Introduction
Cancer has become one of the leading causes of
mortality worldwide due to genetic and environmental factors.
However, the exact mechanism of carcinogenesis remains largely
unknown (1). With research
developing, it is becoming clear that the characteristics of
cancer, which are founded on genome instability, include
maintaining proliferative signaling, enabling replicative
immortality, inducing angiogenesis, invasion and metastasis,
reprogramming the energy metabolism, evading growth suppressors,
resisting cell death and evading immune destruction (2). Recently, it has become evident that
genetic variation plays a significant role in the development and
progression of cancer. More studies based on gene polymorphisms
have proved that the polymorphisms may contribute to the cancer
risk (3).
Vascular endothelial growth factor (VEGF) plays a
key role in a number of pathological processes, including
angiogenesis, tumor growth and metastasis. VEGF plays an important
role in tumor angiogenesis through promoting endothelial cell
growth and migration (4). The human
VEGF gene is located on chromosome 6p21.3 and includes a 14-kb
coding region with eight exons and seven introns (5). Certain polymorphisms have been
identified in the VEGF gene. In order to evaluate the association
between the VEGF polymorphism with the various types of cancer
risk, numerous molecular epidemiological studies have been
performed in different populations recently (6). However, there were no clear
conclusions due to inconsistent statistical results. Some of these
conclusions may ascribe to the possible small influence of the gene
polymorphism on the cancer risk and others may be caused by the
relatively small samples in these published studies. Specific
meta-analyses analyzed the association for only one type of cancer,
including colorectal, lung or bladder cancer. Therefore, a
relatively comprehensive meta-analysis was performed, including the
most recent and relevant studies to provide more accurate
statistical evidence for the association between the VEGF
polymorphism and risk of the types of cancer that have been studied
(7–40). Meta-analysis can alleviate the
problems caused by small samples and deficient statistical genetic
studies of multiple traits. Therefore, it can provide more reliable
results than a single case-control study. The aim of the present
study was to utilize the meta-analysis to summarize the relevant
studies regarding the VEGf -2578C/A polymorphism and the risk of
cancer.
Materials and methods
Identification and eligibility of
relevant studies
The relevant studies that were published by August
23, 2013 were identified using the Embase, PubMed, Web of Science
and Chinese National Knowledge Infrastructure databases. The
following terms were used in the search: ‘Genetic polymorphism’,
‘polymorphism’ or ‘genetic variants’; ‘VEGF’ or ‘vascular
endothelial growth factor’; and ‘cancer’ or ‘carcinoma’. The
case-control and cohort studies that explored the association
between the VEGF polymorphism and cancer risk with genotyping data
were included. All the eligible studies were reviewed and only the
published studies were included in the meta-analysis. For studies
in which the data partly overlapped, only the most recent or
complete studies were included. When the same sample was applied to
several different studies, the most integrated data was selected
following careful examination.
Inclusion and exclusion criteria
The studies that were included in the meta-analysis
met the following criteria: i) Case-control studies focused on the
associations between the VEGF -2578C/A polymorphism and cancer
risk; ii) all the patients were diagnosed by pathological or
histological examinations; iii) the frequencies of the genotypes in
cancer cases and controls could be extracted; and iv) published in
English or Chinese. The excluded studies were: i) Not case-control
studies; ii) published in a language other than English or Chinese;
and iii) were letters, reviews, meta-analyses or editorial
studies.
Data extraction
The data were independently extracted from all the
eligible studies by two investigators according to the
aforementioned inclusion criteria. From each study, the following
information was extracted: First author’s name, year of
publication, country, ethnicity, type of cancer, DNA sample, source
of controls, study design, methods and total number of cases and
controls. Ethnicity was categorized as the ‘Caucasian,’ ‘African,’
(including African-Americans) and ‘Asian’ populations. One study
did not state the included ethnic groups according to phenotype,
and therefore, the sample was known as ‘mixed’ (29). However, each control was
individually matched to a case with regards to birth date (±6
months), date of blood collection (±6 months) and ethnicity
(Caucasian, African-American, Hispanic, Asian and other/unknown).
Furthermore, the studies investigating more than one type of cancer
were considered as individual data sets only in the subgroup
analyses by the type of cancer. There was no definition as to the
minimum number of patients that were included in the present
meta-analysis. The studies that reported different ethnic groups
and countries or locations, were considered as separate study
samples for each aforementioned category.
Statistical analysis
The statistical analyses were performed using Stata
software (version 9.0; StataCorp, College Station, TX, USA).
Heterogeneity among the various studies was assessed by the
Q-statistic and quantified by calculating the I2 value.
According to the Q-statistic, heterogeneity was significant when
P<0.10. Among the studies, the I2 value demonstrated
the percentage of variation associated with heterogeneity, instead
of chance. No heterogeneity was observed when I2=0%, and
0–25% accounted for low, 25–50% for moderate and 50–75% for high
heterogeneity. Consistent with published recommendations (29) for the quality assessment in genetic
association meta-analyses, three genetic models were selected:
Allele (C vs. A), for homozygote (CC vs. AA) and heterozygote
comparisons (CA vs. AA); dominant (CC+CA vs. AA); and recessive
models (CC vs. CA+AA), to prevent the use of the wrong genetic
model. For each study, the odds ratio (OR), together with the
corresponding 95% confidence interval (95% CI), was calculated to
assess the association between the VEGF polymorphism and the risk
of cancer. Meta-analysis was performed for the polymorphisms that
had been investigated in at least two studies. The overall estimate
of risk (OR) was calculated by a fixed-effects (Mantel-Haenszel) or
a random-effects model (DerSimonian-Laird) according to the
presence (P<0.10 or I2>50%) or absence (P>0.10
or I2<50%) of heterogeneity, respectively. In
addition to the comparison among all the groups, subgroup analysis
was performed in correlation with the type of cancer and ethnicity.
The significant differences in genotype and allelic frequency
between the two groups were determined using the χ2
test. In order to exclude the allele frequencies in the controls,
deviating greatly from the Hardy-Weinberg equilibrium, the
χ2 test (minimum Pearson χ2 estimate) was
performed in the sensitivity analysis and deviation was considered
when P<0.01. All the statistical tests were two-sided and
P<0.05 was considered to indicate a statistically significant
difference.
Results
Characteristics of studies
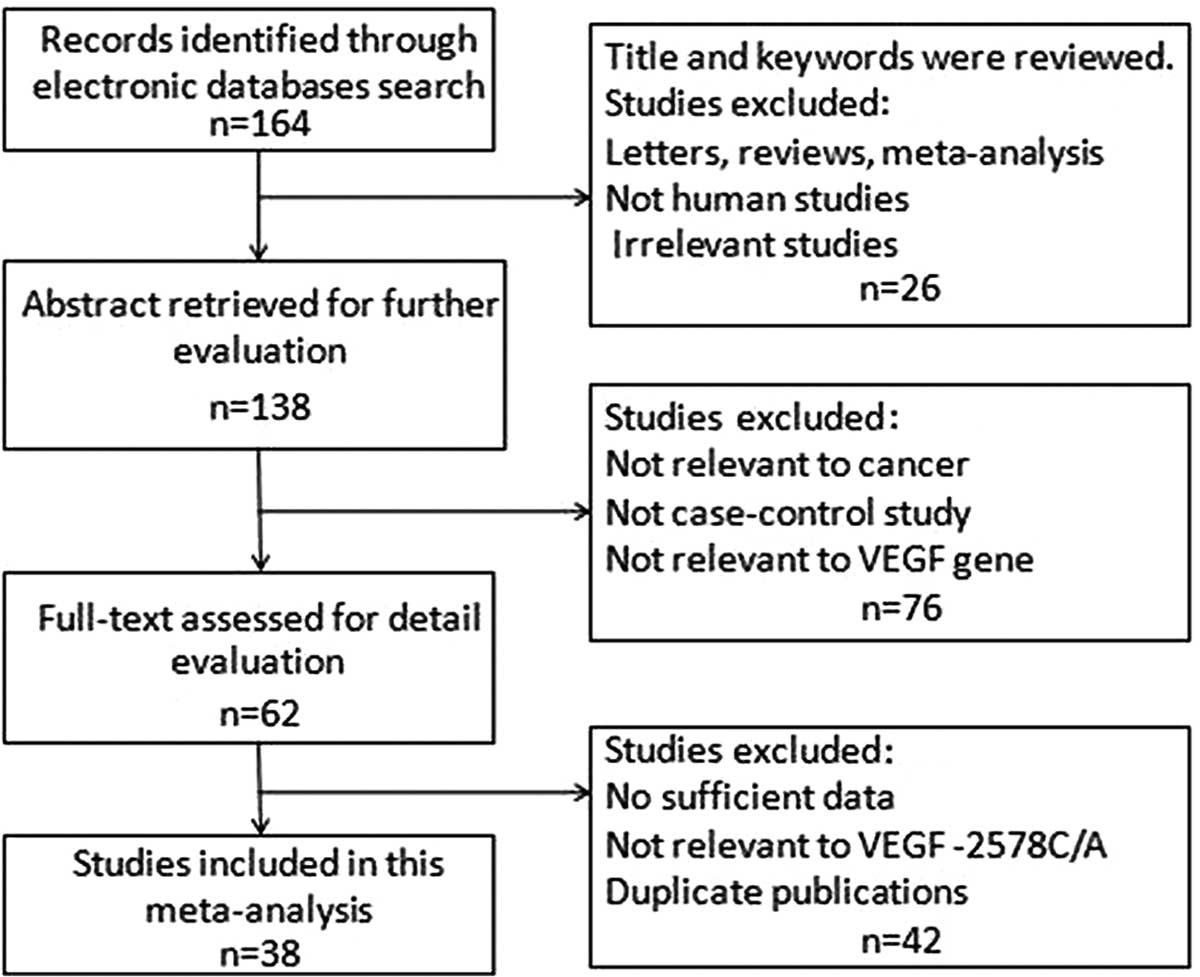
According to the inclusion and exclusion criteria,
37 case-control studies were included that ranged between 2002 and
2013 (Fig. 1). Among these studies,
17 were studies of the Caucasian population and 18 were of the
Asian population. All the patients were diagnosed histologically or
pathologically. Blood samples were used for genotyping in 30
studies and tissue samples were used in eight studies. A total of
20 studies used hospital-based controls, whereas 15 studies used
population-based controls. The polymerase chain
reaction-restriction fragment length polymorphism assay was used
for genotyping in 13 studies, whereas the TaqMan assay was used in
11 studies. The details of the included studies are summarized in
Table I.
 | Table ICharacteristics of the primary studies
in the meta-analysis. |
Table I
Characteristics of the primary studies
in the meta-analysis.
| First author | Year | Country | Ethnicity | Cancer type | DNA sample | Source of
control | Study design | Methods | Case, n | Control, n | (Refs.) |
|---|
| Howell | 2002 | UK | Caucasian | CMM | Tissue | Unknown | Case-cont | Unknown | 134 | 266 | (15) |
| Kim | 2005 | Korea | Asian | Bladder | Blood | HB | Case-cont | TaqMan | 153 | 153 | (41) |
| Jin | 2005 | Poland | Caucasian | Breast | Tissue | PB | Case-cont | PCR-RFLP | 411 | 423 | (21) |
| 2005 | Germany | Caucasian | Breast | Tissue | PB | Case-cont | PCR-RFLP | 153 | 162 | |
| 2005 | Sweden | Caucasian | Breast | Tissue | PB | Case-cont | PCR-RFLP | 939 | 940 | |
| Jacobs | 2006 | America | Mixed | Breast | Blood | PB | Case-cont | TaqMan | 498 | 495 | (18) |
| Hofmann | 2008 | Austria | Caucasian | Colorectal | Blood | PB | Case-cont | TaqMan | 433 | 427 | (14) |
| Nikiteas | 2007 | Greece | Caucasian | Gastric | Blood | HB | Case-cont | PCR | 100 | 100 | (32) |
| Hsiao | 2007 | Taiwan | Asian | Thyroid | Blood | HB | Case-cont | TaqMan | 297 | 249 | (16) |
| Park | 2007 | Korea | Asian | Colorectal | Blood | HB | Case-cont | PCR | 203 | 246 | (33) |
| Pharoah | 2005 | UK | Caucasian | Breast | Unknown | PB | Case-cont | TaqMan | 2015 | 2139 | (34) |
| Nasr | 2008 | Tunisia | African | Nasopharyngeal
carcinoma | Blood | HB | Case-cont | PCR-RFLP | 163 | 169 | (31) |
| Diao | 2009 | China | Asian | Non-Hodgkin’s
lymphoma | Blood | HB | Case-cont | PCR-RFLP | 431 | 172 | (11) |
| Ke | 2008 | China | Asian | Gastric | Blood | PB | Case-cont | PCR-RFLP | 540 | 561 | (23) |
| Dassoulas | 2009 | Greece | Caucasian | Colorectal | Tissue | HB | Case-cont | TaqMan | 312 | 362 | (9) |
| Maltese | 2009 | Italy | Caucasian | Colorectal | Blood | PB | Case-cont | PCR | 302 | 115 | (28) |
| Liang | 2009 | China | Asian | Lung | Blood | HB | Case-cont | PCR-RFLP | 171 | 172 | (27) |
| Wu | 2009 | China | Asian | Hepatocellular
carcinoma | Blood | HB | Case-cont | TaqMan | 92 | 99 | (39) |
| Zhang | 2011 | China | Asian | Colorectal | Blood | HB | Case-cont | PCR-RFLP | 110 | 110 | (40) |
| Li | 2010 | China | Asian | Ovarian | Blood | PB | Case-cont | PCR-RFLP | 303 | 303 | (26) |
| Kämmerer | 2013 | Germany | Caucasian | Oral squamous cell
carcinoma | Blood | HB | Case-cont | RT-PCR | 80 | 40 | (22) |
| VanCleave | 2010 | America | African | Prostate | Blood | HB | Case-cont | TaqMan | 190 | 635 | (37) |
| Wang | 2011 | China | Asian | Nasopharyngeal
carcinoma | Blood | HB | Case-cont | PCR-RFLP | 156 | 161 | (38) |
| Galimberti | 2010 | Italy | Caucasian | Mantle cell
lymphoma | Blood | PB | Case-cont | RT-PCR | 32 | 58 | (12) |
| Kim | 2010 | Korea | Asian | Cervical | Blood | HB | Case-cont | PCR | 199 | 211 | (24) |
| Ajaz | 2011 | Pakistan | Asian | Renal cell
carcinoma | Blood | PB | Case-cont | TaqMan | 143 | 106 | (7) |
| Jang | 2013 | South Korea | Asian | Colorectal | Blood | HB | Case-cont | PCR-RFLP | 390 | 492 | (20) |
| Supic | 2012 | Serbia | Caucasian | Oral squamous cell
carcinoma | Blood | PB | Case-cont | TaqMan | 114 | 126 | (36) |
| Henríquez-
Hernández | 2012 | Spain | Caucasian | Bladder | Blood | HB | Case-cont | PCR-RFLP | 59 | 43 | (13) |
| Sáenz-López | 2013 | Spain | Caucasian | Renal cell
carcinoma | Tissue | HB | Case-cont | RT-PCR | 216 | 272 | (35) |
| Li | 2012 | China | Asian | Lung | Blood | HB | Case-cont | PCR-RFLP | 150 | 150 | (25) |
| Jaiswal | 2013 | India | Asian | Bladder | Blood | HB | Case-cont | PCR-RFLP | 250 | 200 | (19) |
| Moon | 2013 | India | Asian | Colorectal | Blood | PB | Case-cont | PCR | 390 | 492 | (20) |
| Ianni | 2013 | Italy | Caucasian | Prostate | Blood | PB+HB | Case-cont | TaqMan | 224 | 156 | (17) |
|
Martinez-Fierro | 2013 | Mexico | Caucasian | Prostate | Tissue | Unknown | Case-cont | PCR-RFLP | 77 | 172 | (29) |
| Mishra | 2013 | India | Asian | Bladder | Blood | HB | Case-cont | PCR | 195 | 300 | (30) |
| Liang | 2009 | China | Asian | Lung | Blood | PB | Case-cont | PCR | 171 | 172 | (27) |
| Deng | 2014 | China | Asian | Lung | Blood | PB | Case-cont | PCR | 65 | 110 | (10) |
| Antonacopoulou | 2011 | Greece | Caucasian | Colorectal | Tissue | Unknown | Case-cont | PCR | 222 | 263 | (8) |
Quantitative data synthesis
The summary of the meta-analysis of the associations
between VEGF -2578C/A and cancer risk is shown in Table II. The random-effects model was
used when the heterogeneity was evident under the genetic models
(P>0.05), otherwise the fixed-effects models was used. When all
the eligible studies were pooled, no significant association was
observed between the VEGF -2578C/A polymorphism and the risk of
cancer (CC vs. AA: OR, 0.4; 95% CI, 0.86–1.02; P=0 for
heterogeneity) (Fig. 2) or
recessive model (CC vs. CA/AA: OR, 0.97; 95% CI, 0.88–1.08; P=0 for
heterogeneity).
 | Table IIMain results of the pooled ORs in the
meta-analysis. |
Table II
Main results of the pooled ORs in the
meta-analysis.
| | C vs. A | CC vs. AA | CA vs. AA | CC/CA vs. AA | CC vs. CA/AA |
|---|
| |
|
|
|
|
|
|---|
| Variables | n | OR (95% CI) | P-value | OR (95% CI) | P-value | OR (95% CI) | P-value | OR (95% CI) | P-value | OR (95% CI) | P-value |
|---|
| Total | 38 | 0.97
(0.91–1.04) | 0 | 0.94
(0.86–1.02) | 0 | 0.92
(0.80–1.06) | 0 | 0.96
(0.89–1.03) | 0 | 0.97
(0.88–1.08) | 0 |
| Ethnicities |
| Caucasian | 17 | 0.99
(0.89–1.10) | 0 | 0.92
(0.76–1.11) | 0 | 0.93
(0.79–1.11) | 0 | 0.96
(0.88–1.04) | 0.03 | 1.01
(0.85–1.20) | 0 |
| Asian | 18 | 0.93
(0.87–1.00) | 0.07 | 0.78
(0.66–0.94) | 0.07 | 0.82
(0.61–1.11) | 0 | 0.85
(0.72–1.00) | 0.03 | 0.91
(0.78–1.07) | 0 |
| African | 2 | 1.20
(0.97–1.45) | 0.19 | 1.31
(0.67–2.58) | 0.13 | 1.31
(0.67–2.58) | 0.23 | 1.26
(0.86–1.83) | 0.20 | 1.23
(0.92–1.64) | 0.30 |
| Cancer |
| Bladder | 3 | 1.02
(0.87–1.20) | 0.37 | 1.06
(0.74–1.53) | 0.27 | 1.12
(0.59–2.11) | 0.01 | 1.13
(0.70–1.82) | 0.06 | 1.04
(0.70–1.54) | 0.10 |
| Breast | 5 | 1.01
(0.95–1.07) | 0.84 | 1.01
(0.90–1.15) | 0.84 | 1.03
(0.93–1.15) | 0.80 | 1.03
(0.93–1.13) | 0.79 | 0.99
(0.90–1.10) | 0.93 |
| Colorectal | 6 | 0.88
(0.77–1.00) | 0.04 | 0.73
(0.60–0.89) | 0.26 | 0.79
(0.66–0.95) | 0.73 | 0.76
(0.64–0.91) | 0.51 | 0.90
(0.80–1.02) | 0.07 |
| Lung | 4 | 0.91
(0.68–1.23) | 0.10 | 0.33
(0.19–0.60) | 0.88 | 0.26
(0.11–0.64) | 0.09 | 0.32
(0.18–0.57) | 0.57 | 1.08
(0.65–1.79) | 0.01 |
| Sources of
controls |
| PB | 18 | 0.99
(0.89–1.11) | 0 | 0.89
(0.72–1.11) | 0.31 | 0.96
(0.87–1.07) | 0.48 | 0.97
(0.88–1.07) | 0.56 | 1.03
(0.87–1.21) | 0.77 |
| HB | 12 | 0.95
(0.87–1.04) | 0.08 | 0.90
(0.73–1.10) | 0.29 | 0.91
(0.65–1.28) | 0.59 | 0.93
(0.70–1.22) | 0.59 | 0.97
(0.80–1.17) | 0.74 |
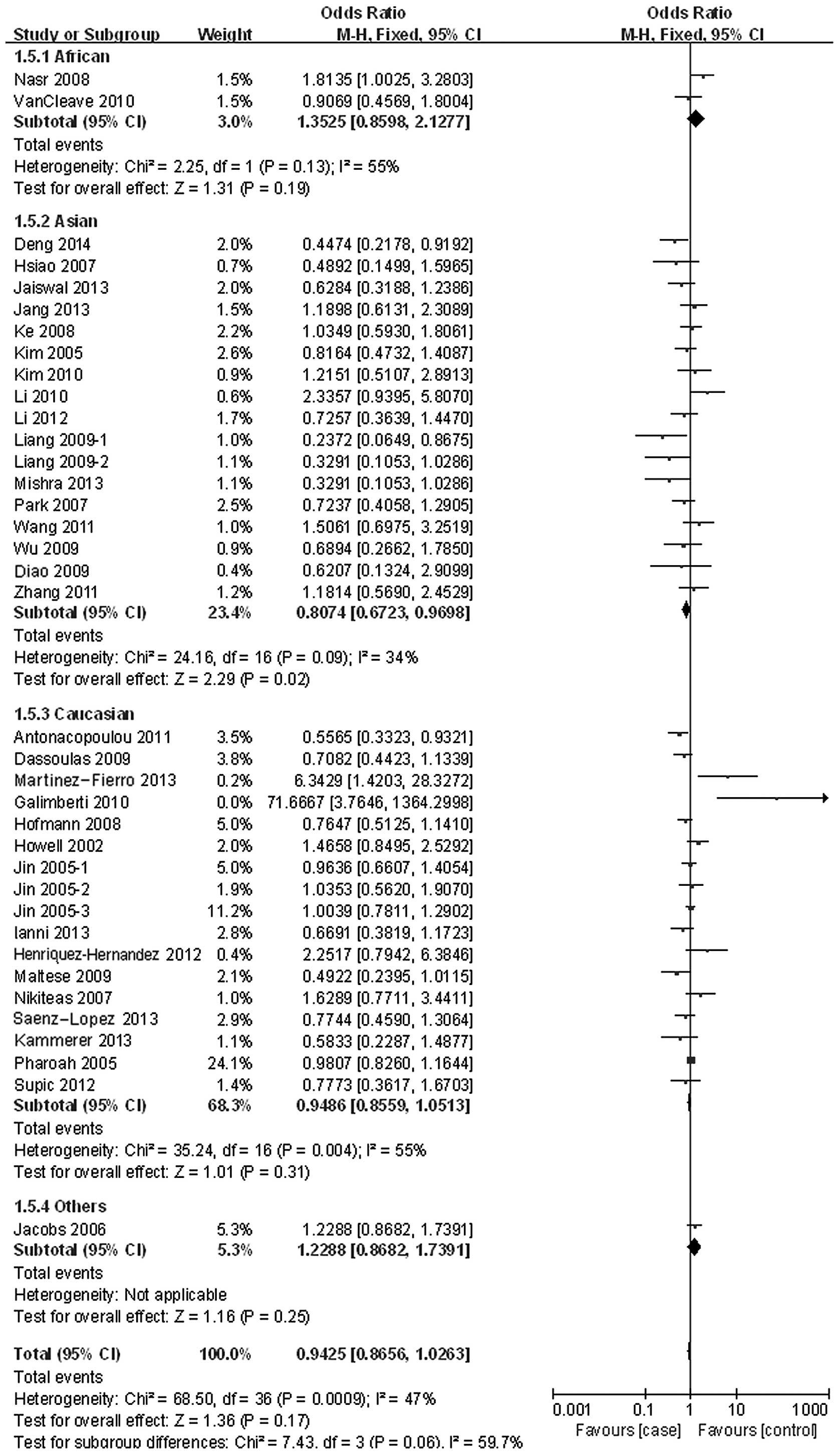
In the stratified analysis by ethnicities, the VEGF
-2578C/A polymorphism was associated with a significant decrease
risk in the Asian population in the three tested models (C vs. A:
OR, 0.93; 95% CI, 0.87–1.00; P=0.07 for heterogeneity; CC vs. AA:
OR, 0.78; 95% CI, 0.66–0.94; P=0.07 for heterogeneity; and dominant
model: OR, 0.85; 95% CI, 0.72–1.00; P=0.03 for heterogeneity)
(Fig. 3), of colorectal cancer in
four tested models (C vs. A: OR, 0.88; 95% CI, 0.77–1.00; P=0.04
for heterogeneity; CC vs. AA: OR, 0.73; 95% CI, 0.60–0.89; P=0.26
for heterogeneity; CA vs. AA: OR, 0.79; 95% CI, 0.66–0.95; P=0.73
for heterogeneity; and dominant model: OR, 0.76; 95% CI, 0.64–0.91;
P=0.51 for heterogeneity) (Fig. 2)
and of lung cancer in three tested models (CC vs. AA: OR, 0.33; 95%
CI, 0.19–0.60; P=0.88 for heterogeneity; CA vs. AA: OR, 0.26; 95%
CI, 0.11–0.64; P=0.09 for heterogeneity; and dominant model: OR,
0.32; 95% CI, 0.18–0.57; P=0.57 for heterogeneity) (Fig. 2). However, no associations were
found with the other types of cancer (Table II).
Sensitivity analysis
Sensitivity analysis was performed to explore the
influence of individual studies on the pooled results. No
individual study was shown to affect the pooled OR significantly,
as no substantial change was found.
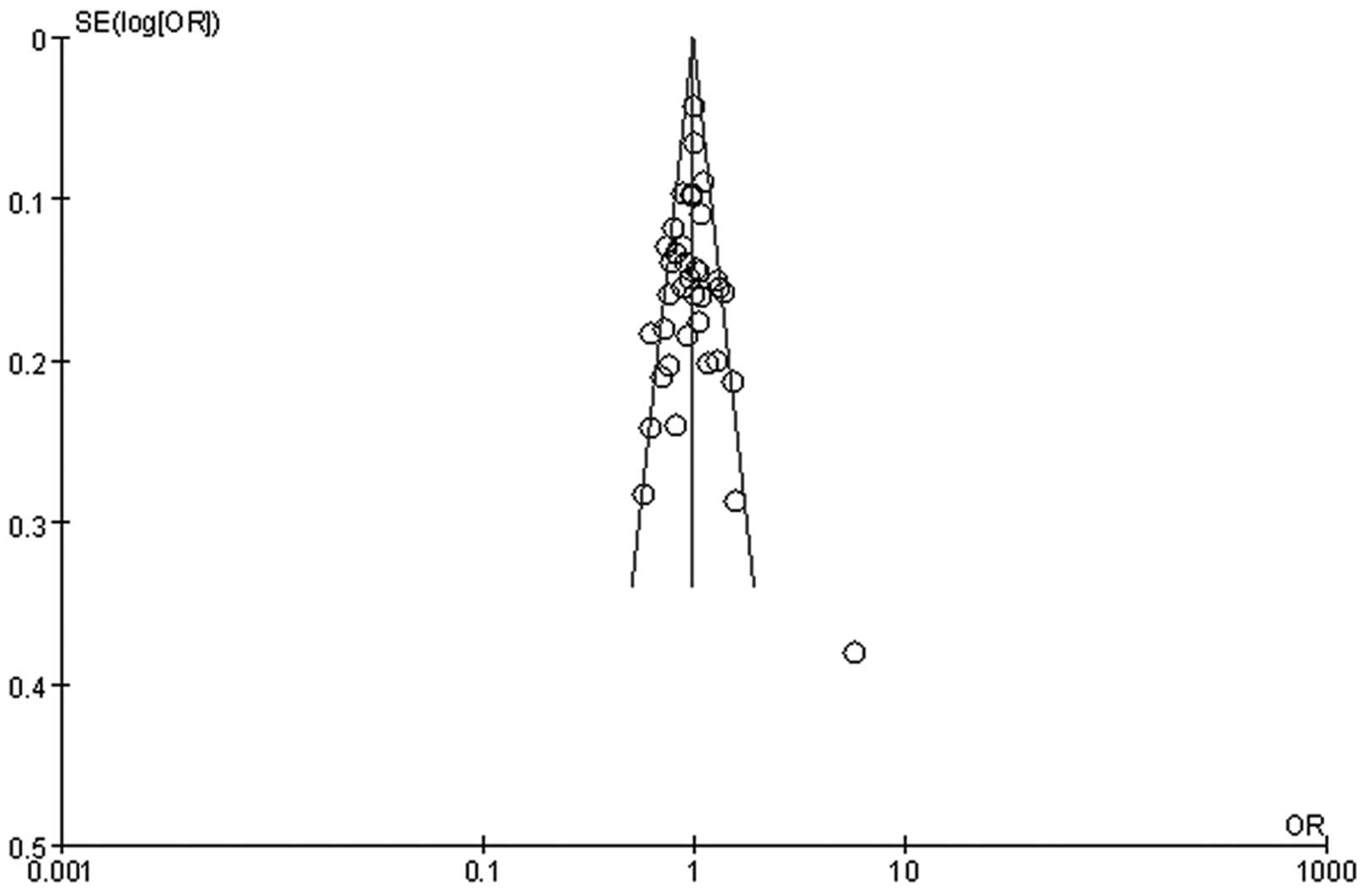
Publication bias
Publication bias of the studies was assessed by
Begg’s funnel plot and Egger’s test (Fig. 4). The arrangement of the data points
did not reveal any evidence of clear asymmetry. Formal evaluation
using Egger’s regression asymmetry tests for the homozygote
comparison did not show any evidence of publication bias (t=1.77
P=0.086).
Discussion
Based on 37 cases-control studies, the present
meta-analysis included 11,083 cases and 11,822 controls, and
indicated that there was no association between the VEGF -2578C/A
polymorphism and the risk of malignancy in the pooled analyses.
Subgroup analyses with regards to the type of cancer showed
specific positive associations. The VEGF -2578C/A polymorphism
decreased the risk of colorectal and lung cancers under the
codominant, dominant and recessive models. For future studies, the
VEGF polymorphism could act as a predictive marker to develop a
novel antiangiogenesis medicine. VEGF -2578C/A-targeted therapy
could be applied to patients who want to receive an individualized
treatment. However, in the stratified analysis by ethnicity and
type of cancer, no evident co-association was observed. A different
mechanism of carcinogenesis or various functions of the gene
polymorphism may contribute to this phenomenon.
For the risk of colorectal and lung cancers, the
present findings correspond to certain preceding studies that
evaluated the influence of the VEGF -2578C/A polymorphism on the
risk of these types of cancer (10). Significant heterogeneity did not
exist in the present study, despite performing a careful search,
establishing strict criteria, accurate data extraction and
comprehensive analysis. Therefore, the subgroup analyses were
performed to minimize the effects of heterogeneity in the following
way: Ethnicity, type of cancer and sources of control.
Some of the limitations that existed in the present
meta-analysis were inherent for all the previous meta-analysis that
focused on single-nucleotide polymorphisms, and others were caused
by artificial factors. Specific limitations in these studies should
be carefully explained. First, ethnicity, multifarious types of
cancer and various control sources gave rise to clear
heterogeneity. Second, the number of certain cancer subgroups,
including thyroid and ovarian cancers, was too small to investigate
the potential existence of a correlation between the VEGF -2578C/A
polymorphism and the corresponding cancer risk. Third, publication
bias may have arisen due to the search languages that contain
studies published only in English and Chinese. Fourth, the
exclusion of unattained data generally contributed to a false
estimation of the true effect. Regardless of the aforementioned
limitations, advantages of the present meta-analysis were evidently
facilitative to the final outcomes, which included a comprehensive
searching method, strict analytical procedures and a significant
conclusion that may contribute to individual therapy in the
future.
In conclusion, the present study indicates that the
VEGF -2578C/A polymorphism may have a particular association with
certain types of cancer, including lung and colorectal cancers, and
no evident association with breast cancer. More large scale
samples, including various types of cancer, particularly in single
studies, and different populations should be analyzed in a future
meta-analysis to obtain a more conclusive understanding with
regards to the function of the VEGF -2578C/A polymorphism in cancer
development. More information, including medical history, exposure
history, profession or even the climatic environment, should be
obtained in future individual studies to assess the possible
environmental-genome interaction.
References
|
1
|
Babashah S and Soleimani M: The oncogenic
and tumour suppressive roles of microRNAs in cancer and apoptosis.
Eur J Cancer. 47:1127–1137. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: the next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lacko M, Braakhuis BJ, Sturgis EM, et al:
Genetic susceptibility to head and neck squamous cell carcinoma.
Int J Radiat Oncol Biol Phys. 89:38–48. 2014. View Article : Google Scholar
|
|
4
|
Kaumaya PT and Foy KC: Peptide vaccines
and targeting HER and VEGF proteins may offer a potentially new
paradigm in cancer immunotherapy. Future Oncol. 8:961–987. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Vincenti V, Cassano C, Rocchi M and
Persico G: Assignment of the vascular endothelial growth factor
gene to human chromosome 6p21.3. Circulation. 93:1493–1495. 1996.
View Article : Google Scholar
|
|
6
|
Yang Y, Zhang X, Song D and Wei J:
Association between vascular endothelial growth factor gene
polymorphisms and bladder cancer risk. Mol Clin Oncol. 2:501–505.
2014.PubMed/NCBI
|
|
7
|
Ajaz S, Khaliq S, Abid A, et al:
Association of a single-nucleotide polymorphism in the promoter
region of the VEGF gene with the risk of renal cell carcinoma.
Genet Test Mol Biomarkers. 15:653–657. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Antonacopoulou AG, Kottorou AE,
Dimitrakopoulos FI, et al: VEGF polymorphisms may be associated
with susceptibility to colorectal cancer: a case-control study.
Cancer Biomark. 10:213–217. 2011.PubMed/NCBI
|
|
9
|
Dassoulas K, Gazouli M, Rizos S, et al:
Common polymorphisms in the vascular endothelial growth factor gene
and colorectal cancer development, prognosis, and survival. Mol
Carcinog. 48:563–569. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
10
|
Deng ZC, Cao C, Yu YM, Ma HY and Ye M:
Vascular endothelial growth factor -634G/C and vascular endothelial
growth factor -2578C/A polymorphisms and lung cancer risk: a
case-control study and meta-analysis. Tumour Biol. 35:1805–1811.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Diao LP, Yu XM, Gao YH, et al: Association
of VEGF genetic polymorphisms with the clinical characteristics of
non-Hodgkin’s lymphoma. J Cancer Res Clin Oncol. 135:1473–1481.
2009.PubMed/NCBI
|
|
12
|
Galimberti S, Nagy B, Palumbo GA, et al:
Vascular endothelial growth factor polymorphisms in mantle cell
lymphoma. Acta Haematol. 123:91–95. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Henríquez-Hernández LA, Navarro P, Luzardo
OP, et al: Polymorphisms of glutathione S-transferase μ and θ, MDR1
and VEGF genes as risk factors of bladder cancer: a case-control
study. Urol Oncol. 30:660–665. 2012.
|
|
14
|
Hofmann G, Langsenlehner U, Renner W, et
al: Common single nucleotide polymorphisms in the vascular
endothelial growth factor gene and colorectal cancer risk. J Cancer
Res Clin Oncol. 134:591–595. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Howell WM, Bateman AC, Turner SJ, Collins
A and Theaker JM: Influence of vascular endothelial growth factor
single nucleotide polymorphisms on tumour development in cutaneous
malignant melanoma. Genes Immun. 3:229–232. 2002. View Article : Google Scholar
|
|
16
|
Hsiao PJ, Lu MY, Chiang FY, Shin SJ, Tai
YD and Juo SH: Vascular endothelial growth factor gene
polymorphisms in thyroid cancer. J Endocrinol. 195:265–270. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ianni M, Porcellini E, Carbone I, et al:
Genetic factors regulating inflammation and DNA methylation
associated with prostate cancer. Prostate Cancer Prostatic Dis.
16:56–61. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jacobs EJ, Feigelson HS, Bain EB, et al:
Polymorphisms in the vascular endothelial growth factor gene and
breast cancer in the Cancer Prevention Study II cohort. Breast
Cancer Res. 8:R222006. View
Article : Google Scholar : PubMed/NCBI
|
|
19
|
Jaiswal PK, Tripathi N, Shukla A and
Mittal RD: Association of single nucleotide polymorphisms in
vascular endothelial growth factor gene with bladder cancer risk.
Med Oncol. 30:5092013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Jang MJ, Jeon YJ, Kim JW, et al:
Association of VEGF and KDR single nucleotide polymorphisms with
colorectal cancer susceptibility in Koreans. Mol Carcinog. 52(Suppl
1): E60–E69. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
21
|
Jin Q, Hemminki K, Enquist K, et al:
Vascular endothelial growth factor polymorphisms in relation to
breast cancer development and prognosis. Clin Cancer Res.
11:3647–3653. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kämmerer PW, Koch FP, Schiegnitz E, et al:
Associations between single-nucleotide polymorphisms of the VEGF
gene and long-term prognosis of oral squamous cell carcinoma. J
Oral Pathol Med. 42:374–381. 2013.
|
|
23
|
Ke Q, Liang J, Wang LN, et al: Potentially
functional polymorphisms of the vascular endothelial growth factor
gene and risk of gastric cancer. Mol Carcinog. 47:647–651. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kim YH, Kim MA, Park IA, et al: VEGF
polymorphisms in early cervical cancer susceptibility,
angiogenesis, and survival. Gynecol Oncol. 119:232–236. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li Y, Liang J, Liu X, et al: Correlation
of polymorphisms of the vascular endothelial growth factor gene and
the risk of lung cancer in an ethnic Han group of North China. Exp
Ther Med. 3:673–676. 2012.PubMed/NCBI
|
|
26
|
Li Y, Wang Y, Kang S, et al: Association
of vascular endothelial growth factor gene polymorphisms with
susceptibility to epithelial ovarian cancer. Int J Gynecol Cancer.
20:717–723. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Liang J, Yu X, Liu X, et al: Vascular
endothelial growth factor polymorphisms and risk of lung cancer.
Chin Ger J Clin Oncol. 8:269–272. 2009. View Article : Google Scholar
|
|
28
|
Maltese P, Canestrari E, Ruzzo A, et al:
VEGF gene polymorphisms and susceptibility to colorectal cancer
disease in Italian population. Int J Colorectal Dis. 24:165–170.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Martinez-Fierro ML, Garza-Veloz I,
Rojas-Martinez A, et al: Positive association between vascular
endothelial growth factor (VEGF) -2578 C/A variant and prostate
cancer. Cancer Biomark. 13:235–241. 2013.PubMed/NCBI
|
|
30
|
Mishra K, Behari A, Kapoor VK, Khan MS,
Prakash S and Agrawal S: Vascular endothelial growth factor
single-nucleotide polymorphism in gallbladder cancer. J
Gastroenterol Hepatol. 28:1678–1685. 2013.PubMed/NCBI
|
|
31
|
Nasr HB, Chahed K, Bouaouina N and
Chouchane L: Functional vascular endothelial growth factor -2578
C/A polymorphism in relation to nasopharyngeal carcinoma risk and
tumor progression. Clin Chim Acta. 395:124–129. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Nikiteas NI, Tzanakis N, Theodoropoulos G,
et al: Vascular endothelial growth factor and endoglin (CD-105) in
gastric cancer. Gastric Cancer. 10:12–17. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Park HM, Hong SH, Kim JW, et al:
Gender-specific association of the VEGF -2578C > A polymorphism
in Korean patients with colon cancer. Anticancer Res. 27:2535–2539.
2007.PubMed/NCBI
|
|
34
|
Pharoah PD, Tyrer J, Dunning AM, Easton DF
and Ponder BA: SEARCH Investigators: Association between common
variation in 120 candidate genes and breast cancer risk. PLoS
Genet. 3:e422005. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Sáenz-López P, Vazquez F, Cozar JM,
Carretero R, Garrido F and Ruiz-Cabello F: VEGF polymorphisms are
not associated with an increased risk of developing renal cell
carcinoma in Spanish population. Hum Immunol. 74:98–103. 2013.
|
|
36
|
Supic G, Jovic N, Zeljic K, Kozomara R and
Magic Z: Association of VEGF-A genetic polymorphisms with cancer
risk and survival in advanced-stage oral squamous cell carcinoma
patients. Oral Oncol. 48:1171–1177. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
VanCleave TT, Moore JH, Benford ML, et al:
Interaction among variant vascular endothelial growth factor (VEGF)
and its receptor in relation to prostate cancer risk. Prostate.
70:341–352. 2010.PubMed/NCBI
|
|
38
|
Wang K, Liu L, Zhu ZM, Shao JH and Xin L:
Five polymorphisms of vascular endothelial growth factor (VEGF) and
risk of breast cancer: a meta-analysis involving 16,703
individuals. Cytokine. 56:167–173. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Wu LM, Xie HY, Zhou L, Yang Z, Zhang F and
Zheng SS: A single nucleotide polymorphism in the vascular
endothelial growth factor gene is associated with recurrence of
hepatocellular carcinoma after transplantation. Arch Med Res.
40:565–570. 2009. View Article : Google Scholar
|
|
40
|
Zhang L, Zhang G, Wang P, Gong J, Cao Y
and Tang L: Association of vascular endothelial growth factor
-2578C/A gene polymorphism in Chinese patients with colon cancer.
Genet Test Mol Biomarkers. 15:117–121. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Kim EJ, Jeong P, Quan C, et al: Genotypes
of TNF-alpha, VEGF, hOGG1, GSTM1, and GSTT1: useful determinants
for clinical outcome of bladder cancer. Urology. 65:70–75. 2005.
View Article : Google Scholar : PubMed/NCBI
|