Introduction
Head and neck squamous cell carcinoma (HNSCC) is the
sixth most common type of cancer in the world (1). It is an epithelial cancer arising in the
upper aerodigestive tract, including the pharynx, larynx and oral
cavity (2). Furthermore, the head and
neck region contains several distinct structures, such as the lips,
nasopharynx, oropharynx and hypopharynx, which result in the large
heterogeneity of HNSCC (2,3). In total, >600,000 novel cases of
HNSCC are diagnosed annually (1).
Currently, chemotherapy or radiotherapy with locoregional treatment
is used for HNSCC patients (4,5). However,
the survival rate of this disease is only 40–50% within 5 years
after diagnosis and treatment (6).
Numerous studies have explored the pathological
mechanism underlying the development of HNSCC (7,8). Several
genes have been identified to participate in the progression of
HNSCC. For example, Zhang et al (9) reported that fos-related activator-1
could be used as a potential therapeutic target gene in oral
squamous cell carcinoma, while transgelin 2 has an oncogenic
function and may be regulated by the tumor suppressor microRNA-1 in
HNSCC (7). Aberrant promoter
methylation of the Nei endonuclease VIII-like 1 gene has a
critical role in the progression and development of HNSCC (8). Certain signaling pathways have also been
demonstrated to be important in HNSCC. For example, Pedrero et
al (10) reported that
dysregulation of the phosphatidylinositol-4,5-bisphosphate
3-kinase/AKT/phosphatase and tensin homolog signaling pathway may
contribute to early HNSCC tumorigenesis. In addition,
cyclooxygenase-2 (COX-2) signaling pathway is closely associated
with tumor angiogenesis in HNSCC, and COX-2 overexpression predicts
a shorter survival in patients with head and neck cancer (11). The coactivation of the
mitogen-activated protein kinase and IκB kinase signaling pathways
may suppress the mechanism of signal transduction by regulating the
secretion of interleukin-8 and vascular endothelial growth factor
in human HNSCC (12). Although
numerous factors have been identified to contribute to HNSCC, the
pathogenic mechanisms of HNSCC remain to be clearly demonstrated in
order to identify potential target genes for the treatment of
HNSCC.
In the present study, the differentially expressed
genes (DEGs) between HNSCC and normal samples were analyzed to gain
a better insight of HNSCC. Gene Ontology (GO) and Kyoto
Encyclopedia of Genes and Genomes (KEGG) enrichment analyses of
DEGs were performed, and the protein-protein interaction (PPI)
network of these DEGs was constructed. The purpose of the present
study was to explore the underlying mechanisms of HNSCC and to
identify novel potential target genes for HNSCC therapy.
Materials and methods
Affymetrix microarray data
Gene Expression Omnibus (GEO; www.ncbi.nlm.nih.gov/geo/) is a database repository of
high throughput gene expression data, which segregates data into
three principle components: Platform (GPL), series (GSE) and sample
(GSM). The array data of GSE6631, based on the GPL8300 Affymetrix
Human Genome U95 Version 2 Array platform (Affymetrix, Inc., Santa
Clara, CA, USA) was downloaded from the GEO database, which was
deposited by Kuriakose et al (13). The dataset was generated from paired
(from the same patient) samples of tumor and normal tissues from 22
patients with histologically confirmed HNSCC by Kuriakose et
al (13).
Data preprocessing and DEGs
analysis
The original probe-level data in CEL files (raw
probe level data) were converted into gene expression values. Data
were normalized using the Bioconductor R package affy version
1.32.0 (Affymetrix, Inc., Santa Clara, CA, USA) (14). Nonspecific probes were filtered. If
multiple probes corresponded to the same gene, the average
expression value was calculated to represent the expression levels
of that gene. The samr package (version 2.0; cran.r-project.org/web/packages/samr/index.html) in R
(www.r-project.org/) (15) was applied to identify DEGs between
HNSCC and normal samples. ∆=1.3 and fold-change >1.5 were used
as the cutoff criteria, based on the experience of the present
authors.
Functional enrichment analysis of
DEGs
The GO database (geneontology.org/page/go-database) (16) is a collection of numerous gene
annotation terms. The knowledge contained in the KEGG database
(www.genome.jp/kegg/) (17) was applied to identify functional and
metabolic pathways. The Database for Annotation, Visualization and
Integrated Discovery (DAVID) version 6.7 (National Cancer Institute
at Frederick, Frederick, MD, USA) (18) was used as a gene functional enrichment
analysis tool to understand the biological meaning of the results
of bioinformatics analysis. GO and KEGG enrichment analyses for the
upregulated and downregulated identified DEGs were performed with
DAVID. P<0.05 and false discovery rate <0.01 were selected as
the cutoff criteria.
Construction of PPI network and
disease enrichment analysis
The Search Tool for the Retrieval of Interacting
Genes/Proteins (version 9.05; string-db.org)
(19) is an online database that
contains comprehensive information of proteins. This online tool
was applied to analyze the interactions of protein pairs. PPI
network of DEGs was constructed using Cytoscape software (version
3.0.1; Cytoscape Consortium San Diego, CA, USA) (20). The degree of connectivity was analyzed
and used to obtain the hub proteins in the PPI network.
Results
Identification of DEGs
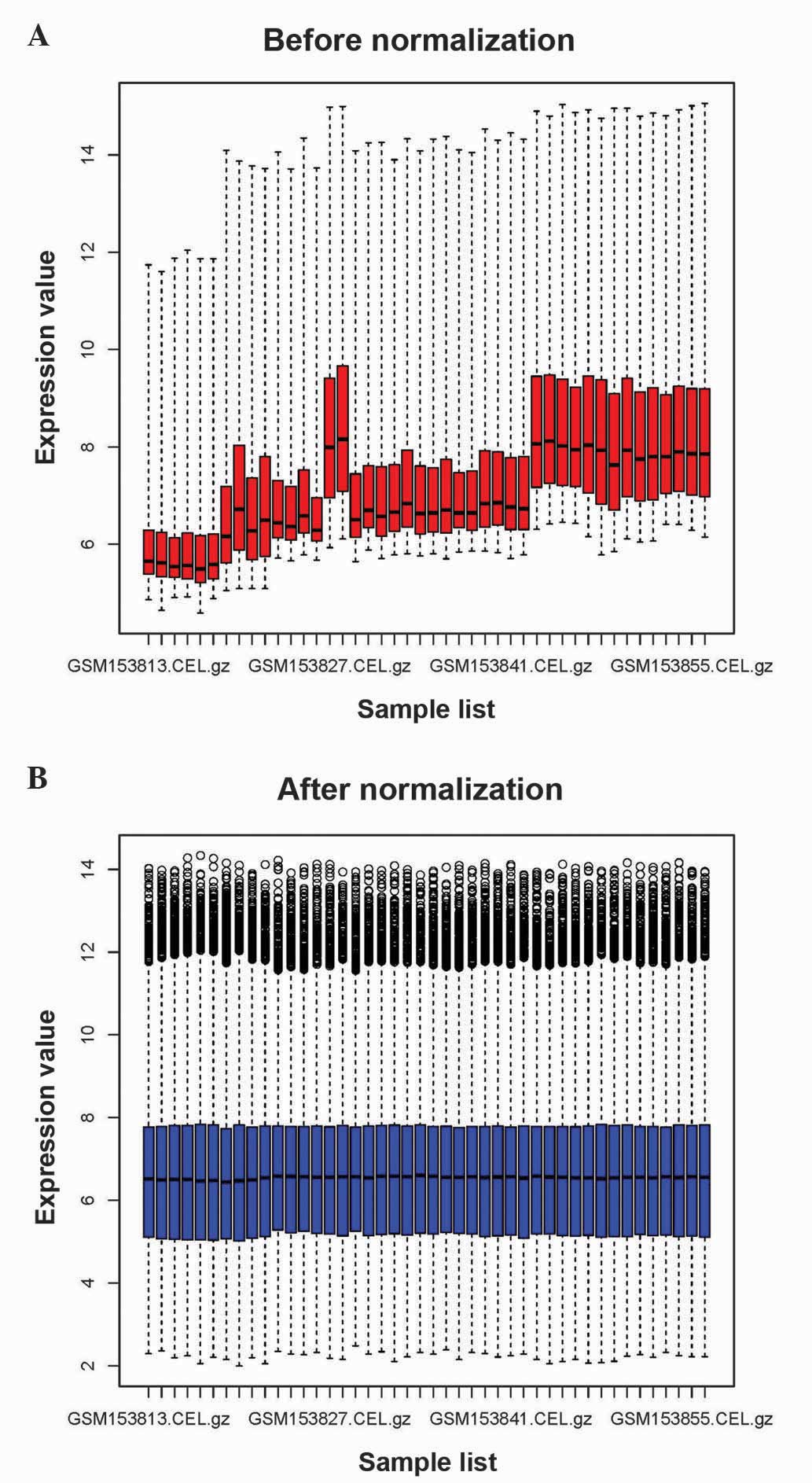
As represented in Fig.
1, the raw expression data were preprocessed and normalized. A
total of 419 DEGs were identified between HNSCC and normal samples,
including 196 upregulated genes and 223 downregulated genes.
Functional enrichment analysis of
DEGs
A total of 39 GO terms of upregulated and
downregulated DEGs were obtained. The top 5 GO terms of upregulated
and downregulated genes are indicated in Table I. The upregulated DEGs were
significantly associated with cell adhesion, extracellular matrix
(ECM) organization, collagen metabolic process and proteinaceous
ECM, while the downregulated genes were mainly involved in
epidermis development, ectoderm development and epidermal cell
differentiation.
 | Table I.GO terms most frequently enriched by
upregulated and downregulated DEGs in head and neck squamous cell
carcinoma. |
Table I.
GO terms most frequently enriched by
upregulated and downregulated DEGs in head and neck squamous cell
carcinoma.
| Category | Term | Counta | P-value | FDR |
|---|
| Upregulated DEGs |
|
|
|
|
|
GOTERM_BP | GO:0007155~cell
adhesion | 42 | 4.92E-17 | 8.08E-14 |
|
GOTERM_BP | GO:0022610~biological
adhesion | 42 | 5.18E-17 | 8.50E-14 |
|
GOTERM_BP |
GO:0030198~extracellular matrix
organization | 17 | 2.12E-13 | 3.48E-10 |
|
GOTERM_BP |
GO:0043062~extracellular structure
organization | 19 | 2.20E-12 | 3.62E-09 |
|
GOTERM_BP | GO:0032963~collagen
metabolic process | 10 | 3.75E-11 | 6.15E-08 |
|
GOTERM_CC |
GO:0005578~proteinaceous extracellular
matrix | 40 | 1.88E-26 | 2.48E-23 |
|
GOTERM_CC |
GO:0031012~extracellular matrix | 40 | 3.23E-25 | 4.27E-22 |
|
GOTERM_CC |
GO:0044420~extracellular matrix part | 25 | 5.83E-22 | 7.70E-19 |
|
GOTERM_CC |
GO:0044421~extracellular region part | 56 | 4.52E-21 | 5.97E-18 |
|
GOTERM_CC |
GO:0005581~collagen | 14 | 3.91E-16 | 5.88E-13 |
|
GOTERM_MF |
GO:0005201~extracellular matrix structural
constituent | 15 | 3.11E-12 | 4.32E-09 |
|
GOTERM_MF |
GO:0050840~extracellular matrix
binding | 8 | 3.24E-08 | 4.50E-05 |
|
GOTERM_MF |
GO:0005198~structural molecule
activity | 27 | 9.82E-08 | 1.36E-04 |
|
GOTERM_MF | GO:0005509~calcium
ion binding | 32 | 4.02E-07 | 5.58E-04 |
|
GOTERM_MF | GO:0005518~collagen
binding | 7 | 5.28E-06 | 7.33E-03 |
| Downregulated
DEGs |
|
|
|
|
|
GOTERM_BP |
GO:0008544~epidermis development | 20 | 1.27E-11 | 2.11E-08 |
|
GOTERM_BP | GO:0007398~ectoderm
development | 20 | 5.04E-11 | 8.39E-08 |
|
GOTERM_BP |
GO:0009913~epidermal cell
differentiation | 13 | 3.26E-10 | 5.42E-07 |
|
GOTERM_BP |
GO:0030855~epithelial cell
differentiation | 15 | 8.05E-09 | 1.34E-05 |
|
GOTERM_BP |
GO:0030216~keratinocyte
differentiation | 11 | 2.64E-08 | 4.40E-05 |
|
GOTERM_CC |
GO:0001533~cornified envelope | 9 | 3.27E-10 | 4.24E-07 |
|
GOTERM_MF |
GO:0005198~structural molecule
activity | 26 | 5.36E-06 | 7.55E-03 |
The pathways of these upregulated and downregulated
genes are indicated in Table II. The
upregulated genes were mainly involved in ECM-receptor interaction,
focal adhesion and small cell lung cancer. Genes such as
fibronectin 1 (FN1), epidermal growth factor receptor (EGFR) and
collagen type I alpha 1 (COL1A1) were identified in the focal
adhesion pathway. By contrast, the downregulated DEGs were enriched
in drug metabolism. Cytochrome P450 3A5 (CYP3A5) was identified in
the drug metabolism pathway.
 | Table II.KEGG pathway enrichment analysis of
upregulated and downregulated differentially expressed genes. |
Table II.
KEGG pathway enrichment analysis of
upregulated and downregulated differentially expressed genes.
| KEGG pathway
term | Counta | Genes | P-value | FDR |
|---|
| Upregulated
genes |
|
|
|
|
|
ECM-receptor interaction | 23 | COL1A1, COL4A1,
TNC | 1.24E-20 | 1.35E-17 |
| Focal
adhesion | 26 | FN1, EGFR,
COL1A1 | 4.22E-15 | 4.59E-12 |
| Small
cell lung cancer | 11 | FN1, CKS1B, LAMB3,
COL4A2 | 2.11E-06 | 0.00229 |
| Downregulated
genes |
|
|
|
|
| Drug
metabolism | 9 | CYP3A5, CYP2C18,
FMO2, MAOB | 6.49E-06 | 0.0071 |
PPI network construction and disease
enrichment analysis
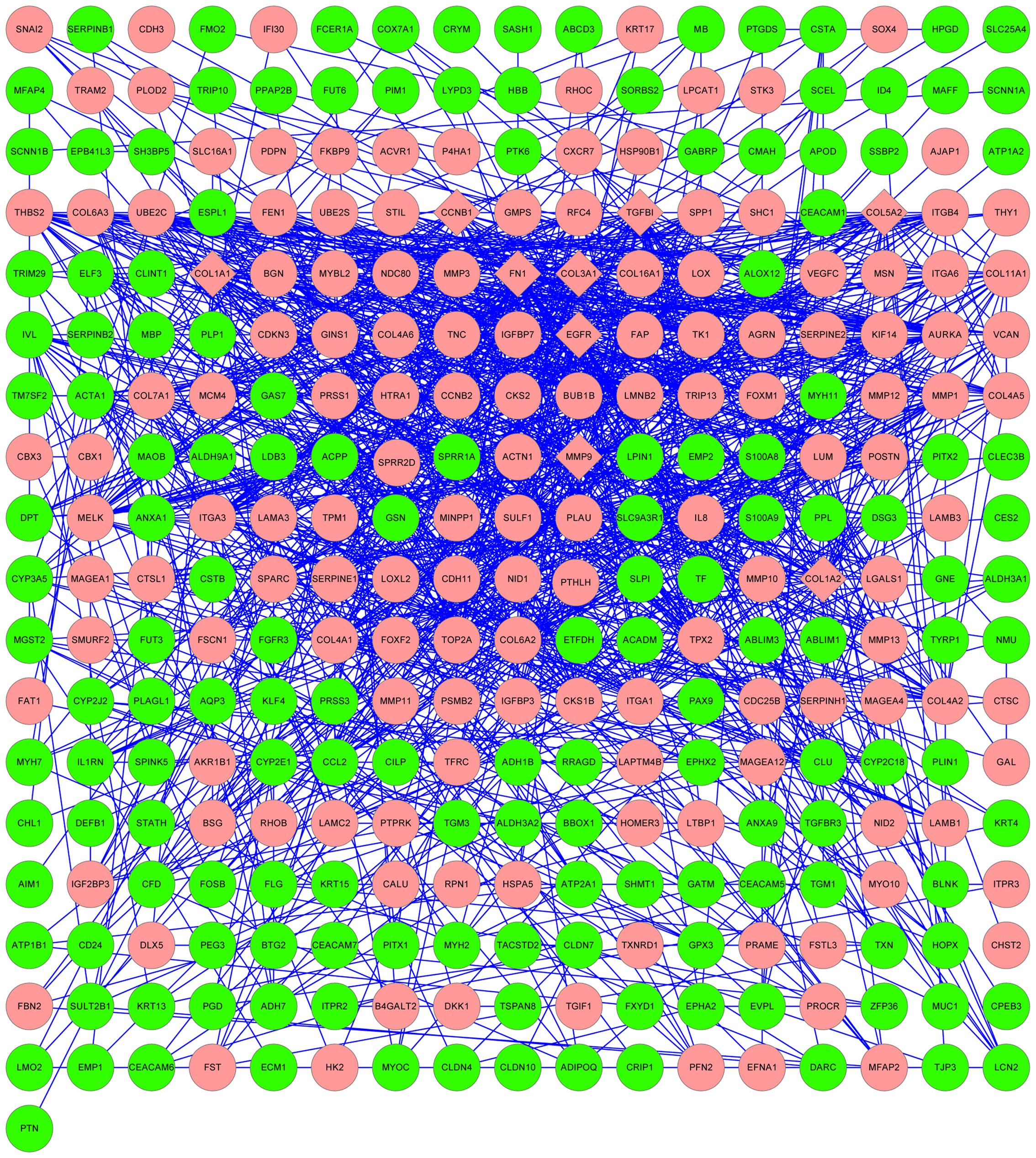
The results of the PPI network analysis are
represented in Fig. 2. The
upregulated genes FN1, EGFR, COL1A1, matrix metallopeptidase-9
(MMP-9), COL5A2, COL1A2, COL3A1, transforming growth factor,
beta-induced and cyclin B1 were selected as hub nodes.
Discussion
In the present study, gene expression profile data
were downloaded from the GEO database to identify DEGs in HNSCC
using bioinformatics analysis. A total of 419 DEGs between HNSCC
and normal samples, including 196 upregulated and 223 downregulated
genes, were selected. The results of functional enrichment analysis
revealed that the upregulated genes, including FN1, EGFR and
COL1A1, were associated with GO term of cell adhesion, while the
downregulated DEGs, including CYP3A5, were enriched in drug
metabolism pathways. According to the results of the PPI network
analysis, FN1, EGFR, COL1A1 and MMP-9 were identified as hub nodes.
Therefore, these DEGs and their interacting patners may be involved
in HNSCC development.
Cell adhesion is the process of binding of a cell to
a surface or substrate, such as the ECM or another cell (21). In the present study, the majority of
the upregulated DEGs were enriched in pathways of ECM-receptor
interaction and focal adhesion. Previous studies have indicated
that ECM-receptor interaction and focal adhesion were associated
with cell adhesion (22). Recent
evidence suggests that cell adhesion is mediated by several genes,
including FN1, EGFR and COL4A1 (23–25). FN1
is an ECM glycoprotein (26) involved
in cell adhesion (27), which
corresponds to the pathway identified in the present study. It was
previously reported that FN1 acts as a tumor suppressor gene,
playing a critical role in migration and invasion of laryngeal
carcinoma (23), which is the most
common type of HNSCC (28). EGFR was
also indicated to be associated with HNSCC (29). EGFR is the cell-surface receptor of
the EGF family (30). In the present
study, EGFR was enriched in GO terms of cell adhesion and pathway
of focal adhesion, which was consistent with previous studies that
reported that EGFR contributed to transduce extracellular signals
to intracellular responses, thus influencing adhesion and
proliferation in tumor cells (24,31). Rubin
Grandis et al (32) reported
that EGFR was overexpressed in HNSCC. High expression levels of
EGFR have been associated with reduced survival and increased risk
of recurrence in HNSCC (33). COL4A1
is a member of the collagen family, and is also associated with
cell adhesion (25). The adhesion of
cells to collagen is mediated by fibronectin (25). Tanaka et al (34) indicated that the differential
expression of type IV collagen chains was associated with the
invasive potential of cell carcinoma. The results of the present
study indicated that FN1, EGFR and COL4A1 were upregulated genes in
HNSCC and hub nodes in the PPI network, which suggests that FN1,
EGFR and COL4A1 may regulate cell adhesion in HNSCC. Thus, cell
adhesion may participate in HNSCC through multiple genes, including
FN1, EGFR and COL4A1, which may be potential therapeutic target
genes in HNSCC.
In the present study, the downregulated DEGs such as
CYP3A5, were significantly enriched in the pathway of drug
metabolism (P=6.49E-06). CYP3A5 encodes a member of the cytochrome
P450 superfamily of enzymes (34). It
has been reported that cytochrome P450 proteins catalyze multiple
reactions, including drug metabolism (35). Olivieri et al (36) reported that cytochrome P450 gene
polymorphisms were important in the tumorigenesis and progression
of HNSCC (36). These results
suggested that CYP3A5 may regulate HNSCC development through the
drug metabolism pathway. Therefore, this pathway may be associated
with HNSCC progression.
In addition to FN1, EGFR and COL4A1, MMP-9 was also
identified as a hub node in the present PPI network analysis. MMP-9
is an enzyme that belongs to the MMP family (35). It has been reported that MMPs
participate in cancer invasion and metastasis (37). In the present study, MMP-9 was an
upregulated gene, which was consistent with previous studies
(38,39). For example, Riedel et al
(38) reported that the expression
levels of MMP-9 were significantly higher in HNSCC patients than in
healthy individuals. MMP-9 regulates cell proliferation through
modulating the nuclear factor-κB signaling pathway in HNSCC
(40). Furthermore, MMP-9 was
associated with cancer in the present study. Thus, MMP-9 may be a
potential target gene for the treatment of HNSCC.
In conclusion, a total of 419 DEGs were identified
between HNSCC and normal samples, and the present study indicates
that cell adhesion and drug metabolism may be closely associated
with HNSCC development. Genes such as FN1, EGFR, COL4A1 and MMP-9
may be potential therapeutic target genes in HNSCC. However,
further studies are required to confirm the present results.
References
|
1
|
Ferlay J, Shin HR, Bray F, Forman D,
Mathers C and Parkin DM: Estimates of worldwide burden of cancer in
2008: GLOBOCAN 2008. Int J Cancer. 127:2893–2917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Rothenberg SM and Ellisen LW: The
molecular pathogenesis of head and neck squamous cell carcinoma. J
Clin Invest. 122:1951–1957. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hasina R, Whipple ME, Martin LE, Kuo WP,
Ohno-Machado L and Lingen MW: Angiogenic heterogeneity in head and
neck squamous cell carcinoma: Biological and therapeutic
implications. Lab Invest. 88:342–353. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Belcher R, Hayes K, Fedewa S and Chen AY:
Current treatment of head and neck squamous cell cancer. J Surg
Oncol. 110:551–574. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Colevas AD: Chemotherapy options for
patients with metastatic or recurrent squamous cell carcinoma of
the head and neck. J Clin Oncol. 24:2644–2652. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Leemans CR, Braakhuis BJ and Brakenhoff
RH: The molecular biology of head and neck cancer. Nat Rev Cancer.
11:9–22. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Nohata N, Sone Y, Hanazawa T, Fuse M,
Kikkawa N, Yoshino H, Chiyomaru T, Kawakami K, Enokida H, Nakagawa
M, et al: miR-1 as a tumor suppressive microRNA targeting TAGLN2 in
head and neck squamous cell carcinoma. Oncotarget. 2:29–42.
2011.PubMed/NCBI
|
|
8
|
Chaisaingmongkol J, Popanda O, Warta R,
Dyckhoff G, Herpel E, Geiselhart L, Claus R, Lasitschka F, Campos
B, Oakes CC, et al: Epigenetic screen of human DNA repair genes
identifies aberrant promoter methylation of NEIL1 in head and neck
squamous cell carcinoma. Oncogene. 31:5108–5116. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhang L, Pan HY, Zhong LP, Wei KJ, Yang X,
Li J, Shen GF and Zhang Z: Fos-related activator-1 is overexpressed
in oralsquamous cell carcinoma and associated with tumor lymph node
metastasis. J Oral Pathol Med. 39:470–476. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Pedrero JM, Carracedo DG, Pinto CM,
Zapatero AH, Rodrigo JP, Nieto CS and Gonzalez MV: Frequent genetic
and biochemical alterations of the PI3-K/AKT/PTEN pathway in head
and neck squamous cell carcinoma. Int J Cancer. 114:242–248. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Gallo O, Masini E, Bianchi B, Bruschini L,
Paglierani M and Franchi A: Prognostic significance of
cyclooxygenase-2 pathway and angiogenesis in head and neck squamous
cell carcinoma. Hum Pathol. 33:708–714. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Bancroft CC, Chen Z, Dong G, Sunwoo JB,
Yeh N, Park C and Van Waes C: Coexpression of proangiogenic factors
IL-8 and VEGF by human head and neck squamous cell carcinoma
involves coactivation by MEK-MAPK and IKK-NF-kappaB signal
pathways. Clin Cancer Res. 7:435–442. 2001.PubMed/NCBI
|
|
13
|
Kuriakose MA, Chen WT, He ZM, Sikora AG,
Zhang P, Zhang ZY, Qiu WL, Hsu DF, McMunn-Coffran C, Brown SM, et
al: Selection and validation of differentially expressed genes in
head and neck cancer. Cell Mol Life Sci. 61:1372–1383. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gautier L, Cope L, Bolstad BM and Irizarry
RA: affy - analysis of Affymetrix GeneChip data at the probe level.
Bioinformatics. 20:307–315. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tusher VG, Tibshirani R and Chu G:
Significance analysis of microarrays applied to the ionizing
radiation response. Proc Natl Acad Sci USA. 98:5116–5121. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: The Gene Ontology Consortium: Gene Ontology: Tool for the
unification of biology. Nat Genet. 25:25–29. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Altermann E and Klaenhammer TR:
PathwayVoyager: Pathway mapping using the Kyoto Encyclopedia of
Genes and Genomes (KEGG) database. BMC Genomics. 6:602005.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Huang DW, Sherman BT, Tan Q, Kir J, Liu D,
Bryant D, Guo Y, Stephens R, Baseler MW, Lane HC and Lempicki RA:
DAVID Bioinformatics Resources: Expanded annotation database and
novel algorithms to better extract biology from large gene lists.
Nucleic Acids Res. 35(Web Server issue): W169–W175. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Szklarczyk D, Franceschini A, Kuhn M,
Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork
P, et al: The STRING database in 2011: Functional interaction
networks of proteins, globally integrated and scored. Nucleic Acids
Res. 39(Database issue): D561–D568. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Smoot ME, Ono K, Ruscheinski J, Wang PL
and Ideker T: Cytoscape 2.8: New features for data integration and
network visualization. Bioinformatics. 27:431–432. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Gumbiner BM: Cell adhesion: The molecular
basis of tissue architecture and morphogenesis. Cell. 84:345–357.
1996. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Albelda SM and Buck CA: Integrins and
other cell adhesion molecules. FASEB J. 4:2868–2880.
1990.PubMed/NCBI
|
|
23
|
Wang F, Song G, Liu M, Li X and Tang H:
miRNA-1 targets fibronectin 1 and suppresses the migration and
invasion of the HEp2 laryngeal squamous carcinoma cell line. FEBS
Lett. 585:3263–3269. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Goerner M, Seiwert TY and Sudhoff H:
Molecular targeted therapies in head and neck cancer - an update of
recent developments. Head Neck Oncol. 2:82010. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kleinman HK, Martin GR and Fishman PH:
Ganglioside inhibition of fibronectin-mediated cell adhesion to
collagen. Proc Natl Acad Sci USA. 76:3367–3371. 1979. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Pankov R and Yamada KM: Fibronectin at a
glance. J Cell Sci. 115:3861–3863. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Soikkeli J, Podlasz P, Yin M, Nummela P,
Jahkola T, Virolainen S, Krogerus L, Heikkilä P, von Smitten K,
Saksela O and Hölttä E: Metastatic outgrowth encompasses COL-I,
FN1, and POSTN up-regulation and assembly to fibrillar networks
regulating cell adhesion, migration, and growth. Am J Pathol.
177:387–403. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Mao L, Hong WK and Papadimitrakopoulou VA:
Focus on head and neck cancer. Cancer Cell. 5:311–316. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Erjala K, Sundvall M, Junttila TT, Zhang
N, Savisalo M, Mali P, Kulmala J, Pulkkinen J, Grenman R and
Elenius K: Signaling via ErbB2 and ErbB3 associates with resistance
and epidermal growth factor receptor (EGFR) amplification with
sensitivity to EGFR inhibitor gefitinib in head and neck squamous
cell carcinoma cells. Clin Cancer Res. 12:4103–4111. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Herbst RS: Review of epidermal growth
factor receptor biology. Int J Radiat Oncol Biol Phys. 59(Suppl 2):
S21–S26. 2004. View Article : Google Scholar
|
|
31
|
Rocha-Lima CM, Soares HP, Raez LE and
Singal R: EGFR targeting of solid tumors. Cancer Control.
14:295–304. 2007.PubMed/NCBI
|
|
32
|
Grandis Rubin J, Melhem MF, Gooding WE,
Day R, Holst VA, Wagener MM, Drenning SD and Tweardy DJ: Levels of
TGF-alpha and EGFR protein in head and neck squamous cell carcinoma
and patient survival. J Natl Cancer Inst. 90:824–832. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Bei R, Budillon A, Masuelli L, Cereda V,
Vitolo D, Di Gennaro E, Ripavecchia V, Palumbo C, Ionna F, Losito
S, et al: Frequent overexpression of multiple ErbB receptors by
head and neck squamous cell carcinoma contrasts with rare antibody
immunity in patients. J Pathol. 204:317–325. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Tanaka K, Iyama K, Kitaoka M, Ninomiya Y,
Oohashi T, Sado Y and Ono T: Differential expression of·alpha
1(IV), alpha 2(IV), alpha 5(IV) and·alpha 6(IV) collagen chains in
the basement membrane of basal cell carcinoma. Histochem J.
29:563–570. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Bains RK, Kovacevic M, Plaster CA,
Tarekegn A, Bekele E, Bradman NN and Thomas MG: Molecular diversity
and population structure at the cytochrome P450 3A5 gene in Africa.
BMC Genet. 14:342013. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Olivieri EH, da Silva SD, Mendonça FF,
Urata YN, Vidal DO, Faria MA, Nishimoto IN, Rainho CA, Kowalski LP
and Rogatto SR: CYP1A2*1C, CYP2E1*5B, and GSTM1 polymorphisms are
predictors of risk and poor outcome in head and neck squamous cell
carcinoma patients. Oral Oncol. 45:e73–e79. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Curran S and Murray GI: Matrix
metalloproteinases: Molecular aspects of their roles in tumour
invasion and metastasis. Eur J Cancer. 36:1621–1630. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Riedel F, Götte K, Schwalb J and Hörmann
K: Serum levels of matrix metalloproteinase-2 and −9 in patients
with head and neck squamous cell carcinoma. Anticancer Res.
20:3045–3049. 2000.PubMed/NCBI
|
|
39
|
Sinpitaksakul SN, Pimkhaokham A,
Sanchavanakit N and Pavasant P: TGF-β1 induced MMP-9 expression in
HNSCC cell lines via Smad/MLCK pathway. Biochem Biophys Res Commun.
371:713–718. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Aggarwal S, Takada Y, Singh S, Myers JN
and Aggarwal BB: Inhibition of growth and survival of human head
and neck squamous cell carcinoma cells by curcumin via modulation
of nuclear factor-kappaB signaling. Int J Cancer. 111:679–692.
2004. View Article : Google Scholar : PubMed/NCBI
|