Introduction
Autophagy is an evolutionarily conserved process in
which cytoplasmic contents are degraded through the cell's own
lysosomal machinery. Depending on how autophagic substrates are
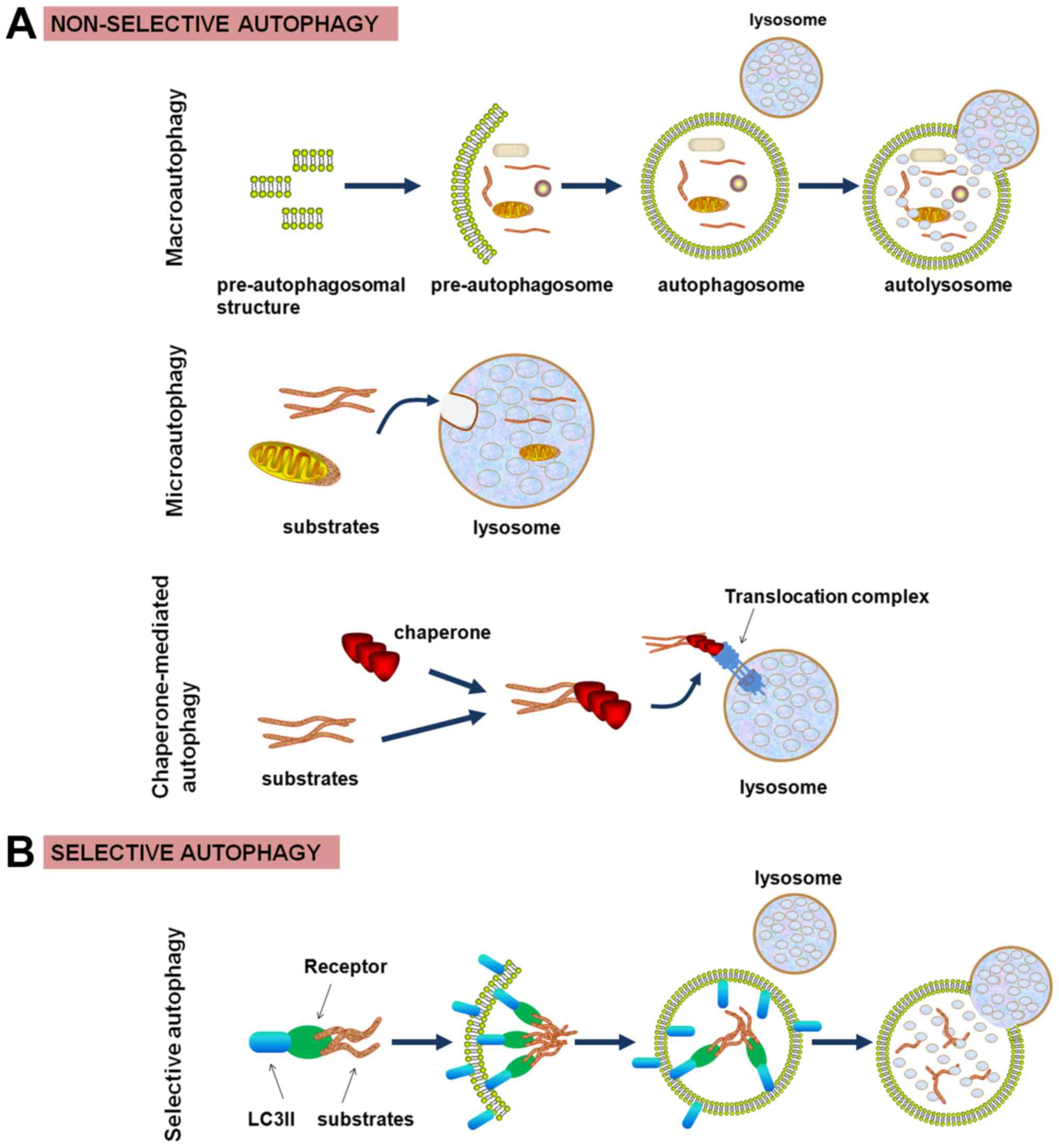
delivered to the lysosome, three different types of autophagy are
identified: macroautophagy, microautophagy, and chaperone-mediated
autophagy (CMA). As the principal pathway, macroautophagy is used
primarily to sequester intact organelles and portions of the
cytosol into the autophagosome, a double-membrane vesicle. After
the completed autophagosome matures via fusing with the lysosome,
its contents are degraded by lysosomal hydrolases. Microautophagy
occurs through the direct engulfment of material into the lysosome.
CMA, a complex and specific pathway, involves the direct
recognition of substrates with the assistance of a lysosomal
chaperone (1). Although the
above-mentioned classic autophagy pathways are generally considered
to be non-selective, accumulating evidence has indicated that
specific cargoes can also be degraded selectively (Fig. 1). The discovery and characterization
of receptor proteins, such as p62/sequestosome 1 (SQSTM1), have
provided pivotal mechanistic insights into this selective
autophagic process (2,3).
Since being coined by de Duve in 1963, autophagy has
been characterized as an adaptive catabolic process that plays a
normal part in cell growth and development, helping to maintain
cellular bioenergetic homeostasis (4). In addition, recent studies have
revealed other roles for the autophagic machinery in regulating a
wide variety of pathological conditions, including tumorigenesis
and cardiovascular diseases, suggesting autophagy modulation may
have therapeutic value (5–7). However, in the development of human
diseases, autophagy has been shown to be a double-edged sword. In
some cases, autophagy is a cytoprotective mechanism, but in others,
autophagy is a pro-death response to stresses, especially
chemotherapy at the cellular and organic levels (8,9). Many
autophagy-related genes (Atgs) have been implicated in controlling
these complicated behaviors during autophagy, and approximately
half of them are evolutionarily conserved from yeast to human
(10). Clearly, the regulation of
autophagy and the resulting downstream effects are complex and very
likely to be cell- and disease type-specific.
Recently, non-coding RNAs, such as long non-coding
RNAs (lncRNAs), were demonstrated to regulate cell autophagy in
vitro and in vivo, further contributing to many of the
hallmarks of disease phenotypes (11) (Fig.
2). The lncRNA highly upregulated liver cancer (HULC) was shown
to significantly inhibit cell apoptosis by activating autophagy,
contributing to the malignant phenotype of gastric cancer (12). The lncRNA maternally expressed gene
3 (MEG3), a novel tumor suppressor, was shown to be downregulated
in bladder cancer and negatively correlated with
microtubule-associated protein 1A/1B-light chain 3 (Atg8/LC3), an
autophagy marker. Inhibition of MEG3 by small-interfering RNA
(siRNA) activated autophagy, increasing cell proliferation and
suppressing cell apoptosis in human bladder cancer cell lines
(13). However, in bacterial
pathogen infection, the MEG3-autophagy axis has as an opposite
effect. In Mycobacterium bovis BCG-infected human
macrophages, MEG3 is clearly decreased. Specifically, in
silico assays link MEG3 to protein kinase B (Akt) and mammalian
target of rapamycin (mTOR) signaling pathways, which can both
downregulate cell autophagy. Furthermore, knockdown of MEG3
resulted in the induction of autophagy, which helped to combat the
intracellular BCG infection. Upon treatment with IFN-γ, increased
autophagy led to sustained and significant MEG3 downregulation,
reinforcing the effective eradication of mycobacteria (14). These investigations have identified
a significant relationship between lncRNA and cell autophagy. This
review mainly provides an overview of lncRNA biology in autophagy
modulation and discusses specific studies that have provided new
insight into the underlying mechanism of the lncRNA-autophagy axis
for therapeutic intervention.
The lncRNA-miRNA axis and autophagy
LncRNA-miRNA axis and autophagy
Accumulating studies have highlighted the importance
of the non-coding genome in autophagy. Based on length, non-coding
RNA can be categorized into several types, such as microRNAs
(miRNAs, approximately 22 nucleotides) and lncRNAs (>200
nucleotides). miRNAs have emerged as critical mediators of gene
expression through binding to targeted mRNA, including Atgs
(15,16). As the dysregulated expression of
Atgs has been shown to have a far reaching impact on human
diseases, miRNAs are being examined as potential novel biomarkers
or therapeutic targets (17). One
particular example is the miR-30a family, which is linked to
autophagy and various pathophysiological conditions. miR-30a
negatively regulates BECN1/Atg6 stability via the predicted binding
sequence in its 3′ untranslated region (UTR). Downregulation of
miRNA-30a increased the mRNA and protein levels of BECN1, markedly
initiating basal autophagy in mature adipocytes (18). Similarly, miR-30a inhibition
alleviated ischemia/reperfusion-induced neurological dysfunction
through enhancing BECN1-mediated autophagy (19). Additionally, miRNA-30a-5p can target
metadherin (MTDH) mRNA for cleavage or translational repression. By
targeting MTDH, miRNA-30a-5p downregulated activation of the Akt
signaling pathway, which might regulate autophagy and apoptosis of
cancer cells (20,21). In another study, identical
recognition sites of miR-137 were found at the 3′UTRs of the
selective autophagy receptors NIX/BNIP3L (BCL2/adenovirus E1B 19
kDa interacting protein 3-like) and FUN14 domain containing 1
(FUNDC1). In response to hypoxia, miR-137 reduced the expression of
Nix and FUNDC1, thereby inhibiting mitophagy, a selective autophagy
of impaired mitochondria, without affecting classic autophagy
(22). However, the exact role of
different miRNAs in regulating classic or selective autophagy must
still be elucidated. Gibbings et al demonstrated that
selective autophagy can in turn modulate miRNA homeostasis. The
miRNA-processing enzyme Dicer and the miRNA effector Argonaute 2
(Ago2) are critical components responsible for miRNA synthesis.
Under the control of the autophagy receptor nuclear dot protein 52
kDa (NDP52), Dicer and Ago2 were targeted for degradation by
selective autophagy, which inhibited homeostasis and activity of
the targeted miRNAs (23,24). These data have profound implications
for the association of miRNA-levels with dysregulated
autophagy.
The role of miRNAs in the regulation
of autophagy by lncRNA
Bioinformatics analysis of molecular interactions
has identified miRNA recognition elements (MREs) on lncRNA
sequences, demonstrating the direct transcriptional regulation of
miRNA by lncRNA (25). Furthermore,
perturbation of lncRNA expression disrupted the balance of
lncRNA-miRNA regulatory paradigms, contributing to the pathogenesis
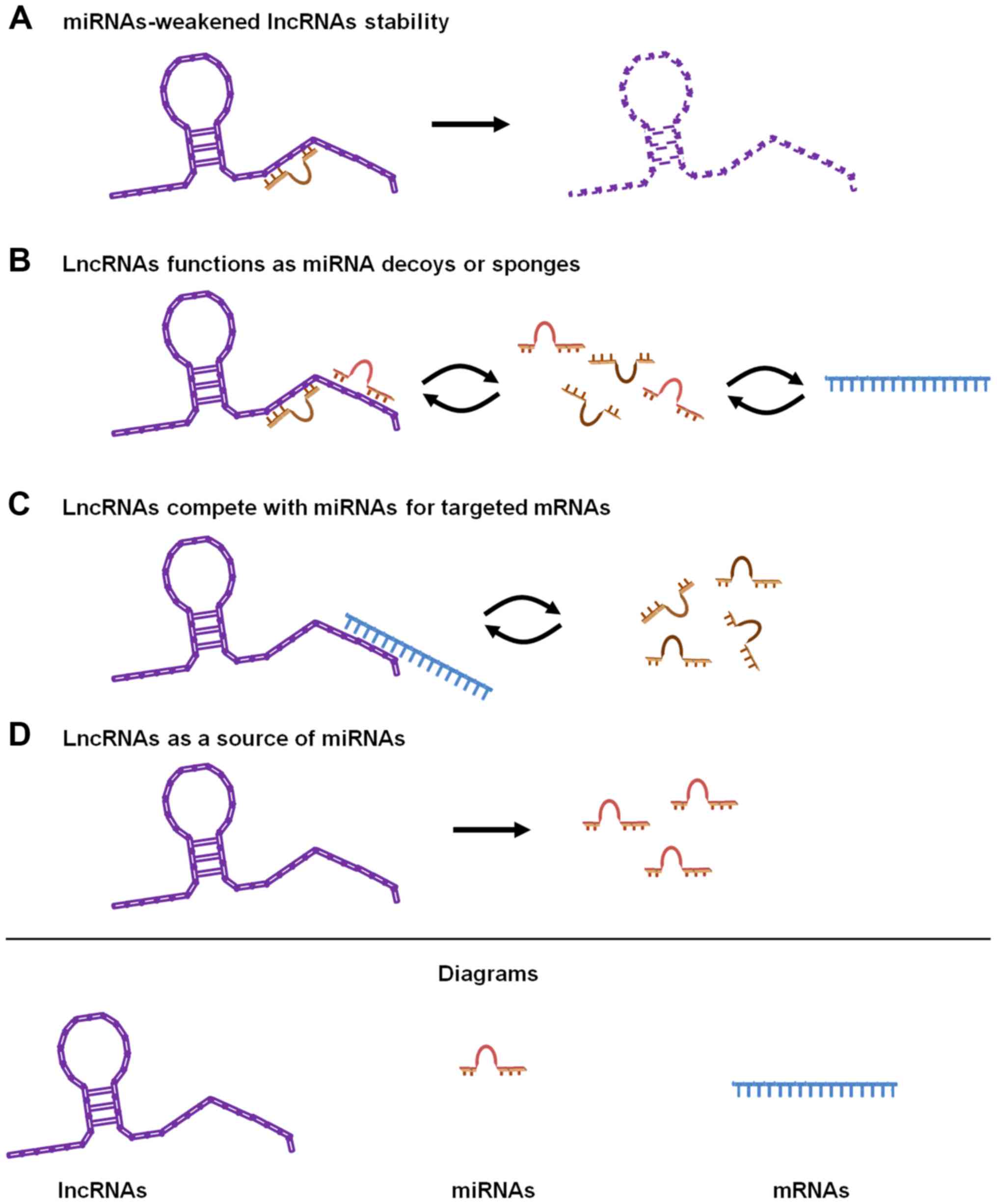
of various diseases, including cancers. To date, four forms of
functional cross-regulation have been artificially identified
(26,27) (Fig.
3). The stability of lncRNAs can be weakened through
interaction with specific miRNAs. Yoon et al found that the
stability of lncRNA p21 was controlled by the miRNA let-7b.
Overexpression of precursor-let-7 decreased the half-life of lncRNA
p21 and accelerated its degradation (28). Likewise, another lncRNA, HOX
antisense intergenic RNA (HOTAIR), was shown to be enriched and
stable after using an antagomir to inhibit endogenous let-7i in
human cervical carcinoma HeLa cells (29). LncRNAs can function as miRNA decoys
or sponges, also known as competitive endogenous RNAs (ceRNAs). The
function of lncRNAs as ceRNAs significantly expands their
regulatory capacity; ceRNAs can compete with mRNA for the binding
of miRNA and thus repress the target gene at the
post-transcriptional level. Furthermore, numerous lines of
bioinformatics evidence from several well-known groups have
supported the ceRNA function of lncRNAs, including the database of
human long non-coding RNA acting as competing endogenous RNA
(lnCeDB) (30), the
lncRNA-associated competing triplet database (LncACTdb) (31) and the database of cancer somatic
mutations altering microRNA-ceRNA interactions (SomamiR 2.0)
(32). LncRNAs can compete with
miRNAs for interaction with shared target mRNAs. Franklin et
al revealed that the long non-coding N-ras functional RNA
(ncNRFR) competes with let-7 and reverses the repression of let-7
site-bearing mRNAs (33). Faghihi
et al found a miR-485-5p binding site in the mRNA of β-site
amyloid precursor protein cleaving enzyme 1 (BACE1). By blocking
access of miR-485-5p, the lncRNA BACE1-antisense transcript
prevented miRNA-induced suppression of BACE1 mRNA (34). LncRNAs are a source of miRNAs for
inducing target mRNA silencing. Exon 1 of lncRNA H19 encodes two
conserved miRNAs, miR-675-3p and miR-675-5p. Exogenous miR-675-3p
and miR-675-5p both rescue abnormal skeletal muscle regeneration
induced by H19 depletion (35). In
summary, the lncRNA-miRNA axis controls gene expression patterns
driving many cellular processes, such as cell development,
proliferation, and cell autophagy.
It is well known that mTOR, a central
negative-regulator of cell autophagy, plays a key role in
regulating the balance between cell growth and death in response to
stress signals. Modulating abnormal mTOR signaling is important for
numerous physiological and pathological conditions (36). After treatment with doxorubicin,
mTOR activation via suppression of ELK3 impaired autophagy, and
thus enhanced cell viability in the breast cancer cell line
MDA-MB-231 (37). Using molecular
docking prediction, Ge et al identified a novel mTOR
activator,
3-benzyl-5-((2-nitrophenoxy)methyl)-dihydrofuran-2(3H)-one (3BDO).
They found that activation of mTOR induced by 3BDO increased
phosphorylation of T-cell-restricted intracellular antigen-1
(TIA1), which is responsible for processing FLJ11812, a nuclear
lncRNA derived from the 3′UTR of transforming growth factor β2
(TGFβ2). Acting as a ceRNA, FLJ11812 competed for binding with
miR-4459, resulting in the increase of its target Atg13 and
promotion of autophagy (38). Apart
from miR-4459, FLJ11812 also acts as the molecular decoy of
miR-3960 and miR-4488, elevating the expression of miRNA targets
ceramide synthase 1 (CERS1) and N-acetyltransferase 8-like (NAT8L),
respectively, two molecules participating in mitophagy (39). In addition, phospholipase D (PLD)
inhibitor stimulates cell autophagy and exhibits attractive
antitumorigenic effects by destabilizing mTOR. The lncRNA antisense
non-coding RNA in the INK4 locus (ANRIL) has been shown to be
responsible for the functions of PLD inhibitor (40). In sleeping beauty (SB)-baculovirus
(BV) systems, ectopic expression of the lncRNA phosphatase and
tensin (PTEN) homolog pseudogene 1 (PTENP1) in hepatocellular
carcinoma suppressed the Akt/mTOR signaling pathway by promoting
PTEN transcription, thus inducing autophagic associated cell death.
Moreover, PTENP1 also acted as a decoy for miR-17, miR-19b, and
miR-20a, which enhanced autophagy of their target genes Atg7 and
Unc-51 like autophagy activating kinase 1 (ULK1) (41). While most current studies mainly
focus on the function of lncRNAs in the nucleus, Liu et al
recently uncovered a function of the cytoplasmic lncRNA neighbor of
BRCA1 gene 2 (NBR2). Energy stress could promote the colocalization
of NBR2 and AMPK in the cytoplasm, potentiating AMPK activation.
Finally, knockdown of NBR2 attenuated stress-induced mTOR
inactivation by repressing the AMPK kinase signal, thus altering
the autophagy response and increasing tumor development (42). Together, these results indicate that
mTOR activation can inhibit cell autophagy via lncRNA-miRNA
signaling, leading to participation in autophagy-mediated
processes, such asembryogenesis (43), atherosclerosis (44), and tumorigenesis (41).
Apart from mTOR, the lncRNA-miRNA axis also directly
affects Atgs modulation. As a direct target of miR-21, the lncRNA
growth arrest-specific 5 (GAS5) is upregulated during
osteoarthritis. Overexpression of GAS5 inhibited cell autophagy, as
indicated by downregulation of the LC3-II/LC3-I ratio and BECN1,
and stimulated the death of articular chondrocytes (45). Recently, Liu et al defined
another growth-promoted lncRNA in hepatocellular carcinoma,
HNF1A-AS1 (antisense transcript of HNF1A). Oncogenic HNF1A-AS1
sequestered miR-30b in a ceRNA-dependent manner and stimulated
autophagy flux via promoting the formation of the Atg12 complex,
thus contributing to hepatocarcinogenesis (46). In ischemia/reperfusion-induced
myocardial infarction, Atg7, a specific target of miR-188-3p, is
upregulated, leading to autophagic cell death. Through lncRNA array
analysis, Wang et al reported a novel lncRNA, autophagy
promoting factor (APF). APF regulated the expression of Atg7 and
the consequent autophagy by sponging miR-188-3p (47). Together, these data indicate
critical roles of lncRNAs in cell autophagy as well as
autophagic-associated diseases.
Other lncRNAs on autophagy regulation
The human PVT1 gene is a lncRNA on chromosome 8q24
and is critical in the regulation of cell growth, and
differentiation. Many studies have demonstrated that the biological
function of PVT1 mainly depends on two mechanisms. First, PVT1
inhibits gene promoter activity. Zhang et al found PVT1 was
an oncogenic gene in cervical cancer. Ectopic expression of PVT1
recruited enhancer of zeste homolog 2 (EZH2), a member of the
polycomb-group (PcG) family, to the miR-200b promoter and repressed
miR-200b expression by methylating Lys-27 (H3K27me) of histone H3
on the miR-200b promoter (48).
Similarly, PVT1 repressed large tumor suppressor kinase 2 (LATS2)
transcription by recruiting EZH2 to the LASTS2 promoter,
facilitating cell proliferation in non-small cell lung cancer
(49). Secondly, PVT1 regulates
protein stability. As the PVT1 locus is adjacent to the c-Myc
locus, PVT1 is required for high Myc protein levels in cancer
cells. According to a study by Tseng et al, PVT1 controlled
Myc levels through regulation of protein stability (50). Furthermore, knockdown of PVT1 by RNA
interference led to inhibition of cancer cell proliferation by
suppressing the Myc protein level (51). Additionally, another group studied
PVT1-mediated autophagy in cognitive impairment of diabetes
mellitus. They found that autophagy inhibitor 3-methyladenine
(3-MA) could significantly induce apoptotic cell death by
decreasing hippocampal PVT1 expression, which subsequently
heightened cognitive impairment (52). This was the first time that lncRNA
homeostasis was shown to be controlled by cell autophagy, but the
specific mechanisms need to be further elucidated.
BRAF-activated lncRNA (BANCR), a 693-bp transcripton
chromosome 9, is a useful biomarker or future therapeutic target in
various cancers, including melanoma (53), papillary thyroid carcinoma (54), and lung cancer (55). Flockhart et al and Wajapeyee
et al performed massively parallel cDNA sequencing (RNA-seq)
and found that the expression of BANCR was regulated by mutant
BRAFV600E protein, the most prevalent mutation of the BRAF gene in
the circulating tumor DNA of a patient (56,57). A
recent study demonstrated that the autophagy inhibitors VATG-027
and VATG-032, which are acridine and tetrahydroacridine derivatives
of the antimalarial agent chloroquine, significantly enhanced the
sensitivity of cancer cells to the BRAF V600E inhibitor vemurafenib
(58). Moreover, selective
BRAFV600E inhibitors induced autophagy by activating AMPK signaling
in colorectal cancer cells (59).
As BANCR is shown to be strongly linked with BRAFV600E mutation,
BANCR may have a possible role in autophagy modulation. As
expected, overexpression of BANCR in papillary thyroid carcinoma
enhanced the ratio of LC3-II/LC3-I, a marker for autophagy,
resulting in increased cell proliferation and inhibited apoptosis
(60).
Concerning cardiac remodeling, Viereck and
colleagues performed global lncRNA expression profiling and
identified the lncRNA cardiac hypertrophy-associated transcript
(Chast) as a potential influence on cardiomyocyte hypertrophy in
vivo. GapmeR-mediated Chast silencing upregulates Atg5 and
increases the LC3-II/LC3-I ratio, driving cell autophagy capacity
and impeding the pathologic remodeling processes. Bioinformatics
analysis of gene sequence revealed partial overlaps between Chast
and the antisense strand of pleckstrin homology domain-containing
protein family M member 1 (plekhm1), a regulator of autophagy,
suggesting that Chast might downregulate cell autophagy by a
plekhm1-dependent mechanism (61).
Conclusions and future directions
Advances in sequencing technologies have identified
numerous lncRNAs. Though previously considered ‘junk sequences’ in
our genomes, the epigenetic role of lncRNAs promises to be another
exciting frontier for disease research and therapy. The discovery
of lncRNAs and their functions has shed light on novel molecular
mechanisms of autophagy regulation, potentially improving the
diagnosis and treatment of human autophagy-associated diseases
(Table I). Although the precise
cellular mechanism of lncRNA is not completely understood, recent
studies have revealed the existence of stable lncRNAs protected
from endogenous RNases in blood and other bodily fluids of
patients, the so-called circulating lncRNAs (62). Due to growing interest in the
function of lncRNAs as diagnostic and prognostic biomarkers,
lncRNAs are increasingly gaining the attention of scientists and
clinicians. However, two critical issues must still be solved:
uniformity of sample preparation and data normalization (63). In addition, lncRNAs in extracellular
vesicles, such as exosomes, could offer a novel, enriched source of
lncRNAs. These exosomes have been found to harbor disease-derived
specific lncRNAs, which were significantly different between
patients and normal controls (64).
Qu et al found that the exosome-transmitted lncRNA activated
in renal cell carcinoma with sunitinib resistance (ARSR)
potentially affects the chemosensitivity of cancer cells (65). In addition, the regulation of cell
autophagy by circulating lncRNAs under pathophysiological
conditions needs to be discussed. The exciting potential of these
emerging lncRNAs as biomarkers could be an important advance in
disease management.
 | Table I.Examples of lncRNAs associated with
autophagy regulations. |
Table I.
Examples of lncRNAs associated with
autophagy regulations.
| lncRNA | Locus | Role in
autophagy | Related
diseases | Refs. |
|---|
| HULC | Chr6 | Increasing
LC3-II/LC3-I ratio | Gastric cancer | (12) |
| MEG3 | Chr14 | Negatively
correlateing with LC3 | Bladder cancer | (13) |
|
|
| Promoting Akt/mTOR
signal pathways | Bacterial
infection | (14) |
| FLJ11812 | Chr1 | Increasing Atg13
level | VECs
dysfunction | (40) |
| ANRIL | Chr9 | Increasing
LC3-II/LC3-I ratio | Lung cancer | (41) |
| PTENP1 | Chr9 | Suppressing
Akt/mTOR signal pathways | Hepatocellular | (42) |
|
|
| Increasing Atg7 and
ULK1 level | carcinoma |
|
| NBR2 | Chr17 | Activating the
cytoplasmic AMPK | Several
cancers | (43) |
| GAS5 | Chr1 | Suppressing
LC3-II/LC3-I ratio and BECN1 | Osteoarthritis | (46) |
| HNF1A-AS1 | Chr12 | promoting the
formation of Atg12 complex | Hepatocellular
carcinoma | (47) |
| APF | Chr8 | Increasing Atg7
level | Myocardial
infarction | (48) |
| PVT1 | Chr8 | Downregulated by
autophagy inhibitor | Diabetes
mellitus | (53) |
| BANCR | Chr9 | Increasing
LC3-II/LC3-I ratio | Papillary thyroid
carcinoma | (61) |
| Chast | Chr11 | Suppressing
LC3-II/LC3-I ratio, BECN1 and plekhm1 | Cardiomyocyte
hypertrophy | (62) |
Autophagy, a tightly regulated catabolic process of
cellular self-digestion, can be activated by several stresses,
especially pharmacological action, and constitutes a potential
target for disease therapy (66,67).
Studies have indicated that autophagy appears to serve as either a
pro-survival or pro-death response to therapeutic treatment
(68). However, the precise role of
autophagy in modulating cell biological behavior is highly
dependent on the cellular context and its extent. A recent study
found that lncRNAs can also influence drug-sensitivity by
modulating cell autophagy. Overexpression of the lncRNA regulator
of insulin sensitivity and autophagy (Risa) in hepatocytes or
myotubes significantly decreased autophagy, thus attenuating
insulin resistance. Knockdown of Atg7 or Atg5 by siRNAs inhibited
the effect of Risa on insulin resistance in vivo (69). These findings show a potentially
attractive approach to improving the curative effect: modulating
the lncRNA-autophagy axis. Thus, further investigation of lncRNAs
in autophagy regulation may identify novel strategies to enhance
the benefits of pharmacotherapy in the treatment of human
diseases.
Acknowledgements
This study was supported by the National Natural
Science Foundation of China (no. 81572946), the Changsha Science
and Technology Project (no. k1508024-31), and the Clinical and
Rehabilitation Research Foundation of Xiangya Hospital -
Beidaweiming.
References
|
1
|
Jia G and Sowers JR: Autophagy: A
housekeeper in cardiorenal metabolic health and disease. Biochim
Biophys Acta. 1852:219–224. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Wurzer B, Zaffagnini G, Fracchiolla D,
Turco E, Abert C, Romanov J and Martens S: Oligomerization of p62
allows for selection of ubiquitinated cargo and isolation membrane
during selective autophagy. eLife. 4:e089412015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ryter SW, Cloonan SM and Choi AM:
Autophagy: A critical regulator of cellular metabolism and
homeostasis. Mol Cells. 36:7–16. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Galluzzi L, Bravo-San Pedro JM, Blomgren K
and Kroemer G: Autophagy in acute brain injury. Nat Rev Neurosci.
17:467–484. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Roos WP, Thomas AD and Kaina B: DNA damage
and the balance between survival and death in cancer biology. Nat
Rev Cancer. 16:20–33. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lorenzen JM and Thum T: Long noncoding
RNAs in kidney and cardiovascular diseases. Nat Rev Nephrol.
12:360–373. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bento CF, Renna M, Ghislat G, Puri C,
Ashkenazi A, Vicinanza M, Menzies FM and Rubinsztein DC: Mammalian
autophagy: How does it work? Annu Rev Biochem. 85:685–713. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Evangelisti C, Evangelisti C, Chiarini F,
Lonetti A, Buontempo F, Neri LM, McCubrey JA and Martelli AM:
Autophagy in acute leukemias: A double-edged sword with important
therapeutic implications. Biochim Biophys Acta. 1853:14–26. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Chen Y, Henson ES, Xiao W, Huang D,
McMillan-Ward EM, Israels SJ and Gibson SB: Tyrosine kinase
receptor EGFR regulates the switch in cancer cells between cell
survival and cell death induced by autophagy in hypoxia. Autophagy.
12:1029–1046. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Xie Y, Kang R, Sun X, Zhong M, Huang J,
Klionsky DJ and Tang D: Posttranslational modification of
autophagy-related proteins in macroautophagy. Autophagy. 11:28–45.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Choudhry H, Harris AL and McIntyre A: The
tumour hypoxia induced non-coding transcriptome. Mol Aspects Med.
47–48, 35–53. 2016.PubMed/NCBI
|
|
12
|
Zhao Y, Guo Q, Chen J, Hu J, Wang S and
Sun Y: Role of long non-coding RNA HULC in cell proliferation,
apoptosis and tumor metastasis of gastric cancer: A clinical and in
vitro investigation. Oncol Rep. 31:358–364. 2014.PubMed/NCBI
|
|
13
|
Ying L, Huang Y, Chen H, Wang Y, Xia L,
Chen Y, Liu Y and Qiu F: Downregulated MEG3 activates autophagy and
increases cell proliferation in bladder cancer. Mol Biosyst.
9:407–411. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Pawar K, Hanisch C, Vera SE Palma,
Einspanier R and Sharbati S: Down regulated lncRNA MEG3 eliminates
mycobacteria in macrophages via autophagy. Sci Rep. 6:194162016.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hall DP, Cost NG, Hegde S, Kellner E,
Mikhaylova O, Stratton Y, Ehmer B, Abplanalp WA, Pandey R, Biesiada
J, et al: TRPM3 and miR-204 establish a regulatory circuit that
controls oncogenic autophagy in clear cell renal cell carcinoma.
Cancer Cell. 26:738–753. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Piccoli MT, Gupta SK and Thum T: Noncoding
RNAs as regulators of cardiomyocyte proliferation and death. J Mol
Cell Cardiol. 89(Pt A): 59–67. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ouimet M, Koster S, Sakowski E,
Ramkhelawon B, van Solingen C, Oldebeken S, Karunakaran D,
Portal-Celhay C, Sheedy FJ, Ray TD, et al: Mycobacterium
tuberculosis induces the miR-33 locus to reprogram autophagy and
host lipid metabolism. Nat Immunol. 17:677–686. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Deng Y, Xu J, Zhang X, Yang J, Zhang D,
Huang J, Lv P, Shen W and Yang Y: Berberine attenuates autophagy in
adipocytes by targeting BECN1. Autophagy. 10:1776–1786. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wang P, Liang J, Li Y and Li J, Yang X,
Zhang X, Han S, Li S and Li J: Down-regulation of miRNA-30a
alleviates cerebral ischemic injury through enhancing beclin
1-mediated autophagy. Neurochem Res. 39:1279–1291. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li WF, Dai H, Ou Q, Zuo GQ and Liu CA:
Overexpression of microRNA-30a-5p inhibits liver cancer cell
proliferation and induces apoptosis by targeting MTDH/PTEN/AKT
pathway. Tumour Biol. 37:5885–5895. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang RC, Wei Y, An Z, Zou Z, Xiao G,
Bhagat G, White M, Reichelt J and Levine B: Akt-mediated regulation
of autophagy and tumorigenesis through Beclin 1 phosphorylation.
Science. 338:956–959. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Li W, Zhang X, Zhuang H, Chen HG, Chen Y,
Tian W, Wu W, Li Y, Wang S, Zhang L, et al: MicroRNA-137 is a novel
hypoxia-responsive microRNA that inhibits mitophagy via regulation
of two mitophagy receptors FUNDC1 and NIX. J Biol Chem.
289:10691–10701. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Gibbings D, Mostowy S, Jay F, Schwab Y,
Cossart P and Voinnet O: Selective autophagy degrades DICER and
AGO2 and regulates miRNA activity. Nat Cell Biol. 14:1314–1321.
2012. View
Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gibbings D, Mostowy S and Voinnet O:
Autophagy selectively regulates miRNA homeostasis. Autophagy.
9:781–783. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Paraskevopoulou MD, Vlachos IS, Karagkouni
D, Georgakilas G, Kanellos I, Vergoulis T, Zagganas K, Tsanakas P,
Floros E, Dalamagas T, et al: DIANA-LncBase v2: Indexing microRNA
targets on non-coding transcripts. Nucleic Acids Res. 44(D1):
D231–D238. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yoon JH, Abdelmohsen K and Gorospe M:
Functional interactions among microRNAs and long noncoding RNAs.
Semin Cell Dev Biol. 34:9–14. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ballantyne MD, McDonald RA and Baker AH:
lncRNA/MicroRNA interactions in the vasculature. Clin Pharmacol
Ther. 99:494–501. 2016. View
Article : Google Scholar : PubMed/NCBI
|
|
28
|
Yoon JH, Abdelmohsen K, Srikantan S, Yang
X, Martindale JL, De S, Huarte M, Zhan M, Becker KG and Gorospe M:
LincRNA-p21 suppresses target mRNA translation. Mol Cell.
47:648–655. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Yoon JH, Abdelmohsen K, Kim J, Yang X,
Martindale JL, Tominaga-Yamanaka K, White EJ, Orjalo AV, Rinn JL,
Kreft SG, et al: Scaffold function of long non-coding RNA HOTAIR in
protein ubiquitination. Nat Commun. 4:29392013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Das S, Ghosal S, Sen R and Chakrabarti J:
lnCeDB: Database of human long noncoding RNA acting as competing
endogenous RNA. PLoS One. 9:e989652014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wang P, Ning S, Zhang Y, Li R, Ye J, Zhao
Z, Zhi H, Wang T, Guo Z and Li X: Identification of
lncRNA-associated competing triplets reveals global patterns and
prognostic markers for cancer. Nucleic Acids Res. 43:3478–3489.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Bhattacharya A and Cui Y: SomamiR 2.0: A
database of cancer somatic mutations altering microRNA-ceRNA
interactions. Nucleic Acids Res. 44(D1): D1005–D1010. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Franklin JL, Rankin CR, Levy S, Snoddy JR,
Zhang B, Washington MK, Thomson JM, Whitehead RH and Coffey RJ:
Malignant transformation of colonic epithelial cells by a
colon-derived long noncoding RNA. Biochem Biophys Res Commun.
440:99–104. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Faghihi MA, Zhang M, Huang J, Modarresi F,
van der Brug MP, Nalls MA, Cookson MR, St-Laurent G III and
Wahlestedt C: Evidence for natural antisense transcript-mediated
inhibition of microRNA function. Genome Biol. 11:R562010.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Dey BK, Pfeifer K and Dutta A: The H19
long noncoding RNA gives rise to microRNAs miR-675-3p and
miR-675-5p to promote skeletal muscle differentiation and
regeneration. Genes Dev. 28:491–501. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Gallagher LE, Williamson LE and Chan EY:
Advances in autophagy regulatory mechanisms. Cells. 5:242016.
View Article : Google Scholar :
|
|
37
|
Park JH, Kim KP, Ko JJ and Park KS:
PI3K/Akt/mTOR activation by suppression of ELK3 mediates
chemosensitivity of MDA-MB-231 cells to doxorubicin by inhibiting
autophagy. Biochem Biophys Res Commun. 477:277–282. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Ge D, Han L, Huang S, Peng N, Wang P,
Jiang Z, Zhao J, Su L, Zhang S, Zhang Y, et al: Identification of a
novel MTOR activator and discovery of a competing endogenous RNA
regulating autophagy in vascular endothelial cells. Autophagy.
10:957–971. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Huang S, Lu W, Ge D, Meng N, Li Y, Su L,
Zhang S, Zhang Y, Zhao B and Miao J: A new microRNA signal pathway
regulated by long noncoding RNA TGFB2-OT1 in autophagy and
inflammation of vascular endothelial cells. Autophagy.
11:2172–2183. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Kang YH, Kim D and Jin EJ: Down-regulation
of phospholipase D stimulates death of lung cancer cells involving
up-regulation of the long ncRNA ANRIL. Anticancer Res.
35:2795–2803. 2015.PubMed/NCBI
|
|
41
|
Chen CL, Tseng YW, Wu JC, Chen GY, Lin KC,
Hwang SM and Hu YC: Suppression of hepatocellular carcinoma by
baculovirus-mediated expression of long non-coding RNA PTENP1 and
microRNA regulation. Biomaterials. 44:71–81. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Liu X, Xiao ZD, Han L, Zhang J, Lee SW,
Wang W, Lee H, Zhuang L, Chen J, Lin HK, et al: LncRNA NBR2 engages
a metabolic checkpoint by regulating AMPK under energy stress. Nat
Cell Biol. 18:431–442. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Lu W, Han L, Su L, Zhao J, Zhang Y, Zhang
S, Zhao B and Miao J: A 3′ UTR-associated RNA, FLJ11812 maintains
stemness of human embryonic stem cells by targeting miR-4459. Stem
Cells Dev. 24:1133–1140. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Peng N, Meng N, Wang S, Zhao F, Zhao J, Su
L, Zhang S, Zhang Y, Zhao B and Miao J: An activator of mTOR
inhibits oxLDL-induced autophagy and apoptosis in vascular
endothelial cells and restricts atherosclerosis in apolipoprotein
E-/− mice. Sci Rep. 4:55192014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Song J, Ahn C, Chun CH and Jin EJ: A long
non-coding RNA, GAS5, plays a critical role in the regulation of
miR-21 during osteoarthritis. J Orthop Res. 32:1628–1635. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Liu Z, Wei X, Zhang A, Li C, Bai J and
Dong J: Long non-coding RNA HNF1A-AS1 functioned as an oncogene and
autophagy promoter in hepatocellular carcinoma through sponging
hsa-miR-30b-5p. Biochem Biophys Res Commun. 473:1268–1275. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Wang K, Liu CY, Zhou LY, Wang JX, Wang M,
Zhao B, Zhao WK, Xu SJ, Fan LH, Zhang XJ, et al: APF lncRNA
regulates autophagy and myocardial infarction by targeting
miR-188-3p. Nat Commun. 6:67792015. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zhang S, Zhang G and Liu J: Long noncoding
RNA PVT1 promotes cervical cancer progression through
epigenetically silencing miR-200b. APMIS. 124:649–658. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Wan L, Sun M, Liu GJ, Wei CC, Zhang EB,
Kong R, Xu TP, Huang MD and Wang ZX: Long noncoding RNA PVT1
promotes non-small cell lung cancer cell proliferation through
epigenetically regulating LATS2 expression. Mol Cancer Ther.
15:1082–1094. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Tseng YY, Moriarity BS, Gong W, Akiyama R,
Tiwari A, Kawakami H, Ronning P, Reuland B, Guenther K, Beadnell
TC, et al: PVT1 dependence in cancer with MYC copy-number increase.
Nature. 512:82–86. 2014.PubMed/NCBI
|
|
51
|
Zeng C, Yu X, Lai J, Yang L, Chen S and Li
Y: Overexpression of the long non-coding RNA PVT1 is correlated
with leukemic cell proliferation in acute promyelocytic leukemia. J
Hematol Oncol. 8:1262015. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Li Z, Hao S, Yin H, Gao J and Yang Z:
Autophagy ameliorates cognitive impairment through activation of
PVT1 and apoptosis in diabetes mice. Behav Brain Res. 305:265–277.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
McCarthy N: Epigenetics. Going places with
BANCR. Nat Rev Cancer. 12:4512012. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Zheng H, Wang M, Jiang L, Chu H, Hu J,
Ning J, Li B, Wang D and Xu J: BRAF-activated long noncoding RNA
modulates papillary thyroid carcinoma cell proliferation through
regulating thyroid stimulating hormone receptor. Cancer Res Treat.
48:698–707. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Sun M, Liu XH, Wang KM, Nie FQ, Kong R,
Yang JS, Xia R, Xu TP, Jin FY, Liu ZJ, et al: Downregulation of
BRAF activated non-coding RNA is associated with poor prognosis for
non-small cell lung cancer and promotes metastasis by affecting
epithelial-mesenchymal transition. Mol Cancer. 13:682014.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Flockhart RJ, Webster DE, Qu K,
Mascarenhas N, Kovalski J, Kretz M and Khavari PA: BRAFV600E
remodels the melanocyte transcriptome and induces BANCR to regulate
melanoma cell migration. Genome Res. 22:1006–1014. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Wajapeyee N, Serra RW, Zhu X, Mahalingam M
and Green MR: Oncogenic BRAF induces senescence and apoptosis
through pathways mediated by the secreted protein IGFBP7. Cell.
132:363–374. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Goodall ML, Wang T, Martin KR, Kortus MG,
Kauffman AL, Trent JM, Gately S and MacKeigan JP: Development of
potent autophagy inhibitors that sensitize oncogenic BRAF V600E
mutant melanoma tumor cells to vemurafenib. Autophagy.
10:1120–1136. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Sueda T, Sakai D, Kawamoto K, Konno M,
Nishida N, Koseki J, Colvin H, Takahashi H, Haraguchi N, Nishimura
J, et al: BRAF V600E inhibition stimulates AMP-activated protein
kinase-mediated autophagy in colorectal cancer cells. Sci Rep.
6:189492016. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Wang Y, Guo Q, Zhao Y, Chen J, Wang S, Hu
J and Sun Y: BRAF-activated long non-coding RNA contributes to cell
proliferation and activates autophagy in papillary thyroid
carcinoma. Oncol Lett. 8:1947–1952. 2014.PubMed/NCBI
|
|
61
|
Viereck J, Kumarswamy R, Foinquinos A,
Xiao K, Avramopoulos P, Kunz M, Dittrich M, Maetzig T, Zimmer K,
Remke J, et al: Long noncoding RNA Chast promotes cardiac
remodeling. Sci Transl Med. 8:326ra222016. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Qi P, Zhou XY and Du X: Circulating long
non-coding RNAs in cancer: Current status and future perspectives.
Mol Cancer. 15:392016. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Tiberio P, Callari M, Angeloni V, Daidone
MG and Appierto V: Challenges in using circulating miRNAs as cancer
biomarkers. BioMed Res Int. 2015:7314792015. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Dong L, Lin W, Qi P, Xu MD, Wu X, Ni S,
Huang D, Weng WW, Tan C, Sheng W, et al: Circulating long RNAs in
serum extracellular vesicles: Their characterization and potential
application as biomarkers for diagnosis of colorectal cancer.
Cancer Epidemiol Biomarkers Prev. 25:1158–1166. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Qu L, Ding J, Chen C, Wu ZJ, Liu B, Gao Y,
Chen W, Liu F, Sun W, Li XF, et al: Exosome-transmitted lncARSR
promotes sunitinib resistance in renal cancer by acting as a
competing endogenous RNA. Cancer Cell. 29:653–668. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Rubinsztein DC, Codogno P and Levine B:
Autophagy modulation as a potential therapeutic target for diverse
diseases. Nat Rev Drug Discov. 11:709–730. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Cheng Y, Ren X, Hait WN and Yang JM:
Therapeutic targeting of autophagy in disease: Biology and
pharmacology. Pharmacol Rev. 65:1162–1197. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Høyer-Hansen M and Jäättelä M: Autophagy:
An emerging target for cancer therapy. Autophagy. 4:574–580. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Wang Y, Hu Y, Sun C, Zhuo S, He Z, Wang H,
Yan M, Liu J, Luan Y, Dai C, et al: Down-regulation of Risa
improves insulin sensitivity by enhancing autophagy. FASEB J.
30:3133–3145. 2016. View Article : Google Scholar : PubMed/NCBI
|