Introduction
Rheumatoid arthritis (RA) is a chronic debilitating
systemic autoimmune disease that affects 1% of the population and
may lead to permanent joint destruction resulting insignificant
morbidity (1). Early and accurate
diagnosis is important for selecting an effective treatment,
maintaining the quality of life and improving the survival rate of
RA. The current conventional methods for RA diagnosis primarily
include the American College of Rheumatology (ACR) classification
criteria (2). However, The ACR RA
classification criteria, including clinical and laboratory
characteristics, have various disadvantages regarding early
diagnosis (3), such as subjective
clinical diagnosis, the need for refinement using objective and
more sensitive imaging modalities, and the relatively low
sensitivity of anti-citrullinated protein antibodies (ACPA) for RA
(72%). Therefore, identifying appropriate novel biomarkers for
early diagnosis of RA is urgently required.
Novel diagnostic methods, such as linear RNAs,
including microRNAs (miRNAs) and long non-coding RNAs, have been
demonstrated to be useful as biomarkers for the diagnosis of RA
(4-7). Circular RNAs (circRNAs) are a novel
class of RNAs that form from the covalent linkage of the 3′ and 5′
ends to produce a closed loop (8,9).
Accumulating evidence indicates that several circRNAs regulate
various physiological and pathological processes by acting as
competing endogenous RNAs (ceRNAs) to restrain the activity of
miRNAs (10-12). circRNAs differ from linear RNAs in
that they can resist RNase digestion and exhibit high stability,
which allows circRNAs to be selectively enriched during sample
processing and makes them more suitable biomarkers compared with
linear RNAs (13,14). Recent evidence has indicated that
circRNAs may serve as novel biomarkers in the diagnosis and
prognosis of numerous diseases (15,16). In 2017, Ouyang et al
(17) confirmed that five
circRNAs (092516, 003524, 103047, 104871 and 101873) that were
found to be increased in peripheral blood mononuclear cells (PBMCs)
from patients with RA are potential biomarkers for RA diagnosis.
Our previous 2018 study revealed that peripheral blood
hsa_circ_0044235 may act as a biomarker for RA diagnosis (18). More recently, Tang et al
(19) quantified the elevated
expression of ciRS-7 in PBMC and suggested that it may be a
suitable diagnostic biomarker for RA. However, the diagnostic value
of these reported circRNAs is not optimal, due to the low or
moderate area under the receiver operating characteristic (ROC)
curve (AUC). Therefore, novel circRNA biomarkers for early
diagnosis and prognosis of RA must be identified. Our previous
study revealed that certain circRNAs in PBMCs were differentially
expressed between patients with systemic lupus erythematosus (SLE),
which is a common autoimmune disease, and healthy controls (HC)
(20). Considering that RA is
also a common autoimmune disease, three upregulated
(hsa_circ_0082689, hsa_circ_0087798, hsa_circ_0000175) and three
downregulated (hsa_circ_0008410, hsa_circ_0049356 and
hsa_circ_0032959) circRNAs, which were found to be significantly
aberrant in both peripheral blood mononuclear cells (PBMCs) from
patients with SLE in our previous study (20), and T cells from patients with SLE
in previous literature (21),
were selected to explore the possibility of being used as
biomarkers for diagnosis of RA.
Materials and methods
Patient variables
Patients (n=87) who fulfilled the revised ACR 2010
criteria for RA (2) were
consecutively enrolled from The First Affiliated Hospital of
Nanchang University (Jiangxi, China) between September 2018 and
March 2019. Those RA patients accompanied by other autoimmune,
inflammatory or hormonal disease, cancer or mental disorder were
excluded. All patients had new-onset RA and had not received
corticosteroids or immunosuppressive drugs prior to recruitment.
Then, 9 new-onset RA cases were administered therapeutic regimens
with corticosteroids and immunosuppressive drugs for at least 15
days. The information on disease activity score 28 (DAS28), swollen
joint count (SJC), tender joint count (TJC), patient visual
analogue scale (VAS), disease activity, erythrocyte sedimentation
rate (ESR), C-reactive protein (CRP), ACPA, rheumatoid factor (RF),
white blood cell count (WBC), red blood cell count (RBC),
hemoglobin, hematocrit (HCT), platelet count (PLT), mean platelet
volume (MPV), plateletcrit (PCT), platelet distribution width
(PDW), lymphocyte count (L), lymphocyte percentage (L%), monocyte
count (M), monocyte percentage (M%), neutrophil count (N),
neutrophil percentage (N%), neutrophil-to-lymphocyte ratio (NLR)
and platelet-to-lymphocyte ratio (PLR) was collected. HC (n=45)
without autoimmune or inflammatory diseases and who were also
unrelated to patients with autoimmune diseases were randomly
enrolled from the First Affiliated Hospital of Nanchang University
between September 2018 and March 2019. As a disease control, 91
patients with autoimmune diseases [50 SLE and 41 ankylosing
spondylitis (AS) cases] clinically confirmed by the diagnostic
criteria of SLE (22) and the
diagnostic criteria of AS (23)
after excluding RA, were also enrolled from the First Affiliated
Hospital of Nanchang University during the same period. All study
protocols complied with the principles outlined in the Declaration
of Helsinki and were approved by the Ethics Committee of the First
Affiliated Hospital of Nanchang University (ethics no. 019). All
participants provided signed informed consent.
Collection of PBMCs and total RNA
extraction
Blood samples (2 ml) were collected from all
subjects in K2-EDTA tubes, and PBMCs were isolated by
density-gradient centrifugation using Ficoll-Paque Plus (GE
Healthcare Life Sciences) at 25°C. TRIzol® reagent
(Invitrogen; Thermo Fisher Scientific, Inc.) was added to the PBMCs
and stored at −80°C. Total RNA extraction from PBMC specimens was
carried out according to the manufacturer's instructions. The
concentration and integrity of the RNA was assessed by a NanoDrop
ND-1000 spectrophotometer (Agilent Technologies, Inc.).
Reverse transcription-quantitative PCR
(RT-qPCR) analysis
RT and qPCR were performed with PrimeScript™ RT kit
(Takara Bio Inc.) and SYBR Premix Ex Taq™ II (Takara Bio Inc.),
respectively. RT-qPCR was performed on an ABI 7500 Real Time PCR
system (Applied Biosystems; Thermo Fisher Scientific, Inc.) with
the following PCR thermocycler protocol: Initial denaturation step
at 95°C for 5 min, followed by 40 cycles of 95°C for 15 sec
(denaturation), 60°C for 1 min (annealing and elongation) and 72°C
for 2 min (final extension). The selected circRNAs for RT-qPCR
analysis are shown in Table I.
β-actin was used as an internal control. The primers used in
RT-qPCR are listed in Table II.
The data were analyzed using the 2−ΔΔCq method (15).
 | Table IDetails of the selected circRNAs. |
Table I
Details of the selected circRNAs.
| circRNAs | Source | Chrom | Strand | circRNA_type | GeneSymbol |
|---|
|
hsa_circ_0082689 | circBase | chr7 | − | Exonic | PARP12 |
|
hsa_circ_0087798 | circBase | chr9 | + | Exonic | NIPSNAP3A |
|
hsa_circ_0000175 | circBase | chr1 | − | Exonic | ELK4 |
|
hsa_circ_0008410 | circBase | chr17 | + | Intronic | PGS1 |
|
hsa_circ_0049356 | circBase | chr19 | + | Exonic | CARM1 |
|
hsa_circ_0032959 | circBase | chr14 | − | Exonic | CCDC88 |
 | Table IISpecific circRNAs primers used for
RT-qPCR analysis. |
Table II
Specific circRNAs primers used for
RT-qPCR analysis.
| circRNAs | Primer sequence
(F) | Primer sequence
(R) |
|---|
|
hsa_circ_0082689 |
GTCCCCAAACACTCTTAGCCA |
CACACTCAGGTTGTGTTCGG |
|
hsa_circ_0087798 |
GCAGTTCATGTTCTTTGGTGGA |
CTGGGTCCCGTAGCAAAAGA |
|
hsa_circ_0000175 |
GCCCATTTTCCCCAGACCTAC |
GGAACTGCCACAGGGTGATA |
| hsa_circ_0008410
C |
TGCTTTTGTCCTTGAAGCCAG |
CACCAGCTCCGTGAAGAAGTC |
| hsa_circ_0049356
C |
ACCAAGGCCAACTTCTGGTA |
CGGTCCGTCAGGTTGTTACT |
|
hsa_circ_0032959 |
ACAGCTGGACATTGAGACCC |
TTTCCTCTCACACTGGACAGC |
| β-actin |
TACTGCCCTGGCTCCTAGCA |
TGGACAGTGAGGCCAGGATAG |
Statistical analysis
Statistical analysis and graphic presentation were
carried out with GraphPad Prism 5.0 (GraphPad Software, Inc.) and
SPSS version 17.0 (SPSS Inc.). The equation
n=Z2×б2d2=1.282×0.52/0.12=40.96
was used to calculate minimal sample size. A Student's t-test was
used between two groups where the samples passed the normality
test; otherwise, the non-parametric Mann-Whitney test was used to
analyze the data. The paired t-test was performed for evaluation of
changes in treatment. Kruskal-Wallis test was used for statistical
analysis between three or more groups followed by a Dunn's post-hoc
test for subsequent analyses between individual groups. Spearman's
rho method was used for correlation analysis. Multivariate analysis
(logistic regression) was used to analyze the risk factors. ROC
curves were constructed to evaluate the diagnostic value of
circRNAs that were dysregulated in the PBMCs of patients with RA
compared with HC, SLE and AS. P<0.05 was considered to indicate
statistically significant differences.
Results
Characteristics of the study
subjects
A total of 223 subjects were enrolled in the present
study, including 87 patients with RA, 50 patients with SLE, 45 HC
and 41 patients with AS. RA patients were classified into screening
and validation cohorts. The screening cohort included 24 RA
patients and 24 HC. An independent cohort consisting of 63 RA
patients and 21 HC were enrolled in the validation set for
evaluation of abnormal circRNAs. The characteristics of the study
subjects are summarized in Table
III. There were no significant differences between RA patients
and HC regarding age or sex. Due to the differences in age and sex
among RA, AS and SLE patients (the incidence of RA was high among
women of 50-60 years, the incidence of AS was high among young male
patients and the incidence of SLE was high among women of
childbearing age), patients with RA, AS and SLE were not
age-matched, and patients with RA and AS were not sex-matched in
the present study. No correlation between circRNA levels and age or
sex was observed in AS, SLE, RA or HC (data not shown).
 | Table IIIClinical characteristics of the
patients with RA, HC, SLE and AS. |
Table III
Clinical characteristics of the
patients with RA, HC, SLE and AS.
| Clinical
characteristic | RA | HC | AS | SLE |
|---|
| Number of
subjects | 87 | 45 | 41 | 50 |
| Sex | | | | |
| Male | 16 | 10 | 29b | 4 |
| Female | 71 | 35 | 12b | 46 |
| Age, years | 49.89±12.86 | 45.24±13.43 | 31.02±9.96b | 35.85±15.48c |
| Days since
diagnosis |
1,400.01±2,268.53 | | | |
| DAS28-ESR | 5.99±1.51 | | | |
| DAS28-CRP | 5.35±1.38 | | | |
| SJC | 10.83±7.65 | | | |
| PJC | 12.53±7.54 | | | |
| VAS | 48.44±31.39 | | | |
| RF, IU/ml | 436.02±558.63 | | | |
| ACPA, RU/ml | 911.04±972.24 | | | |
| ESR, mm/h | 51.04±33.32 | | 20.08±20.85b | 63.49±36.74 |
| CRP, mg/l | 27.54±32.13 | | 11.08±16.04b | 17.86±33.18 |
| WBC,
109/l | 7.97±2.39a | 5.74±0.83 | 7.40±1.66 | 6.32±3.66c |
| RBC,
1012/l | 4.37±0.51a | 4.56±0.37 | 4.78±0.71b | 3.75±0.82c |
| HGB, g/l |
124.60±19.99a | 138.87±11.37 |
140.78±20.59b |
117.66±77.67c |
| HCT, l/l | 0.38±0.05a | 0.41±0.03 | 0.43±0.06b | 0.33±0.08c |
| PLT,
109/l |
329.36±121.63a | 244.20±51.96 | 319.88±69.42 |
201.50±105.79c |
| MPV, fl | 10.22±1.41a | 10.91±0.89 | 9.46±1.02b | 10.63±1.27 |
| PCT, % | 0.33±0.11a | 0.26±0.05 | 0.30±0.06 | 0.25±0.09c |
| PDW, fl | 12.61±3.29a | 12.9±2.48 | 14.06±2.62b | 12.95±2.67 |
| L,
109/l | 1.64±0.58a | 1.84±0.31 | 2.08±0.57b | 1.25±0.64c |
| L, % | 21.57±8.51a | 32.41±5.31 | 28.51±7.13b | 23.37±10.46 |
| M,
109/l | 0.43±0.17a | 0.35±0.07 | 0.44±0.15 | 0.49±0.39 |
| M, % | 5.63±2.04a | 6.12±1.30 | 6.07±1.96 | 7.77±3.24c |
| N,
109/l | 5.74±2.25a | 3.42±0.71 | 4.70±1.38b | 4.51±3.16c |
| N, % | 70.47±10.45a | 59.04±5.95 | 62.98±7.53b | 67.80±11.87 |
| PLR |
237.62±181.47a | 136.03±36.19 |
164.94±57.32b |
191.23±136.05c |
| NLR | 4.12±2.73a | 1.90±0.50 | 2.43±0.98b | 4.26±3.84 |
PBMC hsa_circ_0000175 and
hsa_circ_0008410 expression is abnormal in RA patients
The expression of hsa_circ_0082689,
hsa_circ_0087798, hsa_circ_0000175, hsa_circ_0008410,
hsa_circ_0049356 and hsa_circ_0032959 in PBMCs was first detected
in the screening cohort including 24 RA patients and 24 HC using
RT-PCR. Compared with HC, the expression of hsa_circ_0000175 and
hsa_circ_0008410 was significantly different in the PBMCs from
patients with RA (all P<0.05, Fig.
1), while the expression of hsa_circ_0082689, hsa_circ_0087798,
hsa_circ_0049356 and hsa_circ_0032959 did not significantly differ
between the two groups.
Validation of PBMC hsa_circ_0000175 and
hsa_circ_0008410 expression in the second stage
Subsequently, the expression of hsa_circ_0000175 and
hsa_circ_0008410 in PBMCs was verified in an independent cohort,
including 63 RA patients and 21 HC. In accordance with the
screening results, the expression of hsa_circ_0000175 in PBMCs from
63 RA patients was significantly decreased compared with that in 21
HC (P<0.0001, Fig. 2A),
whereas the expression of hsa_circ_0008410 was significantly
increased in the PBMCs from 63 RA patients compared with that in 21
HC (P<0.0001, Fig. 2B). When
considering the data from all 87 RA patients and 45 HC, the same
patterns were observed in hsa_circ_0000175 and hsa_circ_0008410
between RA patients and HC (both P<0.0001, Fig. 2C and D).
Correlation of PBMC hsa_circ_0000175 and
hsa_circ_0008410 expression with clinical characteristics of
RA
To investigate whether the expressions of PBMC
hsa_circ_0000175 and hsa_circ_0008410 in RA patients could be
considered as biomarkers for the activity and severity of the
disease, Spearman's rho method was used to assess the association
between the expression of PBMC hsa_circ_0000175 and
hsa_circ_0008410 in RA patients with clinical characteristics,
including DAS28, TJC, TJC, VAS, disease duration, ACPA, RF, ESR,
CRP, WBC, RBC, HGB, HCT, PLT, MPV, PCT, PDW, L, L%, M, M%, N, N%,
NLR and PLR. As shown in Fig. 3,
the expression of PBMC hsa_circ_0000175 in RA patients was
correlated with ACPA (rs=−0.294, P=0.0090), WBC
(rs=−0.246, P=0.0216), L (rs=0.226,
P=0.0356), L% (rs=0.350, P=0.0009), N
(rs=−0.295, P=0.0056), N% (rs=−0.343,
P=0.0011) and NLR (rs=−0.367, P=0.0005), and the
expression of PBMC hsa_circ_0008410 in RA patients was correlated
with TJC (rs=0.213, P=0.0471), disease duration
(rs=0.211, P=0.0498), PLT (rs=0.241,
P=0.0247) and PCT (rs=0.267, P=0.0138), which indicated
the activity and severity of RA.
 | Figure 3Correlation of PBMC hsa_circ_0000175
and hsa_circ_0008410 expression with clinical characteristics of
RA. (A) The expression of PBMC hsa_circ_0000175 in RA patients was
negatively correlated with ACPA (Spearman's method). (B) The
expression of PBMC hsa_circ_0000175 in RA patients was negatively
correlated with WBC (Spearman's method). (C) The expression of PBMC
hsa_circ_0000175 in RA patients was negatively correlated with N
(Spearman's method). (D) The expression of PBMC hsa_circ_0000175 in
RA patients was negatively correlated with N% (Spearman's method).
(E) The expression of PBMC hsa_circ_0000175 in RA patients was
positively correlated with L (Spearman's method). (F) The
expression of PBMC hsa_circ_0000175 in RA patients was positively
correlated with L% (Spearman's method). (G) The expression of PBMC
hsa_circ_0000175 in RA patients was negatively correlated with NLR
(Spearman's method). (H) The expression of PBMC hsa_circ_0008410 in
RA patients was positively correlated with TJC (Spearman's method).
(I) The expression of PBMC hsa_circ_0008410 in RA patients was
positively correlated with disease duration (Spearman's method).
(J) The expression of PBMC hsa_circ_0008410 in RA patients was
positively correlated with PLT (Spearman's method). (K) The
expression of PBMC hsa_circ_0008410 in RA patients was positively
correlated with PCT (Spearman's method). ACPA, anti-citrullinated
protein antibodies; L, lymphocyte count; L%, lymphocyte percentage;
N, neutrophil count; N%, neutrophil percentage; NLR,
neutrophil-to-lymphocyte ratio; PBMC, peripheral blood mononuclear
cells; PCT, platelet-crit; PLT, platelet count; TJC, tender joint
count; RA, rheumatoid arthritis; WBC, white blood cell count. |
Subsequently, the expression of PBMC
hsa_circ_0000175 and hsa_circ_0008410 was detected in 9 new-onset
RA cases pre- and post-treatment; however, there was no difference
between pre- and post-treatment levels (data not shown).
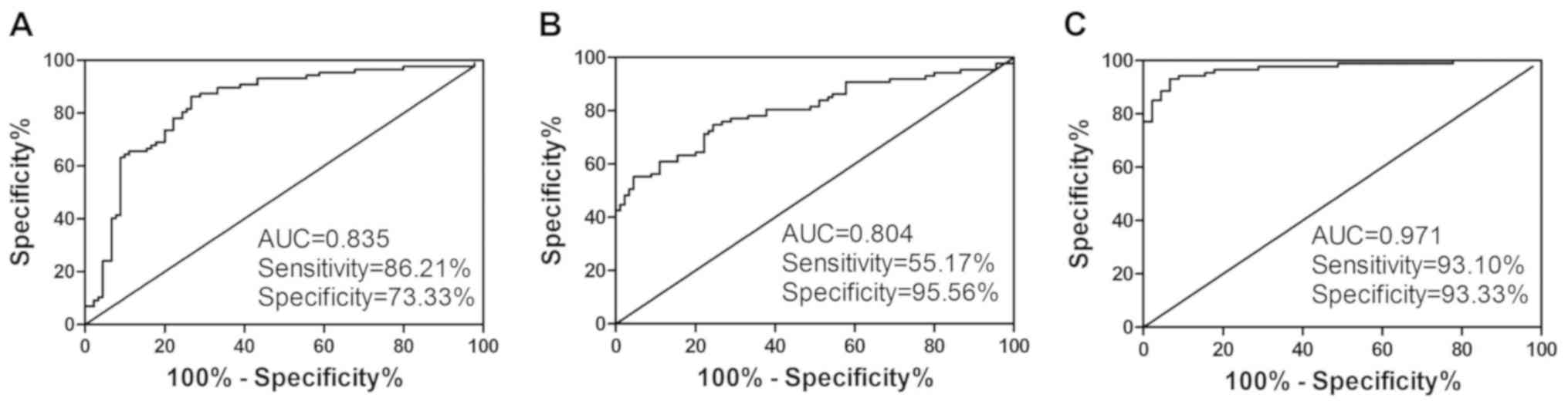
Diagnostic value of PBMC hsa_circ_0000175
and hsa_circ_0008410 expression in RA patients
Next, ROC curves were produced to investigate the
diagnostic value of PBMC hsa_circ_0000175 and hsa_circ_0008410
expression in RA. The data indicated that PBMC hsa_circ_0000175
expression had a moderate ability to distinguish RA patients from
HC, with an AUC of 0.835, a cut-off of <0.936, a sensitivity of
86.21% and a specificity of 73.33% (Fig. 4A), PBMC hsa_circ_0008410
expression also had a moderate ability to distinguish RA patients
from HC, with an AUC of 0.804, a cut-off of >2.685, a
sensitivity of 55.17% and a specificity of 95.56% (Fig. 4B). Moreover, the logistic
regression model revealed that the combination of PBMC
hsa_circ_0000175 and hsa_circ_0008410 expression exhibited an
improved ability to distinguish RA patients from HC, with an AUC of
0.971, a sensitivity of 93.10% and a specificity of 93.33%
(Fig. 4C). These results
demonstrated that the combination of PBMC hsa_circ_0000175 and
hsa_circ_0008410 expression may be a useful biomarker in RA.
PBMC hsa_circ_0000175 and
hsa_circ_0008410 expression in RA, SLE and AS patients
As shown in Fig.
5A, there was significant differences between the expression of
PBMC hsa_circ_0000175 in RA, SLE, and AS patients. The expression
of PBMC hsa_circ_0000175 was significantly increased in RA patients
compared with that in SLE patients, but markedly decreased compared
with that in AS patients. In addition, the expression of PBMC
hsa_circ_0008410 was markedly increased in RA patients compared
with that in SLE and AS patients (Fig. 5B).
 | Figure 5ROC curve analysis of the risk score
of PBMC hsa_circ_0000175 and hsa_circ_0008410. (A) There was a
significant difference between RA, SLE, and AS patients in the
expression of PBMC hsa_circ_0000175 (Kruskal-Wallis test). The
expression of PBMC hsa_circ_0000175 was markedly increased in RA
compared with SLE patients (Dunn's post-hoc test), while the
expression of PBMC hsa_circ_0000175 was markedly decreased in RA
compared with AS patients (Dunn's post-hoc test). (B) There was a
significant difference between RA, SLE, and AS patients in the
expression of PBMC hsa_circ_0008410 (Kruskal-Wallis test). The
expression of PBMC hsa_circ_0008410 was markedly increased in RA
compared with SLE (Dunn's post-hoc test) and AS patients (Dunn's
post-hoc test). (C) ROC curve analysis of PBMC hsa_circ_0000175 in
RA vs. SLE patients. (D) ROC curve analysis of PBMC
hsa_circ_0008410 in RA vs. SLE patients. (E) ROC curve analysis of
PBMC hsa_circ_0000175 in RA vs. AS patients. (F) ROC curve analysis
of PBMC hsa_circ_0008410 in RA vs. AS patients. (G) ROC curve
analysis of PBMC hsa_circ_0000175 in RA patients vs. controls (HC +
SLE + AS). (H) ROC curve analysis of PBMC hsa_circ_0008410 in RA
patients vs. controls (HC + SLE + AS). (I) ROC curve analysis of
combined PBMC hsa_circ_0000175 and hsa_circ_0008410 in RA patients
vs. controls (HC + SLE + AS). AUC, area under the curve; AS,
ankylosing spondylitis; HC, healthy controls; K-W test,
Kruskal-Wallis test; PBMC, peripheral blood mononuclear cells; RA,
rheumatoid arthritis; ROC, receiver operating characteristics; SLE,
systemic lupus erythematosus. |
Next, ROC curves based on PBMC hsa_circ_0000175 and
hsa_circ_0008410 were further analyzed in RA and SLE patients. The
AUC for PBMC hsa_circ_0000175 in RA and SLE patients was 0.642,
with a sensitivity of 60.92% and a specificity of 66.00% (Fig. 5C), and the AUC for PBMC
hsa_circ_0008410 in RA and SLE patients was 0.675, with a
sensitivity of 66.67% and a specificity of 65.31% (Fig. 5D). The expression of PBMC
hsa_circ_0000175 and hsa_circ_0008410 were also successful in
distinguishing RA from AS patients. The AUC for PBMC
hsa_circ_0000175 in RA and AS patients was 0.810, with a
sensitivity of 64.37% and a specificity of 90.24% (Fig. 5E), and the AUC for PBMC
hsa_circ_0008410 in RA and AS patients was 0.682, with a
sensitivity of 57.47% and a specificity of 82.93% (Fig. 5F). In addition, the expression of
PBMC hsa_circ_0000175 and hsa_circ_0008410 distinguished RA
patients from all controls (HC + SLE + AS). The AUC for PBMC
hsa_circ_0000175 in RA patients and all controls (HC + SLE + AS)
was 0.652, with a sensitivity of 83.97% and a specificity of 47.06%
(Fig. 5G), the AUC for PBMC
hsa_circ_0008410 in RA patients and all controls (HC + SLE + AS)
was 0.720, with a sensitivity of 55.17% and a specificity of 83.70%
(Fig. 5H), and the AUC for the
combination of PBMC hsa_circ_0000175 and hsa_circ_0008410 in RA
patients and all controls (HC + SLE + AS) was 0.805, with a
sensitivity of 83.76% and a specificity of 62.22% (Fig. 5I).
Expression levels of PBMC
hsa_circ_0000175 and hsa_circ_0008410 are risk factors for RA
The aforementioned results demonstrated that the
expression levels of PBMC hsa_circ_0000175 and hsa_circ_0008410 in
RA were different from HC, SLE and AS, and were associated with the
activity and severity of RA. Thus, to investigate whether the
expression labels of PBMC hsa_circ_0000175 and hsa_circ_0008410
were risk factors for RA, the 'enter method' of logistic regression
was used. As shown in Table IV,
the equation on the expression of hsa_circ_0000175 and
hsa_circ_0008410 in PBMC was as follows: Y=−2.019 X1
(hsa_circ_0000175) + 0.550 X2 (hsa_circ_0008410) - 0.292. The
expression levels of PBMC hsa_circ_0000175 and hsa_circ_0008410
were identified as risk factors for RA (all P<0.0001).
 | Table IVExpression of PBMC hsa_circ_0000175
and hsa_circ_0008410 in equation. |
Table IV
Expression of PBMC hsa_circ_0000175
and hsa_circ_0008410 in equation.
| Item | B | S.E | Wald | df | P | Exp (B) |
|---|
| hsa_circ_0000175
- | 2.019 | 0.443 | 20.742 | 1 | <0.0001 | 0.133 |
|
hsa_circ_0008410 | 0.550 | 0.106 | 26.877 | 1 | <0.0001 | 1.733 |
| Constant | −0.292 | 0.351 | 0.694 | 1 | 0.4050 | 0.747 |
hsa_circ_0000175/miRNA,
hsa_circ_0008410/miRNA interaction analysis
To confirm the function of hsa_circ_0000175 and
hsa_circ_0008410, potential miRNA targets of the circRNAs were
predicted by aligning with the miRNA response elements (MREs) using
Arraystar's home-made miRNA target prediction software based on
TargetScan (24) and miRanda
(25). Three putative miRNA
targets of hsa_circ_0000175 were identified, including hsa-miR-608,
hsa-miR-654-3p and hsa-miR-3652. Three putative miRNA targets of
hsa_circ_0008410 were also identified, including hsa-miR-6776-3p,
hsa-miR-412-3p and hsa-miR-4697-3p. The molecular interactions
between these two circRNAs and the target miRNAs mentioned above
are illustrated in Fig. 6.
Discussion
It has been previously reported that PBMC circRNAs
may be associated with RA. In 2017, Ouyang et al (17) and Zheng et al (26) simultaneously investigated
differentially expressed circRNAs in PBMCs from RA patients by
microarray and RT-qPCR. In addition, Tang et al (19) recently demonstrated that the
expression of ciRS-7 was increased in PBMCs from RA patients.
However, little was known regarding the expression of
hsa_circ_0000175 and hsa_circ_0008410 in PBMCs from RA. The present
study demonstrated that PBMC hsa_circ_0000175 and hsa_circ_0008410
were markedly decreased and increased, respectively, in RA patients
compared with HC.
As shown in Table
III, the clinical manifestations (DAS28, TJC, SJC and VAS) and
laboratory indicators (autoantibodies, inflammation markers and
immune cells) in RA were different from the controls, indicating
the activity and severity of RA. Spearman's rho correlation
analysis revealed that PBMC hsa_circ_0000175 was correlated with
the ACPA titer, which is a biomarker of RA and reflects its
activity (27). In addition, PBMC
hsa_circ_0000175 was correlated with inflammatory markers,
including WBC, N, N%, L, L% and NLR, which reflect the activity and
severity of RA (28). Moreover,
PBMC hsa_circ_0008410 was correlated with TJC, disease duration and
PLT, which are associated with the development of peripheral
neuropathy and are correlated with DAS28 (29,30), and PCT, which is associated with
the treatment of RA (31). Thus,
these data indicated that PBMC hsa_circ_0000175 and
hsa_circ_0008410 were associated with the activity and severity of
RA. Importantly, logistic regression analysis revealed that PBMC
hsa_circ_0000175 and hsa_circ_0008410 were risk factors for RA,
suggesting that hsa_circ_0000175 and hsa_circ_0008410 may be
involved in the pathogenesis of RA.
Recent evidence has indicated that circRNAs may
serve as novel biomarkers in the diagnosis of RA (17-19). To explore whether PBMC
hsa_circ_0000175 and hsa_circ_0008410 constitute biomarkers for the
diagnosis of RA, they were analyzed in larger patient cohorts using
ROC curves. The cut-off values of hsa_circ_0000175 and
hsa_circ_0008410 that best distinguished RA patients from HC were
determined. Hsa_circ_0000175 had an AUC of 0.835, a specificity of
73.33% and a sensitivity of 86.21% for RA, whereas hsa_circ_0008410
had an AUC of 0.804, a specificity of 95.56% and a sensitivity of
55.17%. The logistic regression model using both targets in
combination exhibited an improved ability for distinguishing RA
patients from HC, with an AUC of 0.971, a sensitivity of 93.10% and
a specificity of 93.33%, indicating the additive effects of the two
circRNAs in terms of diagnostic value. The diagnostic potential
appears to be superior to that of blood biomarkers of RA reported
previously, particularly in terms of AUC and specificity (17-19). Subsequently, the ability of PBMC
hsa_circ_0000175 and hsa_circ_0008410 to effectively distinguish RA
from other autoimmune diseases, such as SLE and AS, was assessed.
As aforementioned, PBMC hsa_circ_0000175 and hsa_circ_0008410 were
successful in distinguishing RA patients from SLE patients, AS
patients and HC.
It is well-known that circRNAs may act as miRNA
sponges and regulate target genes to alter the course of disease
development. Li et al (32) reported that hsa_circ_0001859
regulated ATF2 expression by acting as an miR-204/211 sponge in
human RA. Our previous study (18) demonstrated that hsa-miR-892a, an
miRNA target of hsa_circ_0044235, was increased in RA patients,
while hsa_circ_0044235 was decreased, indicating that
hsa_circ_0044235 may play a role in RA by interacting with
hsa-miR-892a. Multiple miRNAs are implicated in the occurrence and
development of RA (33,34). Bioinformatics predicted that
hsa-miR-608, hsa-miR-654-3p and hsa-miR-3652 may be potential
targets of hsa_circ_0000175. Hsa_circ_0008410 was shown to
potentially bind hsa-miR-6776-3p, hsa-miR-412-3p and
hsa-miR-4697-3p. Although most potential miRNAs interacting with
circRNAs have been predicted for RA, no study on the function of
these miRNAs in RA has been published to date, to the best of our
knowledge.
However, several limitations of the present study
should be acknowledged. Firstly, six circRNAs that were
differentially expressed in both PBMC from SLE patients in our
previous study and T cells from SLE patients in previous literature
were selected, and the possibility of using them as biomarkers for
diagnosis of RA was explored. Although RA and SLE exhibit different
pathogenies, these two diseases have similar pathological and
immunological abnormalities. Furthermore, there were significant
differences in the levels of PBMC hsa_circ_0000175 and
hsa_circ_0008410 between RA and SLE patients. Thus, the possibility
of hsa_circ_0000175 and hsa_circ_0008410 being used as biomarkers
for diagnosis of RA require further exploration. Secondly, the
sample size of the patients with new-onset RA, SLE, AS and HC was
relatively small, and the samples were sourced from only one
hospital, which may limit the universality of the results.
Therefore, the current findings require confirmation in larger and
more diverse samples.
In conclusion, the PBMC hsa_circ_0000175,
hsa_circ_0008410, and combination of hsa_circ_0000175 and
hsa_circ_0008410, may improve the diagnostic accuracy for RA. In
addition, the expression levels of PBMC hsa_circ_0000175 and
hsa_circ_0008410 were found to be associated with the activity and
severity of RA.
Funding
The present was supported by the Key Research and
Development Plan Project of Jiangxi Province (grant no.
20181BBG70013), the Science and Technology Plan Project of the
Education Department of Jiangxi Province (grant no. 170008), the
National Natural Science Foundation of China (grant nos. 81360459
and 81660277), the Jiangxi Provincial Natural Science Foundation of
China (grant nos. 20151BAB215031 and 20171BAB205113), the Science
and Technology Project of Health and Family Planning Commission of
Jiangxi Province of China (grant no. 20165094) and the Foundation
for Distinguished Young Scientists of Jiangxi Province of China
(grant no. 20171BCB23087).
Availability of data and materials
The data used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
QL, LLZ and LBZ performed the experiments. JYR, LZ,
YG, ZKH and JML analyzed and interpreted the data. QL and JML made
substantial contributions to the design and supervision of the
present study, and wrote the manuscript. All authors have reviewed
the results and approved the final version of the manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of the First Affiliated Hospital of Nanchang University.
All participants provided informed consent prior to inclusion in
the study.
Patient consent for publication
Not applicable.
Competing interests
The sauthors declare that they have no competing
interests.
Acknowledgments
The authors would like to acknowledge Dr Rui Wu
(Department of Rheumatology, the First Affiliated Hospital of
Nanchang University, Jiangxi, China) for their help in screening
patients and collecting specimens.
References
|
1
|
McInnes IB and Schett G: Cytokines in the
pathogenesis of rheumatoid arthritis. Nat Rev Immunol. 7:429–442.
2007. View
Article : Google Scholar : PubMed/NCBI
|
|
2
|
Aletaha D, Neogi T, Silman AJ, Funovits J,
Felson DT, Bingham CO III, Birnbaum NS, Burmester GR, Bykerk VP,
Cohen MD, et al: 2010 rheumatoid arthritis classification criteria:
An American College of Rheumatology/European League Against
Rheumatism collaborative initiative. Ann Rheum Dis. 69:1580–1588.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Cohen S and Emery P: The American College
of Rheumatology/European League Against Rheumatism criteria for the
classification of rheumatoid arthritis: A game changer. Arthritis
Rheum. 62:2592–2594. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Abdul-Maksoud RS, Sediq AM, Kattaia A,
Elsayed W, Ezzeldin N, Abdel Galil SM and Ibrahem RA: Serum miR-210
and miR-155 expression levels as novel biomarkers for rheumatoid
arthritis diagnosis. Br J Biomed Sci. 7:1–5. 2017.
|
|
5
|
Wang W, Zhang Y, Zhu B, Duan T, Xu Q, Wang
R, Lu L and Jiao Z: Plasma microRNA expression profiles in Chinese
patients with rheumatoid arthritis. Oncotarget. 6:42557–42568.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lu MC, Yu HC, Yu CL, Huang HB, Koo M, Tung
CH and Lai NS: Increased expression of long noncoding RNAs
LOC100652951 and LOC100506036 in T cells from patients with
rheumatoid arthritis facilitates the inflammatory responses.
Immunol Res. 64:576–583. 2016. View Article : Google Scholar
|
|
7
|
Zhang HJ, Wei QF, Wang SJ, Zhang HJ, Zhang
XY, Geng Q, Cui YH and Wang XH: LncRNA HOTAIR alleviates rheumatoid
arthritis by targeting miR-138 and inactivating NF-κB pathway. Int
Immunopharmacol. 50:283–290. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Qu S, Yang X, Li X, Wang J, Gao Y, Shang
R, Sun W, Dou K and Li H: Circular RNA: A new star of noncoding
RNAs. Cancer Lett. 365:141–148. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Chen LL and Yang L: Regulation of circRNA
biogenesis. RNA Biol. 12:381–388. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Fischer JW and Leung AK: CircRNAs: A
regulator of cellular stress. Crit Rev Biochem Mol Biol.
52:220–233. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wu HJ, Zhang CY, Zhang S, Chang M and Wang
HY: Microarray expression profile of circular RNAs in heart tissue
of mice with myocardial infarction-induced heart failure. Cell
Physiol Biochem. 39:205–216. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kulcheski FR, Christoff AP and Margis R:
Circular RNAs are miRNA sponges and can be used as a new class of
biomarker. J Biotechnol. 238:42–51. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Salzman J, Chen RE, Olsen MN, Wang PL and
Brown PO: Cell-type specific features of circular RNA expression.
PLoS Genet. 9:e10037772013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Rybak-Wolf A, Stottmeister C, Glazar P,
Jens M, Pino N, Giusti S, Hanan M, Behm M, Bartok O, Ashwal-Fluss
R, et al: Circular RNAs in the mammalian brain are highly abundant,
conserved, and dynamically expressed. Mol Cell. 58:870–885. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Huang Z, Su R, Qing C, Peng Y, Luo Q and
Li J: Plasma circular RNAs hsa_circ_0001953 and hsa_circ_0009024 as
diagnostic biomarkers for active tuberculosis. Front Microbiol.
9:20102018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bach DH, Lee SK and Sood AK: Circular RNAs
in cancer. Mol Ther Nucleic Acids. 16:118–129. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ouyang Q, Wu J, Jiang Z, Zhao J, Wang R,
Lou A, Zhu D, Shi GP and Yang M: Microarray expression profile of
circular RNAs in peripheral blood mononuclear cells from rheumatoid
arthritis patients. Cell Physiol Biochem. 42:651–659. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Luo Q, Zhang L, Li X, Fu B, Deng Z, Qing
C, Su R, Xu J, Guo Y, Huang Z and Li J: Identification of circular
RNAs hsa_circ_0044235 in peripheral blood as novel biomarkers for
rheumatoid arthritis. Clin Exp Immunol. 194:118–124. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Tang X, Wang J, Xia X, Tian J, Rui K, Xu H
and Wang S: Elevated expression of ciRS-7 in peripheral blood
mononuclear cells from rheumatoid arthritis patients. Diagn Pathol.
14:112019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Luo Q, Zhang L, Li X, Fu B, Guo Y, Huang Z
and Li J: Identification of circular RNAs hsa_circ_0044235 and
hsa_circ_0068367 as novel biomarkers for systemic lupus
erythematosus. Int J Mol Med. 44:1462–1472. 2019.PubMed/NCBI
|
|
21
|
Li LJ, Zhu ZW, Zhao W, Tao SS, Li BZ, Xu
SZ, Wang JB, Zhang MY, Wu J, Leng RX, et al: Circular RNA
expression profile and potential function of hsa_circ_0045272 in
systemic lupus erythematosus. Immunology. 155:137–149. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hochberg MC: Updating the American College
of Rheumatology revised criteria for the classification of systemic
lupus erythematosus. Arthritis Rheum. 40:17251997. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
van der Linden S, Valkenburg HA and Cats
A: Evaluation of diagnostic criteria for ankylosing spondylitis, A
proposal for modification of the New York criteria. Arthritis
Rheum. 27:361–368. 1984. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Enright A, John B, Gaul U, Tuschl T,
Sander C and Marks DS: MicroRNA targets in Drosophila. Genome Biol.
5:R12003. View Article : Google Scholar
|
|
25
|
Pasquinelli AE: MicroRNAs and their
targets: Recognition, regulation and an emerging reciprocal
relationship. Nat Rev Genet. 13:271–282. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zheng F, Yu X, Huang J and Dai Y: Circular
RNA expression profiles of peripheral blood mononuclear cells in
rheumatoid arthritis patients, based on microarray chip technology.
Mol Med Rep. 16:8029–8036. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hafkenscheid L, de Moel E, Smolik I,
Tanner S, Meng X, Jansen BC, Bondt A, Wuhrer M, Huizinga TWJ, Toes
REM, et al: N-linked glycans in the variable domain of ACPA-IgG
predict the development of rheumatoid arthritis. Arthritis
Rheumatol. 71:1626–1633. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sargin G, Senturk T, Yavasoglu I and Kose
R: Relationship between neutrophil-lymphocyte, platelet-lymphocyte
ratio and disease activity in rheumatoid arthritis treated with
rituximab. Int J Rheum Dis. 21:2122–2127. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li Y, Jiang L, Zhang Z, Li H, Jiang L,
Wang L and Li Z: Clinical characteristics of rheumatoid arthritis
patients with peripheral neuropathy and potential related risk
factors. Clin Rheumatol. 38:2099–2107. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zhou L, Xiao DM, Qin W, Xie BH, Wang TH,
Huang H, Zhao BJ, Han X, Sun QQ, Wu XD and Cen H: The clinical
value of hematological markers in rheumatoid arthritis patients
treated with tocilizumab. J Clin Lab Anal. 19:e228622019.
|
|
31
|
Liu J, Cao Y, Huang C, Wang Y, Chen X,
Zhang W, Wang G, Fan H, Ge Y, Chen R, et al: Use of xinfeng capsule
to treat abar-ticular pathologic changes in patients with
rheumatoid arthritis. J Tradit Chin Med. 34:532–538. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li B, Li N, Zhang L, Li K, Xie Y, Xue M
and Zheng Z: Hsa_circ_0001859 regulates ATF2 expression by
functioning as an MiR-204/211 sponge in human Rheumatoid Arthritis.
J Immunol Res. 2018:94123872018. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Qu Y, Zhang YP, Wu J, Jie LG, Deng JX,
Zhao DB and Yu QH: Downregulated microRNA-135a ameliorates
rheumatoid arthritis by inactivation of the phosphatidylinositol
3-kinase/AKT signaling pathway via phosphatidylinositol 3-kinase
regulatory subunit 2. J Cell Physiol. 234:17663–17676. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Philippe L, Alsaleh G, Bahram S, Pfeffer S
and Georgel P: The miR-17~92 Cluster: A key player in the control
of inflammation during Rheumatoid Arthritis. Front Immunol.
4:702013. View Article : Google Scholar
|