed in glioma tumor tissues. Resent studies show that CAPG
are highly expressed in lung cancer (19), pancreatic cancer (20), breast cancer (21,22),
prostate cancer (23) and gastric
cancer (24,25), and are closely related with
metastasis-CAPG is obviously highly expressed in tumor tissues with
metastasis. The expression level of CAPG in glioma might be related
to age, sex, metastasis, pathological stage and tissue type. There
are two studies on glioma which could be retrieved at present. Xing
and Zeng (26) reported a study in
2015, on tissue samples from 90 gliomas and 90 adjacent brain
tissues and 5 pairs with grade I–IV gliomas. RNA was extracted and
expressed on Affymetrix Human Gen U133 Plus 2.0 Array. The results
showed that 1,725 genes were differentially expressed in glioma,
and their expression levels were related to pathological grade, and
were not related to age and subtype, and the mRNA expression level
of CAPG was increasing from stage II to IV. Thus proving that CAPG
protein expression in human glioma was related. The survival
analysis also showed that patients with high expression of CAPG had
a short survival time, which was consistent with our study. Yun
et al (27) published a study
in 2018, which was based on the TCGA database. It also pointed out
that the mRNA and protein levels of CAPG in human glioma were
significantly increased, and the high expression of CAPG was an
independent prognostic factor for poor prognosis. Multivariate COX
analysis showed that the median survival was short in glioma
patients over 60 years of age with high CAPG expression. The
results showed that overall survival was shortened under the high
expression of CAPG.
As can be seen from the above studies, the
expression of CAPG in glioma is indeed high, and it is highly
expressed with the severity of the disease, and it is obviously
related to the prognosis. Therefore, CAPG could be used as a
biomarker for pathological grade and prognosis of glioma. However,
the related studies are not consistent on the expression of
different sex and ages, and further study is needed.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
QF, MS and QZ were responsible for RT-qPCR. SL and
YK collected and analyzed general data of patients. YD and BL were
responsible for statistical analysis. QF wrote the manuscript. The
final version was read and approved by all the authors.
Ethics approval and consent to
participate
The study was approved by the Ethics Committee of
the First Affiliated Hospital of Xinjiang Medical University
(Urumqi, China). Patients who participated in this research had
complete clinical data. The signed informed consents were obtained
from the patients or the guardians.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Oh J, Kim Y, Che L, Kim JB, Chang GE,
Cheong E, Kang SG and Ha Y: Regulation of cAMP and GSK3 signaling
pathways contributes to the neuronal conversion of glioma. PLoS
One. 12:e01788812017. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Yu X, Sun NR, Jang HT, Guo SW and Lian MX:
Associations between EGFR gene polymorphisms and susceptibility to
glioma: a systematic review and meta-analysis from GWAS and
case-control studies. Oncotarget. 8:86877–86885. 2017.PubMed/NCBI
|
|
3
|
Zhao P, Chen A, Qi Q, Zhou W, Feng Z, Wang
J, Yang N, Li X, Wang J, Huang Q, et al: Impact of VEGFA
polymorphisms on glioma risk in Chinese. Oncotarget. 8:83712–83722.
2017.PubMed/NCBI
|
|
4
|
Gao L, Chen B, Li J, Yang F, Cen X, Liao Z
and Long X: Wnt/β-catenin signaling pathway inhibits the
proliferation and apoptosis of U87 glioma cells via different
mechanisms. PLoS One. 12:e01813462017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Abdul AURM, De Silva B and Gary RK: The
GSK3 kinase inhibitor lithium produces unexpected
hyperphosphorylation of β-catenin, a GSK3 substrate, in human
glioblastoma cells. Biol Open. 7:72018. View Article : Google Scholar
|
|
6
|
Biau J, Chautard E, De Schlichting E,
Dupic G, Pereira B, Fogli A, Müller-Barthélémy M, Dalloz P, Khalil
T, Dillies AF, et al: Radiotherapy plus temozolomide in elderly
patients with glioblastoma: A ‘real-life’ report. Radiat Oncol.
12:1972017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chen Q, Ye L, Fan J, Zhang X, Wang H, Liao
S, Song P, Wang Z, Wang S, Li Y, et al: Autophagy suppression
potentiates the anti-glioblastoma effect of asparaginase in vitro
and in vivo. Oncotarget. 8:91052–91066. 2017.PubMed/NCBI
|
|
8
|
Andronesi OC, Esmaeili M, Borra RJH,
Emblem K, Gerstner ER, Pinho MC, Plotkin SR, Chi AS, Eichler AF,
Dietrich J, et al: Early changes in glioblastoma metabolism
measured by MR spectroscopic imaging during combination of
anti-angiogenic cediranib and chemoradiation therapy are associated
with survival. NPJ Precis Oncol. Jun 12–2017.(Epub ahead of print).
doi: 10.1038/s41698-017-0020-3. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Costa RB, Costa R, Kaplan J, Cruz MR, Shah
H, Matsangou M and Carneiro B: A rare case of glioblastoma
multiforme with osseous metastases. Case Rep Oncol Med.
2017:29383192017.PubMed/NCBI
|
|
10
|
Hays EM, Duan W and Shigdar S: Aptamers
and glioblastoma: Their potential use for imaging and therapeutic
applications. Int J Mol Sci. 18:182017. View Article : Google Scholar
|
|
11
|
Van den Abbeele A, De Corte V, Van Impe K,
Bruyneel E, Boucherie C, Bracke M, Vandekerckhove J and Gettemans
J: Downregulation of gelsolin family proteins counteracts cancer
cell invasion in vitro. Cancer Lett. 255:57–70. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Seifabadi S, Vaseghi G, Javanmard SH,
Omidi E, Tajadini M and Zarrin B: The cytotoxic effect of memantine
and its effect on cytoskeletal proteins expression in metastatic
breast cancer cell line. Iran J Basic Med Sci. 20:41–45.
2017.PubMed/NCBI
|
|
14
|
Vitali E, Boemi I, Rosso L, Cambiaghi V,
Novellis P, Mantovani G, Spada A, Alloisio M, Veronesi G, Ferrero
S, et al: FLNA is implicated in pulmonary neuroendocrine tumors
aggressiveness and progression. Oncotarget. 8:77330–77340. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Glaser J, Neumann MH, Mei Q, Betz B, Seier
N, Beyer I, Fehm T, Neubauer H, Niederacher D and Fleisch MC:
Macrophage capping protein CapG is a putative oncogene involved in
migration and invasiveness in ovarian carcinoma. BioMed Res Int.
2014:3798472014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Aevermann BD, Pickett BE, Kumar S, Klem
EB, Agnihothram S, Askovich PS, Bankhead A III, Bolles M, Carter V,
Chang J, et al: A comprehensive collection of systems biology data
characterizing the host response to viral infection. Sci Data.
1:1400332014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Welzel TM, Graubard BI, Zeuzem S, El-Serag
HB, Davila JA and McGlynn KA: Metabolic syndrome increases the risk
of primary liver cancer in the United States: a study in the
SEER-Medicare database. Hepatology. 54:463–471. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Tomczak K, Czerwińska P and Wiznerowicz M:
The Cancer Genome Atlas (TCGA): An immeasurable source of
knowledge. Contemp Oncol (Pozn) 19 (1A). A68–A77. 2015.
|
|
19
|
Zhu WY, Hunag YY, Liu XG, He JY, Chen DD,
Zeng F, Zhou JH and Zhang YK: Prognostic evaluation of CapG,
gelsolin, P-gp, GSTP1, and Topo-II proteins in non-small cell lung
cancer. Anat Rec (Hoboken). 295:208–214. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tonack S, Patel S, Jalali M, Nedjadi T,
Jenkins RE, Goldring C, Neoptolemos J and Costello E:
Tetracycline-inducible protein expression in pancreatic cancer
cells: Effects of CapG overexpression. World J Gastroenterol.
17:1947–1960. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Van Impe K, Bethuyne J, Cool S, Impens F,
Ruano-Gallego D, De Wever O, Vanloo B, Van Troys M, Lambein K,
Boucherie C, et al: A nanobody targeting the F-actin capping
protein CapG restrains breast cancer metastasis. Breast Cancer Res.
15:R1162013. View
Article : Google Scholar : PubMed/NCBI
|
|
22
|
Westbrook JA, Cairns DA, Peng J, Speirs V,
Hanby AM, Holen I, Wood SL, Ottewell PD, Marshall H, Banks RE, et
al: CAPG and GIPC1: Breast cancer biomarkers for bone metastasis
development and treatment. J Natl Cancer Inst. 108:1082016.
View Article : Google Scholar
|
|
23
|
Li BK, Guo K, Li CY, Li HL, Zhao PP, Chen
K and Liu CX: Influence of suppression of CapG gene expression by
siRNA on the growth and metastasis of human prostate cancer cells.
Genet Mol Res. 14:15769–15778. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Xiang G: Relationship between macrophage
capping protein and gastric cancer cell's proliferation and
migration ability. Beijing Da Xue Xue Bao Yi Xue Ban. 49:489–494.
2017.(In Chinese). PubMed/NCBI
|
|
25
|
Ichikawa H, Kanda T, Kosugi S, Kawachi Y,
Sasaki H, Wakai T and Kondo T: Laser microdissection and
two-dimensional difference gel electrophoresis reveal the role of a
novel macrophage-capping protein in lymph node metastasis in
gastric cancer. J Proteome Res. 12:3780–3791. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Xing W and Zeng C: An integrated
transcriptomic and computational analysis for biomarker
identification in human glioma. Tumour Biol. 37:7185–7192. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yun DP, Wang YQ, Meng DL, Ji YY, Chen JX,
Chen HY and Lu DR: Actin-capping protein CapG is associated with
prognosis, proliferation and metastasis in human glioma. Oncol Rep.
39:1011–1022. 2018.PubMed/NCBI
|