Gastrointestinal cancer is among the most prevalent
and predominant malignancies globally, consisting mainly of gastric
cancer (GC) and colorectal cancer (CRC), accounting for 26% of the
global cancer incidence and 35% of all cancer-associated
mortalities in 2018 (1). At least
~1/3 (30–35%) of cases of gastrointestinal cancer are associated
with imbalanced diet and lack of exercise; ~1/4 (20–30%) are
associated with tobacco products; and ~1/5 (15–20%) are triggered
by infectious agents (2).
Currently, fiberoptic gastroscopy and biopsy are the
major invasive techniques used to diagnose gastrointestinal cancer
(3). Not only can the diagnostic
methods of gastrointestinal cancer be harmful, the diagnosis period
of this disease is also mostly diagnosed at an advanced stage,
which seriously affects the survival and prognosis of patients
(3,4). The treatment of gastrointestinal
cancer, which mostly involves the removal of the primary cancer
tumor or the administration of medications such as cetuximab,
paclitaxel and sorafenib, has likewise seen slow development in
research (5). However, patients
with surgically treated gastrointestinal cancer have some
complications and a high recurrence rate, and patients treated with
drugs will also develop a certain degree of drug resistance
(5,6). Therefore, research into safer and more
effective diagnostic and treatment approaches is urgently needed.
Recent studies have found that targeted therapies are effective in
treating certain types of cancers (7,8). In
order to increase the survival rate of patients with
gastrointestinal cancer, it is crucial to research innovative
targeted molecules related to this illness.
Immune cells, stromal cells, extracellular matrix,
substances released by cells (such growth factors and extracellular
vesicles), lymphatic arteries, vascular networks and extracellular
vesicles frequently exist in the tumor microenvironment (9). These elements interact in the tumor
microenvironment to control the emergence, growth and metastasis of
malignancies (9,10). Exosomes are small membranous
vesicles that are produced when internal multivesicular bodies fuse
with the cell membrane and are deposited in the extracellular
matrix (11,12). A growing number of studies in recent
years have revealed that exosomes can not only receive a wide range
of neurotransmission and endocrine signaling, but also serve as a
signaling platform for intercellular communication and coordinate
various autocrine and paracrine functions, thereby influencing the
microenvironment and development of tumors (13,14).
In addition to proteins, mRNAs, microRNAs
(miRNAs/miRs) and circular RNAs (circRNAs) are also important
components of exosomes (15).
CircRNAs are abnormally expressed in cancer cells, as evidenced by
the growing body of research, which has an impact on processes
including cancer growth and treatment resistance (16,17).
CircRNAs are stabler compared with other RNAs and vary from
conventional linear RNAs in that they have a covalent closed-loop
structure (18). Exosomal circRNAs
have been shown to be useful as biomarkers for gastrointestinal
cancer diagnosis, prognosis and treatment targets in a number of
investigations (19,20). For example, Xing et al
confirm that hsa_circ_0004831 is highly expressed in blood
extracellular vesicles in patients with CRC. The results of this
bioinformatics analysis identified miRNAs involved in the
regulation of this circRNA, such as hsa-miR-4326, which may be
related to the development of CRC (21).
The major traits and purposes of exosomes and
circRNAs are reviewed in the present study, along with the
biological significance and putative molecular pathways of exosomal
circRNAs in the development of gastrointestinal cancer.
Additionally, the present study provides an overview of the
prospects and problems in this area and highlights novel
developments in exosomal circRNAs as potential diagnostic
biomarkers and therapeutic targets for gastrointestinal cancer.
Exosomes are a type of extracellular vesicle (EVs),
while ectosomes are the other type (22). Compared with ectosomes, the
biological origin and particle diameter of exosomes are different.
After forming an outward bud, ectosomes sever the surface of the
plasma membrane, and their diameters range from 50 nm to 1 mm
(23). However, exosomes are
nanoscale vesicles originating from endosomes with diameters of
40–160 nm (average, ~100 nm) (23).
All cells secrete exosomes in normal and pathological conditions,
which maintain cell homeostasis and mediate intercellular
communication (24). Exosomes
contain numerous cellular components, including nucleic acids,
amino acids, lipids, glycans, metabolites, cytoplasm and cell
surface proteins (24). In exosomes
from distinct cell types, 9,769 proteins, 3,408 mRNAs, 2,838 miRNAs
and 1,116 lipids have so far been discovered (25). Exosomes have been the subject of an
increasing number of research in recent years due to their
potential use as non-invasive liquid biopsy instruments for the
detection and treatment of a variety of disorders as well as their
potential for usage in prescription drugs (23,26).
Exosomes were discovered for the first time in sheep
reticulocytes by Pan and Johnstone in 1983 (27). Initially considered as cellular
debris, exosomes were later found to have independent structures
(27). The occurrence of exosomes
starts from the vesicles on the cell membrane, which turn inward to
form exosomes, and the cytoplasmic membrane invaginates to enclose
some specific extracellular biological macromolecules and cell
membrane proteins to form intracellular multivesicular bodies
(MVBs) (28). After the formation
of MVBs, cells may fuse with autophagosomes or lysosomes, during
which the previously encapsulated substances may be degraded or
released through the fusion of the endosome with the plasma
membrane to form exosomes (29). At
present, several possibilities have been suggested for cargo
sorting into exosomes, but the specific mechanism remains unclear
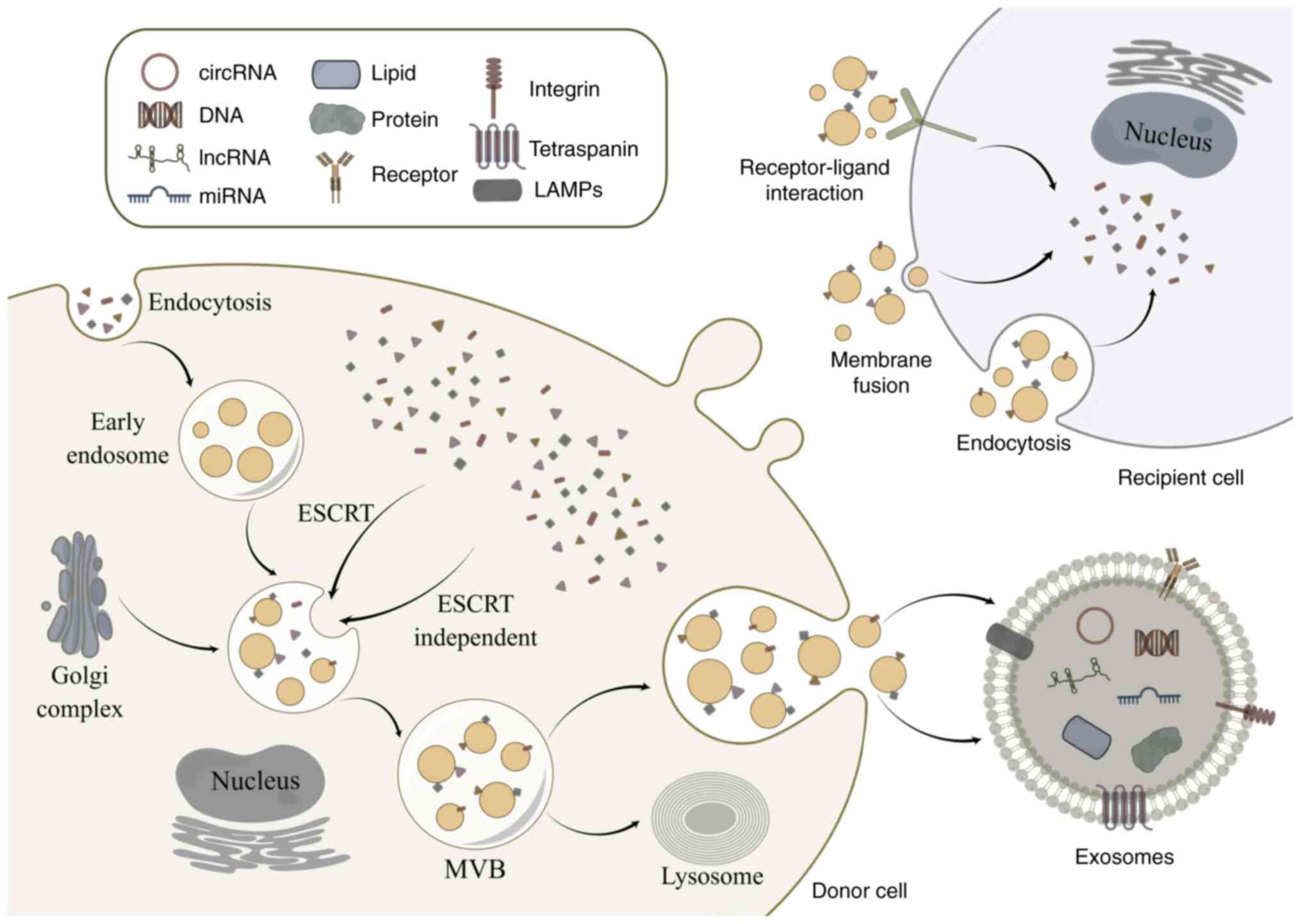
(30). Exosome release and
biomolecule packing into exosomes are reportedly regulated by the
endosomal sorting complex required for transport (ESCRT) pathway,
and ESCRT proteins import ubiquitin proteins into MVBs (30). A total of five protein-protein
complexes make up the ESCRT system: i) ESCRT-0; ii) ESCRT-I; iii)
ESCRT-II; iv) ESCRT-III; and v) Vps4-Vta1 (31). In summary, ESCRT-0 gathers
ubiquitinated cargo to start the process; protein complexes in a
saddle shape are created when the ESCRT-I and ESCRT-II complexes
bind cargo; the ESCRT-III complex promotes vesicle contraction and
maturation; and Vps4-Vta1 promotes membrane fracture (32). In addition, some ESCRT-independent
pathways also play important roles in exosome genesis. Lipid rafts,
such as flotillins and caveolins, play a key role in
ESCRT-independent intraluminal vesicles formation (33,34).
Exosome production has also been linked to flotillin, caveolin-1,
cholesterol and tetraspanins (Fig.
1) (35–38).
As aforementioned, exosomes were initially
considered waste products excreted by cells. However, exosomes can
control the proliferation, migration and invasion of tumor cells by
information transfer, as more and more studies have demonstrated
(39,40). For instance, He et al
revealed that pancreatic cancer cells release exosomes that are
abundant in FGD5 antisense RNA 1, which facilitates the evolution
of pancreatic cancer by way of tumor-associated alternatively
activated (M2) macrophage polarization (41). Exosomes have also been revealed to
modulate immune responses (42).
According to Yang et al, exosomal hsa_circ_0085361 is
abnormally expressed in bladder cancer tissues and controls T cell
depletion by influencing downstream miRNAs (43). Exosomes can also be crucial in a
number of biological processes of the neurological system (44). According to studies, damaging
proteins such β amyloid peptide, superoxide dismutase and α
synuclein are present in the exosomes of the central nervous system
and help advance disorders (44–47).
For example, studies have found that α synuclein, which is secreted
by cells through exosomal calcium-dependent mechanisms, helps
amplify and spread Parkinson's disease-related pathology (47). Exosomes also contribute to the
development of metabolic and other disorders, such as
cardiovascular ailments (23).
Notably, there are several potential uses for exosomes in the
treatment, identification and assessment of tumors. According to Xu
et al, the exosome-related gene fibroblast growth factor 9
can be downregulated and exploited as a novel diagnostic and
prognostic target for ovarian cancer (48). Because a number of components of
exosomes are related to diseases, research on exosomes has
increased (49,50).
CircRNAs are closed circular RNA molecules found in
a variety of specimen types, including bone marrow cells,
platelets, white blood cells and red blood cells (51). Unlike conventional linear RNA,
circRNA backsplicing joins the upstream 3′ splice acceptor site and
the downstream 5′ splice donor site to create a single-stranded
covalent closed loop (52).
Circular RNA is more durable compared with linear RNA and resistant
to exonuclease disintegration because it lacks the 5′ and 3′ ends
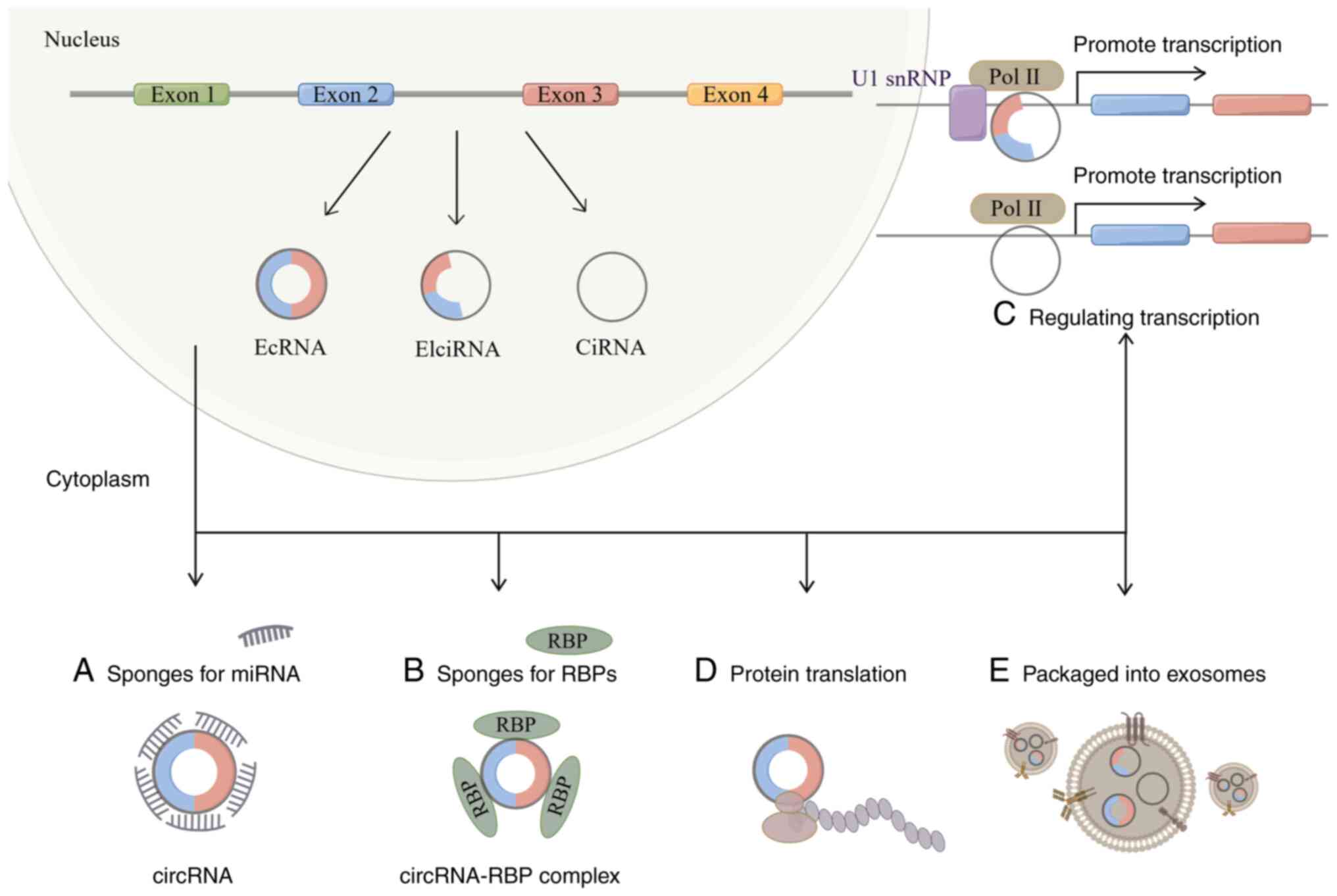
(52). According to the various
genome constituent sequences, circRNAs can be divided into three
different categories: Exonic circular RNAs, intronic circular RNAs
(ciRNAs) and exon-intron circular RNAs (EIciRNAs) (Fig. 2) (53). The majority of them are present in
the nucleus, including ciRNAs and EIciRNAs; however, there are few
miRNA targets for ciRNAs (53,54).
Due to its distinctive structure, circRNAs have a
wide range of biological roles and are crucial in numerous
illnesses, according to recent studies (55,56).
CircRNAs can control gene expression by interacting with miRNAs, as
a number of studies have demonstrated (57,58).
For example, Huang et al has revealed that hsa_circ_104348
can regulate the expression of miR-187-3p, thereby affecting the
progression of hepatocellular carcinoma (59). The exonic region of RNase P RNA
component H27 is the source of circRNA-002178, which has been shown
to be overexpressed in esophageal cancer cells (60). CircRNA-002178 can act as a sponge
for miR-34a and miR-28-5p to reduce their inhibitory effects on
their target genes PDL1 and PD1 in cancer cells and CD8 T cells
(60,61). Circular RNAs can also function by
attaching to RNA-binding proteins. A new circRNA has been
identified by Chen et al in the friend leukemia virus
integration 1 promoter, and it is reported to encourage breast
cancer spread by controlling the enzymes methylcytosine dioxygenase
TET1 and DNA methyl transferase (62). Due to the absence of the 5′ cap, it
has also been proposed that circRNAs translate in a cap-independent
manner (54,63).
At present, the following circRNAs that can be
translated into proteins have been studied: circ-AKT3 (64), circSHPRH (65) and circ-ZNF609 (66). For example, circ-AKT3 is expressed
at a lower level in glioblastoma tissue compared with normal brain
tissue, and it can encode AKT3-174aa (64). The tumorigenicity of glioblastoma
can be affected by influencing the expression of circ-AKT3 and
AKT3-174aa (64). As circRNA
research advances, these types of molecules may be employed as
tumor diagnostic markers. For example, Lu et al demonstrated
that hsa_circ_0001789 is underexpressed in GC tissues and plasma,
with an area under receiver operating curve (AUC) of 0.82; this
circRNA may become a novel diagnostic biomarker for GC, and its
efficacy is better compared with that of traditional tumor markers
(67). According to a previous
study, the hepatocellular carcinoma diagnostic biomarkers
hsa_circ_0004001, hsa_circ_0004123, hsa_circ_0075792 and the
combination of the three exhibit greater sensitivity and
specificity compared with the three circRNAs separately (68).
According to previous studies, circRNAs influence
various biological processes, are persistent and are concentrated
in exosomes (69,70). Exosomal circPRRX1, as the competing
endogenous RNA of miR-596, has been revealed to promote the
proliferation, migration and invasion of GC cells, and decreases
their radiation sensitivity through the upregulation of
NF-κB-activated protein (71).
Since a number of cell types can receive exosomes, exosomes can act
as messengers in intercellular communication (72). In the occurrence and development of
tumors, circRNA can be packaged into exosomes and sent to receptor
cells to play a certain role, affecting the tumor microenvironment
and thus influencing the progression of the disease (73). As shown in Fig. 1, exosomes transfer circRNAs into
recipient cells mainly via three processes: Receptor-ligand
interaction, membrane fusion and endocytosis (23). However, the specific mechanism of
selective packaging of circRNAs into exosomes is still unclear.
According to Yang et al, the exosome circ-133 produced by
hypoxia cells is substantially expressed in the plasma of CRC
patients and can be transferred to healthy cells to advance the
disease (74). Notably, numerous
other studies have found a connection between exosomes and the
removal of intracellular circRNAs (75,76).
For example, it has been revealed that exosomes can be further
eliminated by the reticuloendothelial system or produced by the
kidney and liver (75). Moreover,
previous research has revealed that, compared with healthy cells,
CRC cells can actively release more circ-rho-related BTB
domain-containing 3 (circRHOBTB3) to sustain colorectal cancer cell
fitness (76). In another study,
Chen et al demonstrated that circRHOBTB3 inhibits the
progression of CRC by regulating intracellular reactive oxygen
species and metabolic pathways (76). In addition, this study advanced a
novel theory of tumor escape known as ‘the tumor exosome escape
mechanism’, which postulates that tumor cells secrete tumor
inhibitory circRNAs in order to maintain the fitness of cancer
cells. The mechanism of tumor exosome escape is of great
significance for understanding and treating cancer (76). Furthermore, the expression levels of
circRNA in some patients with tumors are significantly different
compared with that of the healthy population, such as
hsa_circ_001783 and circRNA_102231, and because of its strong
stability, long half-life and anti-degradation, circRNA can be used
as a tumor diagnosis and molecular marker (77,78).
Exosomal circRNAs have been implicated in the
emergence and progression of gastrointestinal cancer in a growing
number of studies over the past several years (79,80).
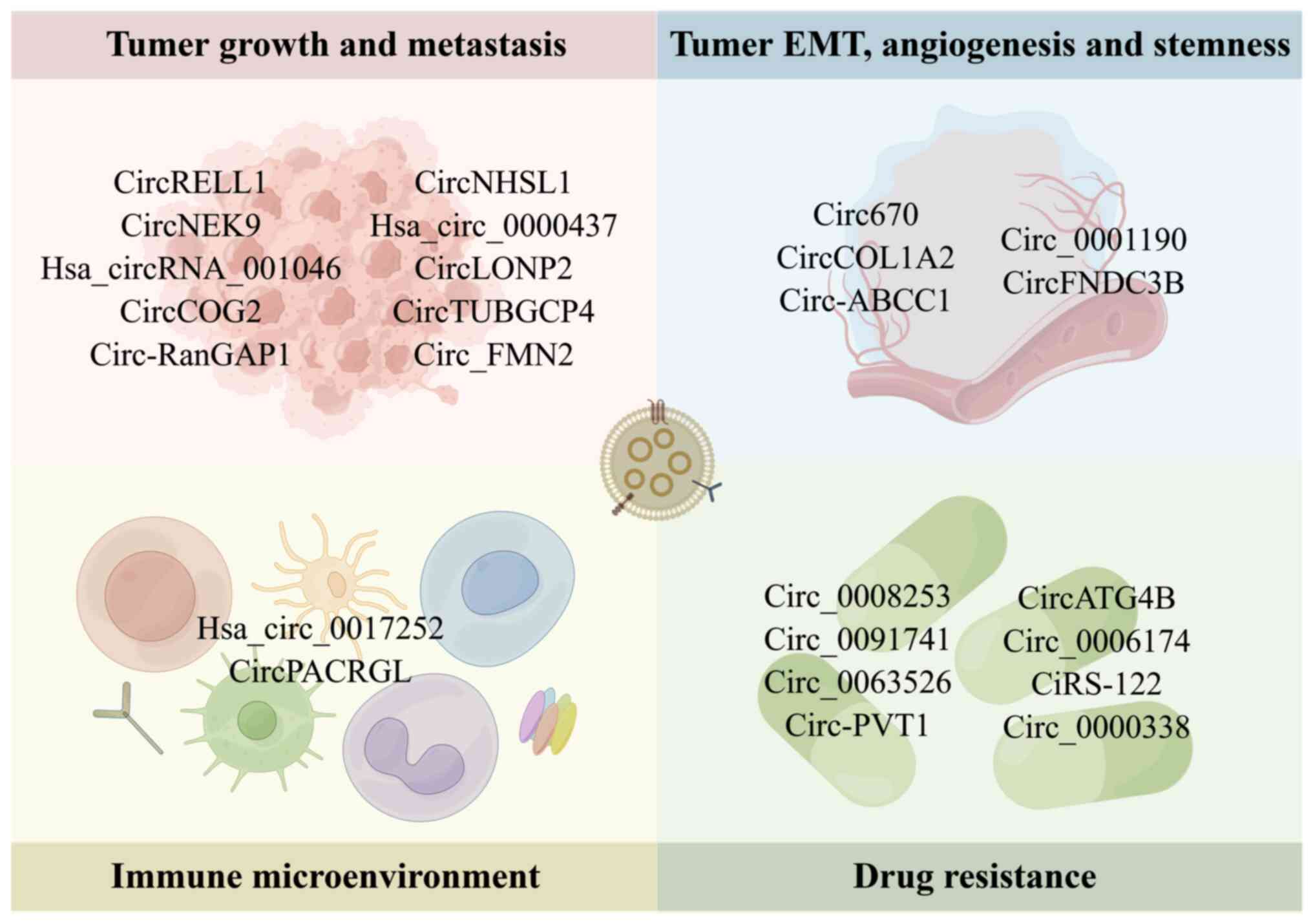
By binding to miRNA, receptor proteins, or other regulatory
mechanisms, exosomal circRNAs influence tumor growth, metastasis,
stemness, angiogenesis, the immunological microenvironment and
treatment resistance (Fig. 3 and
Table I).
Exosomal circRNAs have been shown to control the
microenvironment and surrounding tissue cells, which has been
demonstrated to have an impact on how gastrointestinal cancer
progresses (81). It has been
reported that exosomal circRNAs promote the occurrence and
development of gastrointestinal cancer by affecting tumor
proliferation, metastasis, stemness and angiogenesis (81). Sang et al demonstrated that
circRNA RELL1 is significantly downregulated in GC tissues and
transmitted by exosomes (82).
CircRELL1 inhibits GC progression through the miR-637/ephrin type-B
receptor 3 axis, which may be a novel diagnostic marker and
therapeutic target for GC (82). Yu
et al revealed that circRNA NIMA-related kinase 9
accelerates GC progression by targeting the miR-409-3p/microtubule
associated protein 7 axis, which can be reflected in malignant
behaviors such as migration and invasion (83). A possible therapeutic target for CRC
has been identified in research as the cancer-derived exosomal
hsa_circRNA_001046, which plays an oncogenic role in the
progression of the disease (84).
According to Gao et al, circRNA component of oligomeric
Golgi complex 2 (COG2) may be transported from cancer cells with a
high metastatic potential to cancer cells with a low metastatic
potential via exosomes (85).
circCOG2 has been demonstrated to be enhanced in CRC tissue and
plasma exosomes. By controlling downstream miRNA and associated
proteins, circCOG2 aids in the development of the malignant
phenotype of CRC. According to this, circCOG2 may be both a
therapeutic target and a predictive factor for CRC (85).
In addition, there is growing evidence that exosomes
released by various living cells promote the metastasis of
gastrointestinal cancer by delivering circRNAs (86,87).
For example, circ-RanGAP1 is found in plasma exosomes from
preoperative patients with GC, according to the research of Lu
et al (86). In addition,
the ability of GC cells to migrate and invade is improved by the
plasma exosomes that were obtained from these individuals. This
research raises the possibility that circ-RanGAP1 might serve as a
predictive biomarker and a therapeutic target for the treatment of
GC (86). Hui et al
demonstrated that circNHSL1 knockdown represses migration, invasion
and glutaminolysis and inhibits tumor growth by downregulating the
miR-149-5p/tyrosine 3-monooxygenase axis in GC, implying an
underlying therapy target for GC treatment (87). Shen et al also revealed that
hsa_circ_0000437 regulates malignant behaviors such as migration of
human lymphatic endothelial cells by targeting serine and arginine
rich splicing factor 3 and inhibiting programmed cell death factor
4, as well as lymph node metastasis (LNM) in the popliteal LNM
model (88). In CRC, circLONP2 is
abnormally highly expressed in metastatic primary CRC tissues and
can be transferred between cells by regulating miR-17 (89). This study suggests that circLONP2
may be a potential anti-metastatic therapeutic target for CRC
(89). The findings of Chen et
al determine that CRC cells are rich in exosomal circTUBGCP4,
which promotes CRC metastasis through upregulation of pyruvate
dehydrogenase kinase 2 (90).
Additionally, Yu et al revealed that the exosomal circ_FMN2
is abundant in CRC cells and that its overexpression can encourage
the spread of CRC cells and the development of colorectal tumors
(91).
The incidence, invasion, metastasis and treatment
resistance of gastrointestinal cancer are all assumed to be
influenced by epithelial-mesenchymal transition (EMT), a
fundamental cellular process (92).
CircRNAs are crucial to EMT in gastrointestinal cancer (93,94).
Exosomal circ670 from the tissues of patients with GC has been
demonstrated to alter the EMT process of GC stem cells, hence
boosting the growth of GC, according to a previous study (95). According to Miao et al,
exosomal circRNA collagen type I α 2 chain (circCOL1A2) is
significantly expressed in CRC cells, and miR-665 can impact EMT
and other cellular properties when combined with circCOL1A2 or LIM
and SH3 Protein 1 (96). Moreover,
this study indicated that circCOL1A2 is a potential novel
therapeutic target for CRC (96).
Additionally, it has been shown that a number of
exosomal circRNAs are associated with the stemness and angiogenesis
of gastrointestinal tumor cells (97). Zhao et al demonstrated that
the exosomes of CD133 cells are rich in hsa_circ_0000677, which can
mediate the stemness of CRC cells and promote the progression of
CRC, and is expected to become a potential therapeutic target for
this cancer (97). Liu et al
also revealed that the expression of circ_0001190 in the exosome
source of GC is low, and overexpression of circ_0001190 can inhibit
the angiogenesis ability of cells, thus inhibiting the development
of GC (98). Additionally, Zeng
et al demonstrated that circFNDC3B has a low expression
level in CRC tissues and cells and that it has the ability to stop
the growth of the disease by preventing the angiogenic features of
the disease (99). These studies
indicate that exosomal circRNAs can promote or inhibit the
progression of gastrointestinal cancer.
Exosomal circRNAs can affect the function of immune
cells in the tumor microenvironment and then affect the occurrence
and development of gastrointestinal cancer. For instance, exosomal
hsa_circ_0017252 secreted by GC cells slows the growth of the
disease by inhibiting M2-like polarization of macrophages (100). Exosome-derived circPACRGL is
significantly expressed in CRC cells. By controlling downstream
miRNAs, circPACRGL promotes the malignant activity of CRC cells and
the differentiation of N1 to N2 neutrophils. To the best of our
knowledge, this research is the first to demonstrate that exosomal
circPACRGL contributes to CRC carcinogenesis and may have promise
as a biomarker (101). These
findings suggest that exosomal circRNAs can affect the progression
of gastrointestinal cancer through immune cells.
It has been revealed that circRNAs cannot only
affect the progression of gastrointestinal cancer, but also
participate in their chemoresistance (102). A number of circRNAs have a role in
drug resistance in GC, such as circ 0008253 and circ_0091741
(103,104). According to a recent study, circ
0008253 is abundant in M2-polarized exosomes that are produced by
macrophages and can be delivered to GC cells via exosomes to
increase their resistance to oxaliplatin (OXA) (103). Chen et al also demonstrated
that the exosomal circ_0091741 derived from GC cells increases the
expression of tripartite motif containing 14, thereby enhancing OXA
resistance (104). In addition,
circ_0063526 is highly expressed in GC tissues and cells, and
cisplatin resistance is enhanced by regulating the expression of
serine hydroxymethyltransferase 2 (105). Moreover, exosome-derived circ-PVT1
is underexpressed in GC cells and serum, and contributes to DDP
resistance by regulating miR-30a-5p; therefore, Exosomal circ-PVT1
may be a potential therapeutic target for GC (106). In CRC, a novel support for a
possible therapeutic target for oxaliplatin resistance is provided
by the exosomal circATG4B, which contributes to the lower
chemosensitivity of CRC cells (107). Zhang et al revealed that
circ_0006174 is upregulated in exosomes of doxorubicin
(DOX)-resistant CRC cells and can enhance their resistance through
the intercellular transfer of exosomes (108). Another study has revealed that
hsa_circ_0005963 can be transferred from chemotherapy-resistant CRC
cells to sensitive cells via exosomes, enhancing drug resistance in
sensitive cells (109). A study
suggests that hsa_circ_0005963 may be a potential therapeutic
target for drug-resistant CRC (109). Moreover, Zhao et al
revealed that miR-217 and miR-485-3p are controlled by
exosome-mediated circ_0000338 transfer, which increases
3-fluorouracil resistance in CRC (110). These results suggest that exosomal
circRNAs can enhance the resistance of oxaliplatin, 3-fluorouracil,
cisplatin and DOX in gastrointestinal cancer. The application of
antitumor drugs and specific circRNA inhibitors can improve the
therapeutic effect of gastrointestinal tumors.
Because most of the early symptoms of
gastrointestinal cancer are not obvious, early detection and
diagnosis are challenging, and most patients are at an advanced
stage when they have obvious symptoms, and their survival rate is
low (111). Therefore, the search
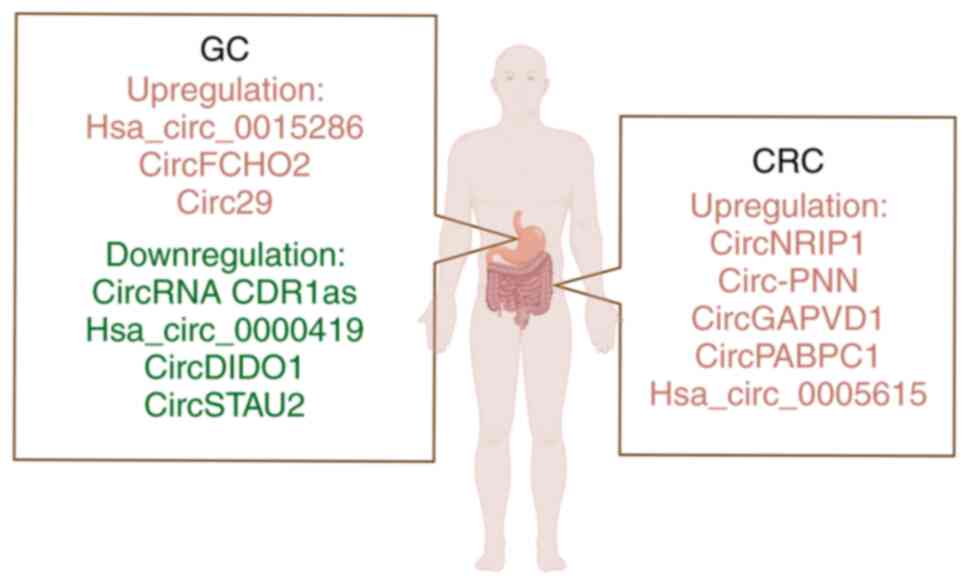
for new biomarkers for gastrointestinal cancer is crucial. Exosomes
exist stably in body fluids and can act as messengers between
cells, which are considered as promising biomarkers (Fig. 4 and Table II) (112,113). CircRNAs have been found to be
stable and abundant in exosomes, making it simple to detect their
levels (114,115). In addition, circRNAs can be
specifically and differentially expressed across tissues and body
fluids under various pathological conditions, so circRNAs can be
used as a novel biomarker for a number of diseases (116,117). A previous study has shown that the
circulating levels of exosomal circRNAs can not only predict tumor
progression, but also serve as potential diagnostic biomarkers.
Zheng et al demonstrated that hsa_circ_0015286 is
significantly highly expressed in GC cells, tissues and plasma
compared with normal controls and the AUC is 0.778, which is higher
compared with the AUC of traditional tumor markers CA 19-9 and CEA
(118). In addition, the AUC can
be as high as 0.843 when the three indicators are combined. This
study shows that exosomal hsa_circ_0015286 can be used in
combination with traditional tumor markers CEA and CA 19-9 for high
diagnostic efficacy, and it can also be diagnosed alone (118). Li et al examined the high
expression of CDR1 antisense RNA (CDR1as) in the plasma and
exosomes of patients with GC, and heat shock protein family E
member 1 may be a key protein in the regulation of miRNA by CDR1as
(119). These authors evaluated
the AUC of tissue, plasma and exosomal derived CDR1as and revealed
that tissue-derived CDR1as has the highest AUC of 0.782. The AUC of
plasma CDR1as combined with conventional tumor markers is higher
compared with that of tissue-derived CDR1as. This study suggests
that CDR1as may be a potential diagnostic biomarker for GC
(119). Tao et al showed
that hsa_circ_0000419 is stably present in plasma exosomes and has
low expression in GC tissues and cells. It regulates the
development of GC by interacting with hsa-miR-589-3p and
hsa-miR-141-5p. When the critical value of hsa_circ_0000419 is
4.90, the specificity and sensitivity of plasma hsa_circ_0000419
reaches 0.884 and 0.682, respectively (120).
Furthermore, there are differences in the expression
of exosomal circRNAs in CRC. Pan et al revealed that
exosome-derived hsa-circ-0004771 is lowly expressed in CRC cells
and tissues, but highly expressed in the serum of patients with CRC
(121). The authors suggest that
hsa-circ-000477 may bind to RNA-binding proteins and be transported
to exosomes, which may be a mechanism for circRNA clearance. In
different periods of CRC, the AUC of hsa-circ-000477 is >0.8,
indicating that hsa-circ-000477 can be used as a novel potential
diagnostic biomarker for CRC (121). According to Xie et al,
circ-PNN is considerably overexpressed and strongly correlated with
the prognosis of patients with CRC (122). The authors revealed that circ-PNN
level has high performance in being able to differentiate patients
with CRC from healthy controls; in addition, this study further
investigated the possible clinical importance of serum exosomal
circ-PNN. Circ-PNN might therefore be used as a diagnostic
biomarker for CRC (122). Li et
al revealed that circGAPVD1 is highly expressed in the plasma
exosomes of CRC, with an AUC area of 0.7662 and diagnostic
specificity and sensitivity of 71.79 and 75.64%, respectively
(123). These results indicate
that circGAPVD1 is expected to be a diagnostic marker for CRC
(123). The aforementioned
findings indicate that exosomal circRNAs are expected to be a novel
biomarker for the prediction and diagnosis of gastrointestinal
cancer.
Currently, the first-line treatment strategy for
gastrointestinal cancer is chemotherapy, but cancer cells may
become resistant to anticancer drugs or their targets. Finding
novel treatment strategies at the molecular level is therefore
crucial. Increasing evidence suggests that exosomal circRNAs are
expected to be promising therapeutic targets for gastrointestinal
cancer (124,125). Zhang et al revealed that
circFCHO2 is highly expressed in serum exosomes of patients with GC
(126). CircFCHO2 regulates the
JAK1/STAT3 pathway by binding to miR-194-5p, which affects some
malignant behaviors of GC cells; therefore, circFCHO2 may be a
novel therapeutic target and diagnostic biomarker for GC (126). Li et al also demonstrated
that exosomal-derived circ29 is substantially expressed in GC,
which influences the GC formation by regulating the
miR-29a/vascular endothelial growth factor axis, and is anticipated
to emerge as a novel tumor marker and promising therapeutic target
(127). In addition, Li et
al demonstrated that exosome-derived circPABPC1 is highly
expressed in CRC tissues and exist in the cytoplasm (128). CircPABPC1 promotes CRC progression
by promoting the expression of important regulators of liver
metastasis in CRC such as a disintegrin and metalloproteinase 19
and bone morphogenetic protein 4. Therefore, exosome-derived
circPABPC1 is expected to be a novel biomarker and anti-metastatic
therapeutic target for CRC (128).
These circRNAs promote the malignant transformation of cancer
cells, and inhibition of these exosomal circRNAs may help to
suppress gastrointestinal cancers. In addition, it has been
reported that exosome-mediated circRNA delivery with anti-tumor
effects is considered to be an effective therapeutic method
(109,129). For example, the results of Guo
et al show that circDIDO1 inhibits GC progression by
regulating the miR-1307-3p/suppressor of cytokine signaling 2 axis
(130). Systemic administration of
RGD-modified circDIDO1-loaded exosomes inhibits GC tumorigenicity
and invasiveness in vitro and in vivo, suggesting
that RGD-Exo-circDIDO1 can be used as a viable nanofat for GC
treatment (130). Zhang et
al demonstrated that exosome-delivered circSTAU2 can act as a
tumor suppressor and inhibit GC progression through the
miR-589/capping actin protein of muscle Z-line subunit α 1 axis,
which indicates that circSTAU2 may be a potential therapeutic
target for GC (131). Moreover, in
CRC, it has been shown that circ_0005615 from serum exosomes of
patients with CRC can modulate CRC malignant progression by
controlling fos-like antigen 2 expression through sponging
miR-873-5p, which reveals circ_0005615 as a new candidate target
for CRC treatment (132). In
summary, the treatment of gastrointestinal cancer mediated by
exosomal circRNAs is still in its infancy, and the specific
mechanism and safety of this strategy need to be further
studied.
Exosomal circRNAs have a significant role in
gastrointestinal cancers, as research has revealed in recent years
(133,134). It has been demonstrated that
circRNA has an impact on the proliferation, metastasis, stemness,
angiogenesis, immune microenvironment and drug resistance of
gastrointestinal tumors, and may develop into a useful therapeutic
target for gastrointestinal malignancies as well as a potential
biomarker (135). Although more
studies have focused on the application of exosome-derived circRNAs
in gastrointestinal tumors, there are still a number of problems to
be solved.
First of all, existing studies lack technologies
for rapid extraction and detection of exosomes and exosomal
circRNAs. Currently, ultracentrifugation, density gradient
centrifugation, ultrafiltration, immunoaffinity and polymer
precipitation are the major methods used to isolate exosomes
(136–138). Ultracentrifugation, the most
commonly employed technique, is time-consuming and inconvenient and
can lead to the destruction of exosomes. Exosomes obtained by
ultracentrifugation have low purity and low yield due to the
contamination of non-exosome components (136). So far, there is no clear method to
directly extract circRNA, but total RNA can be digested with RNase
R to enrich circRNA. Second, the secretion mechanism of
exosomes-derived circRNAs remains unclear. Future research should
concentrate on how exosomes discharge circRNAs to act on the
desired cells. In addition, the limited number of circRNAs with
clear mechanisms of action in gastrointestinal cancer requires
further research and exploration. Finally, there is still a lack of
a large number of prospective studies on exosome-derived circRNA as
a diagnostic marker and targeted therapy molecule for
gastrointestinal tumors. It is important to further assess the
sensitivity, specificity and diagnostic impact of exosomal circRNAs
paired with current tumor markers as a diagnostic marker. Exosomal
circRNAs, which is a targeted therapeutic molecule, encounters more
difficulties with regard to its efficacy, safety, potential side
effects, delivery method, biodistribution and effective dose.
In conclusion, exosomal circRNAs have promise as
novel diagnostic biomarkers and therapeutic targets and play a
critical role in the development of gastrointestinal malignancies.
It will be beneficial for its clinical use if the study of exosomal
circRNAs and gastrointestinal cancers becomes more in-depth and
cutting-edge technologies, including exosomal circRNA separation
and detection, are developed. The present study hypothesizes that
diagnostic and therapeutic strategies based on exosomal circRNAs
can be effectively applied to clinical practice in the future.
Not applicable.
This work was supported by the National Natural Science
Foundation of China (grant no. 81602883), the Project of Social
Development in Zhenjiang (grant no. SH2021045), the Technology
Development Foundation of Jiangsu University (grant no. 20220516),
the Postgraduate Research & Practice Innovation Program of
Jiangsu Province (grant no. KYCX23_3765) and the Student Innovation
Training Program of Jiangsu University (grant nos. 202310299410X
and 202310299476X).
Not applicable.
ZL, JJ and HQ designed research and wrote the
paper. YX, XuZ, XiZ, JS, XZ and ZG have made contributions to
analysis and interpretation of data. XuZ, XiZ, JS, XZ and ZG have
made contributions to the production of tables and figures. YX, JH
and XiZ contributed to the writing and revisions. All authors read
and approved the final manuscript. Data authentication is not
applicable.
Not applicable.
Not applicable.
All authors declare that they have no competing
interests.
|
1
|
Arnold M, Abnet CC, Neale RE, Vignat J,
Giovannucci EL, McGlynn KA and Bray F: Global Burden of 5 major
types of gastrointestinal cancer. Gastroenterology.
159:335–349.e15. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Anand P, Kunnumakkara AB, Sundaram C,
Harikumar KB, Tharakan ST, Lai OS, Sung B and Aggarwal BB: Cancer
is a preventable disease that requires major lifestyle changes.
Pharm Res. 25:2097–2116. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kashyap S, Pal S, Chandan G, Saini V,
Chakrabarti S, Saini NK, Mittal A, Thakur VK, Saini AK and Saini
RV: Understanding the cross-talk between human microbiota and
gastrointestinal cancer for developing potential diagnostic and
prognostic biomarkers. Semin Cancer Biol. 86((Pt 3)): 643–651.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yang Z, Wang D, Zhang C, Liu H, Hao M, Kan
S, Liu D and Liu W: The applications of gold nanoparticles in the
diagnosis and treatment of gastrointestinal cancer. Front Oncol.
11:8193292022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Jiang DM, Chan KKW, Jang RW, Booth C, Liu
G, Amir E, Mason R, Everest L and Elimova E: Anticancer drugs
approved by the Food and Drug Administration for gastrointestinal
malignancies: Clinical benefit and price considerations. Cancer
Med. 8:1584–1593. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Singh D, Dheer D, Samykutty A and Shankar
R: Antibody drug conjugates in gastrointestinal cancer: From lab to
clinical development. J Control Release. 340:1–34. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhang M, Hu S, Liu L, Dang P, Liu Y, Sun
Z, Qiao B and Wang C: Engineered exosomes from different sources
for cancer-targeted therapy. Signal Transduct Target Ther.
8:1242023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mantovani A, Allavena P, Marchesi F and
Garlanda C: Macrophages as tools and targets in cancer therapy. Nat
Rev Drug Discov. 21:799–820. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Senthebane DA, Rowe A, Thomford NE,
Shipanga H, Munro D, Mazeedi MAMA, Almazyadi HAM, Kallmeyer K,
Dandara C, Pepper MS, et al: The role of tumor microenvironment in
chemoresistance: To survive, keep your enemies closer. Int J Mol
Sci. 18:15862017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Roma-Rodrigues C, Mendes R, Baptista PV
and Fernandes AR: Targeting tumor microenvironment for cancer
therapy. Int J Mol Sci. 20:8402019. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
van Niel G, D'Angelo G and Raposo G:
Shedding light on the cell biology of extracellular vesicles. Nat
Rev Mol Cell Biol. 19:213–228. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Liu Y, Gu Y and Cao X: The exosomes in
tumor immunity. Oncoimmunology. 4:e10274722015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Sung BH, von Lersner A, Guerrero J,
Krystofiak ES, Inman D, Pelletier R, Zijlstra A, Ponik SM and
Weaver AM: A live cell reporter of exosome secretion and uptake
reveals pathfinding behavior of migrating cells. Nat Commun.
11:20922020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Pluchino S and Smith JA: Explicating
exosomes: Reclassifying the rising stars of intercellular
communication. Cell. 177:225–227. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lu Y, Huang W, Li M and Zheng A:
Exosome-Based carrier for RNA delivery: Progress and challenges.
Pharmaceutics. 15:5982023. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Meng X, Xiao W, Sun J, Li W, Yuan H, Yu T,
Zhang X and Dong W: CircPTK2/PABPC1/SETDB1 axis promotes
EMT-mediated tumor metastasis and gemcitabine resistance in bladder
cancer. Cancer Lett. 554:2160232023. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Shen Y, Zhang N, Chai J, Wang T, Ma C, Han
L and Yang M: CircPDIA4 induces gastric cancer progression by
promoting ERK1/2 activation and enhancing biogenesis of oncogenic
circRNAs. Cancer Res. 83:538–552. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Long F, Lin Z, Li L, Ma M, Lu Z, Jing L,
Li X and Lin C: Comprehensive landscape and future perspectives of
circular RNAs in colorectal cancer. Mol Cancer. 20:262021.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li X, Lin YL, Shao JK, Wu XJ, Li X, Yao H,
Shi FL, Li LS, Zhang WG, Chang ZY, et al: Plasma exosomal
hsa_circ_0079439 as a novel biomarker for early detection of
gastric cancer. World J Gastroenterol. 29:3482–3496. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Huang XJ, Wang Y, Wang HT, Liang ZF, Ji C,
Li XX, Zhang LL, Ji RB, Xu WR, Jin JH and Qian H: Exosomal
hsa_circ_000200 as a potential biomarker and metastasis enhancer of
gastric cancer via miR-4659a/b-3p/HBEGF axis. Cancer Cell Int.
23:1512023. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Xing L, Xia M, Jiao X and Fan L:
Hsa_circ_0004831 serves as a blood-based prognostic biomarker for
colorectal cancer and its potentially circRNA-miRNA-mRNA regulatory
network construction. Cancer Cell Int. 20:5572020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Cocucci E and Meldolesi J: Ectosomes and
exosomes: Shedding the confusion between extracellular vesicles.
Trends Cell Biol. 25:364–372. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kalluri R and LeBleu VS: The biology,
function, and biomedical applications of exosomes. Science.
367:eaau69772020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wortzel I, Dror S, Kenific CM and Lyden D:
Exosome-Mediated Metastasis: Communication from a Distance. Dev
Cell. 49:347–360. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kok VC and Yu CC: Cancer-Derived Exosomes:
Their role in cancer biology and biomarker development. Int J
Nanomedicine. 15:8019–8036. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Huang X, Wu W, Jing D, Yang L, Guo H, Wang
L, Zhang W, Pu F and Shao Z: Engineered exosome as targeted lncRNA
MEG3 delivery vehicles for osteosarcoma therapy. J Control Release.
343:107–117. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Pan BT and Johnstone RM: Fate of the
transferrin receptor during maturation of sheep reticulocytes in
vitro: Selective externalization of the receptor. Cell. 33:967–978.
1983. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Doyle LM and Wang MZ: Overview of
extracellular vesicles, their origin, composition, purpose, and
methods for exosome isolation and analysis. Cells. 8:7272019.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kahlert C and Kalluri R: Exosomes in tumor
microenvironment influence cancer progression and metastasis. J Mol
Med (Berl). 91:431–437. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zhang X, Zhang H, Gu J, Zhang J, Shi H,
Qian H, Wang D, Xu W, Pan J and Santos HA: Engineered extracellular
vesicles for cancer therapy. Adv Mater. 33:e20057092021. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tallon C, Hollinger KR, Pal A, Bell BJ,
Rais R, Tsukamoto T, Witwer KW, Haughey NJ and Slusher BS: Nipping
disease in the bud: nSMase2 inhibitors as therapeutics in
extracellular vesicle-mediated diseases. Drug Discov Today.
26:1656–1668. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Henne WM, Buchkovich NJ and Emr SD: The
ESCRT pathway. Dev Cell. 21:77–91. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Skotland T, Hessvik NP, Sandvig K and
Llorente A: Exosomal lipid composition and the role of ether lipids
and phosphoinositides in exosome biology. J Lipid Res. 60:9–18.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Dawson G: Isolation of lipid rafts
(Detergent-Resistant Microdomains) and comparison to extracellular
vesicles (Exosomes). Methods Mol Biol. 2187:99–112. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Parton RG, McMahon KA and Wu Y: Caveolae:
Formation, dynamics, and function. Curr Opin Cell Biol. 65:8–16.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Kwiatkowska K, Matveichuk OV, Fronk J and
Ciesielska A: Flotillins: At the Intersection of Protein
S-Palmitoylation and lipid-mediated signaling. Int J Mol Sci.
21:22832020. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Ikonen E and Zhou X: Cholesterol transport
between cellular membranes: A balancing act between interconnected
lipid fluxes. Dev Cell. 56:1430–1436. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Kummer D, Steinbacher T, Schwietzer MF,
Thölmann S and Ebnet K: Tetraspanins: integrating cell surface
receptors to functional microdomains in homeostasis and disease.
Med Microbiol Immunol. 209:397–405. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Lee YJ, Shin KJ, Jang HJ, Ryu JS, Lee CY,
Yoon JH, Seo JK, Park S, Lee S, Je AR, et al: GPR143 controls
ESCRT-dependent exosome biogenesis and promotes cancer metastasis.
Dev Cell. 58:320–334.e8. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Qi R, Bai Y, Li K, Liu N, Xu Y, Dal E,
Wang Y, Lin R, Wang H, Liu Z, et al: Cancer-associated fibroblasts
suppress ferroptosis and induce gemcitabine resistance in
pancreatic cancer cells by secreting exosome-derived
ACSL4-targeting miRNAs. Drug Resist Updat. 68:1009602023.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
He Z, Wang J, Zhu C, Xu J, Chen P, Jiang
X, Chen Y, Jiang J and Sun C: Exosome-derived FGD5-AS1 promotes
tumor-associated macrophage M2 polarization-mediated pancreatic
cancer cell proliferation and metastasis. Cancer Lett.
548:2157512022. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Xu Z, Zeng S, Gong Z and Yan Y:
Exosome-based immunotherapy: A promising approach for cancer
treatment. Mol Cancer. 19:1602020. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Yang C, Wu S, Mou Z, Zhou Q, Dai X, Ou Y,
Chen X, Chen Y, Xu C, Hu Y, et al: Exosome-derived circTRPS1
promotes malignant phenotype and CD8+ T cell exhaustion in bladder
cancer microenvironments. Mol Ther. 30:1054–1070. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Rajendran L, Honsho M, Zahn TR, Keller P,
Geiger KD, Verkade P and Simons K: Alzheimer's disease beta-amyloid
peptides are released in association with exosomes. Proc Natl Acad
Sci USA. 103:11172–11177. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Fevrier B, Vilette D, Archer F, Loew D,
Faigle W, Vidal M, Laude H and Raposo G: Cells release prions in
association with exosomes. Proc Natl Acad Sci USA. 101:9683–9688.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Gomes C, Keller S, Altevogt P and Costa J:
Evidence for secretion of Cu, Zn superoxide dismutase via exosomes
from a cell model of amyotrophic lateral sclerosis. Neurosci Lett.
428:43–46. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Emmanouilidou E, Melachroinou K,
Roumeliotis T, Garbis SD, Ntzouni M, Margaritis LH, Stefanis L and
Vekrellis K: Cell-produced alpha-synuclein is secreted in a
calcium-dependent manner by exosomes and impacts neuronal survival.
J Neurosci. 30:6838–6851. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Xu Z, Cai Y, Liu W, Kang F, He Q, Hong Q,
Zhang W, Li J, Yan Y and Peng J: Downregulated exosome-associated
gene FGF9 as a novel diagnostic and prognostic target for ovarian
cancer and its underlying roles in immune regulation. Aging (Albany
NY). 14:1822–1835. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Li B, Cao Y, Sun M and Feng H: Expression,
regulation, and function of exosome-derived miRNAs in cancer
progression and therapy. FASEB J. 35:e219162021. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Sun Z, Yang S, Zhou Q, Wang G, Song J, Li
Z, Zhang Z, Xu J, Xia K, Chang Y, et al: Emerging role of
exosome-derived long non-coding RNAs in tumor microenvironment. Mol
Cancer. 17:822018. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Vea A, Llorente-Cortes V and de
Gonzalo-Calvo D: Circular RNAs in Blood. Adv Exp Med Biol.
1087:119–130. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Xie Y, Shao Y, Sun W, Ye G, Zhang X, Xiao
B and Guo J: Downregulated expression of hsa_circ_0074362 in
gastric cancer and its potential diagnostic values. Biomark Med.
12:11–20. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Li Z, Huang C, Bao C, Chen L, Lin M, Wang
X, Zhong G, Yu B, Hu W, Dai L, et al: Exon-intron circular RNAs
regulate transcription in the nucleus. Nat Struct Mol Biol.
22:256–264. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Fu L, Jiang Z, Li T, Hu Y and Guo J:
Circular RNAs in hepatocellular carcinoma: Functions and
implications. Cancer Med. 7:3101–3109. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Peng D, Luo L, Zhang X, Wei C, Zhang Z and
Han L: CircRNA: An emerging star in the progression of glioma.
Biomed Pharmacother. 151:1131502022. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Zhang Y, Liu F, Feng Y, Xu X, Wang Y, Zhu
S, Dong J, Zhao S, Xu B and Feng N: CircRNA circ_0006156 inhibits
the metastasis of prostate cancer by blocking the ubiquitination of
S100A9. Cancer Gene Ther. 29:1731–1741. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zhang D, Ni N, Wang Y, Tang Z, Gao H, Ju
Y, Sun N, He X, Gu P and Fan X: CircRNA-vgll3 promotes osteogenic
differentiation of adipose-derived mesenchymal stem cells via
modulating miRNA-dependent integrin α5 expression. Cell Death
Differ. 28:283–302. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Zhang W, Liu H, Jiang J, Yang Y, Wang W
and Jia Z: CircRNA circFOXK2 facilitates oncogenesis in breast
cancer via IGF2BP3/miR-370 axis. Aging (Albany NY). 13:18978–18992.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Huang G, Liang M, Liu H, Huang J, Li P,
Wang C, Zhang Y, Lin Y and Jiang X: CircRNA hsa_circRNA_104348
promotes hepatocellular carcinoma progression through modulating
miR-187-3p/RTKN2 axis and activating Wnt/β-catenin pathway. Cell
Death Dis. 11:10652020. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Su H, Lin F, Deng X, Shen L, Fang Y, Fei
Z, Zhao L, Zhang X, Pan H, Xie D, et al: Profiling and
bioinformatics analyses reveal differential circular RNA expression
in radioresistant esophageal cancer cells. J Transl Med.
14:2252016. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Wang J, Zhao X, Wang Y, Ren F, Sun D, Yan
Y, Kong X, Bu J, Liu M and Xu S: circRNA-002178 act as a ceRNA to
promote PDL1/PD1 expression in lung adenocarcinoma. Cell Death Dis.
11:322020. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Chen N, Zhao G, Yan X, Lv Z, Yin H, Zhang
S, Song W, Li X, Li L, Du Z, et al: A novel FLI1 exonic circular
RNA promotes metastasis in breast cancer by coordinately regulating
TET1 and DNMT1. Genome Biol. 19:2182018. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Pamudurti NR, Bartok O, Jens M,
Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E,
Perez-Hernandez D, Ramberger E, et al: Translation of CircRNAs. Mol
Cell. 66:9–21.e7. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Xia X, Li X, Li F, Wu X, Zhang M, Zhou H,
Huang N, Yang X, Xiao F, Liu D, et al: A novel tumor suppressor
protein encoded by circular AKT3 RNA inhibits glioblastoma
tumorigenicity by competing with active phosphoinositide-dependent
Kinase-1. Mol Cancer. 18:1312019. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Begum S, Yiu A, Stebbing J and Castellano
L: Novel tumour suppressive protein encoded by circular RNA,
circ-SHPRH, in glioblastomas. Oncogene. 37:4055–4057. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Legnini I, Di Timoteo G, Rossi F, Morlando
M, Briganti F, Sthandier O, Fatica A, Santini T, Andronache A, Wade
M, et al: Circ-ZNF609 Is a Circular RNA that can be translated and
functions in myogenesis. Mol Cell. 66:22–37.e9. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Lu H, Zhang Z, Yuan X, Song H and Li P:
The role of circular RNA hsa_circ_0001789 as a diagnostic biomarker
in gastric carcinoma. Scand J Gastroenterol. 58:248–253. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Sun XH, Wang YT, Li GF, Zhang N and Fan L:
Serum-derived three-circRNA signature as a diagnostic biomarker for
hepatocellular carcinoma. Cancer Cell Int. 20:2262020. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J,
Chen D, Gu J, He X and Huang S: Circular RNA is enriched and stable
in exosomes: A promising biomarker for cancer diagnosis. Cell Res.
25:981–984. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Shabaninejad Z, Vafadar A, Movahedpour A,
Ghasemi Y, Namdar A, Fathizadeh H, Pourhanifeh MH, Savardashtaki A
and Mirzaei H: Circular RNAs in cancer: New insights into functions
and implications in ovarian cancer. J Ovarian Res. 12:842019.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
He Y, Zheng L, Yuan M, Fan J, Rong L, Zhan
T and Zhang J: Exosomal circPRRX1 functions as a ceRNA for miR-596
to promote the proliferation, migration, invasion, and reduce
radiation sensitivity of gastric cancer cells via the upregulation
of NF-κB activating protein. Anticancer Drugs. 33:1114–1125. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Lasda E and Parker R: Circular RNAs
co-precipitate with extracellular vesicles: A possible mechanism
for circRNA clearance. PLoS One. 11:e01484072016. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Huang XY, Huang ZL, Huang J, Xu B, Huang
XY, Xu YH, Zhou J and Tang ZY: Exosomal circRNA-100338 promotes
hepatocellular carcinoma metastasis via enhancing invasiveness and
angiogenesis. J Exp Clin Cancer Res. 39:202020. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Yang H, Zhang H, Yang Y, Wang X, Deng T,
Liu R, Ning T, Bai M, Li H, Zhu K, et al: Hypoxia induced exosomal
circRNA promotes metastasis of Colorectal Cancer via targeting
GEF-H1/RhoA axis. Theranostics. 10:8211–8226. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Choi H and Lee DS: Illuminating the
physiology of extracellular vesicles. Stem Cell Res Ther. 7:552016.
View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Chen C, Yu H, Han F, Lai X, Ye K, Lei S,
Mai M, Lai M and Zhang H: Tumor-suppressive circRHOBTB3 is excreted
out of cells via exosome to sustain colorectal cancer cell fitness.
Mol Cancer. 21:462022. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Yuan G, Ding W, Sun B, Zhu L, Gao Y and
Chen L: Upregulated circRNA_102231 promotes gastric cancer
progression and its clinical significance. Bioengineered.
12:4936–4945. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Liu Z, Zhou Y, Liang G, Ling Y, Tan W, Tan
L, Andrews R, Zhong W, Zhang X, Song E and Gong C: Circular RNA
hsa_circ_001783 regulates breast cancer progression via sponging
miR-200c-3p. Cell Death Dis. 10:552019. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Yang K, Zhang J and Bao C: Exosomal
circEIF3K from cancer-associated fibroblast promotes colorectal
cancer (CRC) progression via miR-214/PD-L1 axis. BMC Cancer.
21:9332021. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Zhang X, Wang S, Wang H, Cao J, Huang X,
Chen Z, Xu P, Sun G, Xu J, Lv J and Xu Z: Circular RNA circNRIP1
acts as a microRNA-149-5p sponge to promote gastric cancer
progression via the AKT1/mTOR pathway. Mol Cancer. 18:202019.
View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Vakhshiteh F, Hassani S, Momenifar N and
Pakdaman F: Exosomal circRNAs: New players in colorectal cancer.
Cancer Cell Int. 21:4832021. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Sang H, Zhang W, Peng L, Wei S, Zhu X,
Huang K, Yang J, Chen M, Dang Y and Zhang G: Exosomal circRELL1
serves as a miR-637 sponge to modulate gastric cancer progression
via regulating autophagy activation. Cell Death Dis. 13:562022.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Yu L, Xie J, Liu X, Yu Y and Wang S:
Plasma Exosomal CircNEK9 accelerates the progression of gastric
cancer via miR-409-3p/MAP7 axis. Dig Dis Sci. 66:4274–4289. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Fan L, Li W and Jiang H: Circ_0000395
Promoted CRC Progression via Elevating MYH9 Expression by
Sequestering miR-432-5p. Biochem Genet. 61:116–137. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Gao L, Tang X, He Q, Sun G, Wang C and Qu
H: Exosome-transmitted circCOG2 promotes colorectal cancer
progression via miR-1305/TGF-β2/SMAD3 pathway. Cell Death Discov.
7:2812021. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Lu J, Wang YH, Yoon C, Huang XY, Xu Y, Xie
JW, Wang JB, Lin JX, Chen QY, Cao LL, et al: Circular RNA
circ-RanGAP1 regulates VEGFA expression by targeting miR-877-3p to
facilitate gastric cancer invasion and metastasis. Cancer Lett.
471:38–48. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Hui C, Tian L and He X: Circular RNA
circNHSL1 contributes to gastric cancer progression through the
miR-149-5p/YWHAZ axis. Cancer Manag Res. 12:7117–7130. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Shen X, Kong S, Ma S, Shen L, Zheng M, Qin
S, Qi J, Wang Q, Cui X and Ju S: Hsa_circ_0000437 promotes
pathogenesis of gastric cancer and lymph node metastasis. Oncogene.
41:4724–4735. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Han K, Wang FW, Cao CH, Ling H, Chen JW,
Chen RX, Feng ZH, Luo J, Jin XH, Duan JL, et al: CircLONP2 enhances
colorectal carcinoma invasion and metastasis through modulating the
maturation and exosomal dissemination of microRNA-17. Mol Cancer.
19:602020. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Chen C, Liu Y, Liu L, Si C, Xu Y, Wu X,
Wang C, Sun Z and Kang Q: Exosomal circTUBGCP4 promotes vascular
endothelial cell tipping and colorectal cancer metastasis by
activating Akt signaling pathway. J Exp Clin Cancer Res. 42:462023.
View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Yu Q, Zhang Y, Tian Y, Peng A, Cui X, Ding
B, Yang L, Liu Y, Ju Y and Gao C: Exosomal Circ_FMN2 derived from
the serum of colorectal cancer patients promotes cancer progression
by miR-338-3p/MSI1 Axis. Appl Biochem Biotechnol. Mar 30–2023.(Epub
ahead of print). View Article : Google Scholar
|
|
92
|
Wei C, Yang C, Wang S, Shi D, Zhang C, Lin
X, Liu Q, Dou R and Xiong B: Crosstalk between cancer cells and
tumor associated macrophages is required for mesenchymal
circulating tumor cell-mediated colorectal cancer metastasis. Mol
Cancer. 18:642019. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Li YF, Pei FL and Cao MZ: CircRNA_101951
promotes migration and invasion of colorectal cancer cells by
regulating the KIF3A-mediated EMT pathway. Exp Ther Med.
19:3355–3361. 2020.PubMed/NCBI
|
|
94
|
Zhou LH, Yang YC, Zhang RY, Wang P, Pang
MH and Liang LQ: CircRNA_0023642 promotes migration and invasion of
gastric cancer cells by regulating EMT. Eur Rev Med Pharmacol Sci.
22:2297–2303. 2018.PubMed/NCBI
|
|
95
|
Liang ZF, Zhang Y, Guo W, Chen B, Fang S
and Qian H: Gastric cancer stem cell-derived exosomes promoted
tobacco smoke-triggered development of gastric cancer by inducing
the expression of circ670. Med Oncol. 40:242022. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Miao Z, Zhao X and Liu X: Exosomal
circCOL1A2 from cancer cells accelerates colorectal cancer
progression via regulating miR-665/LASP1 signal axis. Eur J
Pharmacol. 950:1757222023. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Zhao H, Chen S and Fu Q: Exosomes from
CD133+ cells carrying circ-ABCC1 mediate cell stemness and
metastasis in colorectal cancer. J Cell Biochem. 121:3286–3297.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Liu C, Yang J, Zhu F, Zhao Z and Gao L:
Exosomal circ_0001190 regulates the progression of gastric cancer
via miR-586/SOSTDC1 Axis. Biochem Genet. 60:1895–1913. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Zeng W, Liu Y, Li WT, Li Y and Zhu JF:
CircFNDC3B sequestrates miR-937-5p to derepress TIMP3 and inhibit
colorectal cancer progression. Mol Oncol. 14:2960–2984. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Song J, Xu X, He S, Wang N, Bai Y, Li B
and Zhang S: Exosomal hsa_circ_0017252 attenuates the development
of gastric cancer via inhibiting macrophage M2 polarization. Hum
Cell. 35:1499–1511. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Shang A, Gu C, Wang W, Wang X, Sun J, Zeng
B, Chen C, Chang W, Ping Y, Ji P, et al: Exosomal circPACRGL
promotes progression of colorectal cancer via the
miR-142-3p/miR-506-3p-TGF-β1 axis. Mol Cancer. 19:1172020.
View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Zheng Y, Li Z, Wang Y, Chen W, Lin Y, Guo
J and Ye G: CircRNA: A new class of targets for gastric cancer drug
resistance therapy. Pathol Oncol Res. 29:16110332023. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Yu D, Chang Z, Liu X, Chen P, Zhang H and
Qin Y: Macrophage-derived exosomes regulate gastric cancer cell
oxaliplatin resistance by wrapping circ 0008253. Cell Cycle.
22:705–717. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Chen Y, Liu H, Zou J, Cao G, Li Y, Xing C
and Wu J: Exosomal circ_0091741 promotes gastric cancer cell
autophagy and chemoresistance via the
miR-330-3p/TRIM14/Dvl2/Wnt/β-catenin axis. Hum Cell. 36:258–275.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Yang G, Tan J, Guo J, Wu Z and Zhan Q:
Exosome-mediated transfer of circ_0063526 enhances cisplatin
resistance in gastric cancer cells via regulating miR-449a/SHMT2
axis. Anticancer Drugs. 33:1047–1057. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Yao W, Guo P, Mu Q and Wang Y:
Exosome-Derived Circ-PVT1 contributes to cisplatin resistance by
regulating autophagy, invasion, and apoptosis Via miR-30a-5p/YAP1
axis in gastric cancer cells. Cancer Biother Radiopharm.
36:347–359. 2021.PubMed/NCBI
|
|
107
|
Pan Z, Zheng J, Zhang J, Lin J, Lai J, Lyu
Z, Feng H, Wang J, Wu D and Li Y: A novel protein encoded by
exosomal CircATG4B induces oxaliplatin resistance in colorectal
cancer by promoting autophagy. Adv Sci (Weinh). 9:e22045132022.
View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Zhang Y, Tan X and Lu Y: Exosomal transfer
of circ_0006174 contributes to the chemoresistance of doxorubicin
in colorectal cancer by depending on the miR-1205/CCND2 axis. J
Physiol Biochem. 78:39–50. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Wang X, Zhang H, Yang H, Bai M, Ning T,
Deng T, Liu R, Fan Q, Zhu K, Li J, et al: Exosome-delivered circRNA
promotes glycolysis to induce chemoresistance through the
miR-122-PKM2 axis in colorectal cancer. Mol Oncol. 14:539–555.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Zhao K, Cheng X, Ye Z, Li Y, Peng W, Wu Y
and Xing C: Exosome-Mediated Transfer of circ_0000338 Enhances
5-Fluorouracil resistance in colorectal cancer through regulating
MicroRNA 217 (miR-217) and miR-485-3p. Mol Cell Biol. 41:e00517–20.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Kobayashi H, Enomoto A, Woods SL, Burt AD,
Takahashi M and Worthley DL: Cancer-associated fibroblasts in
gastrointestinal cancer. Nat Rev Gastroenterol Hepatol. 16:282–295.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Xu R, Greening DW, Zhu HJ, Takahashi N and
Simpson RJ: Extracellular vesicle isolation and characterization:
Toward clinical application. J Clin Invest. 126:1152–1162. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Colombo M, Raposo G and Théry C:
Biogenesis, secretion, and intercellular interactions of exosomes
and other extracellular vesicles. Annu Rev Cell Dev Biol.
30:255–289. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Kristensen LS, Andersen MS, Stagsted LVW,
Ebbesen KK, Hansen TB and Kjems J: The biogenesis, biology and
characterization of circular RNAs. Nat Rev Genet. 20:675–691. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Wang S, Zhang K, Tan S, Xin J, Yuan Q, Xu
H, Xu X, Liang Q, Christiani DC, Wang M, et al: Circular RNAs in
body fluids as cancer biomarkers: The new frontier of liquid
biopsies. Mol Cancer. 20:132021. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Li XN, Wang ZJ, Ye CX, Zhao BC, Li ZL and
Yang Y: RNA sequencing reveals the expression profiles of circRNA
and indicates that circDDX17 acts as a tumor suppressor in
colorectal cancer. J Exp Clin Cancer Res. 37:3252018. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Chen RX, Chen X, Xia LP, Zhang JX, Pan ZZ,
Ma XD, Han K, Chen JW, Judde JG, Deas O, et al: N6-methyladenosine
modification of circNSUN2 facilitates cytoplasmic export and
stabilizes HMGA2 to promote colorectal liver metastasis. Nat
Commun. 10:46952019. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Zheng P, Gao H, Xie X and Lu P: Plasma
Exosomal hsa_circ_0015286 as a potential diagnostic and prognostic
biomarker for gastric cancer. Pathol Oncol Res. 28:16104462022.
View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Li R, Tian X, Jiang J, Qian H, Shen H and
Xu W: CircRNA CDR1as: a novel diagnostic and prognostic biomarker
for gastric cancer. Biomarkers. 28:448–457. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Tao X, Shao Y, Lu R, Ye Q, Xiao B, Ye G
and Guo J: Clinical significance of hsa_circ_0000419 in gastric
cancer screening and prognosis estimation. Pathol Res Pract.
216:1527632020. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Pan B, Qin J, Liu X, He B, Wang X, Pan Y,
Sun H, Xu T, Xu M, Chen X, et al: Identification of Serum Exosomal
hsa-circ-0004771 as a novel diagnostic biomarker of colorectal
cancer. Front Genet. 10:10962019. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Xie Y, Li J, Li P, Li N, Zhang Y, Binang
H, Zhao Y, Duan W, Chen Y, Wang Y, et al: RNA-Seq profiling of
serum exosomal circular RNAs Reveals Circ-PNN as a potential
biomarker for human colorectal cancer. Front Oncol. 10:9822020.
View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Li T, Zhou T, Wu J, Lv H, Zhou H, Du M,
Zhang X, Wu N, Gong S, Ren Z, et al: Plasma exosome-derived
circGAPVD1 as a potential diagnostic marker for colorectal cancer.
Transl Oncol. 31:1016522023. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Wang H, Zeng X, Zheng Y, Wang Y and Zhou
Y: Exosomal circRNA in digestive system tumors: the main player or
coadjuvants? Front Oncol. 11:6144622021. View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Lu L, Fang S, Zhang Y, Jin L, Xu W and
Liang Z: Exosomes and Exosomal circRNAs: The rising stars in the
progression, diagnosis and prognosis of gastric cancer. Cancer
Manag Res. 13:8121–8129. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Zhang Z, Sun C, Zheng Y and Gong Y:
circFCHO2 promotes gastric cancer progression by activating the
JAK1/STAT3 pathway via sponging miR-194-5p. Cell Cycle.
21:2145–2164. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Li S, Li J, Zhang H, Zhang Y, Wang X, Yang
H, Zhou Z, Hao X, Ying G and Ba Y: Gastric cancer derived exosomes
mediate the delivery of circRNA to promote angiogenesis by
targeting miR-29a/VEGF axis in endothelial cells. Biochem Biophys
Res Commun. 560:37–44. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Li Y, Hu J, Wang M, Yuan Y, Zhou F, Zhao
H, Qiu T and Liang L: Exosomal circPABPC1 promotes colorectal
cancer liver metastases by regulating HMGA2 in the nucleus and
BMP4/ADAM19 in the cytoplasm. Cell Death Discov. 8:3352022.
View Article : Google Scholar : PubMed/NCBI
|
|
129
|
Yi Q, Yue J, Liu Y, Shi H and Sun W, Feng
J and Sun W: Recent advances of exosomal circRNAs in cancer and
their potential clinical applications. J Transl Med. 21:5162023.
View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Guo Z, Zhang Y, Xu W, Zhang X and Jiang J:
Engineered exosome-mediated delivery of circDIDO1 inhibits gastric
cancer progression via regulation of MiR-1307-3p/SOCS2 Axis. J
Transl Med. 20:3262022. View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Zhang C, Wei G, Zhu X, Chen X, Ma X, Hu P,
Liu W, Yang W, Ruan T, Zhang W, et al: Exosome-Delivered circSTAU2
inhibits the progression of gastric cancer by targeting the
miR-589/CAPZA1 Axis. Int J Nanomedicine. 18:127–142. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Yu L, Zhang F and Wang Y: Circ_0005615
regulates the progression of colorectal cancer through the
miR-873-5p/FOSL2 signaling pathway. Biochem Genet. 61:2020–2041.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Meng X, Yang D, Zhang B, Zhao Y, Zheng Z
and Zhang T: Regulatory mechanisms and clinical applications of
tumor-driven exosomal circRNAs in cancers. Int J Med Sci.
20:818–835. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Lei T, Zhang Y, Wang X, Liu W, Feng W and
Song W: Integrated analysis of the functions and clinical
implications of exosome circRNAs in colorectal cancer. Front
Immunol. 13:9190142022. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Zhang F, Jiang J, Qian H, Yan Y and Xu W:
Exosomal circRNA: Emerging insights into cancer progression and
clinical application potential. J Hematol Oncol. 16:672023.
View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Lin S, Yu Z, Chen D, Wang Z, Miao J, Li Q,
Zhang D, Song J and Cui D: Progress in microfluidics-based exosome
separation and detection technologies for diagnostic applications.
Small. 16:e19039162020. View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Chen L, Wang L, Zhu L, Xu Z, Liu Y, Li Z,
Zhou J and Luo F: Exosomes as drug carriers in anti-cancer therapy.
Front Cell Dev Biol. 10:7286162022. View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Ding L, Yang X, Gao Z, Effah CY, Zhang X,
Wu Y and Qu L: A holistic review of the state-of-the-art
microfluidics for exosome separation: An overview of the current
status, existing obstacles, and future outlook. Small.
17:e20071742021. View Article : Google Scholar : PubMed/NCBI
|