Introduction
Liver fibrosis that progresses by excessive
deposition of the extracellular matrix, is a frequent consequence
of chronic liver damage following hepatitis B virus (HBV) infection
(1,2).
Among liver cells, human stellate cells (HSCs) serve a key role in
the development of liver fibrosis (2).
While the molecular mechanism of fibrosis induction by hepatitis C
virus (HCV) is well described and is mostly attributed to core and
NS3 proteins (3), the responsible HBV
proteins and underlying mechanisms remain obscure. The direct
influence of HBV proteins in the fibrosis process requires further
investigation (4).
In spite of some reports concerning fibrotic roles
of HCV core protein (5–8), the impact of the HBV core protein (HBc)
on HSCs is described poorly. The HBc is the primary structural unit
of the capsid, consisting of 185 amino acids. The carboxyl terminal
region (aa 150–185) has previously been demonstrated to interact
closely with the viral RNA pre-genome or DNA genome (9). The HBV core protein has been identified
to exhibit a tendency for inflammation in the liver site, with
previous studies demonstrating that truncation of the core by
removing this carboxyl end reduces inflammation (10–12).
The research concerning the effect of the HBc
protein on fibrosis is limited. Sequencing data using patients
sample suggested that core protein mutations of the C-terminal
domain contribute to the stage of fibrosis due to functional
alterations (13). The protein
structure of core and HBe proteins are homologous. The exception is
the pre-core sequence in HBe, which helps protein to be secreted
from host cells. A recent study stated that HBe gene exhibited
fibrosis when transfected into a rat stellate cell line (14).
As hepatic stellate cells demonstrated to be a
transient supportive site of HBV infection/replication in
vitro (4) it is noticeable if
endogenous pre-core expression in human HSCs establish a fibrosis
process or not. Several previous studies provided direct evidence
that HBV could replicate and express structural proteins in HSCs
cells (4). As plausible mechanism of
fibrosis, mechanism behind endogenous expression of core and
related derivatives may clarify molecular paths for fibrosis as
well as new targets. Furthermore, the exact role pre-core protein
needed to be investigated.
Materials and methods
Cell culture and plasmids
transfection
The immortalized human stellate cell line, LX-2, was
provided by Professor Scott Friedman (Mount Sinai School of
Medicine, New York City, NY, USA). It displays typical features of
partially active stellate cells that can be employed as a sustained
model for HSC research (15). The
cells were cultured and maintained on 25 cm2 plastic
tissue culture flasks in low glucose Dulbecco's modified Eagle's
medium (DMEM; Gibco; Thermo Fisher Scientific, Inc., Waltham, MA,
USA) supplemented with 2 mM L-glutamine, 100 U/ml
penicillin-streptomycin (Gibco; Thermo Fisher Scientific, Inc.), 5%
fetal bovine serum (Gibco; Thermo Fisher Scientific, Inc.) and
incubated at 37°C in 5% CO2 air humidified
atmosphere.
The pCAGGS plasmids expressing pre-core and HBxAg
proteins was kindly provided by Dr. Gloria Gonzalez-Aseguinolaza
[Centro de Investigación Médica Aplicada (CIMA), Universidad de
Navarra, Pamplona, Spain] and were named pCAG-pre-core and pCAG-X,
respectively. The expression analysis of both plasmids has
previously been performed at this center by western blot analysis
or reverse transcription-quantitative polymerase chain reaction
(RT-qPCR) assays (unpublished data).
LX-2 cells (4×105 cells) were seeded in
six-well plate and, following 24 h, when it reached 80% confluence,
they were transfected with 1 µg DNA plasmid using Lipofectamine
2000™ reagent (Invitrogen; Thermo Fisher Scientific, Inc.) in serum
free DMEM medium, according to the manufacturer's instructions. At
6 h following transfection, the medium was changed with fresh
complete DMEM medium and the plate was left for further 18 h. A
group of cells (equal number to test groups) was treated by 75
ng/ml human leptin, a pro-fibrotic hormone (Sigma-Aldrich; Merck
KGaA, Darmstadt, Germany), was used as a positive control. Hence,
cells that were transfected by empty pCAG plasmid in a separate
well enrolled as negative control group and designated as the
‘plasmid group’.
Cell proliferation assay
The MTT assay was performed to determine the effect
of different components on activated LX-2 cell proliferation as
described before (16). In brief,
7.5×103 cells were seeded into 96-well plates, before
being transfected by Lipofectamine reagent. The cells were
incubated at 37°C in humidified CO2 incubator for 48 h.
At 2 days following incubation, supernatants were changed to fresh
medium and 10µl MTT (Sigma-Aldrich; Merck KGaA) at a concentration
of 5 mg/ml. Dimethylsulfoxide (CinnaGen, Inc., Tehran, Iran) was
added to each well and absorbance was determined by dual
wavelengths of 570 and 630 nm on a microplate ELISA reader (BioTek
Elx 808; BioTek Instruments, Inc., Winooski, VT, USA). Percentage
inhibition of proliferation was calculated as follows: % inhibition
= 100 - [(absorbance of test/absorbance of control) × 100].
RNA extraction and reverse
transcription
Total RNA was extracted from treated cells 36 h post
TGF-β activation, with the help of TRIzol reagent (Invitrogen;
Thermo Fisher Scientific, Inc.) according to manufacturer's
protocol. Total RNA was treated with RQ1 DNase (Promega
Corporation, Madison, WI, USA) to avoid residual DNA contamination.
The cDNA synthesis was made by RocketScript RT Premix kit (Bioneer
Corporation, Daejon, Korea) based on manufacturer's protocol. The
prepared cDNA was stored at −20°C condition until use.
RT-qPCR expression analysis
The primer pairs for α-smooth muscle actin (α-SMA),
collagen type I, α1 chain (COL1A1), TIMP metalloproteinase
inhibitor 1 (TIMP-1) and GAPDH quantification were described and
listed previously (17). The GAPDH
gene was used as reference of normalization in all samples while
H2O as a negative control and RNA mixture were included
in each run. The mRNA levels were relatively quantified using
SYBR-Green I master mix (Takara Bio, Inc., Otsu, Japan) by the help
of StepOnePlus Real-Time PCR system (Applied Biosystems; Thermo
Fisher Scientific, Inc.). The reaction mixtures prepared according
to recommended protocol and amplification reaction was performed in
40 cycles of 10 sec at 94°C for denaturation and 25 sec at 58°C for
the annealing step. The threshold value for each sample was
normalized to the GAPDH signal, and then the 2−∆∆Cq
equation was used for relative expression assay of mRNAs, which
finally was presented as fold-change (18).
ELISA assay for TGF-β1
measurement
At ~36 h following LX-2 cell activation with TGF-β,
released bioactive TGF-β1 in supernatant was measured by the help
of the Human-Mouse TGF-β ELISA kit (eBioscience, Inc.; Thermo
Fisher Scientific, Inc.) according to the manufacturer's
instruction. All the samples were first acid-activated before
starting the method and background TGF-β1 level (due to adding
TGF-β for HSC activation) was deleted from final analysis. The
absorbance was measured at 450 nm by ELISA reader (BioTek
Instruments, Inc.; Elx 808) and then data was calculated against
standard curve and adjusted to pg/ml of culture medium.
Statistical analysis
Statistical analysis was conducting using GraphPad
Prism (version 5; GraphPad Prism Software, Inc., La Jolla, CA, USA)
that used a one-way analysis of variance to evaluate the
differences between means. The statistical significance between
controls and test groups was evaluated by Tukey's post hoc test and
P<0.05 was considered to indicate a statistically significant
difference.
Results
No significant proliferation was
induced by the pre-core gene in LX-2 cells
To compare the anti-proliferative effects of plasmid
expression, LX2 cell viability was measured by MTT assay. Whereas
the proliferation inhibitory impact of all constructs was
detectable on activated HSCs, the significant impact was not
detectable for all constructs. (Data not shown).
Pre-core sequence induced a negligible
fibrotic effect
The mRNA expression levels for pro-fibrotic genes
including TIMP-1, COL1A1, α-SMA were quantified by RT-qPCR
following exposure. As presented in Fig.
1, activated LX-2 (the cells incubated with leptin as positive
control) displayed an increase in production of the extracellular
matrix including α-SMA, COL1A1 and TIMP-1 by 4.3-, 3.9- and
3.1-fold, respectively. Hence, it was demonstrated that the HBX
gene, another HBV protein, also exhibited a significant fibrotic
impact (P<0.01) by 2.2-, 2.3- and 3-fold compared to control
plasmid group or untreated cells. In contrast, the pre-core protein
exhibited no significant effect with an expression upregulation of
1.3, 1.2 and 0.9 for a-SMA, collagen type 1 and TIMP-1,
respectively (all P>0.05) when compared to both plasmid control
group and untreated cell group, as depicted in Fig. 1.
 | Figure 1.The results of gene expression
analysis in LX-2 cells, investigating (A) α-SMA, (B) COL1A1 and (C)
TIMP-1. The pre-core sequence exhibited no significant change in
expression levels of fibrotic molecules in LX-2 cells, when
compared with the plasmid control and untreated cell groups. HBx
significantly presented its fibrotic role, as expression of
pro-fibrotic molecules increased when compared with other groups,
such as pre-core group (P<0.01). Each bar is representative of
the mean ± standard deviation for at least 3 measurements and the
result presents the fold increase/decrease. *P<0.05, **P<0.01
and ***P<0.001. α-SMA, α-smooth muscle actin; COL1A1, collagen
type I, α1 chain; TIMP-1, TIMP metalloproteinase inhibitor-1; NS,
not significant. |
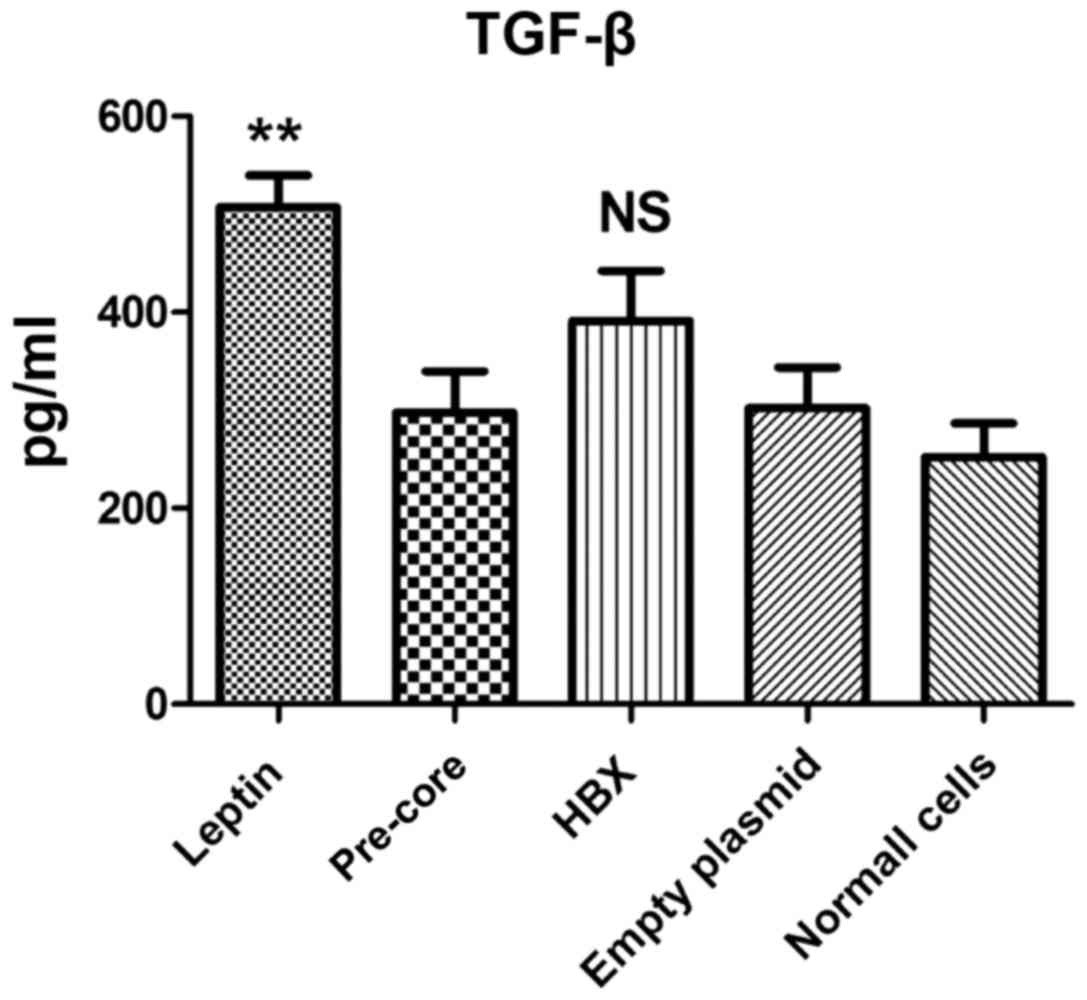
Pre-core sequence did not up-regulate
TGF-β1 production
Following plasmid transfection, TGF-β1 cytokine
production was evaluated in culture supernatants by ELISA assay
(Fig. 2). The result indicated that,
with the exception of leptin treatment, all constructs left no
significant impact on TGF-β1 production on LX-2. In addition, the
data indicated that the cell with no treatment also produced a
measurable level of TGF-β1 (230 pg/ml).
Discussion
Fibrosis is an outcome of elongated liver damage,
arising from HBV chronic infection (19). Similarly, prolonged infection of HBV in
a mouse model may consequently develop fibrosis in liver (20). Hepatic HSCs accounted as the most
important components in the establishment and progression of
fibrosis. The alteration of their gene expression patterns are
responsible for extracellular matrix subversion, which ultimately
leads to fibrosis progression (21,22). The
molecular mechanism of fibrosis by HBV is not clearly clarified and
the direct role of corresponding viral proteins in the process of
fibrosis is also debated (23).
Therefore, the study of the interaction of HBV proteins with this
lineage of cells may facilitate the understanding of causes and
give new opportunity to take control of it.
In one study, HBV-like particles were identified in
the LX-2 cellular cytoplasm with electron microscopy and, while
HSCs were permissive for virus replication, productive replication
of HBV does not occur (4). Regarding
the transient supportive role of HSCs in HBV replication (4), it would be interesting to understand
whether endogenous pre-core expression in human HSCs establishes
fibrosis. A previous study provided direct evidence that HBV could
replicate and express structural proteins in HSCs cells (4). In spite of different reports concerning
the profound role of HCV core protein in liver fibrosis, the impact
of HBV core protein remains to be elucidated in HSCs (5–8).
Some previous studies demonstrated the definitive
fibrotic role of the HBX protein on HSCs in vitro (24–26).
However, the fibrotic role of other HBV proteins is not understood
and requires more investigation. The core protein presented an
inflammatory action in the liver, therefore it may be predicted
that it serves a role in the fibrosis process. Furthermore,
previous works demonstrated that truncation of the core from the
carboxyl end reduces inflammatory property of protein (10,27). The
carboxyl terminal region (aa 150–185) interacts closely with the
viral RNA pre-genome or the viral DNA genome and regulates virus
replication (9). Sequencing data also
suggests that core protein mutations of C-terminal domain
contribute to further progression of liver fibrosis due to function
alteration (13). A recent study by
Zan et al (14) claimed that
the HBe protein, the homologous antigen of core protein exhibited a
fibrotic role in vitro. To the best of the authors'
knowledge, no reports about the pre-core gene have been published
thus far.
The present study demonstrated that, similar to
previous efforts, HBX upregulates α-SMA, COLIA1 and TIMP-1
expression to a level comparable with leptin, a fibrosis inducer
(24–26). Those reports employed HBX protein from
a transfected hepatocyte supernatant, however, the current study
investigated the endogenous HBX expression effect on fibrosis genes
following direct HSC transfection.
The primary outcome of the paper involved the
investigation into the possible effect of pre-core on LX-2 cell
fibrotic gene expression. Assessment of endogenous expression of
the HBV pre-core protein in HSCs and evaluation of subsequent
fibrosis has not been researched prior. Some methodological
limitations should be considered when interpreting these results.
For example, the authors could not analyze protein expression or
relevant secreted forms from supernatant.
These data indicated that the pre-core protein
presented a negligible fibrotic property when compared to the
control plasmid. In comparison to normal LX-2 cells, transfected
cells using different plasmids demonstrated little fibrotic effect
due to DNA structure. Gene expression analysis of COL1A1, α-SMA and
TIMP-1 demonstrated that expression of the pre-core protein
exhibited a similar pattern of fibrosis to control groups. The
authors conclude that the pre-core gene does not perform a fibrotic
role during the fibrosis process of HBV. However, to further
confirm of these in vitro achievements, more investigation in a
suitable animal model is required.
Acknowledgements
The authors would like to thank the Laboratory staff
of the Gastroenterohepatology Research Center, Shiraz University of
Medical Sciences. The current study was funded by Shiraz University
of Medical Sciences (grant no. 94-01-01-9968).
References
|
1
|
Gutierrez-Reyes G, Gutierrez-Ruiz MC and
Kershenobich D: Liver fibrosis and chronic viral hepatitis. Arch
Med Res. 38:644–651. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Mormone E, George J and Nieto N: Molecular
pathogenesis of hepatic fibrosis and current therapeutic
approaches. Chem Biol Interact. 193:225–231. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Khanizadeh S, Ravanshad M, Hosseini SY,
Davoodian P, Zadeh AN, Sabahi F, Sarvari J, Khanlari Z and
Hasani-Azad M: The possible role of NS3 protease activity of
hepatitis C virus on fibrogenesis and miR-122 expression in hepatic
stellate cells. Acta Virol. 60:242–248. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Liu X, Zhu ST, You H, Cong M, Liu TH, Wang
BE and Jia JD: Hepatitis B virus infects hepatic stellate cells and
affects their proliferation and expression of collagen type I. Chin
Med J (Engl). 122:1455–1461. 2009.PubMed/NCBI
|
|
5
|
Bataller R, Paik YH, Lindquist JN,
Lemasters JJ and Brenner DA: Hepatitis C virus core and
nonstructural proteins induce fibrogenic effects in hepatic
stellate cells. Gastroenterology. 126:529–540. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Coenen M, Nischalke HD, Krämer B, Langhans
B, Glässner A, Schulte D, Körner C, Sauerbruch T, Nattermann J and
Spengler U: Hepatitis C virus core protein induces fibrogenic
actions of hepatic stellate cells via toll-like receptor 2. Lab
Invest. 91:1375–1382. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Shin JY, Hur W, Wang JS, Jang JW, Kim CW,
Bae SH, Jang SK, Yang SH, Sung YC, Kwon OJ, et al: HCV core protein
promotes liver fibrogenesis via up-regulation of CTGF with
TGF-beta1. Exp Mol Med. 37:138–145. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wu CF, Lin YL and Huang YT: Hepatitis C
virus core protein stimulates fibrogenesis in hepatic stellate
cells involving the obese receptor. J Cell Biochem. 2012.
|
|
9
|
Roseman AM, Berriman JA, Wynne SA, Butler
PJ and Crowther RA: A structural model for maturation of the
hepatitis B virus core. Proc Natl Acad Sci USA. 102:pp.
15821–15826. 2005; View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lee BO, Tucker A, Frelin L, Sallberg M,
Jones J, Peters C, Hughes J, Whitacre D, Darsow B, Peterson DL and
Milich DR: Interaction of the hepatitis B core antigen and the
innate immune system. J Immunol. 182:6670–6681. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Manigold T, Böcker U, Chen J, Gundt J,
Traber P, Singer MV and Rossol S: Hepatitis B core antigen is a
potent inductor of interleukin-18 in peripheral blood mononuclear
cells of healthy controls and patients with hepatitis B infection.
J Med Virol. 71:31–40. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Vanlandschoot P, van Houtte F, Serruys B
and Leroux-Roels G: The arginine-rich carboxy-terminal domain of
the hepatitis B virus core protein mediates attachment of
nucleocapsids to cell-surface-expressed heparan sulfate. J Gen
Virol. 86:75–84. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mohamadkhani A, Jazii FR, Poustchi H,
Nouraein O, Abbasi S, Sotoudeh M and Montazeri G: The role of
mutations in core protein of hepatitis B virus in liver fibrosis.
Virol J. 6:2092009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zan Y, Zhang Y and Tien P: Hepatitis B
virus e antigen induces activation of rat hepatic stellate cells.
Biochem Biophys Res Commun. 435:391–396. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Xu L, Hui AY, Albanis E, Arthur MJ,
O'Byrne SM, Blaner WS, Mukherjee P, Friedman SL and Eng FJ: Human
hepatic stellate cell lines, LX-1 and LX-2: New tools for analysis
of hepatic fibrosis. Gut. 54:142–151. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Hosseini SY, Kalantar K, Shahin K, Ghayour
M, Rajabibazl M, Fattahi MR, et al: Comparison of the In Vitro
Antifibrogenic Effects of Silymarin, Silybin A and 18α-Glycyrrhizin
on Activated Hepatic Stellate Cells. Jundishapur J Nat Pharm Prod
Inpress. e402852017.
|
|
17
|
Khanizadeh S, Ravanshad M, Hosseini S,
Davoodian P, Zadeh A Nejati and Sarvari J: Blocking of SMAD4
expression by shRNA effectively inhibits fibrogenesis of human
hepatic stellate cells. Gastroenterol Hepatol Bed Bench. 8:262–269.
2015.PubMed/NCBI
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Guo XL, Liang B, Wang XW, Fan FG, Jin J,
Lan R, et al: Glycyrrhizic acid attenuates CCl4-induced hepatocyte
apoptosis in rats via a p53-mediated pathway. World journal of
gastroenterology. World J Gastroenterol. 19:37812013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ye L, Yu H, Li C, Hirsch ML, Zhang L,
Samulski RJ, Li W and Liu Z: Adeno-Associated Virus Vector Mediated
Delivery of the HBV Genome Induces Chronic Hepatitis B Virus
Infection and Liver Fibrosis in Mice. PLoS One. 10:e01300522015.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Reeves HL and Friedman SL: Activation of
hepatic stellate cells - a key issue in liver fibrosis. Front
Biosci. 7:d808–d826. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kisseleva T and Brenner DA: Hepatic
stellate cells and the reversal of fibrosis. J Gastroenterol
Hepatol. 21 Suppl 3:S84–S87. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Lin J, Wu JF, Zhang Q, Zhang HW and Cao
GW: Virus-related liver cirrhosis: Molecular basis and therapeutic
options. World J Gastroenterol. 20:6457–6469. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Martín-Vílchez S, Sanz-Cameno P,
Rodríguez-Muñoz Y, Majano PL, Molina-Jiménez F, López-Cabrera M,
Moreno-Otero R and Lara-Pezzi E: The hepatitis B virus X protein
induces paracrine activation of human hepatic stellate cells.
Hepatology. 47:1872–1883. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Guo GH, Tan DM, Zhu PA and Liu F:
Hepatitis B virus X protein promotes proliferation and upregulates
TGF-beta1 and CTGF in human hepatic stellate cell line, LX-2.
Hepatobiliary Pancreat Dis Int. 8:59–64. 2009.PubMed/NCBI
|
|
26
|
Chen HY, Chen ZX, Huang RF, Lin N and Wang
XZ: Hepatitis B virus X protein activates human hepatic stellate
cells through upregulating TGFbeta1. Genetics and molecular
research. GMR. 13:8645–8656. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wu JF, Ni YH, Chen HL, Hsu HY and Chang
MH: The impact of hepatitis B virus precore/core gene carboxyl
terminal mutations on viral biosynthesis and the host immune
response. J Infect Dis. 209:1374–1381. 2014. View Article : Google Scholar : PubMed/NCBI
|