Introduction
More than 30 years ago, eukaryotic translation
initiation factor 6 (eIF6) was first identified as a protein in
wheat germ (1). This protein
functions as an anti-association factor to interact with the 60S
ribosome, and prevent the assembly of the 60S and 40S subunits in
the cytoplasm (2–5). eIF6 is found in yeast and mammalian
cells, and the majority of eIF6 is located in the cytoplasm of
mammalian cells (4,5). Originally, eIF6 was first observed in
the proliferating compartment of the colonic epithelium and stem
cells (6), and is also highly
expressed in epithelial and embryonic tissues (7–9).
Furthermore, an increasing number of studies have demonstrated that
eIF6 is overexpressed in human cancer (6,10–15). Accumulating evidence suggests that
eIF6 is a useful biomarker in cancer diagnosis, and that it serves
as an anti-cancer molecular target. However, the specific role of
eIF6 in tumorigenesis remains to be elucidated.
The present review focuses on eIF6-associated
studies, particularly those pertaining to its subcellular location,
phosphorylation and dephosphorylation, roles in cancer and
molecular mechanisms in oncogenesis.
Subcellular localization of eIF6
eIF6, also known as integrin β4 binding protein,
p27BBP or β4 integrin interactor, is a remarkably conserved protein
from yeast to mammals (7,13–14). In
yeast, eIF6 is primarily localized in the nucleolus (9–10). By
contrast, in mammalian cells, the majority of eIF6 is present in
the cytoplasm, with a smaller but significant fraction (~30%)
located in the nucleus (4,16–18).
Notably, eIF6 is located in the nucleolus of certain cell lines,
such as HeLa, A431, NIH/3T3 fibroblasts and Jurkat T cells
(8), in addition to neoplastic
tissues, including colonic adenoma and carcinoma (6). Previous studies have demonstrated that
eIF6 functions as a component of the preribosomal particles in the
nucleolus, thus serving an important role in 60S ribosome
biogenesis (8,18). In the cytoplasm, eIF6 functions as a
translation factor (9), therefore,
subcellular localization is crucial for the functional regulation
of eIF6.
Phosphorylation and dephosphorylation of
eIF6
In mammalian cells, eIF6 regulates ribosomal
assembly and biogenesis, thus controlling the binding of 40S and
60S ribosomal subunits and participating in 80S assembly (7–9,16). As described above, eIF6 is present in
the nucleus and cytoplasm (9,17). Notably, it is reported that
nucleocytoplasmic shuttling is caused by the phosphorylation of
eIF6, in line with the well-established hypothesis that
phosphorylation is able to regulate the biological activity of
numerous proteins (9). Therefore,
phosphorylation is likely to modulate eIF6 activity, and three
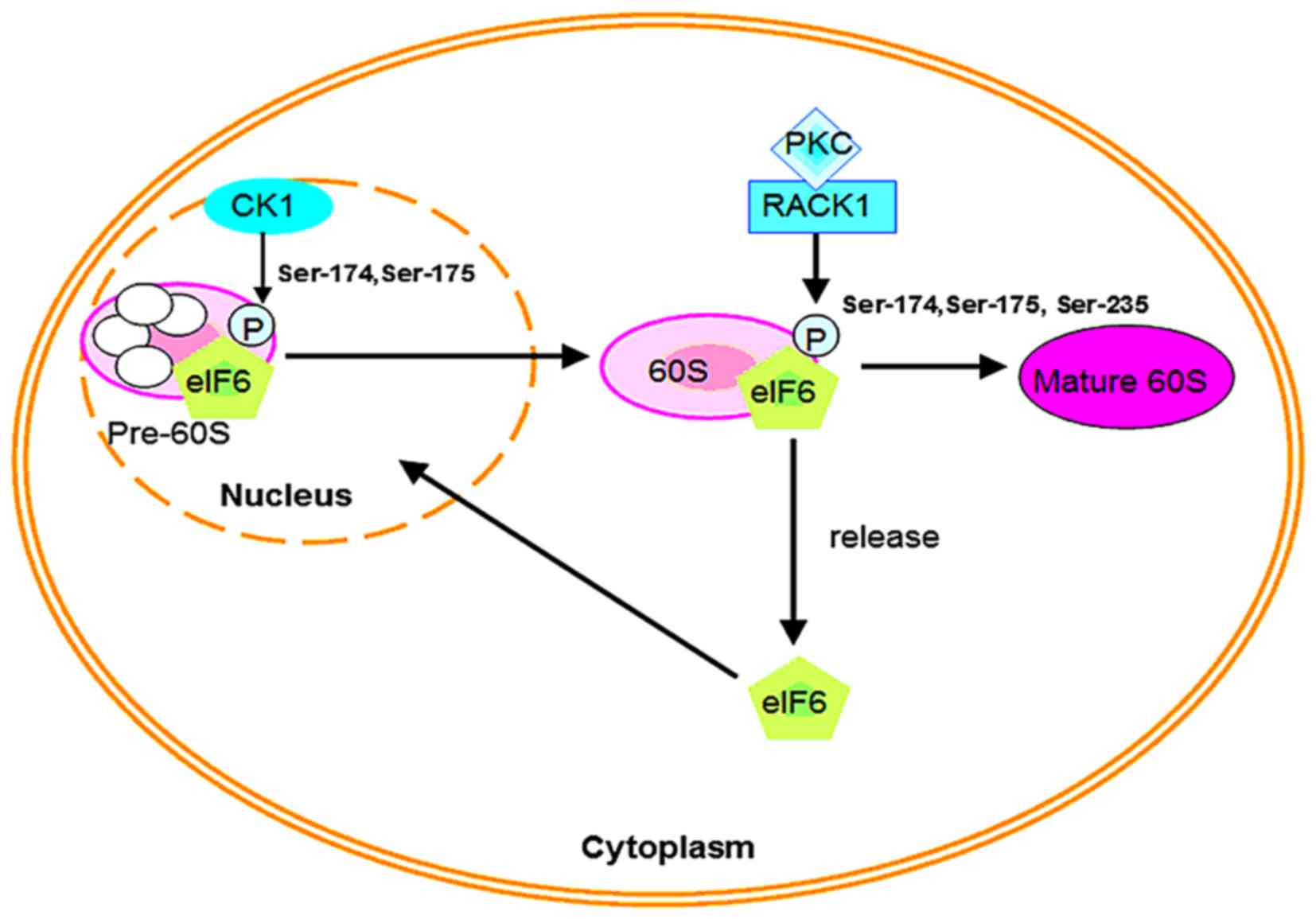
potential phosphorylation sites have been identified (18). For nuclear export in mammalian cells,
eIF6 is phosphorylated in vitro at Ser-175 and Ser-174 by
the nuclear isoforms of casein kinase (CK) CK1α or CK1δ, thereby
promoting the formation of pre-60S ribosomal particles in the
cytoplasm. In addition, the Ca2+/calmodulin-dependent
protein phosphatase calcineurin mediates dephosphorylation, which
facilitates migration of eIF6 back to the nucleolus and continues
60S ribosome biogenesis (18). Such
evidence implies that CK1 controls the subcellular distribution of
eIF6.
Although CK1 is widely found in the nucleus,
cytoplasm, cell membrane and cytoskeleton of mammalian and yeast
cells (19–21), it is unclear whether extranuclear CK1
enters the nucleus to regulate the export of eIF6. It should be
noted that cytoplasmic eIF6 in mammalian cells is also
phosphorylated by receptors for activated C kinase 1
(RACK1)/protein kinase C (PKC) signaling at positions Ser-174,
Ser-175 and Ser-235 (Fig. 1)
(16,22). These procedures result in dissociation
of eIF6 from the 60S subunit, thus aiding its maturation (18). Recent research demonstrates that
GTPase elongation factor-like 1 (EFL1) is involved in the
cytoplasmic maturation of the ribosomal 60S subunit (3). SBDS, the protein mutated in
Shwachman-Bodian-Diamond syndrome, and EFL1 release the
anti-association factor eIF6 from the surface of the 60S subunit
(2,5).
In addition, the Ser235 PKC phosphorylation site has also been
identified in the Xenopus eIF6 protein (23).
However, there is little or no evidence to verify
whether CK1 and the RACK1-PKC complex phosphorylate the Ser-174 and
Ser-175 sites of eIF6 at the same time. Moreover, an increasing
number of studies have demonstrated that eIF6 is highly
overexpressed in tumor cells (8–11). The
C-terminal of eIF6 is subject to RACK1-PKCβII complex
phosphorylation at Ser-235, which modulates the protumorigenic
activity of eIF6 (16), whereas
mutation of the phosphorylation site at Ser235 of eIF6 in mouse
models reduces translation and lymphomagenesis (4). A previous study demonstrated that the
Ras cascade, which regulates phosphorylation of eIF6, is triggered
by agonists of phorbol esters (16).
Therefore, it may be speculated that the Ras cascade recruits
PKCβII and phosphorylates eIF6 at Ser235, and the activity of eIF6
leads to increased translation and tumorigenesis.
Overexpression of eIF6 in human
carcinoma
Numerous studies have demonstrated highly aberrant
expression of eIF6 in human cancer (10–15).
Although the function of eIF6 is not fully understood, differential
expression of eIF6 is correlated with cancer pathogenesis, and eIF6
functions as a regulator in cancer development (6,10–15). The cancer tissues and cell lines in
which eIF6 is overexpressed are presented in Table I. In this section, the potential of
eIF6 as a cancer biomarker is discussed.
 | Table I.Overexpression of eIF6 in various
cancer tissues and cell lines. |
Table I.
Overexpression of eIF6 in various
cancer tissues and cell lines.
| Type | Overexpression of
eIF6 |
|---|
| Cancer tissues | Colorectal cancer,
head and neck carcinoma, NSCLC, ovarian serous adenocarcinoma |
| Cancer cell
lines | A2780 ovarian
cancer cells, WM793 primary melanoma cells, SW480 colorectal cancer
cells |
Colorectal cancer
eIF6 is regarded as a nuclear matrix protein that
accumulates in nucleoli (8), and is
found in the cytoplasm of glandular crypt cells in the human
colonic epithelium (6). However,
higher levels of eIF6 are observed in colorectal carcinoma compared
with colorectal precancer and normal mucosa (6–7,16). Consequently, there is a progressive
increase of eIF6 from normal tissue to dysplastic adenoma and
carcinoma. This raises the question of which mechanisms are
involved in the increased expression of eIF6 protein. It is
hypothesized that eIF6 is upregulated at the transcriptional level,
such that the mRNA coding for eIF6 is highly concentrated in tumors
relative to normal colorectal tissues (6). mRNA translation controls distinct
cellular processes, including tumorigenesis, cell migration,
adhesion and growth, and cell-cycle control (24). Notably, gross gene expression of eIF6
is less well known. Therefore, further research is required to
understand the underlying reasons for this.
As a marker of cell proliferation, the distribution
of argyrophilic nucleolar organizer region (AgNOR)-associated
proteins in the nucleolus and cell correspond to proteins located
in the nucleolar organizer regions. Previously, nucleolar staining
by AgNORs was considered to be a prognostic marker of malignancy
(25). Moreover, the value of AgNORs
as proliferation markers has been reported in various forms of
cancer, such as breast, ovarian, cervical, prostate,
hepatocellular, papillary thyroid, gastric and bladder cancer
(25–31). Certain studies have demonstrated a
correlation between AgNOR count in tumors and various clinical
parameters, including tumor size and staging, and distant
metastasis (28–32). Therefore, eIF6 may be used as a
diagnostic tool on the basis of the function of AgNORs. In
addition, differentially-expressed eIF6 may serve a critical
function in colon carcinogenesis and provide a novel marker in
surgical pathology.
Head and neck carcinoma
eIF6 is overexpressed in colorectal cancer (6). Similarly, in head and neck carcinoma,
the expression of eIF6 is also higher than that observed in normal
mucosa (13). Additionally, nucleolar
overexpression of eIF6 has been detected in head and neck
metastatic carcinoma (13). Head and
neck cancer has previously been reported as the sixth leading type
of cancer worldwide, accounting for ~6% of all tumors, of which
>90% are head and neck squamous cell carcinoma (33,34).
Despite advances in treatment, the prognosis remains poor.
Therefore, the discovery of molecular markers is not only important
for understanding the pathogenesis of head and neck cancer, but may
also provide further insight into tumor biology, diagnosis,
therapeutic perspectives and prognosis (34,35).
eIF6 is highly concentrated in nucleoli, is easily
observed and its overexpression is not difficult to measure. eIF6
may function as a molecular marker for use in surgical pathological
diagnosis. Notably, a larger 52-kDa protein, detected by eIF6
antibody, is also observed in lymph node metastases (13). This larger protein has tissue
specificity due to its absence in samples of colorectal carcinoma,
parotid gland adenocarcinoma and leiomyosarcoma of the larynx
(13). Consequently, this 52-kDa band
is able to be utilized by head and neck surgeons and surgical
pathologists during diagnosis.
Non-small cell lung cancer
(NSCLC)
Lung cancer is an extraordinarily malignant tumor
with the highest morbidity and mortality, of which the most common
variant is NSCLC (36,37). The primary features of this cancer are
invasion and metastasis (36–39). eIF6 interacts with the cytoplasmic
integrin β4 subunit, and in a previous study, positive eIF6
staining was observed in 82.5% (66/80) of NSCLC specimens (37). Therefore, eIF6 is likely to be present
at a high concentration in NSCLC (6,13).
Integrin β4 subunit, α6β4, the receptor for the basement membrane
protein laminin-5, is an important cellular adhesion molecule, and
is closely associated with tumor invasion and metastasis (6,40). α6β4
integrin is expressed in invasive breast carcinomas and is a
potential indicator of poor prognosis (40). Taken together, a large increase in
eIF6 is apparent in NSCLC, which may promote the migration of NSCLC
cells; however, further study is required to confirm this.
Ovarian serous adenocarcinoma
Ovarian serous adenocarcinoma is the most prevalent
form of epithelial ovarian cancer and a fatal type of gynecological
malignancy (41,42). Human eIF6 is located on chromosome
20q12, which is an amplified chromosomal region (20q12-12) in
ovarian cancer (43). This suggests
that increased eIF6 may be a consequence of increased protein
turnover in rapidly proliferating malignant cells based upon its
role in ribosome assembly. Notably, eIF6, Dicer and RNaseIII
endonuclease, which are essential components of miRNA machinery,
are overexpressed in ovarian serous adenocarcinoma and associated
with its clinicopathological features (15). miRNAs are a class of small, noncoding
RNAs that affect the post-transcriptional control of mRNA and
contribute to human tumorigenesis (44–46). Low
eIF6 expression has increasingly been associated with reduced the
probability of disease-free survival (15). Therefore, it is not inconceivable that
downregulated expression of miRNAs and eIF6 could be useful
biomarkers for the prediction of ovarian serous adenocarcinoma.
Additionally, eIF6 and proteins of the miRNA machinery are closely
related to future RNA interference-based therapy.
Upstream modulator of eIF6
As aforementioned, eIF6 is overexpressed in human
colorectal cancer (6), head and neck
cancer (13), lung cancer (14) and ovarian serous adenocarcinoma
(15). Therefore, it is necessary to
determine which oncogenes in the transcriptional network control
eIF6 expression during tumorigenesis. Previous studies have
established that the transcription factor complex GA-binding
protein (GABP) regulates eIF6 expression, as the eIF6 promoter
region contains GABP-binding sites (47). GABP is an E26 transformation-specific
sequence (ETS) transcription factor, which contains an unrelated
GABP protein, an ETS DNA-binding domain and a nuclear localization
signal (48). The transcription of
nuclear genes involved in mitochondrial respiration is controlled
by the GABP complex (48). Moreover,
certain ribosomal proteins are also GABP targets (49,50). For
these reasons, GABP may be essential in regulating the
transcription of ribosomal genes. The activity of the eIF6 promoter
could be inhibited through blocking endogenous GABP activity. To
date, a possible function for GABP in tumorigenesis remains to be
described. Accounting for the fact that GABP could be vital in
mediating the proliferative response, it may be useful to determine
whether certain oncogenes directly affect GABP expression.
It is worth noting that the Notch-1 receptor has
been demonstrated to directly regulate transcription of the eIF6
gene (12). The Notch-1 receptor
belongs to the Notch family of transmembrane proteins in mammals
(51). The highly-conserved Notch
signaling pathway is essential in the regulation of various
physiological processes, including cell development,
differentiation and proliferation (52–55). In
particular, activation of the canonical Notch-1 pathway is of major
significance in human tumorigenesis (12,51,56). A
high level of Notch-1 expression has been observed in salivary
adenoid cystic carcinoma (56) and
breast cancer (57). Notably, it was
reported that Notch-1 activation resulted in a 2 to 3-fold
overexpression of eIF6, thus enhancing the invasiveness of A2780
cells (12,57). Therefore, it is conceivable that the
Notch-1 signal is key to control the expression of eIF6. Notch-1
functions as an upstream regulator of eIF6, which directly
regulates eIF6 expression via a recombinant binding protein
suppressor of Hairless (RBP-Jκ)-dependent mechanism. In other
words, the Notch-1/RBP-Jκ signaling pathway stimulates eIF6
promoter activity, resulting in abnormal expression of eIF6
(57,12). Overexpression of eIF6 enhances cell
migration and invasiveness, but it is noteworthy that it does not
affect proliferation (12).
Downstream regulation of eIF6
eIF6 and the canonical Wnt/β-catenin
signaling pathway
In previous studies, eIF6 has primarily been used to
control translation through regulation of ribosome biogenesis and
assembly (3,4,9,18). Further research using yeast two-hybrid
assays has demonstrated that eIF6 interacts with the C terminus of
β-catenin, functioning as a transcriptional activation domain
(7,55). In addition, the Wnt signal
transduction cascade, with β-catenin as a major transducer, is a
canonical cellular pathway in cell adhesion and proliferation
during embryogenesis in animals (58–61). In
general, the Wnt signal is absent in normal cells or tissues.
However, the aberrant activation of Wnt/β-catenin signaling leads
to the dysregulation of cellular growth and development, and
contributes to human tumorigenesis (58,60). The
targets of β-catenin transcription are also overexpressed in
various types of carcinoma (59,63).
Although the molecular mechanism remains to be clarified, previous
research has demonstrated that dysregulation of Wnt/β-catenin
signaling results in large accumulation of β-catenin in the nucleus
(62,63). Subsequently, combined with T cell
factor/lymphoid enhancing factor (TCF/LEF), transcription of target
genes, including c-Myc (64) and
cyclin D1 (65), may be activated
resulting in carcinogenesis. Previous research has demonstrated
that eIF6 serves as a factor participating in Wnt/β-catenin
signaling and the distribution of eIF6 and β4 is altered in colonic
adenoma and carcinoma (6).
Furthermore, in SW480 cells transfected with full-length eIF6, the
level of activated β-catenin was reduced compared with controls
(66). The question may therefore be
raised as to whether eIF6 has the same effect as Dickkopf
antagonists on the Wnt/β-catenin signaling pathway. However, this
is problematic to answer as eIF6 is overexpressed in colorectal
carcinoma (6). Moreover, MG132, a
proteasome-specific inhibitor, fails to inhibit the decrease in
β-catenin that occurs upon overexpression of yellow fluorescent
protein-eIF6 in SW480 cells (66).
Consequently, despite the fact that β-catenin functions as a
downstream effector of eIF6, eIF6 expression does not directly
regulate the level of β-catenin, indicating that downregulation of
β-catenin may only exist in certain vectors transfected with cell
lines overexpressing eIF6.
Downstream effector of eIF6: Cell
division control protein 42 (Cdc42)
In a previous study, several membrane-associated
proteins differed in abundance upon eIF6 overexpression in A2780
ovarian cancer cells (11). This
effect is particularly notable in Cdc42 (11), a small GTPase belonging to the Ras
homolog family (67–69). A number of studies have established
that Cdc42 regulates cell differentiation, cell cycle progression,
cell polarity, cell fate determination, and cell motility and
adhesion (68,70). Aberrant expression of Cdc42 is pivotal
in tumorigenesis, including that of breast carcinoma (71). Notably, it was reported that Cdc42
expression is disrupted at the post-transcriptional level by
enhanced levels of eIF6 in A2780 ovarian cancer cells (11). In addition, it was observed that
downstream of eIF6 activation, Cdc42 levels are increased by a
post-transcriptional mechanism (11).
Although the underlying mechanisms of eIF6-mediated
Cdc42 expression remain to be elucidated, a possible theory may be
that enhanced levels of eIF6 indirectly control the variation in
the abundance of Cdc42. This is supported by the fact that Cdc42
mRNA expression levels exhibit little or no difference following
eIF6 overexpression (11), which also
demonstrates that eIF6 may target the translation of specific
mRNAs. In A2780 cells overexpressing eIF6, ML-141, a selective and
potent inhibitor of Cdc42 GTPase, has been demonstrated to
significantly decrease migratory activity (11). The tumor-promoting ability of eIF6 is
not restricted to the A2780 cell line; the primary melanoma cell
line WM793 has also been reported to exhibit upregulated Cdc42
expression, in addition to increased motility and invasiveness
(11). Therefore, eIF6 is crucial for
Cdc42 upregulation. As eIF6 affects Cdc42 translation in ovarian
cancer cells, this indicates that the increased expression of eIF6
is more likely to cause Cdc42 activation in ovarian cancer tissue,
which in turn is accountable for increased migration and invasion.
Nevertheless, further studies are required to elucidate the
mechanisms behind these processes.
Conclusions and perspectives
The protein eIF6 possesses a high degree of
evolutionary sequence conservation (1–8), and is
located subcellularly in the nucleolus and cytoplasm.
Phosphorylation of eIF6 regulates nucleocytoplasmic shuttling in
mammalian cells and involves the release of eIF6 from the 60S
ribosome subunit (3,16,17). In
cancer cells, eIF6 is phosphorylated by the RACK1-PKCβII complex,
and thus by the Ras cascade (16,72). eIF6
functions as an important component in gene regulatory networks and
exerts crucial roles in neoplastic progression (10–15).
Nevertheless, the specific molecular mechanisms underlying the role
of eIF6 in these processes remain unclear. Oncogenesis typically
involves several different signaling pathways. For example, the
Ras-extracellular related kinase mitogen-activated protein kinase
pathway and phosphoinositide 3-kinase/AKT/mammalian target of
rapamycin pathway each take part in the phosphorylation of eIF4E,
which is involved in cancer development (73–75).
Consequently, it is possible that these signals are involved in the
mechanisms of eIF6 overexpression in cancer, since eIF4E is a
eukaryotic initiation factor in addition to eIF6.
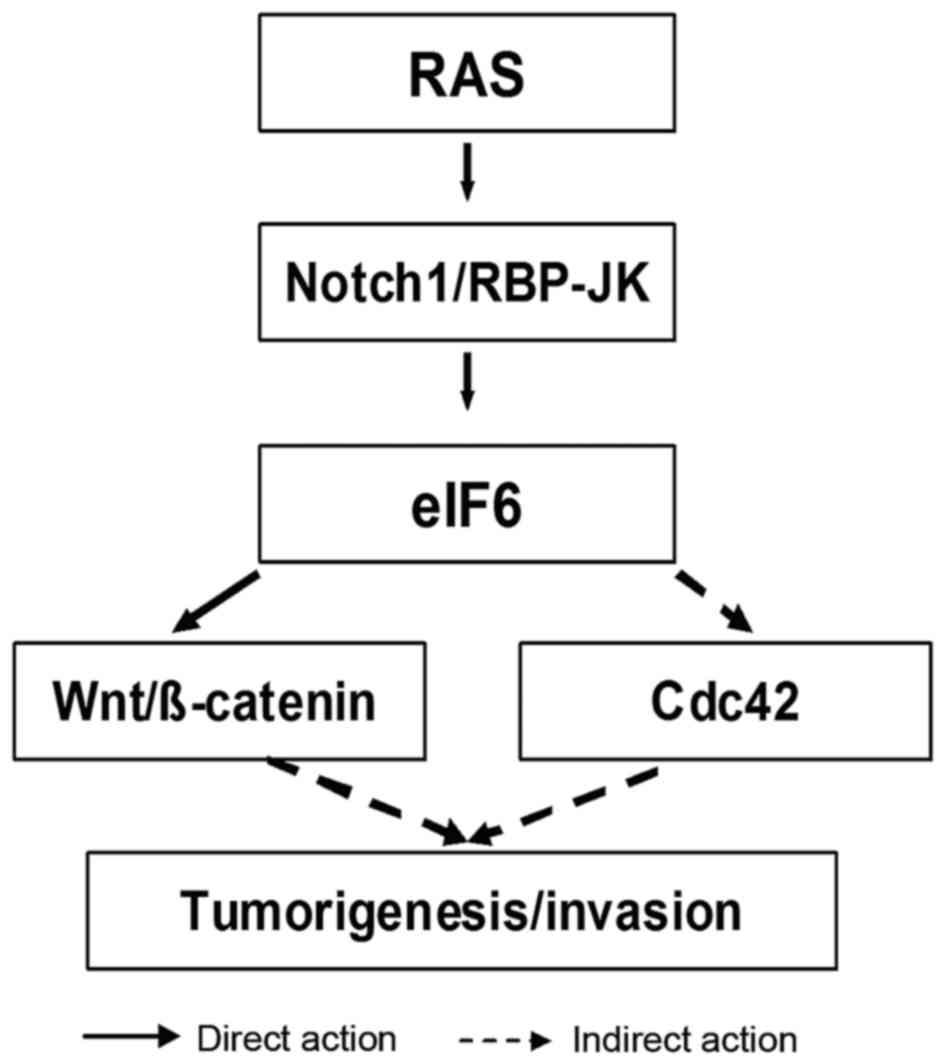
In conclusion, the following hypothesis is proposed
(Fig. 2). Firstly, Notch-1 activated
by oncogenic Ras promotes transcription of the eIF6 gene through an
RBP-Jκ-dependent mechanism. Ras signaling has a key role in
increasing Notch-1 expression in breast carcinoma (75). Secondly, overexpression of eIF6 leads
to aberrant activation of the Wnt/β-catenin signaling pathway.
Similarly, overexpressed eIF6 controls its downstream effector
Cdc42, which in turn affects tumorigenesis. As a consequence,
understanding the signaling network in which eIF6 lies may
contribute novel insights into the pathogenesis of cancer, and
offer a promising target for the development of novel
antineoplastic agents.
Acknowledgements
The authors acknowledge support from the National
Natural Science Foundation of China, (grant no. 81472275) and the
Natural Science Foundation of Guangdong Province, China (grant no.
2014A030313542).
References
|
1
|
Russell DW and Spremulli LL: Mechanism of
action of the wheat germ ribosome dissociation factor: Interaction
with the 60 S subunit. Arch Biochem Biophys. 201:518–526. 1980.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Finch AJ, Hilcenko C, Basse N, Drynan LF,
Goyenechea B, Menne TF, González Fernández A, Simpson P, D'Santos
CS, Arends MJ, et al: Uncoupling of GTP hydrolysis from eIF6
release on the ribosome causes Shwachman-Diamond syndrome. Genes
Dev. 25:917–929. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Gartmann M, Blau M, Armache JP, Mielke T,
Topf M and Beckmann R: Mechanism of eIF6-mediated inhibition of
ribosomal subunit joining. J Biol Chem. 285:14848–14851. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Miluzio A, Beugnet A, Grosso S, Brina D,
Mancino M, Campaner S, Amati B, de Marco A and Biffo S: Impairment
of cytoplasmic eIF6 activity restricts lymphomagenesis and tumor
progression without affecting normal growth. Cancer Cell.
19:765–775. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
García-Márquez A, Gijsbers A, de la Mora E
and Sánchez-Puig N: Defective guanine nucleotide exchange in the
Elongation Factor-Like 1 (EFL1) GTPase by mutations in the
Shwachman-Diamond syndrome protein. J Biol Chem. 290:17669–17678.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Sanvito F, Vivoli F, Gambini S,
Santambrogio G, Catena M, Viale E, Veglia F, Donadini A, Biffo S
and Marchisio PC: Expression of a highly conserved protein, p27BBP,
during the progression of human colorectal cancer. Cancer Res.
60:510–516. 2000.PubMed/NCBI
|
|
7
|
Biffo S, Sanvito F, Costa S, Preve L,
Pignatelli R, Spinardi L and Marchisio PC: Isolation of a novel
beta4 integrin-binding protein (p27 (BBP)) highly expressed in
epithelial cells. J Biol Chem. 272:30314–30321. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sanvito F, Piatti S, Villa A, Bossi M,
Lucchini G, Marchisio PC and Biffo S: The beta4 integrin interactor
p27 (BBP/eIF6) is an essential nuclear matrix protein involved in
60S ribosomal subunit assembly. J Cell Biol. 144:823–837. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Gandin V, Miluzio A, Barbieri AM, Beugnet
A, Kiyokawa H, Marchisio PC and Biffo S: Eukaryotic initiation
factor 6 is rate-limiting in translation, growth and
transformation. Nature. 455:684–688. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Brina D, Grosso S, Miluzio A and Biffo S:
Translational control by 80S formation and 60S availability: The
central role of eIF6, a rate limiting factor in cell cycle
progression and tumorigenesis. Cell Cycle. 10:3441–3446. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Pinzaglia M, Montaldo C, Polinari D,
Simone M, La Teana A, Tripodi M, Mancone C, Londei P and Benelli D:
EIF6 over-expression increases the motility and invasiveness of
cancer cells by modulating the expression of a critical subset of
membrane-bound proteins. BMC Cancer. 15:1312015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Benelli D, Cialfi S, Pinzaglia M, Talora C
and Londei P: The translation factor eIF6 is a Notch-dependent
regulator of cell migration and invasion. PLos One. 7:e320472012.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Rosso P, Cortesina G, Sanvito F, Donadini
A, Di Benedetto B, Biffo S and Marchisio PC: Overexpression of
p27BBP in head and neck carcinomas and their lymph node metastases.
Head Neck. 26:408–417. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Tang CL, Yuan SZ, Yang HP, Wang QL and
Zhang R: Expression and significance of P311 and ITGB4BP in
non-small lung cancer. Zhonghua Zhong Liu Za Zhi. 32:526–528.
2010.(In Chinese). PubMed/NCBI
|
|
15
|
Flavin RJ, Smyth PC, Finn SP, Laios A,
O'Toole SA, Barrett C, Ring M, Denning KM, Li J, Aherne ST, et al:
Altered eIF6 and Dicer expression is associated with
clinicopathological features in ovarian serous carcinoma patients.
Mod Pathol. 21:676–684. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ceci M, Gaviraghi C, Gorrini C, Sala LA,
Offenhäuser N, Marchisio PC and Biffo S: Release of eIF6 (p27BBP)
from the 60S subunit allows 80S ribosome assembly. Nature.
426:579–584. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Biswas A, Mukherjee S, Das S, Shields D,
Chow CW and Maitra U: Opposing action of casein kinase 1 and
calcineurin in nucleo-cytoplasmic shuttling of mammalian
translation initiation factor eIF6. J Biol Chem. 286:3129–3138.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Miluzio A, Beugnet A, Volta V and Biffo S:
Eukaryotic initiation factor 6 mediates a continuum between 60S
ribosome biogenesis and translation. EMBO Rep. 10:459–465. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Cruciat CM: Casein kinase 1 and
Wnt/b-catenin signaling. Curr Opin Cell Biol. 31:46–55. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Schittek B and Sinnberg T: Biological
functions of casein kinase 1 isoforms and putative role in
tumorigenesis. Mol Cancer. 13:2312014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Knippschild U, Kruger M, Richter J, Xu P,
García-Reyes B, Peifer C, Halekotte J, Bakulev V and Bischof J: The
CK1 family: Contribution to cellular stress response and its role
in carcinogenesis. Front Oncol. 4:962014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
de Marco N, Iannone L, Carotenuto R, Biffo
S, Vitale A and Campanella C: p27 (BBP)/eIF6 acts as an
anti-apoptotic factor upstream of Bcl-2 during Xenopus laevis
development. Cell Death Differ. 17:360–372. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
de Marco N, Tussellino M, Vitale A and
Campanella C: Eukaryotic initiation factor 6 (eif6) overexpression
affects eye development in Xenopus laevis. Differentiation.
82:108–115. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Silvera D, Formenti SC and Schneider RJ:
Translational control in cancer. Nat Rev Cancer. 10:254–266. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Atallah AM, Tabll AA, El-Nashar E,
El-Bakry KA, El-Sadany M, Ibrahim T and El-Dosoky I: AgNORs count
and DNA ploidy in liver biopsies from patients with schistosomal
liver cirrhosis and hepatocellular carcinoma. Clin Biochem.
42:1616–1620. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Gottwald L, Danilewic M, Fendler W, Suzin
J, Spych M, Piekarski J, Tylinski W, Chalubinska J,
Topczewska-Tylinska K and Cialkowska-Rysz A: The AgNORs count in
predicting long-term survival in serous ovarian cancer. Arch Med
Sci. 10:84–90. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lewinska A, Adamczyk J, Pajak J, Stoklosa
S, Kubis B, Pastuszek P, Slota E and Wnuk M: Curcumin-mediated
decrease in the expression of nucleolar organizer regions in
cervical cancer (HeLa) cells. Mutat Res Genet Toxicol Environ
Mutagen. 771:43–52. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Winzer KJ, Bellach J and Hufnagl P:
Long-term analysis to objectify the tumour grading by means of
automated microscopic image analysis of the nucleolar organizer
regions (AgNORs) in the case of breast carcinoma. Diagn Pathol.
8:562013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Gupta V, Garg M, Chaudhry M, Singh S, Sen
R, Gill M and Sangwaiya A: Role of cyclin D1 immunoreactivity and
AgNOR staining in the evaluation of benign and malignant lesions of
the prostate. Prostate Int. 2:90–96. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Raĭkhlin NT, Bukaeva IA, Smirnova EA,
Pavlovskaia AI, Brzhezovskiĭ VZh, Bogatyrev VN and Ponomareva MV:
Prognistic value of a study of the expression of argyrophilic
nucleolar organizer region associated proteins in case of papillary
thyroid cancer. Arkh Patol. 72:49–52. 2010.(In Russian). PubMed/NCBI
|
|
31
|
Mondal NK, Roychoudhury S and Ray MR:
Higher AgNOR expression in metaplastic and dysplastic airway
epithelial cells predicts the risk of developing lung cancer in
women chronically exposed to biomass smoke. J Environ Pathol
Toxicol Oncol. 34:35–51. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Alaeddini M, Khalili M, Tirgary F and
Etemad-Moghadam S: Argyrophylic proteins of nucleolar organizer
regions (AgNORs) in salivary gland mucoepidermoid carcinoma and its
relation to histological grade. Oral Surg Oral Pathol Oral Radiol
Endod. 105:758–762. 2008. View Article : Google Scholar
|
|
33
|
Duvvuri U and Myers JN: Cancer of the head
and neck is the sixth most common cancer worldwide. Curr Probl
Surg. 46:114–117. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Leemans CR, Braakhuis BJ and Brakenhoff
RH: The molecular biology of head and neck cancer. Nat Rev Cancer.
11:9–22. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Ferreira MB, De Souza JA and Cohen EE:
Role of molecular markers in the management of head and neck
cancers. Curr Opin Oncol. 23:259–264. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2013. CA Cancer J Clin. 63:11–30. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Peters S, Adjei AA, Gridelli C, Reck M,
Kerr K and Felip E; ESMO Guidelines Working Group, : Metastatic
non-small-cell lung cancer (NSCLC): ESMO Clinical Practice
Guidelines for diagnosis, treatment and follow-up. Ann Oncol.
23:vii56–vii64. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Cetin K, Ettinger DS, Hei YJ and O'Malley
CD: Survival by histologic subtype in stage IV nonsmall cell lung
cancer based on data from the Surveillance, Epidemiology and End
Results Program. Clin Epidemiol. 3:139–48. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhang J, Zhang J, Cui X, Yang Y, Li M, Qu
J, Li J and Wang J: FoxM1: A novel tumor biomarker of lung cancer.
Int J Clin Exp Med. 8:3136–3140. 2015.PubMed/NCBI
|
|
40
|
Diaz LK, Zhou X, Welch K, Sahin A and
Gilcrease MZ: Chromogenic in situ hybridization for alpha6beta4
integrin in breast cancer: Correlation with protein expression. J
Mol Diagn. 6:10–15. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Karlsen MA, Høgdall EV, Christensen IJ,
Borgfeldt C, Kalapotharakos G, Zdrazilova-Dubska L, Chovanec J, Lok
CA, Stiekema A, Mutz-Dehbalaie I, et al: A novel diagnostic index
combining HE4, CA125 and age may improve triage of women with
suspected ovarian cancer-An international multicenter study in
women with an ovarian mass. Gynecol Oncol. 138:640–646. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Tanner MM, Grenman S, Koul A, Johannsson
O, Meltzer P, Pejovic T, Borg A and Isola JJ: Frequent
amplification of chromosomal region 20q12-q13 in ovarian cancer.
Clin Cancer Res. 6:1833–1839. 2000.PubMed/NCBI
|
|
44
|
Katz B, Tropé CG, Reich R and Davidson B:
MicroRNAs in ovarian cancer. Hum Pathol. 46:1245–1256. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
van Jaarsveld MT, Helleman J, Berns EM and
Wiemer EA: MicroRNAs in ovarian cancer biology and therapy
resistance. Int J Biochem Cell Biol. 42:1282–1290. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Chong GO, Jeon HS, Han HS, Son JW, Lee YH,
Hong DG, Lee YS and Cho YL: Differential microRNA expression
profiles in primary and recurrent epithelial ovarian cancer.
Anticancer Res. 35:2611–2617. 2015.PubMed/NCBI
|
|
47
|
Donadini A, Giacopelli F, Ravazzolo R,
Gandin V, Marchisio PC and Biffo S: GABP complex regulates
transcription of eIF6 (p27BBP), an essential trans-acting factor in
ribosome biogenesis. FEBS Lett. 580:1983–1987. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Ristevski S, O'Leary DA, Thornell AP, Owen
MJ, Kola I and Hertzog PJ: The ETS transcription factor GABPalpha
is essential for early embryogenesis. Mol Cell Biol. 24:5844–5849.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Ristola M, Arpiainen S, Shimokawa T, Ra C,
Tienari J, Saleem MA, Holthöfer H and Lehtonen S: Regulation of
nephrin gene by the Ets transcription factor, GA-binding protein.
Nephrol Dial Transplant. 28:846–855. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Yu S, Cui K, Jothi R, Zhao DM, Jing X,
Zhao K and Xue HH: GABP controls a critical transcription
regulatory module that is essential for maintenance and
differentiation of hematopoietic stem/progenitor cells. Blood.
117:2166–2178. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Artavanis-Tsakonas S, Rand MD and Lake RJ:
Notch signaling: Cell fate control and intergration in development.
Science. 284:770–776. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Radtke F, MacDonald HR and
Tacchini-Cottier F: Regulation of innate andadaptive immunity by
Notch. Nat Rev Immunol. 13:427–437. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Ranganathan P, Weaver KL and Capobianco
AJ: Notch signalling in solid tumours: A little bit of everything
but not all the time. Nat Rev Cancer. 11:338–351. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Lin JT, Chen MK, Yeh KT, Chang CS, Chang
TH, Lin CY, Wu YC, Su BW, Lee KD and Chang PJ: Association of high
levels of Jagged-1 and Notch-1 expression with poor prognosis in
head and neck cancer. Ann Surg Oncol. 17:2976–2983. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Ranganathan P, Weaver KL and Capobianco
AJ: Notch signalling in solid tumours: A little bit of everything
but not all the time. Nat Rev Cancer. 11:338–351. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Bell D, Hanna EY, Miele L, Roberts D,
Weber RS and El-Naggar AK: Expression and significance of notch
signaling pathway in salivary adenoid cystic carcinoma. Ann Diagn
Pathol. 18:10–13. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zardawi SJ, Zardawi I, McNeil CM, Millar
EK, McLeod D, Morey AL, Crea P, Murphy NC, Pinese M, Lopez-Knowles
E, et al: High Notch1 protein expression is an early event in
breast cancer development and is associated with the HER-2
molecular subtype. Histopathology. 56:286–296. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Arya M, Thrasivoulou C, Henrique R, Millar
M, Hamblin R, Davda R, Aare K, Masters JR, Thomson C, Muneer A, et
al: Targets of Wnt/ß-catenin transcription in penile carcinoma.
PLos One. 10:e01243952015. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Clevers H and Nusse R: Wnt/b-catenin
signaling and disease. Cell. 149:1192–1205. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Polakis P: Drugging Wnt signaling in
cancer. EMBO J. 31:2737–2746. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Li VS, Ng SS, Boersema PJ, Low TY,
Karthaus WR, Gerlach JP, Mohammed S, Heck AJ, Maurice MM, Mahmoudi
T and Clevers H: Wnt signaling through inhibition of β-catenin
degradation in an intact Axin1 complex. Cell. 149:1245–1256. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Niehrs C and Acebron SP: Mitotic and
mitogenic Wnt signalling. EMBO J. 31:2705–2713. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Klaus A and Birchmeier W: Wnt signaling
and its impact on development and cancer. Nat Rev Cancer.
8:387–398. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Higgs MR, Lerat H and Pawlotsky JM:
Hepatitis C virus-induced activation of β-catenin promotes c-Myc
expression and a cascade of pro-carcinogenetic events. Oncogene.
32:4683–4693. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Wang H, Wang H, Makki MS, Wen J, Dai Y,
Shi Q, Liu Q, Zhou X and Wang J: Overexpression of β-catenin and
cyclinD1 predicts a poor prognosis in ovarian serous carcinomas.
Int J Clin Exp Pathol. 7:264–271. 2013.PubMed/NCBI
|
|
66
|
Ji Y, Shah S, Soanes K, Islam MN, Hoxter
B, Biffo S, Heslip T and Byers S: Eukaryoticinitiation factor 6
selectively regulates Wnt signaling and beta-catenin protein
synthesis. Oncogene. 27:755–762. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Sahai E and Marshall CJ: Rho GTPases and
cancer. Nat Rev Cancer. 2:133–142. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Pedersen E and Brakebusch C: Rho GTPase
function in development: How in vivo models change our view. Exp
Cell Res. 318:1779–1787. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Reymond N, Riou P and Ridley AJ: Rho
GTPases and cancer cell transendothelial migration. Methods Mol
Biol. 827:123–142. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Stengel K and Zheng Y: Cdc42 in oncogenic
transformation, invasion, and tumorigenesis. Cell Signal.
23:1415–1423. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Bray K, Gillette M, Young J, Loughran E,
Hwang M, Sears JC and Vargo-Gogola T: Cdc42 overexpression induces
hyperbranching in the developing mammary gland by enhancing cell
migration. Breast Cancer Res. 15:R912013. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Weijzen S, Rizzo P, Braid M, Vaishnav R,
Jonkheer SM, Zlobin A, Osborne BA, Gottipati S, Aster JC, Hahn WC,
et al: Activation of Notch-1 signaling maintains the neoplastic
phenotype in human Ras-transformed cells. Nat Med. 8:979–986. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Furic L, Rong L, Larsson O, Koumakpayi IH,
Yoshida K, Brueschke A, Petroulakis E, Robichaud N, Pollak M,
Gaboury LA, et al: eIF4E phosphorylation promotes tumorigenesis and
is associated with prostate cancer progression. Proc Natl Acad Sci
USA. 107:pp. 14134–14139. 2010; View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Cope CL, Gilley R, Balmanno K, Sale MJ,
Howarth KD, Hampson M, Smith PD, Guichard SM and Cook SJ:
Adaptation to mTOR kinase inhibitors by amplification of eIF4E to
maintain cap-dependent translation. J Cell Sci. 127:788–800. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Grosso S, Pesce E, Brina D, Beugnet A,
Loreni F and Biffo S: Sensitivity of global translation to mTOR
inhibition in REN cells depends on the equilibrium between eIF4E
and 4E-BP1. PLoS One. 6:e291362011. View Article : Google Scholar : PubMed/NCBI
|