Introduction
Next-generation sequencing (NGS) of nucleic acids is
increasingly being used in rare tumors for diagnosis and actionable
biomarkers identification (1,2). Failed
tumor NGS testing can result in the inability to provide
diagnostic, prognostic, and therapeutic information, thereby
increasing healthcare costs (3-5).
Several factors related to NGS failure have been previously
reported, including tumor type, sampling method, sample size, tumor
volume, tissue type, tumor fraction, DNA yield, decalcifying
procedures and the age of the paraffin blocks (3-6).
The integrity and concentration of nucleic acids, along with
high-quality libraries, are essential to ensuring successful NGS
analysis (7,8). In published studies, NGS failure rates
exhibit considerable variation, ranging from 4.1 to 22.5%, with
discrepancies in the predictive factors for NGS failure reported
across different investigations (3-6).
Traditionally, the NGS testing process is divided into the
pre-analytical, analytical, and post-analytical phases, with the
pre-analytical phase being responsible for NGS failure in 90% of
cases (5). However, assay-related
variables can potentially modify preanalytical causes of NGS
failure. For example, a larger panel size or the use of
hybridization capture can increase DNA or RNA input
requirements.
Most published studies in the field have
predominantly included patients with prevalent tumors and data on
NGS failure rates and their predictors in rare tumors are limited
(3-7).
In rare tumors, the frequent diagnostic challenges and absence of
standardized therapeutic options may prompt physicians to utilize
larger panels, often incorporating RNA sequencing, which can be
particularly useful for precise sarcoma diagnosis. The present
study aimed to assess the NGS failure rate in sarcomas, rare
carcinomas, and melanomas, and analyze its association with
pre-analytical variables and sequencing methods at a single
institution in Oman. The co-primary objectives of the present study
were to estimate the proportion of patients with rare tumors who
experienced NGS failure and the proportion of assays that resulted
in a failed report. The secondary objectives were to estimate the
proportion of patients who achieved successful NGS after repeat
testing and to identify predictors of NGS failure.
Data and methods
Study setting, design and data
collection period
An observational study was conducted at the Sultan
Qaboos Comprehensive Cancer Care and Research Center (SQCCCRC) in
Muscat, Oman. From the institutional molecular pathology database,
the list of patients treated in the Rare Tumors Program who
underwent molecular profiling at SQCCCRC between January 1, 2022,
and October 30, 2023 was retrieved. The study database was updated
every 3-6 months. NGS tests were ordered for selected patients with
rare tumors at the discretion of their physicians, based on their
clinical judgment, and were performed as part of routine patient
care. NGS testing was either performed at a referral laboratory
(CARIS Life Sciences) or at the SQCCCRC molecular pathology
laboratory.
Eligibility
Patients of any sex aged ≥13 years, with a
histopathological diagnosis of sarcomas of any grade, rare
melanomas, carcinomas of unknown primary site, or rare
carcinosarcomas were eligible for inclusion in the study. Ethical
approval for this study was provided by the Sultan Qaboos
Comprehensive Cancer Care and Research Center Institutional Review
Board and Ethics Committee on November 10, 2022, (approval no.
CCCRC-27-2022; Muscat, Oman). Patient consent to participate was
waived due to the retrospective nature of the study.
Procedures
For each patient included in the study, one of the
following two assays was performed: i) Somatic whole
exome/transcriptome sequencing (WETS) using the CARIS MI Profile™
(CARIS Life Sciences) (https://portal.caris.ai/); or ii) targeted sequencing
using the 550 gene panel Oncomine™ Comprehensive Assay Plus (OCA
Plus) (https://www.thermofisher.com/om/en/home/clinical/preclinical-companion-diagnostic-development/oncomine-oncology/oncomine-cancer-research-panel-workflow/oncomine-comprehensive-assay-plus.html).
Both assays were conducted on formalin-fixed
paraffin embedded tissue specimens. Tumor enrichment was performed
using micro-dissection (CARIS MI Profile) or macro-dissection (OCA
Plus). The CARIS MI Profile assay used a hybrid pull-down panel of
baits and sequencing on Illumina sequencing platforms (NextSeq™ or
NovaSeq600™ https://www.illumina.com/systems/sequencing-platforms/novaseq.html).
The in-house OCA Plus assay involved the preparation of libraries
(DNA, 1.4MB across 501 genes; RNA, 49 driver fusions), followed by
sequencing using the Ion S5plus™ platform (Thermo Fisher
Scientific, Inc.) (https://www.thermofisher.com/order/catalog/product/A38195).
Outcomes
The co-primary outcomes were the proportion of
patients and the proportion of reports with failed NGS. The
secondary outcome was the proportion of patients with a successful
NGS after repeated testing.
Data, variables, and statistical
analysis
Clinicopathological and NGS-related data were
extracted from electronic medical records. The storage time or
paraffin block age was calculated as the time difference between
the reporting and sampling dates. Testing was defined as failed
when the reporting laboratory indicated an inability to meet the
laboratory-defined quality requirements or cited an insufficient
quantity or quality of DNA/RNA as stated in the NGS report. Since
CARIS MI reports included results of DNA and RNA sequencing and
immunohistochemistry analysis, an NGS test was defined as failed if
either the DNA or RNA sequencing failed, regardless of the
immunohistochemistry outcome. The study had two units of analysis:
Patients and NGS reports. NGS failure rates were expressed as
proportions. 95% Confidences intervals (CIs) for proportions were
computed by the exact method. Continuous variables were summarized
using the median and interquartile range (IQR). Univariable
logistic regression models were constructed with the outcome
variable NGS failure and the following explanatory variables: assay
type, sampling method, source tissue, and storage time. Odds ratios
(ORs) and 95% CIs were reported. All subgroup analyses were
exploratory. A value of P<0.05 was considered to indicate a
statistically significant difference. SAS (version 9.4; SAS
Institute) and R statistical software (version 4.3; R Foundation
for Statistical Computing) were used for statistical analysis.
Results
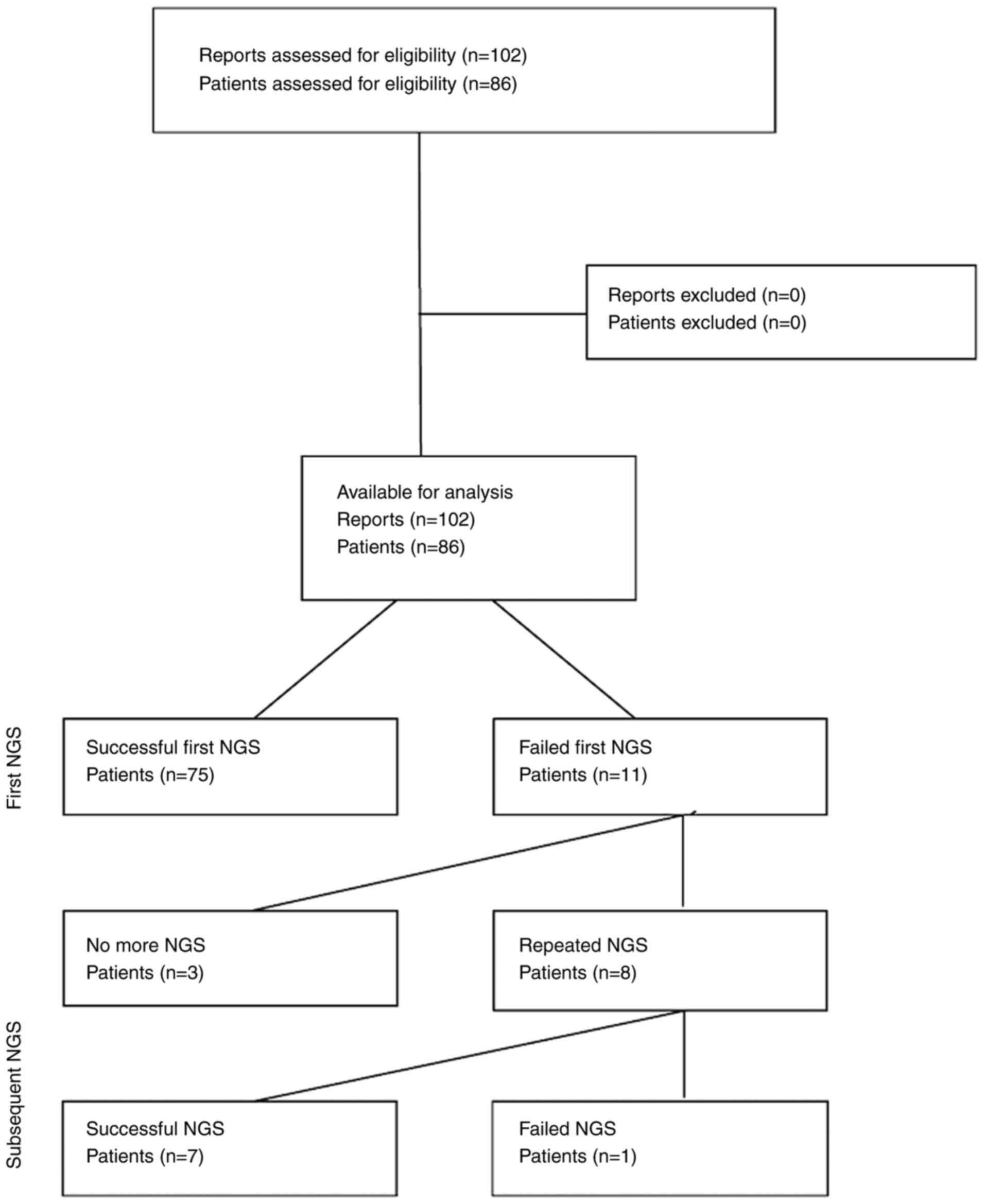
A total of 102 NGS reports from 86 patients with
sarcomas (73.3%), rare carcinomas (16.3%) and melanomas (10.5%) met
the eligibility criteria (Fig. 1 and
Table I). The median age of the
patients was 40 years (IQR, 23-61 years) and 48.8% of the patients
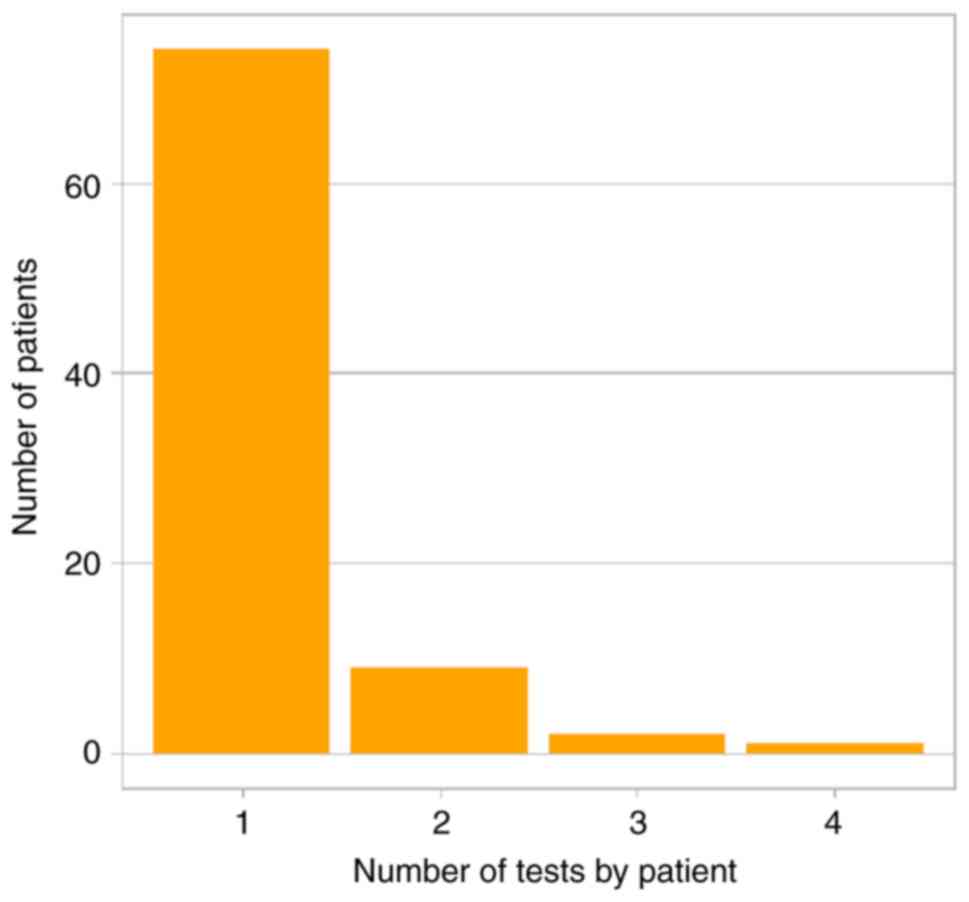
were female. A single NGS test was performed in 83.7% of the
patients, while 16.3% of the patients underwent multiple NGS tests.
Samples were obtained by core biopsy (51%) or surgery (48%) from
soft tissue (92.1%) or bone lesions (7.9%). The median storage time
was 2.5 months (IQR, 1.3-4.6). Targeted sequencing and WETS were
used in 39.2 and 60.8% of reports, respectively (Table II). The sampling method, source
tissue, and storage time were unavailable for 1 out of 102
reports.
 | Table IPatient characteristics. |
Table I
Patient characteristics.
| Characteristic | Value |
|---|
| Age, median
(IQR) | 40 (23-61) |
| Male sex, n (%) | 44/86 (51.2%) |
| Diagnosis, n (%) | |
|
Sarcoma | 63/86 (73.3%) |
|
Carcinoma | 14/86 (16.3%) |
|
Melanoma | 9/86 (10.5%) |
| No. of tests | |
|
Single
test | 72/86 (83.7%) |
|
Repeated
tests | 14/86 (16.3%) |
|
NGS
failure | 4/86 (4.7%) |
 | Table IIAssay and sampling
characteristics. |
Table II
Assay and sampling
characteristics.
| Characteristic | Value |
|---|
| Storage time in
months, median (IQR) | 2.5 (1.3-4.6) |
| Sampling method | |
|
Biopsy | 52/101 (51%) |
|
Surgery | 49/101 (48%) |
|
Assay | |
| Targeted panel | 40/102 (39.2%) |
| WES | 62/102 (60.8%) |
| Tissue | |
|
Soft
tissue | 93/101 (92.1%) |
|
Bone | 8/101 (7.9%) |
|
NGS
failure | 15/102 (14.7%) |
Of the 86 patients, 87.2% had a successful initial
NGS testing, while 12.8% had insufficient quantity or quality of
material. Of the 11 patients with a failed first test, 8 patients
underwent repeat NGS testing up to three times (Fig. 2). Repeated testing was successful in
7 out of 8 (87.5%) patients. NGS failure after any number of tests
was observed in 4 out of the 86 patients (4.7%; 95% CI, 1.3-11.5%),
3 of whom had a single NGS test, and 1 patient underwent repeated
NGS testing (Table SI).
Of the 102 tests, NGS failure due to insufficient
material quantity or quality was observed in 14.7%; (95% CI,
8.5-33.1%), corresponding to 4/86 patients (4.7%; 95% CI,
1.3-11.5%). WETS was significantly associated with a higher
probability of NGS failure due to low material quantity or quality
compared with the targeted panel (OR, 11.4; 95% CI, 1.4-90.4;
P=0.022; Table III). No other
variable significantly predicted NGS failure. The 4 patients who
did not have any successful NGS testing were diagnosed with
sarcoma. The association between paraffin block age, assay type,
and NGS failure is presented Fig.
S1. The supporting anonymized dataset is publicly available at
https://doi.org/10.5281/zenodo.14652764.
 | Table IIIUnivariable logistic regression of
assay sample-related characteristics on the probability of NGS
failure. |
Table III
Univariable logistic regression of
assay sample-related characteristics on the probability of NGS
failure.
| Explanatory
variable | N | Odds ratio | Lower 95% CI | Upper 95% CI | P-value |
|---|
| Assay (WETS vs.
targeted) | 102 | 11.38 | 1.43 | 90.35 | 0.022 |
| Sampling method
(surgery vs. biopsy) | 101 | 1.25 | 0.42 | 3.77 | 0.69 |
| Tissue (bone vs.
soft) | 101 | 2.05 | 0.37 | 11.28 | 0.41 |
| Storage time (1
month)a | 101 | 1.04 | 0.98 | 1.11 | 0.18 |
Discussion
The present study analyzed the NGS testing failure
rate in specimens from patients with rare tumors, including
sarcomas, rare carcinomas and rare melanomas. It was found that NGS
testing in rare tumors had a failure rate of 14.7% in the tested
samples, which is comparable to that reported in more prevalent
hematological and solid tumors (Table
SII). To the best of our knowledge, this is the first study on
NGS failure due to material quantity and quality issues
specifically dedicated to the rare tumors. In our study, the NGS
assay type was the only predictor of NGS failure; however, the
strength of this association was imprecisely estimated, as
reflected by the wide confidence interval around the odds ratio.
Tumor type, sampling site and method, tissue type, and sample age
were not significantly associated with the probability of NGS
failure (9,10).
The small sample size of our study represents its
primary limitation. Other limitations include the lack of
quantification of sample volume and tumor cellularity. In a
previous study on solid and hematolymphoid tumors, insufficient
tumor tissue available for DNA extraction was a major factor
associated with NGS failure (5).
Small samples obtained from minimally invasive procedures, such as
fine-needle aspiration and core needle biopsy, may yield small
quantities of DNA, impacting the success rate of NGS testing.
However, no significant relationship between the sampling method
and the probability of NGS failure was observed in the present
study. The amount of DNA needed for NGS depends on the sequencing
technology, panel size, enrichment method and expected sequencing
depth (9,10). While the Illumina platform
successfully operates with small tissue samples containing 10-70 ng
of input DNA, the Ion Torrent platform has even lower requirement
for the amount of input DNA, needing only 10 ng of DNA for the Ion
PGM cancer hotspot panel (5,11). In the present study, there was a
significant difference between the panel size of the two assays:
This was ~1.4 MB for the OCA Plus and ~30 MB for the CARIS MI
Profile. Additionally, the CARIS MI Profile used the hybrid capture
enrichment method whereas the OCA Plus employed the PCR. Hybrid
capture enrichment has a higher DNA input requirement, particularly
when compared to amplicon-based assays. It was hypothesized that
the differences in the panel size and enrichment strategy
contributed to the variations in failure rates observed between the
two assays in this study. Regarding other potential predictors of
NGS failure, the small sample size in the present study may have
limited its statistical power to identify significant associations
with NGS failure (12,13). Some of the potential predictors of
NGS failure, such as the number of cores and specimen size,
clinical setting of biopsy, the number of cores and specimen size,
tumor cellularity and heterogeneity were mot analyzed in this study
and should be addressed in the future research.
While both the OCA Plus and CARIS MI Profile assays
screen for various types of somatic variants, including single
nucleotide variants, indels, and structural variants, the OCA Plus
assay cannot detect common fusions observed in certain sarcomas
(e.g., Ewing sarcoma and synovial sarcoma), limiting its diagnostic
utility for these subtypes. However, the majority of
therapeutically relevant or actionable fusions, such as
neurotrophic tyrosine receptor kinase (NTRK) 1/2/3, can be detected
using the OCA Plus panel. WETS-based assays are ideal for
identifying any potential gene fusions and actionable variants and
accurately estimating tumor mutation burden. However, these assays
require a higher quantity and quality of DNA. Thus, a customized
approach is required to identify the most appropriate panel for
each patient. The findings of the present study suggest that a
comprehensive targeted gene panel may be preferable to WETS when
the tissue sample is small, and the identification of therapeutic
biomarkers is more critical than diagnosis refinement.
Several studies investigating the frequency of NGS
failure and its predictors in hematologic and solid malignancies
have been published to date. The substantial heterogeneity within
and between study populations, sampling sites and methods, tissue
processing methods (e.g., fresh tissue vs. paraffin-embedded
blocks), and types of NGS assays used makes cross-study comparison
or pooling challenging and may help explain the observed variation
in NGS failure rates and inconsistencies among identified NGS
predictors (Table SII). More
homogeneous populations subjected to uniform interventions should
be considered in the future studies to improve the precision of
estimations and meaningfulness of conclusions.
In conclusion, the results of the present study
suggest that the NGS failure rate in rare tumors is comparable to
previously reported rates in more common neoplasms. The NGS failure
rate was significantly higher with WETS compared to the targeted
panel assay. In the majority of cases, retesting patient samples
successfully overcame the initial NGS testing failure. However, the
findings presented herein require validation in future studies.
Supplementary Material
Association between paraffin block
age, assay type and NGS failure. NGS, next-generation sequencing.
WES, whole exome and transcriptome sequencing.
Repeated NGS testing for 11 patients
with insufficient quantity or quality of material.
Selected studies reporting on NGS
failure.
Acknowledgements
The authors would like to extend their gratitude to
the molecular pathology laboratory workers, Ms. Amna Al Khusaibi,
Mr. Amrou Al Busaidi, Ms. Rahma Al Adawi, Ms. Ruqaya Al Balushi,
Ms. Shahad Al Raisi, Ms. Mariya Al Kalbani, and Ms. Hasna Al Saadi,
SQCCCRC, Muscat, Oman for their technical expertise and
contribution in generating the assay results included in this
study.
Funding
Funding: No funding was received.
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author. The supporting anonymized
dataset ‘NGS Failure dataset supporting the article.xls’ is
publicly available at https://doi.org/10.5281/zenodo.14652764.
Authors' contributions
BI, PAD, SAZ and IAH conceptualized the study. BI,
PAD, AP and DA were responsible for data curation and validation.
HAS and BI performed the formal analysis. BI, HAS, PAD and DA were
responsible for the investigation. BI, HAS and PAD developed the
methodology. BI was the project administrator and was responsible
for data visualization. BI and HAS were responsible for the use of
software. PAD, AP, SAZ and IAH collected resources. SAZ and IAH
supervised the project. BI, HAS, PAD and DA wrote the draft of the
manuscript. All authors have read and approved the final version of
the manuscript. PAD and HAS confirm the authenticity of the raw
data.
Ethics approval and consent to
participate
Ethical approval for the present study was provided
by the Sultan Qaboos Comprehensive Cancer Care and Research Center
Institutional Review Board and Ethics Committee on November 10,
2022 (approval no. CCCRC-27-2022; Muscat, Oman). Patient consent to
participate was waived due to the retrospective nature of the
study.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Use of artificial intelligence tools
During the preparation of this work, AI tools were
used to improve the readability and language of the manuscript or
to generate images, and subsequently, the authors revised and
edited the content produced by the AI tools as necessary, taking
full responsibility for the ultimate content of the present
manuscript.
References
|
1
|
Pestana RC, Moyers JT, Roszik J, Sen S,
Hong DS, Naing A, Herzog CE, Fu S, Piha-Paul SA, Rodon J, et al:
Impact of biomarker-matched therapies on outcomes in patients with
sarcoma enrolled in early-phase clinical trials (SAMBA 101). Clin
Cancer Res. 29:1708–1718. 2023.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Pestana RC, Groisberg R, Roszik J and
Subbiah V: Precision oncology in sarcomas: Divide and conquer. JCO
Precis Oncol. 3(PO.18.00247)2019.PubMed/NCBI View Article : Google Scholar
|
|
3
|
da Silveira Corrêa B, De-Paris F, Viola
GD, Andreis TF, Rosset C, Vianna FSL, da Rosa Rivero LF, de
Oliveira FH, Ashton-Prolla P and de Souza Macedo G: Challenges to
the effectiveness of next-generation sequencing in formalin-fixed
paraffin-embedded tumor samples for non-small cell lung cancer. Ann
Diagn Pathol. 69(152249)2024.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Cho M, Ahn S, Hong M, Bang H, van Vrancken
M, Kim S, Lee J, Park SH, Park JO, Park YS, et al: Tissue
recommendations for precision cancer therapy using next generation
sequencing: A comprehensive single cancer center's experiences.
Oncotarget. 8:42478–42486. 2017.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Al-Kateb H, Nguyen TT, Steger-May K and
Pfeifer JD: Identification of major factors associated with failed
clinical molecular oncology testing performed by next generation
sequencing (NGS). Mol Oncol. 9:1737–1743. 2015.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Goswami RS, Luthra R, Singh RR, Patel KP,
Routbort MJ, Aldape KD, Yao H, Dang HD, Barkoh BA, Manekia J, et
al: Identification of factors affecting the success of
next-generation sequencing testing in solid tumors. Am J Clin
Pathol. 145:222–237. 2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Chen H, Luthra R, Goswami RS, Singh RR and
Roy-Chowdhuri S: Analysis of pre-analytic factors affecting the
success of clinical next-generation sequencing of solid organ
malignancies. Cancers (Basel). 7:1699–1715. 2015.PubMed/NCBI View Article : Google Scholar
|
|
8
|
de Biase D, Fassan M and Malapelle U:
Next-Generation Sequencing in tumor diagnosis and treatment.
Diagnostics (Basel). 10(962)2020.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Song P, Chen SX, Yan YH, Pinto A, Cheng
LY, Dai P, Patel AA and Zhang DY: Selective multiplexed enrichment
for the detection and quantitation of low-fraction DNA variants via
low-depth sequencing. Nat Biomed Eng. 5:690–701. 2021.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Gupta N and Verma VK: Next-Generation
sequencing and its application: Empowering in public health beyond
reality. In: Microbial Technology for the Welfare of Society. Arora
PK (ed.) Springer, Singapore, pp313-341, 2019.
|
|
11
|
Chung J, Son DS, Jeon HJ, Kim KM, Park G,
Ryu GH, Park WY and Park D: The minimal amount of starting DNA for
Agilent's hybrid capture-based targeted massively parallel
sequencing. Sci Rep. 6(26732)2016.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Durães C, Gomes C, Costa J and Quagliata
L: Demystifying the discussion of sequencing panel size in oncology
genetic testing. EMJ. 7:68–77. 2022.
|
|
13
|
Dumur CI, Krishnan R, Almenara JA, Brown
KE, Dugan KR, Farni C, Ibrahim FZ, Sanchez NA, Rathore S, Pradhan D
and Hughes JH: Analytical validation and clinical utilization of
the oncomine comprehensive assay plus panel for comprehensive
genomic profiling in solid tumors. J Mol Pathol. 4:109–127.
2023.
|