Introduction
Cervical cancer (CC) is one of the most common
oncological diseases in women worldwide and one of the leading
causes of female cancer mortality. One etiological factor for CC is
infection with the human papillomavirus of a high carcinogenic risk
(hereafter HR-HPV). Epithelial damage tends to progress slowly
(10–20 years from contact with HPV to the development of invasive
cancer). Therefore, regular cervical screening can allow the
detection of most lesions at early stages and drastically reduce
the risk of CC. At present, the preferred method for primary
cervical screening is cervical cytology, the efficacy of which as a
single screening method is limited due to technical constraints,
human factors as well as limitations of the pathomorphological
classification itself. This results in relatively low and varying
sensitivity (50–80% in different clinical settings) and compromised
specificity of cervical cytology (1). Another problem is the ambiguity of the
prognosis for patients with mild neoplasia (L-SIL) for possible
long- and short-term outcomes, from the complete regression to the
rapid development of invasive cancer.
The value of cervical screening can be improved by
the additional analysis of molecular biomarkers. At present, the
only marker widely used in clinical practice is HR-HPV DNA. HPV
testing has a higher diagnostic sensitivity than the cytological
method for the detection of cervical neoplasia (2,3).
Therefore, it is widely used in cervical screening in combination
with cytology (4,5) or as a method of primary screening
(6,7). However, HR-HPV infection is frequent
in women with no signs of cervical neoplasia even at the age of
risk (26–30 years and older) (8).
In the vast majority of cases, HPV infection is transient and
eliminated spontaneously (9).
Therefore, despite the high diagnostic sensitivity and negative
predictive value (NPV) of HR-HPV testing, its diagnostic
specificity and positive predictive value (PPV) in cervical
screening are relatively low. High viral load of HPV DNA in older
ages is now considered a surrogate marker of the HPV persistence
pointing to an increased risk of malignant transformation but the
PPV of this indicator is insufficient.
In multiple studies, diagnostic relevance of the
wide range of additional molecular biomarkers of dysplastic changes
in the cervix has been reported. These include the integrated form
of HR-HPV DNA, the amplification of telomerase gene subunits, the
levels of various mRNAs and microRNAs, and the aberrant methylation
of the promoters of various genes. Moreover, accumulating evidence
indicates that some morphologically indistinguishable subgroups of
CIN2 and even CIN3 neoplasms have very different long-term chances
of malignant transformation. Such subgroups can be discriminated by
analyzing the content of molecular markers of genetic and/or
epigenetic changes in affected cells [reviewed in ref. (10)].
MicroRNAs play a significant role in the development
of all types of cancer including CC. Cervical lesions are always
accompanied by an increase or decrease in the levels of various
microRNA which are correlated with the severity of the lesion
and/or are characteristic of invasive cancer in comparison with
preinvasive stages (11–20).
MicroRNAs are markedly stable in clinical material,
including cytology specimens. Therefore, they are regarded as
perspective clinical biomarkers. Several recent studies have
demonstrated the feasibility of using microRNA profiling in
cervical samples for diagnostic purposes (18,21,22).
At the same time, the inconsistency of accumulated data concerning
changes in microRNA levels in the above-mentioned studies impedes
the translation of their results into clinical practice. The
reported degree and direction of individual microRNA level changes
in cervical lesions can vary substantially and be even
contradictory in different studies (23–25).
This may be due to differences in the techniques used for
quantification, the characteristics of the cohorts of enrolled
patients, as well as to the different methods of raw data handling.
The importance of proper normalization for quantitative estimates
of microRNAs is undoubted (26–29).
MicroRNAs represent only a small fraction of the total RNA in the
cell; moreover, this fraction can vary significantly between
different types of specimens. The extraction efficiency of these
small molecules can differ significantly from the extraction
efficiency of longer RNAs extracted from sample of the same type by
the same method. Thus, traditionally used housekeeper mRNAs are not
applicable for the normalization of microRNA expression data. At
the same time, profiles of microRNA expression are characterized by
high tissue and cellular specificity (30), and there are no identified microRNA
genes expressed as stably as known protein-coding housekeeping
genes. Due to the above difficulties, the normalizers for microRNA
quantitation in different tissues and specimen types are often
chosen empirically. Depending on the method of reference microRNA
selection, different researchers choose different normalizers. The
use of geometric mean of the group of normalizers instead of a
single reference can reduce the bias introduced by normalization.
Such an approach, called GeNorm (31) makes it possible to rank candidate
reference genes by their expression stability, based on the
calculation of an average pairwise variation between all studied
genes, and to determine the optimum set of reference genes required
for normalization. However, in the case of microRNA analysis this
method either requires all possible normalizers to be analyzed or
faces the problem of rational selection of the normalizers. The
alternative is utilizing the mean expression value of all expressed
microRNAs in a given sample as a normalization factor (27). However, this approach requires a
large set of microRNAs to be profiled in a single specimen, which
may be unacceptable in clinical practice for both technical and
economic reasons.
The aim of the present study was to develop a method
for detecting high-grade cervical intraepithelial neoplasia and CC
in cytological specimens by PCR-based analysis of a small set of
microRNAs.
Materials and methods
Clinical material
The present study was approved by the local Ethics
Committee of the Federal Government Budgetary Institution ‘N.N.
Petrov Research Institute of Oncology’ as of February 13, 2014
(Internal No. 21). The samples were obtained from patients who
underwent examination and treatment at the Oncogynecology
Department of the Oncology Research Institute over the period
2010–2016. Cytological examination of cervical smears and
histological examination of the surgical material were carried out
by specialists at the Cytology Laboratory and Department of
Pathomorphology of the Oncology Research Institute, respectively.
Cytological specimens were obtained from the archives of the
Cytology Laboratory and clinical data were obtained from the
database of the Oncology Research Institute. Before the study, the
clinical material and information were subjected to
anonymization.
The cervical epithelial scrapings were obtained and
prepared by routine methods (Papanicolaou staining). The samples
were classified according to the Bethesda system (32): normal cytology [negative for
intraepithelial lesions or malignancy (NILM)] (n=40, mean age 31),
low-grade squamous intraepithelial lesion (L-SIL) (n=34, mean age
36), high-grade squamous intraepithelial lesion (H-SIL) (n=57, mean
age 44), invasive cervical cancer (CC) (n=43, mean age 53). All
H-SIL and CC diagnoses were histologically verified after
subsequent surgical treatment. The coincidence of cytological and
histological conclusions was observed in 100% of CC cases. Moderate
neoplasia (H-SIL) was confirmed histologically in 88% cases, in the
remaining cases (7 of 57, 12%) intra-epithelial cancer (Ca in
situ) was revealed. In 5 cases of cytologically diagnosed mild
neoplasia (L-SIL), the surgical treatment was prescribed, based on
the clinical specifics of the course of the disease. In all these
cases, a histological study revealed Ca in situ.
Isolation of total RNA and detection
of microRNAs and U6 snRNA by RT-PCR
Isolation of RNA from air-dried cytology
preparations was carried out as previously described (33). From the material of cytological
preparations, we succeeded in obtaining from 5 to 50 µg of total
RNA of satisfactory quality (A 260/280: 1.5–1.8) and in sufficient
concentration for quantitative measurements (120–550 ng/µl). The
microRNA and U6 snRNA expression were analyzed by stem-loop qPCR as
previously described (34). The
list of microRNAs was made based on the meta-analysis of related
literature data. The following 25 microRNA were selected:
hsa-miR-20a-5p (hereinafter referred to as miR-20a), −21-5p,
−23a-3p, −31-5p, −34a-5p, −96-5p, −99a-5p, −106b-5p, −125b-5p,
−126-3p, −145-5p, −143-3p, −146a-5p, −146b-5p, −155-5p, −181b-5p,
−191-5p, −192-5p, −196b-5p, −197-3p, −200b-3p, −203a-3p, −375,
−1246, let-7d. For each sample, the content of each marker molecule
was measured in single repeat. The results corresponding to Cq
>40 were considered negative. Sequences of all oligonucleotides
are available upon request.
Identification, genotyping and
evaluation of HR-HPV viral DNA load
Identification, genotyping and evaluation of HR-HPV
viral DNA load were performed using the ‘RealBest HPV genotype,
quantitative’ kit (AO Vector-Best, Russia) according to the
manufacturer's instructions. The kit is designed to quantify the
viral load of each of 12 HR-HPV genotypes: 16, 18, 31, 33, 35, 39,
45, 51, 52, 56, 58, 59. The viral DNA load, normalized to the
number of copies of the human β-actin gene, was calculated
independently for each genotype and the total viral DNA load was
provided as the sum of loads for different genotypes in the case of
multiple infection. In addition, all samples were tested for HPV
DNA of genotypes 26, 53, 66, 68, 73 and 82 using the kits ‘RealBest
DNA HPV 26/53/66’ and ‘RealBest DNA HPV 68/73/82’ (AO Vector-Best,
Russia). In this case, the viral DNA loads were estimated by the
ΔΔCq method (35), using the Cq
values from amplification of β-actin gene as normalizing
factor.
Content of human DNA
In the isolated sample, the number of HMBS gene
copies was evaluated using a set of reagents ‘RealBest Sample
Validation’ (AO Vector-Best, Russia) in accordance with the
manufacturer's instructions.
Statistical analysis
Data analysis was performed using SciPy library
(36) of Python programming
language. Differences between groups were assessed using the
Mann-Whitney U test. The Bonferroni correction was applied to
correct for multiple testing. All P-values of <0.05 were
considered statistically significant. The receiver operating
characteristic (ROC) curve and logistic regression analysis were
used to assess the performance of high-grade CIN detection.
Classifications were performed using Scikit-learn library (37) of Python programming language. The
linear classification algorithm was used. The stability of
reference genes was estimated by geNorm algorithm (31).
Results
Raw microRNA Cq values in samples from
different lesions
For some microRNAs, the raw Cq values obtained from
amplification curves (shown as box-whisker plots in Fig. 1) differed between the different
cytological diagnoses. No statistically significant difference was
observed for raw Cq values for any microRNA between the L-SIL group
and the NILMs and H-SILs (data not shown). However, for some
microRNAs, the differences between the groups in pairs NILM/CC,
L-SIL/CC and NILM/H-SIL were significant. Thus, the raw Cq values
for 6 microRNAs differed significantly for invasive cancers
compared to both NILM and H-SIL specimens. For two microRNAs, the
significant difference was observed for the NILM group compared to
CCs and/or H-SILs (Table I).
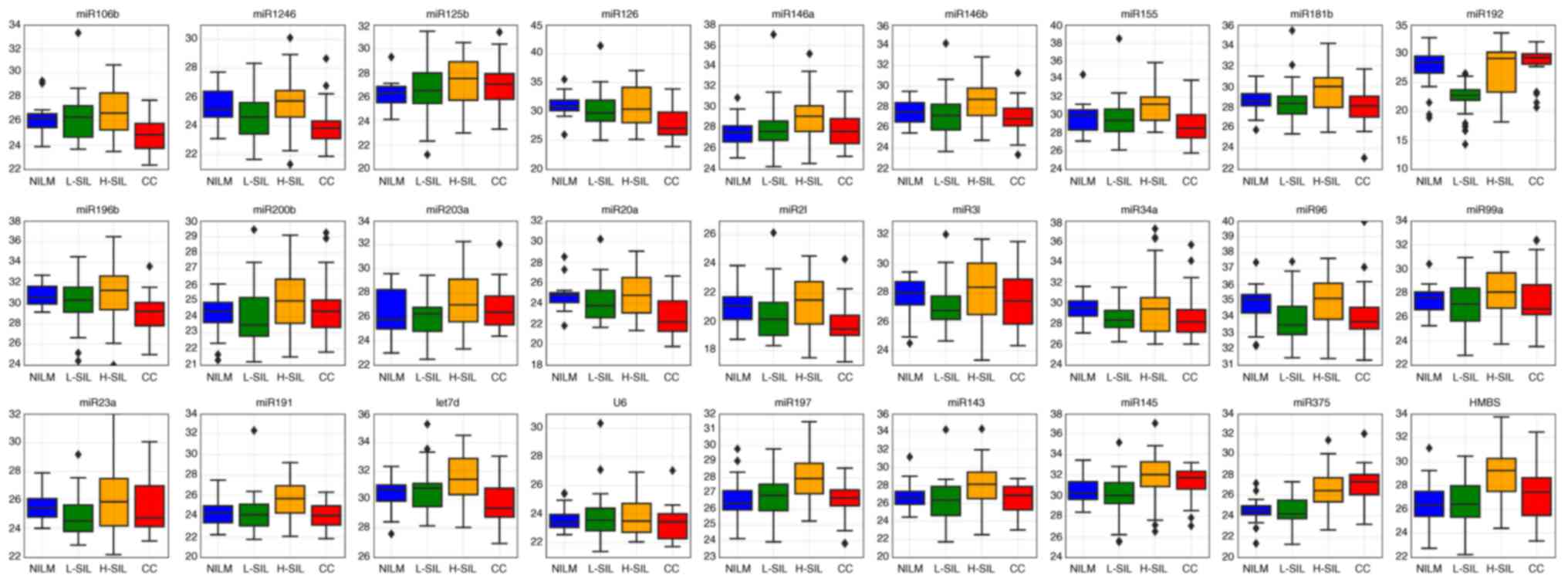 | Figure 1.Box-whisker plots for the raw Cq
values of selected microRNAs and different cytological diagnoses.
Inner lines, median values; box, upper and lower quartiles;
whiskers, non-outlier ranges; diamond, outliers. Red, cervical
cancer; orange, H-SIL; green, L-SIL; blue, NILM. NILM, negative for
intraepithelial lesions or malignancy; H-SIL, high-grade squamous
intraepithelial lesion; L-SIL, low-grade squamous intraepithelial
lesion; CC, cervical cancer. |
 | Table I.Raw Cq values for several microRNAs
that differ between cervical smears from patients with different
cytologic diagnoses. |
Table I.
Raw Cq values for several microRNAs
that differ between cervical smears from patients with different
cytologic diagnoses.
|
| Cq, mean | U test,
P-value |
|---|
|
|
|
|
|---|
|
| NILM | H-SIL | CC | NILM/H-SIL | NILM/CC | H-SIL/CC |
|---|
| miR-106b | 26.13 | 26.85 | 24.72 | 0.13962 | 0.0037b |
0.000131c |
| miR-1246 | 25.42 | 25.53 | 24.01 | 0.71362 |
0.00079c |
0.001344b |
| miR-126 | 30.99 | 30.98 | 27.79 | 0.905053 |
0.00048c |
0.000458c |
| miR-196b | 30.83 | 31.03 | 28.98 | 0.90506 |
0.00062c |
0.001209b |
| miR-20a | 24.74 | 24.97 | 22.52 | 0.783123 |
0.00013c |
0.000093c |
| miR-21 | 21.01 | 21.16 | 19.81 | 0.653008 |
0.00298b |
0.002204b |
| miR-375 | 24.55 | 26.53 | 27.12 |
0.001240b | 4×10−6c | 0.216201 |
| miR-145 | 30.63 | 31.85 | 31.68 |
0.000239c |
0.007866b | 0.296758 |
Estimation of expression stability of
selected microRNAs and U6 snRNA
The initial list of suggested normalizers included:
miR-191 [according to (26), the
expression of this microRNA was the most stable in 13 compared
tissues]; miR-23a [as a normalizer suitable for the analysis of
cervical samples, according to (38)]; U6 (traditionally used as a
stand-alone reference for normalization of microRNA expression
data); and HMBS (as a marker reflecting the input number of
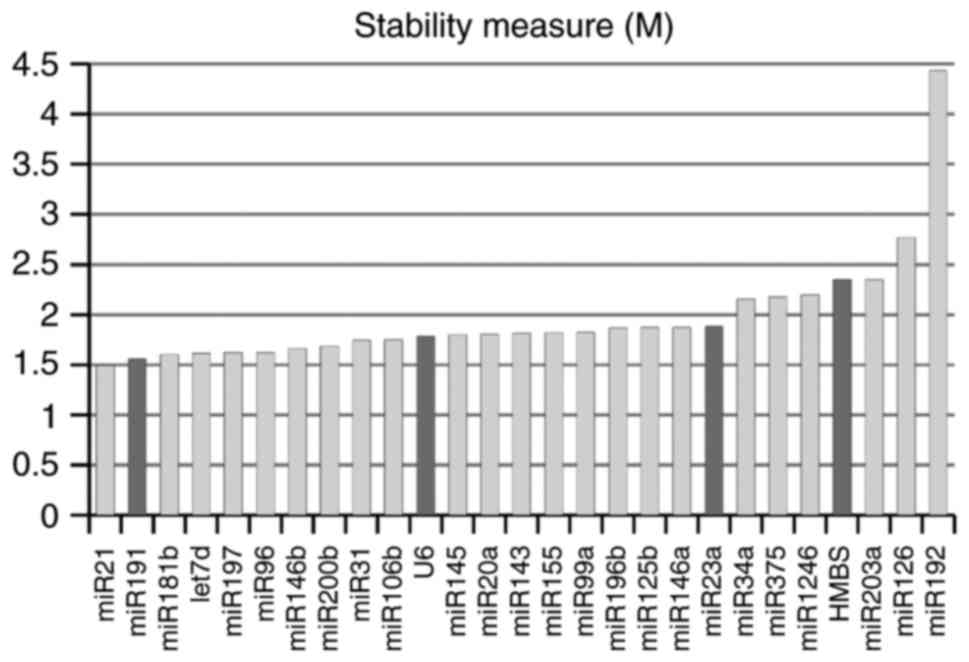
epitheliocytes). Surprisingly, according to geNorm stability
criteria, the most stably expressed was miR-21, which is widely
accepted as an oncomiR, including in CC (39–41).
In our sample, its level was increased in invasive cancers. At the
same time, suggested normalizers U6 and, particularly, miR-23a,
demonstrated relatively low stability, which was comparable to the
known oncogenic (miR-34, miR-20a) and onco-suppressor (miR-375,
miR-143) microRNAs (Fig. 2). All
microRNAs as well as U6 snRNA expectedly demonstrated maximum
stability in NILM specimens compared to other cytological
diagnoses.
Diagnostic utility of paired marker
combinations for detection of cervical lesions in cytological
preparations
According to the geNorm criterion, the most ‘stably
expressed’ among the selected RNAs was not the supposed normalizer
but the oncogene. This suggests that the choice of a normalizer
based on the evaluation of the expression stability may be an
inadequate approach in our case. We evaluated the diagnostic
utility of all possible paired combinations of the 27 selected
markers (25 microRNAs, U6 snRNA, and HMBS copy number). For each
pair of markers A and B, the ΔCqi value (ΔCq = CqA -
CqB) was obtained. The number of possible ΔCqi values in
our case was 27!/(2!25!) = 351. ΔCqi value is
dimensionless, and, on condition of ~100% PCR efficiency, is equal
to log2 of the concentration ratio of two marker molecules in the
pair. This means that the ΔCqi value does not depend on
the amount of input material as each marker in the pair serves as
‘normalizer’ for the other marker.
For each ΔCqi value, 6 different areas
under the ROC curves (ROC AUCs) were calculated. This was done to
estimate the possibility of each paired marker combination to
discriminate the specimens by the cytological diagnosis:
NILM/L-SIL, NILM/H-SIL, NILM/CC, L-SIL/H-SIL, L-SIL/CC and
H-SIL/CC. To calculate the ROC AUCs, the cross-validation strategy
was used. Each time a random subsample including 80% objects of the
original sample was generated. The model was trained on this
subsample, and the ROC AUC was evaluated based on the remaining 20%
objects of the original sample. In this case, the sample was
subdivided into the training and test subsamples in such a way that
the proportion of specimens with the cytological diagnoses was the
same in both. The procedure was repeated 100 times, that is, the
ROC AUC for each ΔCqi value was obtained as a result of
averaging over 100 calculations. The ROC AUC value >0.8 was
considered acceptable for the corresponding paired marker
combination to discriminate the samples with different
cytology.
The utility of different paired marker combinations
for detecting the lesions of different severity varied greatly.
However, the same combinations were among the best at separating
the ‘neighboring’ classes (NILM from L-SIL, L-SIL from H-SIL, H-SIL
from CC). The median and mean ΔCqi values for these
combinations increased or decreased stepwise with the severity of
the lesion.
For none of the ΔCqi values, ROC AUC
exceeded 0.8 at discriminating ‘neighboring’ groups (NILM/L-SIL,
L-SIL/H-SIL and H-SIL/CC) (data not shown). At the same time,
several paired combinations were characterized by high (>0.8)
ROC AUC values at separation of NILMs from H-SILs (n=13), NILMs
from CCs (n=61) or NILMs from both H-SILs and CCs (n=11). In
Fig. 3 a scatter graph is presented
reflecting the ratio between the ROC AUC values for discriminating
NILM specimens from CCs (x-axis) and H-SILs (y-axis) for each
paired marker combination.
For discriminating specimens with different
cytological diagnoses, the highest ROC AUC values were obtained
when the levels of two markers in a pair tended to change in the
opposite way with the increasing lesion severity. At the same time,
combination of the suggested normalizer or the ‘stably expressed’
marker (U6, HMBS, miR-23a, miR-21, miR-191 or miR-181b) with
oncogenic or onco-suppressor microRNA (that is, traditional
normalization) generally resulted in lower ROC AUC values. This is
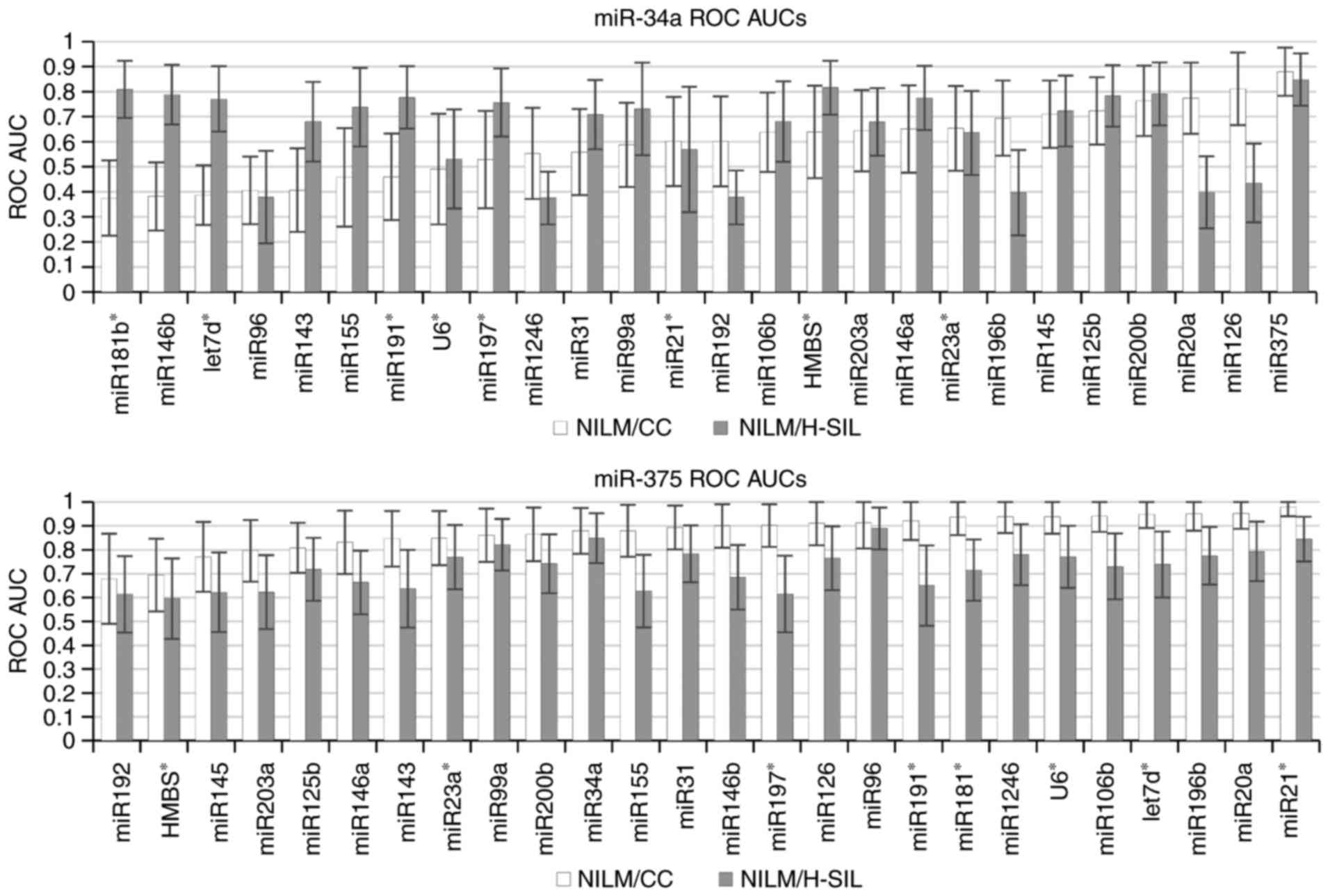
illustrated in Fig. 4, where ROC
AUCs are represented, which are calculated for paired combinations
where the first marker in the pair is oncogenic (miR-34a, above) or
onco-suppressor microRNA (miR-375, below) and the second marker,
serving as ‘normalizer’, is any of the remaining markers from the
selected list. This conclusion remains valid also for the use of
geometric mean 2, 3 and 4 for the most stably expressed microRNAs
(data not shown).
Dependence of ROC AUCs on the number
of markers in linear classifiers
The application of the linear classifier method
involving a larger number of ΔCqi values resulted in a
marked improvement in the quality of classification, compared to
the use of the best single ΔCqi values. Up to a certain
limit, an increase in the number of ΔCqi values included
in the classifier led to an increase in ROC AUC values, after which
the inclusion of additional attributes was no longer significant
(Fig. 5). As the ΔCqi
values included in the classifier changed stepwise with the
severity of lesion, the accuracy in detection of the lesion also
increased with its severity. As can be seen from Fig. 5, the reliability of H-SIL detection
in terms of ROC AUC was lower, compared to CC detection regardless
of the number of ΔCqi values included in the
classifier.
Selection of the best paired marker
combinations for the linear classifier
The results presented in Fig. 5 indicate that the reasonable number
of ΔCqi values for inclusion in the final classifier in
our case did not exceed 8. Thus, we decided to select 8
ΔCqi values for the construction of the final
classifier. The selection was based on the statistical significance
of the observed differences at separation of NILM from CC and NILM
from H-SIL. The non-parametric Mann-Whitney criterion was used.
Taking into account that the statistical criterion was applied
twice (for the separation of the classes NILM/CC and NILM/H-SIL),
the selected significance level was 0.05/2=0.025. Since we dealt
with multiple hypothesis testing (the number of validations in the
case of each classification was 351), we applied the Bonferroni
correction, which is the most conservative for multiple
verification. With this correction, statistically significant
differences at a significance level of 0.025 were observed for 53
ΔCqi values at separating NILM from CC and for 14 values
at separating NILM from H-SIL (8 ΔCqi values were
simultaneously present in these two groups).
We also accepted the required effect size (the
difference in the mean ΔCqi values in groups of
specimens from different cytology diagnoses) to be ≥1. This was
done since in the case when the ΔCqi value did not meet
this requirement (effect size <1) it is comparable to the
characteristic RT-qPCR bias. In this case, despite the statistical
significance of the ΔCqi value, its practical
significance was limited by the analytical variation of the
method.
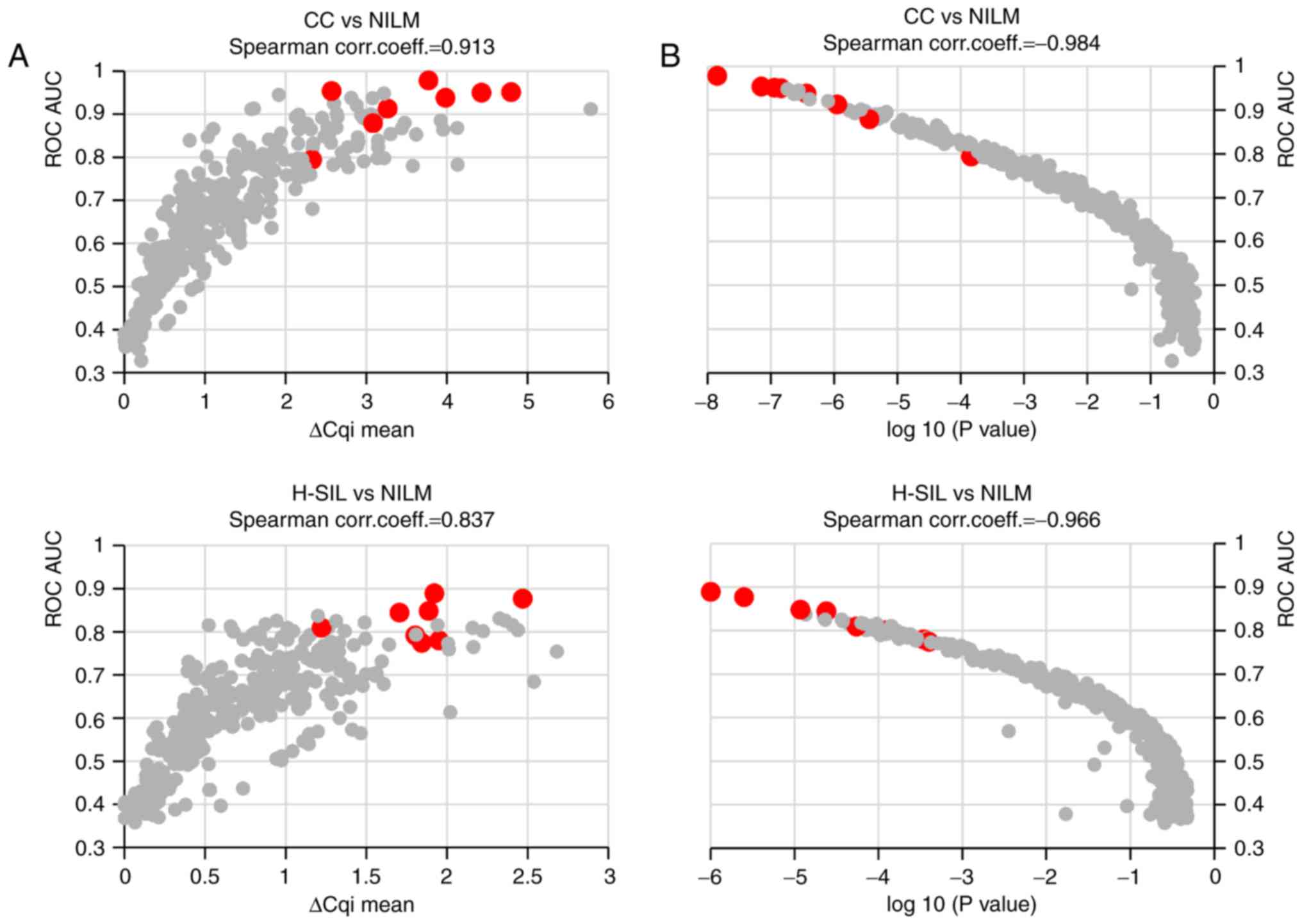
In Fig. 6, the
correlations between the ROC AUC and the effect size for each
ΔCqi value (Fig. 6A) as
well as between the ROC AUC and the achieved significance level
(ASL) of each ΔCqi value (Fig. 6B) are represented as scatter
diagrams.
It can be seen from the figure that the ROC AUC
values were correlated to a greater extent with the ASLs than with
the effect sizes. The highest ROC AUCs corresponded to
ΔCqi values with the lowest ASLs at discriminating the
specimens according to their cytological diagnoses. Thus, the ASL
value was accepted as the primary criterion for selection of the
paired combination. Among the combinations with the lowest ASLs,
the combinations with a larger effect size were further selected.
In our case, for the top 5 paired combinations with the lowest ASL
values (from 1.4x10−8 to
1.8x10−7) at discriminating NILM from CC, the
effect size ranged from 2.6 to 4.8. The top 5-paired combinations
with the lowest ASLs (from 1.0x10−6 to
2.2x10−5) at discriminating NILM from H-SIL
the effect size was smaller (ranged from 1.2 to 2.5) but,
nevertheless, exceeded 1, which suggests the feasibility of their
use for H-SIL detection by a qPCR-based method.
Thus, we selected 6 paired marker combinations with
a high statistical significance. Three of them performed best at
discrimination of NILM from CC: miR21-miR375 (AUC=0.978±0.038;
P=1.42x10−8), miR145-miR196b
(AUC=0.954±0.069; P=7.08x10−8), miR20а-miR375
(AUC=0.951±0.064; P=1.14x10−7). The other
three were selected to separate NILM from H-SIL: miR96-miR375
(AUC=0.889±0.088; P=1.00x10−6), miR1246-HMBS
DNA (AUC=0.877±0.09; P=2.51x10−6),
miR34a-miR375 (AUC=0.848±0.104; P=1.17x10−5).
ΔCqi value for each of these combinations differed
significantly between the specimens with different diagnoses even
with the most conservative Bonferroni correction, and also had
satisfactory effect size at discriminating these specimens. Two
additional combinations were included in the classifier:
miR196b-miR375 and miR375-miR1246. The calculation of these did not
require involvement of additional markers but enabled further
improvement in the classification quality.
In total, 9 markers were selected for the
classifier: 8 microRNAs (miR-20а, miR-21, miR-34a, miR-96, miR-145,
miR-196b, miR-375 and miR-1246) and cellular DNA content. The
difference between invasive cancers and NILMs was statistically
significant even for the raw Cq values for most of these microRNAs
(except for miR-34a, see Table
I).
Absence of correlation between the
microRNA level changes and HR-HPV viral load in cytological
specimens
The proportion of HR-HPV-positive samples with
different cytological diagnoses, as well as the median and mean
viral DNA loads, are provided in Table
II. In all invasive cancers, except three, HPV16 and/or HPV18
DNA was detected. In one of these, only HPV73 DNA was found, in
another - only HPV45, in the third - HPV73 and HPV45
simultaneously. Viral DNA loads in all these cases exceeded
107/108 copies of epithelial cell DNA. It
should be noted that in our sample there were no NILM specimens
with high HR-HPV DNA viral loads, although this is a fairly common
situation even in the risk age group.
 | Table II.Detection of HPV DNA in the different
groups of cytological specimens. |
Table II.
Detection of HPV DNA in the different
groups of cytological specimens.
|
| NILM | L-SIL | H-SIL | CC |
|---|
| No of samples | 40 | 29 | 62 | 43 |
| HR-HPV(+), n
(%) | 14 (35%) | 26 (89.66%) | 61 (98.39%) | 43 (100%) |
| HPV16(+) or
HPV18(+), n (%) | 4 (10%) | 20 (69.0%) | 56 (90.32%) | 40 (93.02%) |
| Median (mean) viral
loada in HR-HPV(+)
samples | 0.34 (39.9) | 436.2
(15,687.3) | 1,405.4
(6,341.8) | 500.3
(4,515.3) |
For none of the microRNAs, regardless of the
normalization method, statistically significant correlation with
the HR-HPV DNA load, genotype, or the number of genotypes in case
of multiple infections was observed. The levels of several
microRNAs notably differed between the HR-HPV-positive L-SILs and
invasive cancers, while the viral loads did not differ
significantly between these groups of specimens (Fig. 7A and B). Important to note, in our
sample the viral load differed significantly in H-SILs and CCs
compared to HR-HPV-positive NILMs (Fig.
7C and D). This may be partly due to the above-mentioned
absence of NILMs with high HR-HPV viral loads. Nevertheless, for
discrimination between subsamples of the CIN ≥2 specimens from the
rest of the sample (CIN <2) some single ΔCqi values
performed as well as the viral DNA load or even surpassed it
(Fig. 7E).
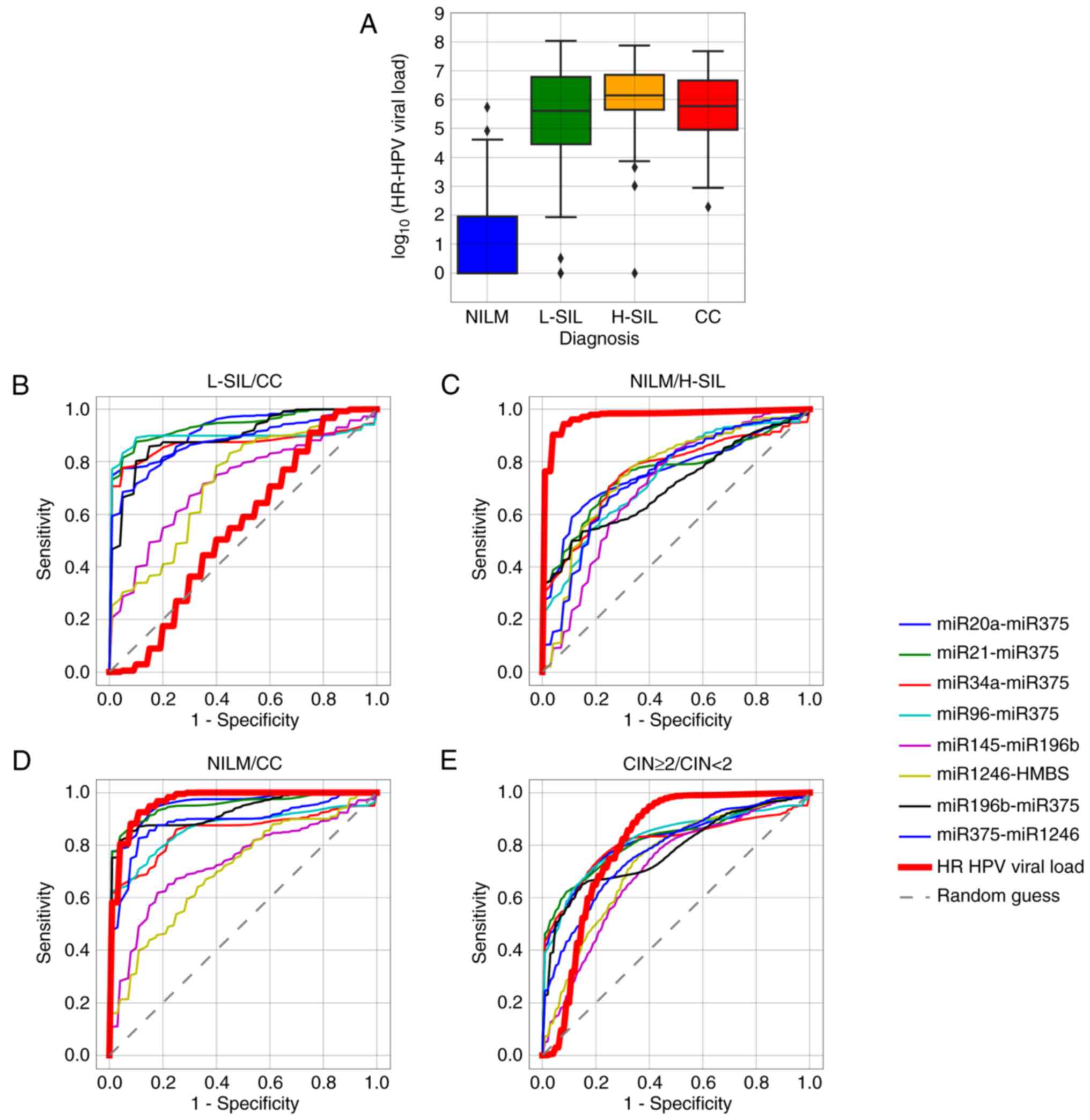 | Figure 7.(A) Box-whisker plots for the HR-HPV
DNA load in samples with different cytological diagnoses. Box,
upper and lower quartiles; inner line, median value; whisker,
non-outlier range; diamond, outliers. Red, invasive cancer; orange,
H-SIL; green, L-SIL; blue, NILM. (B-E) ROC curves corresponding to
8 ΔCqi values selected for the classifier and to the
HR-HPV DNA viral load, for discriminating different classes of
cytological diagnoses. HR-HPV, high-risk human papillomavirus;
NILM, negative for intraepithelial lesions or malignancy; H-SIL,
high-grade squamous intraepithelial lesion; L-SIL, low-grade
squamous intraepithelial lesion; CC, cervical cancer; CIN, cervical
intraepithelial neoplasia. |
Training of the linear classifier
Training of the linear classifier based on 8
ΔCqi values was performed with a sample of 171
specimens. Three CC samples with a highly degraded biomaterial (Cq
values for HMBS and most marker microRNAs close to 40) were
excluded from analysis. The decision function value (hereafter
referred to as DF value) calculated by the classifier is
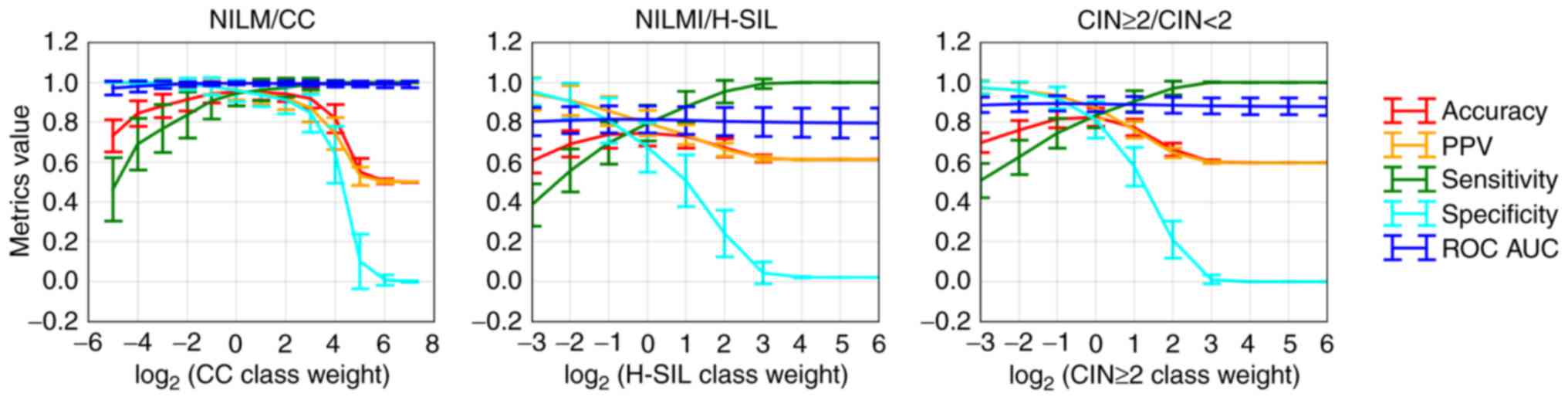
dimensionless. In Fig. 8, the
diagnostic characteristics of the trained classifier for
discrimination between different groups of cervical specimens
depending on the selected cut-offs are presented.
Fig. 9 presents
box-whisker plots for 8 ΔCqi values included in the
classifier and a box-whisker plot for DF values obtained for
discriminating high-grade lesions (CIN ≥2) from the rest of the
sample.
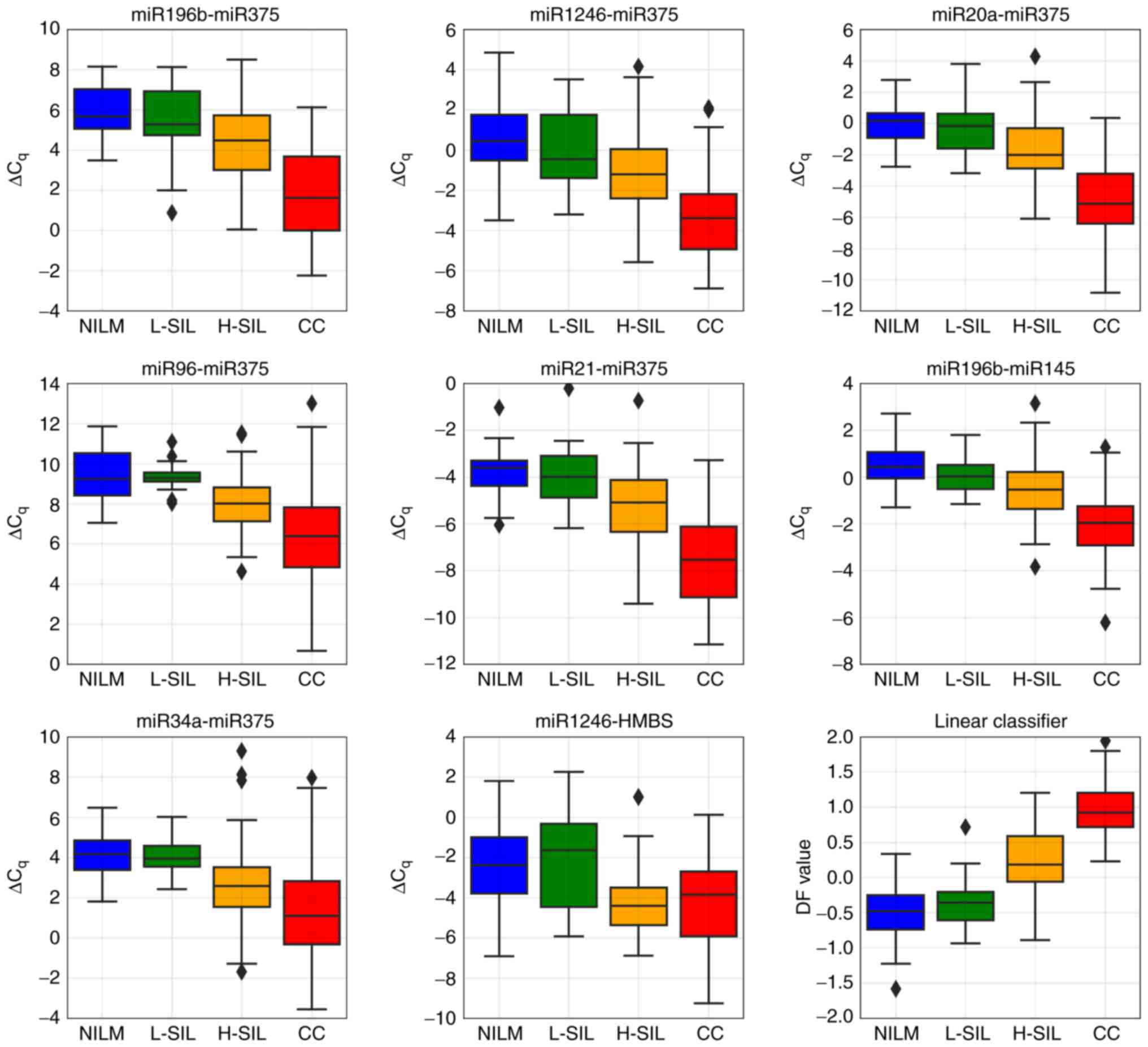 | Figure 9.Box-whisker plots for the
ΔCqi values selected for the classifier and for DF value
calculated by the classifier using these ΔCqi values.
Upper and lower quartiles are shown by the box. Inner line, median
value; whisker, non-outlier range; diamond, outlier. Red, invasive
cancer; orange, H-SIL; green, L-SIL; blue, NILM. NILM, negative for
intraepithelial lesions or malignancy; L-SIL, low-grade squamous
intraepithelial lesion; H-SIL, high-grade squamous intraepithelial
lesion; CC, cervical cancer. |
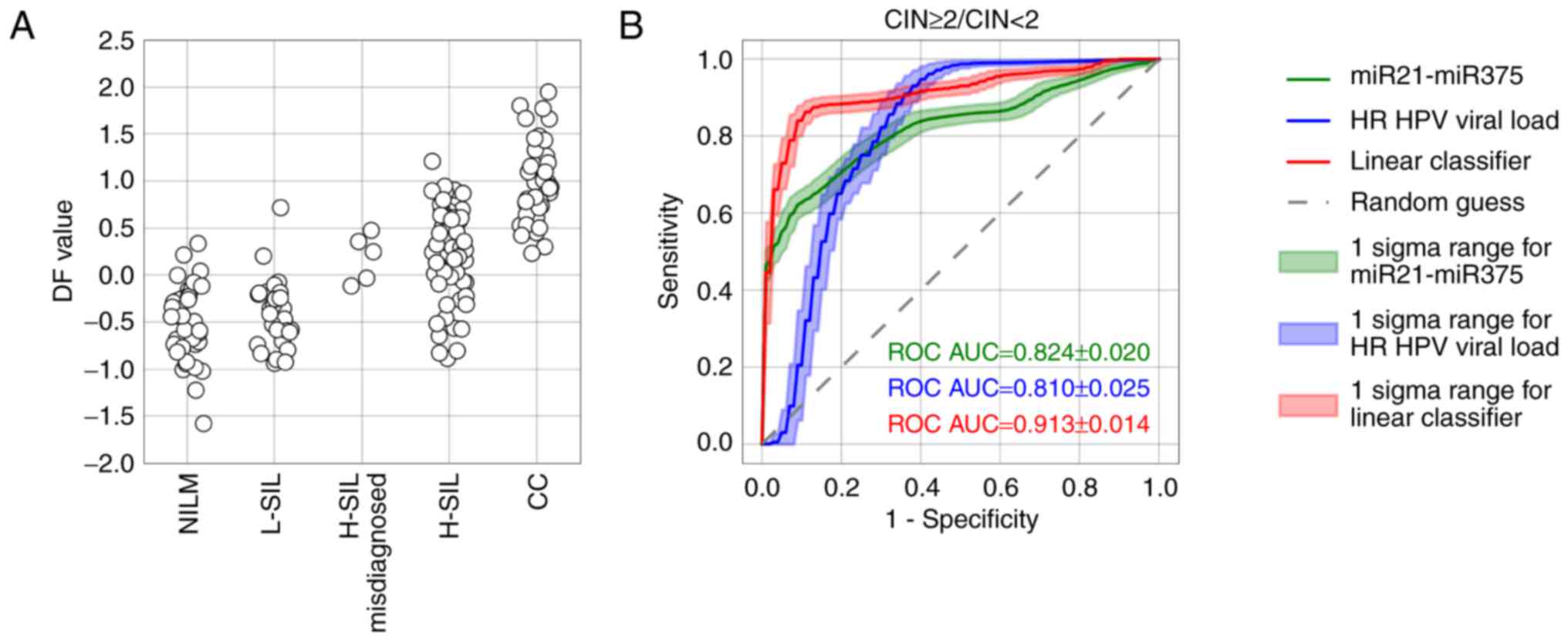
As the marker of high-grade lesions (CIN ≥2), the DF
value performed better than the viral load or any ΔCqi
value (ROC AUC=0.913, diagnostic sensitivity=83.4%, diagnostic
specificity=81.2% at maximum Youden index) (Fig. 10B). Nevertheless, the DF value
ranges sufficiently overlapped between groups of samples with
different pathomorphological diagnoses (Fig. 10A). This overlap may result from
the combined effect of the analytical biases, biological variation
in the marker RNA levels, the cellular heterogeneity of cervical
specimens, and the cytology misclassification.
Estimation of method
reproducibility
The accuracy of DF value measuring results obviously
depends on the analytical variation of RT-PCR. To evaluate the
possible contribution of RT-PCR bias, we re-analyzed extracted
nucleic acid preparations from two cytological specimens (from NILM
and CC patients), in which each marker was analyzed 4-fold and each
Cq value obtained for each marker in the sample was used in
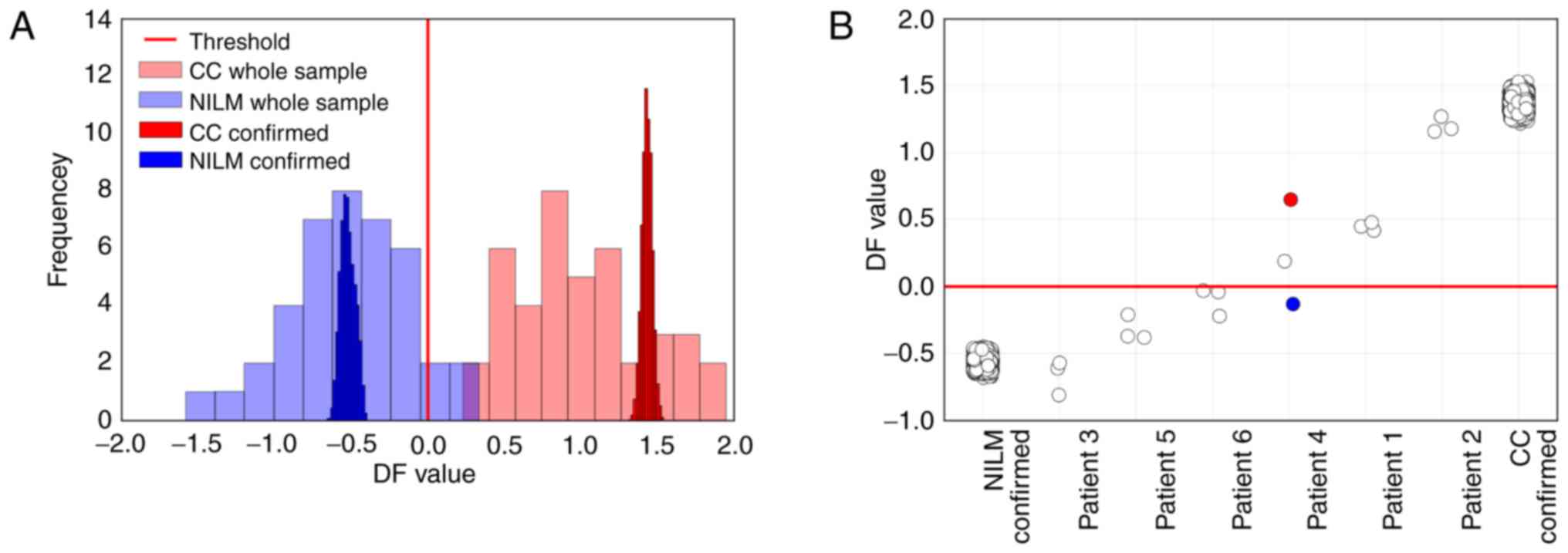
repeated calculations of DF value for this sample. In Fig. 11A, the histograms of DF values for
each specimen (calculated for 10,000 randomly selected combinations
of the obtained Cq values from the possible
4.3x109 combinations) and the histograms of
DF values for the NILMs and CCs are presented. The histograms of DF
values for specimens are several times narrower than the histograms
of DF values for the NILM and CC classes, which means that the
contribution of RT-PCR bias into the accuracy of the DF value
calculation was not high.
For 6 patients with different cytological diagnoses,
3 PAP slides prepared from a single smear were tested. In Fig. 11B, the DF values calculated for the
nucleic acid preparations extracted from each slide are provided
compared to 10,000 DF values for NILM and CC samples calculated as
described above. For 5 out of 6 patients, the variation between the
slides was comparable to the RT-PCR bias. Nevertheless, for patient
no. 4, different classification results for the different slides
were obtained. We suggest that the observed variation was
determined to a greater extent by the procedure of preparing the
slide (which affects the amount and integrity of the material
analyzed) than by the nucleic acid extraction procedure. Thus, in
the case of patient no. 4, the maximum DF value (marked red in
Fig. 11B) corresponded to the
slide for which the HMBS DNA Cq value was ~39 and most Cq values
for microRNA markers were beyond the linear range of qPCR. The
minimum DF value (marked blue) corresponded to the slide from the
same patient, where HMBS DNA Cq was ~33 (~100-fold greater
concentration of input cellular DNA) and Cq values for microRNA
markers were within the qPCR linear range, which suggest that the
classification results for this slide were more reliable.
When comparing the ROC AUCs for DF values calculated
for subsamples of specimens with very different nucleic acid
concentrations (the difference between the max and min Cq values
for HMBS=11.5, the difference between the max and min Cq values for
the most stably expressed miR-21=9.5), we observed no difference.
In addition, we re-classified 3 samples after the 4-fold dilution
of nucleic acid preparation. The differences in the obtained DF
values were within the limits of the variation evaluated in the
previous experiment and in no case led to reclassification of the
sample compared to non-diluted preparation (data not shown). Taken
together, these facts support the lack of a significant
contribution made by PCR efficiency to the classification
reliability.
Discussion
Recent research demonstrates the feasibility of
using microRNAs as biomarkers in cervical cancer (CC) screening and
follow-up. However, the reliable detection of biologically relevant
changes in microRNA levels may be compromised by the variability
introduced by the methodology of analysis and the cellular
heterogeneity of clinical specimens. It is this problem that may
account for the fact that none of the numerous but diverse design
and methodology methods has achieved acceptable diagnostic
characteristics when using microRNA profiling for the diagnosis of
cervical neoplasia.
As mentioned above, the lists of the microRNA
markers, deregulation of which accompanies cervical lesions of
different degrees, as well as the direction of their concentration
changes, often differ between different studies. This can certainly
be attributed to the peculiarities of the microRNA isolation and
quantitation techniques used. Each technology of microRNA profiling
(microarrays, NanoString counting, RNA-seq, TaqMan low density
array) has its own sources of bias. Even the widely used techniques
for microRNA analysis (Exiqon and TaqMan), based on similar RT-qPCR
techniques, demonstrate serious discrepancies in the efficiency of
detecting various microRNAs (42–46).
To search for the most suitable markers for the further design of
microRNA diagnostic tools, researchers typically analyze a limited
sample of patients by microarrays or RNAseq, which allows one to
choose from the great number of microRNAs; then candidate microRNAs
are validated by real-time PCR. This approach may result in
neglecting some relevant microRNAs due to lower efficiency of their
detection by chip-based methods or RNA-seq biases.
The normalization method is another apparent factor
affecting the reliability of microRNA quantification. In cases
where the disease-associated microRNA level changes are relatively
small (as in high-grade CINs), the choice of the normalization
strategy is particularly important, since in such cases the
biologically relevant microRNA expression changes may be comparable
to the biases introduced by the method. Some authors have proposed
individual normalizers for microRNA level measurements in cervical
epithelium by real-time qPCR, e.g., U6 snRNA (18,21,23,47),
miR-23 (22,38) and miR-92a (48). The use of such normalizers by our
research group did not lead to favorable results for the
classification of cervical lesions in cytological preparations.
Moreover, classification results were strongly
normalizer-dependent. With some normalizers, ‘wave-like’ changes in
the content of particular microRNAs with the increase in lesion
severity from NILM to CC were observed (data not shown). Such
changes were not consistent with the known biological functions of
these microRNAs and can be considered artifacts of the analysis. On
the contrary, the use of the classifier based on paired microRNA
combinations selected as described above led to better
classification results than any single normalizer or geometrical
mean of the 3–5 most ‘stably expressed’ microRNAs. This can be
attributed to the following. i) The amplitudes of disease-related
microRNA level changes may be comparable to the analytical
variation of the method used, which hinders the reliable
registration of such changes. The paired marker approach (where the
concentrations of two markers in the pair, the levels of which
change in opposite directions at neoplastic transformation, are
reciprocally normalized) helps to better distinguish biologically
relevant change in microRNA profile from the ‘noise’ (the
fluctuation of measurement results due to analytical variation).
This approach, compared to traditional normalization, better
compensates for the biases caused by the analytical variation of
the method and/or the cellular heterogeneity of the specimen. The
lower the level of physiological changes in the content of the
marker is, the greater the contribution of this compensation.
Similarly to the traditional normalization, this approach also
compensates for the variation associated with the amount and
degradation level of input biomaterial. ii) The use of normalizers
selected by formal criteria can generate a system error. Thus, in
the present study, the most ‘stably expressed’ microRNA was miR-21.
For this microRNA, known as oncogenic in multiple cancers including
CC, one could expect the increase in concentration accompanying
cervical neoplastic transformation. In our case, this was most
likely, as even without any normalization the decline in raw miR-21
Cq values in H-SIL and CC compared to NILM was statistically
significant (Table I). The use of a
normalizer whose concentration itself de facto increases
with the severity of the lesion, will inevitably lower or even mask
the statistically significant concentration changes of the relevant
microRNAs, the levels of which also tend to decline in precancerous
lesions and cancer. iii) The cell heterogeneity of the analyzed
preparation may result in ‘watering down’ the observed microRNA
level changes. The specimen may contain cells corresponding to the
different degree of neoplasia in different proportions. In
addition, the cytological preparation may contain an admixture of
cells not related to the lesion. Thus, the resulting microRNA
profile may appear ‘intermediate’, which will complicate the
classification. In the analysis of cytological specimens, it is
particularly difficult to take into account and compensate for this
source of biases. In addition, it cannot be ruled out that
neoplasms considered as belonging to a single class based on
pathomorphological classification can in fact represent different
subclasses for which microRNA expression profiles differ.
In the present study, the best individual microRNA
marker of cervical neoplasia was miR-375, which was present in 6 of
8-paired combinations included into the classifier. This was the
only microRNA, for which the median ΔCqi values changed
monotonically in the range NILM/L-SIL/H-SIL/CC regardless of the
second marker (‘normalizer’) in the paired combination. The mean
ΔCqi values calculated for the combinations including
this microRNA also demonstrated the greatest difference between the
lesions of different severity. miR-375 is a known tumor suppressor
involved in the development of CC (21,49).
The other microRNAs included in the classifier displayed a tendency
for increase or decrease with the degree of lesion, which, however,
was significantly dependent on the choice of the normalizer.
Thus, we developed a technique for detecting
cervical precancerous lesions and cancer in cytological specimens
using a microRNA-based classifier. The method demonstrated
acceptable diagnostic characteristics and analytical
reproducibility. The analysis can be performed with the same
nucleic acid preparation as used for HPV testing, genotyping, and
the measurement of the HR-HPV viral DNA load. It is now generally
accepted that the detection of HR-HPV DNA has a high NPV for
precancerous cervical lesions while its PPV is low due to the high
frequency of transient HPV carriage without cellular
transformation. The differences in microRNA expression more likely
reflect cellular events related to transformation and, therefore,
may provide a higher PPV if used as a diagnostic marker.
Nevertheless, the NPV of such an analysis can also remain
significant. In our case, in sample no. 129 with a cytological
diagnosis of H-SIL and a histologically confirmed CIN3, in which
HPV DNA was not detected, the DF value corresponded to high-grade
CIN. Our results support the feasibility of using small-scale
microRNA profiling for detection of high-grade cervical
intraepithelial neoplasia. Taking into account the very high NPV of
HPV-testing in cervical screening, the microRNA profiling and the
HR-HPV DNA testing may serve as complementary tools in the
molecular testing for cervical lesions.
Some limitations of our research should be
emphasized. First, the choice of microRNA markers was made from a
limited set, which in itself could fail to be optimal. Secondly,
the sample was relatively small and enriched with invasive cancers,
which could have led to overestimating the diagnostic
characteristics of the classifier. Thirdly, the study was a
single-setting. A separate issue in our case is the interpretation
of the results of the analysis of samples cytologically
characterized as NILM. Since in our case this diagnosis was not
verified, we cannot exclude misclassification of some preparations
from this group. Further clinical validation of the developed
microRNA-based classifier and its use in multicenter and follow-up
studies will additionally substantiate the conclusion concerning
the prospects of its clinical use.
Glossary
Abbreviations
Abbreviations:
|
CIN
|
cervical intraepithelial neoplasia
|
|
HR-HPV
|
high-risk human papillomavirus
|
|
DF value
|
decision funtion value
|
|
NILM
|
negative for intraepithelial lesion or
malignancy
|
|
L-SIL
|
low-grade squamous intraepithelial
lesion
|
|
H-SIL
|
high-grade squamous intraepithelial
lesion
|
|
CC
|
cervical cancer
|
|
PPV
|
positive predictive value
|
|
RT-qPCR
|
quantitative reverse-transcription
real-time PCR
|
|
Cq
|
quantification cycle
|
References
|
1
|
Kitchener HC, Almonte M, Thomson C,
Wheeler P, Sargent A, Stoykova B, Gilham C, Baysson H, Roberts C,
Dowie R, et al: HPV testing in combination with liquid-based
cytology in primary cervical screening (ARTISTIC): A randomised
controlled trial. Lancet Oncol. 10:672–682. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Naucler P, Ryd W, Törnberg S, Strand A,
Wadell G, Elfgren K, Rådberg T, Strander B, Johansson B, Forslund
O, et al: Human papillomavirus and papanicolaou tests to screen for
cervical cancer. N Engl J Med. 357:1589–1597. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Whitlock EP, Vesco KK, Eder M, Lin JS,
Senger CA and Burda BU: Liquid-based cytology and human
papillomavirus testing to screen for cervical cancer: A systematic
review for the U.S. preventive services task force. Ann Intern Med.
155:687–697, W214-W215. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Naucler P, Ryd W, Törnberg S, Strand A,
Wadell G, Elfgren K, Rådberg T, Strander B, Forslund O, Hansson BG,
et al: Efficacy of HPV DNA testing with cytology triage and/or
repeat HPV DNA testing in primary cervical cancer screening. Natl
Cancer Inst. 101:88–99. 2009. View Article : Google Scholar
|
|
5
|
Benoy IH, Vanden Broeck D, Ruymbeke MJ,
Sahebali S, Arbyn M, Bogers JJ, Temmerman M and Depuydt CE: Prior
knowledge of HPV status improves detection of CIN2+ by
cytology screening. Am J Obstet Gynecol. 205:569, e1–e7. 2011.
View Article : Google Scholar
|
|
6
|
Faridi R, Zahra A, Khan K and Idrees M:
Oncogenic potential of human papillomavirus (HPV) and its relation
with cervical cancer. Virol J. 8:2692011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Franceschi S, Denny L, Irwin KL, Jeronimo
J, Lopalco PL, Monsonego J, Peto J, Ronco G, Sasieni P and Wheeler
CM: EUROGIN 2010 roadmap on cervical cancer prevention. Int J
Cancer. 128:2765–2774. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Arbyn M, Sasieni P, Meijer CJ, Clavel C,
Koliopoulos G and Dillner J: Chapter 9: Clinical applications of
HPV testing: A summary of meta-analyses. Vaccine. 24 Suppl
3:S3/78–89. 2006. View Article : Google Scholar
|
|
9
|
de Sanjosé S, Diaz M, Castellsagué X,
Clifford G, Bruni L, Muñoz N and Bosch FX: Worldwide prevalence and
genotype distribution of cervical human papillomavirus DNA in women
with normal cytology: A meta-analysis. Lancet Infect Dis.
7:453–459. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Steenbergen RD, Snijders PJ, Heideman DA
and Meijer CJ: Clinical implications of (epi)genetic changes in
HPV-induced cervical precancerous lesions. Nat Rev Cancer.
14:395–405. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Martinez I, Gardiner AS, Board KF, Monzon
FA, Edwards RP and Khan SA: Human papillomavirus type 16 reduces
the expression of microRNA-218 in cervical carcinoma cells.
Oncogene. 27:2575–2582. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Melar-New M and Laimins LA: Human
papillomaviruses modulate expression of microRNA 203 upon
epithelial differentiation to control levels of p63 proteins. J
Virol. 84:5212–5221. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Greco D, Kivi N, Qian K, Leivonen SK,
Auvinen P and Auvinen E: Human papillomavirus 16 E5 modulates the
expression of host microRNAs. PLoS One. 6:e216462011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zheng ZM and Wang X: Regulation of
cellular miRNA expression by human papillomaviruses. Biochim
Biophys Acta. 1809:668–677. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lajer CB, Garnæs E, Friis-Hansen L,
Norrild B, Therkildsen MH, Glud M, Rossing M, Lajer H, Svane D,
Skotte L, et al: The role of miRNAs in human papilloma virus
(HPV)-associated cancers: Bridging between HPV-related head and
neck cancer and cervical cancer. Br J Cancer. 117:e22017.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Gómez-Gómez Y, Organista-Nava J and
Gariglio P: Deregulation of the miRNAs expression in cervical
cancer: Human papillomavirus implications. Biomed Res Int.
2013:4070522013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Pedroza-Torres A, López-Urrutia E,
Garcia-Castillo V, Jacobo-Herrera N, Herrera LA, Peralta-Zaragoza
O, López-Camarillo C, De Leon DC, Fernández-Retana J, Cerna-Cortés
JF, et al: MicroRNAs in cervical cancer: Evidences for a miRNA
profile deregulated by HPV and its impact on radio-resistance.
Molecules. 19:6263–6281. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wang X, Wang HK, Li Y, Hafner M, Banerjee
NS, Tang S, Briskin D, Meyers C, Chow LT, Xie X, et al: microRNAs
are biomarkers of oncogenic human papillomavirus infections. Proc
Natl Acad Sci USA. 111:pp. 4262–4267. 2014; View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Gocze K, Gombos K, Kovacs K, Juhasz K,
Gocze P and Kiss I: MicroRNA expressions in HPV-induced cervical
dysplasia and cancer. Anticancer Res. 35:523–530. 2015.PubMed/NCBI
|
|
20
|
He Y, Lin J, Ding Y, Liu G, Luo Y, Huang
M, Xu C, Kim TK, Etheridge A, Lin M, et al: A systematic study on
dysregulated microRNAs in cervical cancer development. Int J
Cancer. 138:1312–1327. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Tian Q, Li Y, Wang F, Li Y, Xu J, Shen Y,
Ye F, Wang X, Cheng X, Chen Y, et al: MicroRNA detection in
cervical exfoliated cells as a triage for human
papillomavirus-positive women. J Natl Cancer Inst.
106:pii:dju2412014. View Article : Google Scholar
|
|
22
|
Ribeiro J, Marinho-Dias J, Monteiro P,
Loureiro J, Baldaque I, Medeiros R and Sousa H: miR-34a and
miR-125b expression in HPV infection and cervical cancer
development. Biomed Res Int. 2015:3045842015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Pereira PM, Marques JP, Soares AR, Carreto
L and Santos MA: MicroRNA expression variability in human cervical
tissues. PLoS One. 5:e117802010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Galamb Á, Benczik M, Zinner B, Vígh E,
Baghy K, Jeney C, Kiss A, Lendvai G and Sobel G: Dysregulation of
microRNA expression in human cervical preneoplastic and neoplastic
lesions. Pathol Oncol Res. 21:503–508. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ogechukwu OJ: Discordant reports of miRNA
expression in cervical cancer: An upshot of overlapping factors.
Res Cancer Tumor. 4:15–23. 2015.
|
|
26
|
Peltier HJ and Latham GJ: Normalization of
microRNA expression levels in quantitative RT-PCR assays:
Identification of suitable reference RNA targets in normal and
cancerous human solid tissues. RNA. 14:844–852. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Mestdagh P, Van Vlierberghe P, De Weer A,
Muth D, Westermann F, Speleman F and Vandesompele J: A novel and
universal method for microRNA RT-qPCR data normalization. Genome
Biology. 10:R642009. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Pradervand S, Weber J, Thomas J, Bueno M,
Wirapati P, Lefort K, Dotto GP and Harshman K: Impact of
normalization on miRNA microarray expression profiling. RNA.
15:493–501. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kirschner MB, van Zandwijk N and Reid G:
Cell-free microRNAs: Potential biomarkers in need of standardized
reporting. Front Genet. 4:562013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ludwig N, Leidinger P, Becker K, Backes C,
Fehlmann T, Pallasch C, Rheinheimer S, Meder B, Stähler C, Meese E
and Keller A: Distribution of miRNA expression across human
tissues. Nucleic Acids Res. 44:3865–3877. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Vandesompele J, De Preter K, Pattyn F,
Poppe B, Van Roy N, De Paepe A and Speleman F: Accurate
normalization of real-time quantitative RT-PCR data by geometric
averaging of multiple internal control genes. Genome Biol.
3:RESEARCH00342002. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Solomon D, Davey D, Kurman R, Moriarty A,
O'Connor D, Prey M, Raab S, Sherman M, Wilbur D, Wright T Jr, et
al: The 2001 Bethesda System: Terminology for reporting results of
cervical cytology. JAMA. 287:2114–2119. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kolesnikov NN, Titov SE, Veryaskina YA,
Vladimirova AV, Samsonov RB, Artem'eva AS, Novik VI, Bershteyn LM,
Zhimulev IF, Malek AV, et al: Improvement of accuracy and
diagnostic significance of breast tumor fine-needle aspiration
biopsy by miRNA analysis of material isolated from cytological
smears. Uspehi Molekulârnoj Onkologii. 3:44–52. 2016. View Article : Google Scholar
|
|
34
|
Titov SE, Demenkov PS, Ivanov MK,
Malakhina ES, Poloz TL, Tsivlikova EV, Ganzha MS, Shevchenko SP,
Gulyaeva LF and Kolesnikov NN: Selection and validation of miRNAs
as normalizers for profiling expression of microRNAs isolated from
thyroid fine needle aspiration smears. Oncology Rep. 36:2501–2510.
2016. View Article : Google Scholar
|
|
35
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2ΔΔCT method. Methods. 25:402–408. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Jones E, Oliphant E and Peterson P: SciPy:
Open source scientific tools for python. 2001, http://www.scipy.org/July 24–2017 View Article : Google Scholar
|
|
37
|
Pedregosa F, Varoquaux G, Gramfort A,
Michel V, Thirion B, Grisel O, Blondel M, Prettenhofer P, Weiss R,
Dubourg V, et al: Scikit-learn: Machine learning in python. J Mach
Learn Res. 12:2825–2830. 2011.
|
|
38
|
Shen Y, Li Y, Ye F, Wang F, Wan X, Lu W
and Xie X: Identification of miR-23a as a novel microRNA normalizer
for relative quantification in human uterine cervical tissues. Exp
Mol Med. 43:358–366. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Deftereos G, Corrie SR, Feng Q, Morihara
J, Stern J, Hawes SE and Kiviat NB: Expression of mir-21 and
mir-143 in cervical specimens ranging from histologically normal
through to invasive cervical cancer. PLoS One. 6:e284232011.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Bumrungthai S, Ekalaksananan T, Evans MF,
Chopjitt P, Tangsiriwatthana T, Patarapadungkit N, Kleebkaow P,
Luanratanakorn S, Kongyingyoes B, Worawichawong S and Pientong C:
Up-regulation of miR-21 is associated with cervicitis and human
papillomavirus infection in cervical tissues. PLoS One.
10:e01271092015. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Han Y, Xu GX, Lu H, Yu DH, Ren Y, Wang L,
Huang XH, Hou WJ, Wei ZH, Chen YP, et al: Dysregulation of miRNA-21
and their potential as biomarkers for the diagnosis of cervical
cancer. Int J Clin Exp Pathol. 8:7131–7139. 2015.PubMed/NCBI
|
|
42
|
Git A, Dvinge H, Salmon-Divon M, Osborne
M, Kutter C, Hadfield J, Bertone P and Caldas C: Systematic
comparison of microarray profiling, real-time PCR, and
next-generation sequencing technologies for measuring differential
microRNA expression. RNA. 16:991–1006. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Jensen SG, Lamy P, Rasmussen MH, Ostenfeld
MS, Dyrskjøt L, Orntoft TF and Andersen CL: Evaluation of two
commercial global miRNA expression profiling platforms for
detection of less abundant miRNAs. BMC Genomics. 12:4352011.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Kolbert CP, Feddersen RM, Rakhshan F,
Grill DE, Simon G, Middha S, Jang JS, Simon V, Schultz DA, Zschunke
M, et al: Multi-platform analysis of microRNA expression
measurements in RNA from fresh frozen and FFPE tissues. PLoS One.
8:e525172013. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Mestdagh P, Hartmann N, Baeriswyl L,
Andreasen D, Bernard N, Chen C, Cheo D, D'Andrade P, DeMayo M,
Dennis L, et al: Evaluation of quantitative miRNA expression
platforms in the microRNA quality control (miRQC) study. Nat
Methods. 11:809–815. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Farr RJ, Januszewski AS, Joglekar MV,
Liang H, McAulley AK, Hewitt AW, Thomas HE, Loudovaris T, Kay TW,
Jenkins A and Hardikar AA: A comparative analysis of
high-throughput platforms for validation of a circulating microRNA
signature in diabetic retinopathy. Sci Rep. 5:103752015. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Ben W, Yang Y, Yuan J, Sun J, Huang M,
Zhang D and Zheng J: Human papillomavirus 16 E6 modulates the
expression of host microRNAs in cervical cancer. Taiwan J Obstet
Gynecol. 54:364–370. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Campos-Viguri GE, Jiménez-Wences H,
Peralta-Zaragoza O, Torres-Altamirano G, Soto-Flores DG,
Hernández-Sotelo D, Alarcón-Romero Ldel C, Jiménez-López MA,
Illades-Aguiar B and Fernandez-Tilapa G: miR-23b as a potential
tumor suppressor and its regulation by DNA methylation in cervical
cancer. Infect Agent Cancer. 10:422015. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Wang F, Li Y, Zhou J, Xu J, Peng C, Ye F,
Shen Y, Lu W, Wan X and Xie X: miR-375 is down-regulated in
squamous cervical cancer and inhibits cell migration and invasion
via targeting transcription factor SP1. Am J Pathol. 179:2580–2588.
2011. View Article : Google Scholar : PubMed/NCBI
|