Introduction
The endocytosis of epidermal growth factor receptor
(EGFR) serves as a model for studying ligand-induced
receptor-mediated endocytosis. Upon EGF stimulation, the dimerizd
EGF-EGFR complexes are internalized and transported via
clathrin-coated vesicles to early endosomes. EGFR then recruits and
phosphorylates signaling molecules, leading to the activation of
the MAPK-signal transduction cascade, an important mechanism for
regulating cell growth (1–4). To turn off EGF signaling, the
EGF-EGFR complexes are delivered to the lysosomes for degradation
by a process known as receptor downregulation. Therefore,
endocytosis of EGF-EGFR complexes is closely related to attenuation
of intracellular EGFR signaling. Furthermore, EGFR mediates an
important role in the pathogenesis of different tumors, and
therapies directed at inhibiting EGFR function have potential as
anticancer treatments (5,6).
Gefitinib, a selective EGFR tyrosine kinase
inhibitor, has been shown to block the signal transduction pathways
implicated in the proliferation and survival of cancer cells
(7–10). It was reported previously that of
the 9 non-small cell lung cancer (NSCLC) cell lines examined, the
PC9 cell line was most sensitive to the effect of gefitinib with
respect to EGFR phosphorylation and activation of EGFR downstream
effectors such as AKT and those in the ERK1/2 pathway, which are
required for EGFR-stimulated proliferation (11). In contrast, the other NSCLC lines
such as QG56 and A549 cells showed greater resistance to gefitinib
(11). Consequently, we
hypothesize that the mechanism responsible for determining the
sensitivity of the EGFR endocytic pathway could be useful in
predicting the potential effectiveness of gefitinib in NSCLC
patients. We have previously investigated the endocytosis of Texas
red-labeled EGF in the absence or presence of gefitinib in three
NSCLC cell lines, and then assessed the amounts of internalized
Texas red-EGF or phosphorylated EGFR (pEGFR) by using confocal
immunofluorescence microscopy (12–14).
We showed that an aberration in certain steps of EGF-EGFR/pEGFR
trafficking from the early endosomes to the late
endosomes/lysosomes does occur in the gefitinib-resistant human
lung cancer cell line QG56 and A549, whereas endocytosis of
EGFR/pEGFR is normal in gefitinib-sensitive PC9 cells (12–14).
Accordingly, we suggested that impairment of certain steps of
EGF-EGFR/pEGFR trafficking from early endosomes to late
endosomes/lysosomes might confer gefitinib-resistance in NSCLC cell
lines. Furthermore, we made a novel observation that large amounts
of sorting nexin 1 (SNX1) are localized in the aggregated vesicular
structures of early endosomes where the internalized pEGFR is also
accumulated (13,14). Therefore, we postulate that
impairment of protein function, such as the SNX1 regulation of
EGFR/pEGFR trafficking in the early endocytic pathway, might
perturb EGFR/pEGFR endocytosis, which subsequently leads to
gefitinib-resistance in NSCLC cell lines.
SNX1 was previously demonstrated to be a protein
that interacts with EGFR (15) and
is localized to early endosomes through its phospholipid-binding
motif termed the phox homology (PX) domain (16). SNX1 is homologous to Vps5p, a yeast
protein that is required for endosome-to-Golgi trafficking
(17–19). Previous studies also revealed that
over-expression of SNX1 causes enhanced EGFR degradation and that
deletion mutant of SNX1 blocked EGFR degradation but failed to
inhibit receptor endocytosis (15,20).
Therefore, it was suggested that SNX1 interacts with EGFR and
enhances the degradation of the receptor upon EGF stimulation,
thereby implying that SNX1 plays a role in endosome-lysosome
trafficking. However, recent evidence has failed to support the
intracellular colocalization of SNX1 with EGFR and its direct role
in EGFR degradation, raising the possibility that alternative
mechanisms are involved in the function of SNX1 (21,22).
Consequently, the molecular mechanism underlying EGFR membrane
trafficking remains to be elucidated.
In the present study, we analyzed the intracellular
regulatory function of SNX1 with regard to EGF-induced endocytosis
and downregulation of EGFR/pEGFR using confocal immunofluorescence
microscopy, western blot analysis, and RNAi-mediated knockdown
approaches in gefitinib-sensitive and gefitinib-resistant NSCLC
cell lines. We demonstrated that silencing of endogenous SNX1 by
siRNA stimulates efficient endocytosis of ligand-induced EGFR and
pEGFR in gefitinib-resistant NSCLC cells. We also found that
depletion of SNX1 stimulates the ligand-induced downregulation of
EGFR/pEGFR, while increasing of pEGFR protein expression in
gefitinib-resistant cells. Therefore, we postulate that SNX1 plays
a negative role in the regulation of EGF-dependent down-regulation
of EGFR and its phosphorylation via the early/late endocytic
pathway in human lung cancer cells.
Materials and methods
Materials
Texas red-labeled human transferrin, Texas
red-labeled EGF, and SlowFade anti-fade reagent were purchased from
Molecular Probes (Eugene, OR, USA). DAPI, recombinant human EGF was
purchased from PeproTech (London, UK). Bafilomycin A1, and
cycloheximide (CHX) were obtained from Sigma (St. Louis, MO, USA).
Other chemicals were of reagent grade and were obtained from
commercial sources.
Cell culture
Cell lines PC9, QG56 and A549 (National Kyushu
Cancer Center, Fukuoka, Japan) were cultured in RPMI supplemented
with 10% fetal bovine serum (FBS). Cells were maintained under
standard cell culture conditions at 37°C and 5% CO2 in a
humid environment.
Small interfering RNA
siRNA targeting SNX1 was purchased from Dharmacon
(Boulder, CO, USA). The target sequence of the siRNA was as
follows: 5′-AAGAACAAGACCAAGAGCCAC-3′. Scramble sequence was used as
a control. The 3 NSCLC cell lines were transfected with
Lipofectamine 2000 (Life Technologies, Gaithersburg, MD, USA) in
the presence of 40 nM siRNA targeting SNX1 according to the
manufacturer’s protocol. Knockdown efficiency was determined by
qRT-PCR, and confocal immunofluorescence microscopy analysis.
qRT-PCR analysis
The 3 NSCLC cell lines PC9, QG56 or A549 cells
transfected with siRNA-control or siRNA-SNX1 were stimulated with
EGF (100 ng/ml) at 37°C for 5, 15, or 30 min, and total RNA was
extracted from each cell line using an RNeasy RNA isolation kit
(Qiagen, Hilden, Germany) according to the manufacturer’s
instructions. Transcription into cDNA was done in a 20-μl volume
using ThermoScript RT-PCR System with random hexamer (Invitrogen,
Carlsbad, CA, USA) according to the manufacturer’s instructions.
All PCR reactions were carried out in a final volume of 25 μl and
were performed in the ABI PRISM 7000 Sequence Detection System
(Applied Biosystems, Foster City, CA, USA) according to the
manufacturer’s protocol. Sequence-specific primers were quoted from
an official website ‘PrimerBank’ (http://pga.mgh.harvard.edu/primerbank/) for the
indicated genes (Table I). The
reaction mix consisted of SYBR Premix Ex Taq (2x) (Takara Bio.,
Shiga, Japan) 12.5 μl, ROX Reference Dye (x50) (Takara Bio.) 0.5
μl, 0.2 μM of each specific forward and reverse primer, and 9 μl of
diluted cDNA (equivalent to 0.03–2.85 ng of total RNA).
Amplifications were done under standard conditions (10 sec at 95°C
followed by 40 cycles of 5 sec at 95°C and 31 sec at 60°C). The
number of PCR cycles needed to reach the fluorescence threshold was
determined in triplicate for each cDNA, averaged, and then
normalized to a reference gene (β-actin). A standard curve
generated with serial 3-fold dilutions of a representative cDNA.
For all assays tested, the PCR reaction was linear over the range
studied (20–40 cycles of amplification). All RT-PCR reactions gave
a single band when analyzed by gel electrophoresis.
 | Table IPrimers for qRT-PCR for human SNX1,
EGFR, and β-actin. |
Table I
Primers for qRT-PCR for human SNX1,
EGFR, and β-actin.
| PCR primer name | Nucleotide
sequence |
|---|
| SNX1-forward |
5′-AGCCCCAGCCAACCTATGA-3′ |
| SNX1-reverse |
5′-TCAGGATCAGTTATACCGACTGT-3′ |
| EGFR-forward |
5′-GCGTTCGGCACGGTGTATAA-3′ |
| EGFR-reverse |
5′-GGCTTTCGGAGATGTTGCTTC-3′ |
| β-actin-forward |
5′-CATGTACGTTGCTATCCAGGC-3′ |
| β-actin-reverse |
5′-CTCCTTAATGTCACGCACGAT-3′ |
Antibodies
Alexa 488-labeled goat anti-mouse and goat
anti-rabbit secondary antibodies were obtained from Molecular
Probes. Normal rabbit IgG and normal mouse monoclonal IgG1 were
purchased from Imgenex (San Diego, CA, USA) and Angio-proteomie
(Boston, MA, USA), respectively. Normal goat serum was purchased
from Sigma. Antisera were raised in rabbits (New Zealand white
male) against the native form of LIMPII/LGP85 (23) as described previously. Anti-LIMPII
IgG was affinity-purified by protein A Sepharose CL-4B (Sigma),
followed by immunoaffinity chromatography using antigen-conjugated
Sepharose 4B. Mouse monoclonal antibody to SNX1 was purchased from
BD Biosciences (San Jose, CA, USA). A mouse monoclonal anti-pEGFR
was obtained from Cell Signaling Technology (Beverly, MA, USA), BD
Biosciences and Dako Cytomation (Denmark).
Immunofluorescence microscopy (general
procedures)
Immunofluorescence microscopy was described
previously (12–14,24–26).
Cells were grown for 2 days on glass coverslips in 6-well plates in
RPMI with 10% fetal bovine serum. Cells were fixed with 3.7%
formaldehyde in phosphate-buffered saline (PBS), pH 7.4,
permeabilized in PBS containing 0.1% saponin. After washing with
PBS, cells were blocked with PBS-10% normal goat serum. All
subsequent antibody and wash solutions contained 0.1% saponin. The
PC9, QG56 and A549 cells were incubated with specific primary
antibodies (rabbit anti-LIMPII IgGs, mouse anti-pEGFR mAb, or mouse
anti-SNX1 mAb), for 1 h, followed by washes with PBS containing
0.1% saponin and incubation for 1 h with the secondary antibodies
at 20 μg/ml. Each cell line was stained with DAPI to reveal nuclei.
Controls for antibody specificity were non-immune normal mouse IgG1
or nonimmune normal rabbit IgG. To label early endosomes, cells
were incubated with RPMI without FBS for 3 h at 37°C followed by 20
min incubation in culture medium containing Texas red-conjugated
transferrin, and then cells were fixed and double-stained for SNX1
with anti-SNX1 monoclonal antibody and LIMPII with anti-LIMPII
antibody. Late endosomes/lysosomes were stained with anti-LIMPII
antibody, since the LIMPII protein is distributed within endocytic
organelles and is at its highest concentration in the late
endosomes/lysosomes, as observed for other lysosomal glycoproteins,
namely, lysosomal-associated membrane protein-1 (LAMP-1) and LAMP-2
(23,27,28).
The distribution of the labeled proteins was then analyzed by
confocal immunofluorescence microscopy of the fixed cells. Slides
were mounted with SlowFade anti-fade reagent and observed on a
Zeiss LSM 510 META confocal laser scanning microscope (Carl Zeiss,
Oberkochen, Germany), equipped with krypton/argon laser sources.
For quantification of co-localization between Texas Red-EGF and
LIMPII or pEGFR and LIMPII, merged images as yellow color were
quantified and presented as the percentage of total amounts of
LIMPII-positive vesicles per cell.
Immunofluorescence microscopy (treatment
the cells with EGF)
In order to clarify EGFR internalization, we
followed the uptake of Texas red-conjugated EGF with time in each
cell line. To minimize the contribution of recycling and/or
lysosomal degradation of the internalized EGFR, we quantified the
Texas red-EGF uptake in each cell for time periods of up to 60 min.
At 48 h transfection, PC9, QG56 and A549 cells treated with
siRNA-control or siRNA-SNX1 were starved for 12 h with RPMI without
FBS at 37°C and the serum-starved cells were then incubated with
Texas red-EGF (100 ng/ml) at 37°C for 15, 30, or 60 min, and the
distribution of internalized Texas red-EGF and late
endosomes/lysosomes stained with anti-LIMPII antibody was then
assessed by confocal immunofluorescence microscopy. In some cases,
cells were starved for 3 h with RPMI without FBS at 37°C and then
the phosphorylation of EGFR was induced with EGF (100 ng/ml) for 15
min on ice in binding medium (1 mg/ml BSA in RPMI medium). The
cells were then rinsed with ice-cold PBS, incubated in the presence
of Texas red-transferrin in prewarmed medium, and chased at 37°C
for 60 min. The fixed cells were double-stained for pEGFR with
anti-pEGFR monoclonal antibody and LIMPII with anti-LIMPII
antibody.
Western blot analysis
Protein samples were separated by sodium dodecyl
sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) and then
transferred to polyvinylidene difluoride membranes (Millipore,
Billerica, MA, USA). Following blocking, the membrane was blotted
with the appropriate antibody, and subsequently, horseradish
peroxidase-conjugated anti-mouse or anti-rabbit IgG (GE Healthcare
Bioscience, Tokyo, Japan) was applied. The final signal was
revealed by ECL chemiluminescence (Pierce, Rockford, IL, USA).
Digital images were analyzed with NIH Image software to measure the
density of each band without a saturated signal.
EGF-stimulated EGFR degradation
PC9 and A549 cells were starved for 12 h with RPMI
without FBS at 37°C. The serum-starved cells were then preincubated
for 30 min in the presence of CHX (20 μg/ml) before incubation with
EGF (100 ng/ml) at 37°C for the indicated times. The cells were
then washed with ice-cold-PBS and lysed, followed by SDS-PAGE and
western blot analysis. Bafilomycin A1 (0.17 μM) was added when the
cells were incubated with RPMI.
Statistical analysis
Data are expressed as mean ± SD unless otherwise
noted. Significance (P<0.05) was determined by using Student’s
t-test, since all data met the assumptions for parametric
statistical analysis.
Results
Silencing of SNX1 by specific siRNA
effectively downregulates the expression of endogenous SNX1 protein
levels, and increases expression of EGFR mRNA in the 3 NSCLC cell
lines
To investigate the biological function of SNX1 in
regulating EGFR endocytosis, we used siRNA to knock down the
endogenous of SNX1 in 3 NSCLC cell lines, namely, PC9, QG56 and
A549. Firstly, we examined the depletion of endogenous SNX1 protein
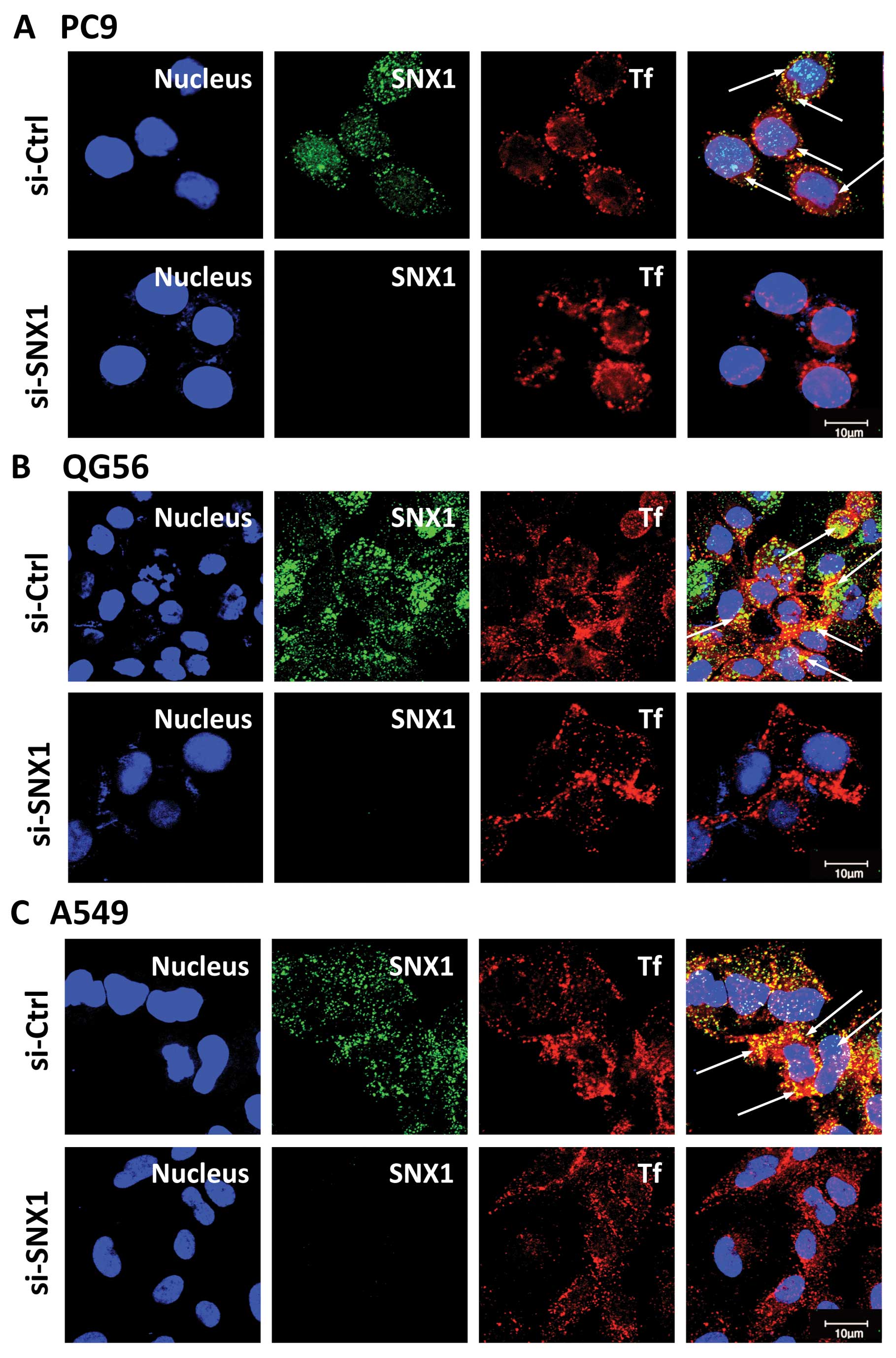
levels by confocal immunofluorescence microscopy. As shown in
Fig. 1, the expression of
endogenous SNX1 was successfully depleted using siRNA in the 3
NSCLC cell lines. Therefore, we used these cells to analyze the
internalization fate of Texas red-EGF or pEGFR over time, as
described in Materials and methods.
Next, we treated the 3 NSCLC cell lines with EGF for
different time periods, and then used qRT-PCR analysis to examine
the endogenous SNX1 mRNA transcript levels. As shown in Fig. 2A, the results revealed that
siRNA-SNX1 reduced SNX1 mRNA in all 3 NSCLC cell lines. Moreover,
it should be noted that a large amount of SNX1 transcript was
observed in the gefitinib-resistant cell lines, QG56 and A549,
compared to the gefitinib-sensitive cell line PC9 (Fig. 2A).
In addition, we found that SNX1 knockdown increased
endogenous expression of EGFR transcript in the 3 NSCLC cell lines
(Fig. 2B). We also found that
endogenous expression of EGFR mRNA transcripts was considerably
higher in the gefitinib-resistant cell lines QG56 and A549,
compared to the gefitinib-sensitive cell line PC9: the expression
level of EGFR transcript in the gefitinib-resistant cell lines QG56
and A549 cells was approximately 6.2-fold and 3.4-fold,
respectively, of the values in the gefitinib-sensitive PC9 cells
(Fig. 2B). These results imply
that SNX1 is involved in the negative regulation of EGFR mRNA
expression in these human lung cancer cells.
Silencing of SNX1 stimulates an efficient
endocytosis of ligand-induced EGFR and phosphorylated EGFR via the
early/late endocytic pathway in gefitinib-resistant NSCLC cell
lines
We recently demonstrated novel evidence that
gefitinib-sensitive cells show efficient endocytosis of EGFR. In
contrast, gefitinib-resistant cells show internalized EGFR
accumulation in the aggregated early endosomes, and this is
associated with SNX1 (13,14). We, therefore, suggest that
impairment of protein function, such as SNX1 in the regulation of
EGFR trafficking in the early endocytic pathway, might cause these
perturbations in EGFR endocytosis, leading to gefitinib-resistance
in NSCLC cell lines.
To further examine the effect of SNX1 silencing on
ligand-induced EGFR via the early/late endocytic pathway, we
studied the fate of internalized Texas red-EGF in early endosomes
or late endosomes/lysosomes in 3 NSCLC cell lines. The
gefitinib-sensitive PC9 cells or gefitinib-resistant QG56 and A549
cells transfected with siRNA-control or siRNA-SNX1 were incubated
with Texas Red-EGF for different time periods and the distribution
of internalized Texas red-EGF was then assessed by confocal
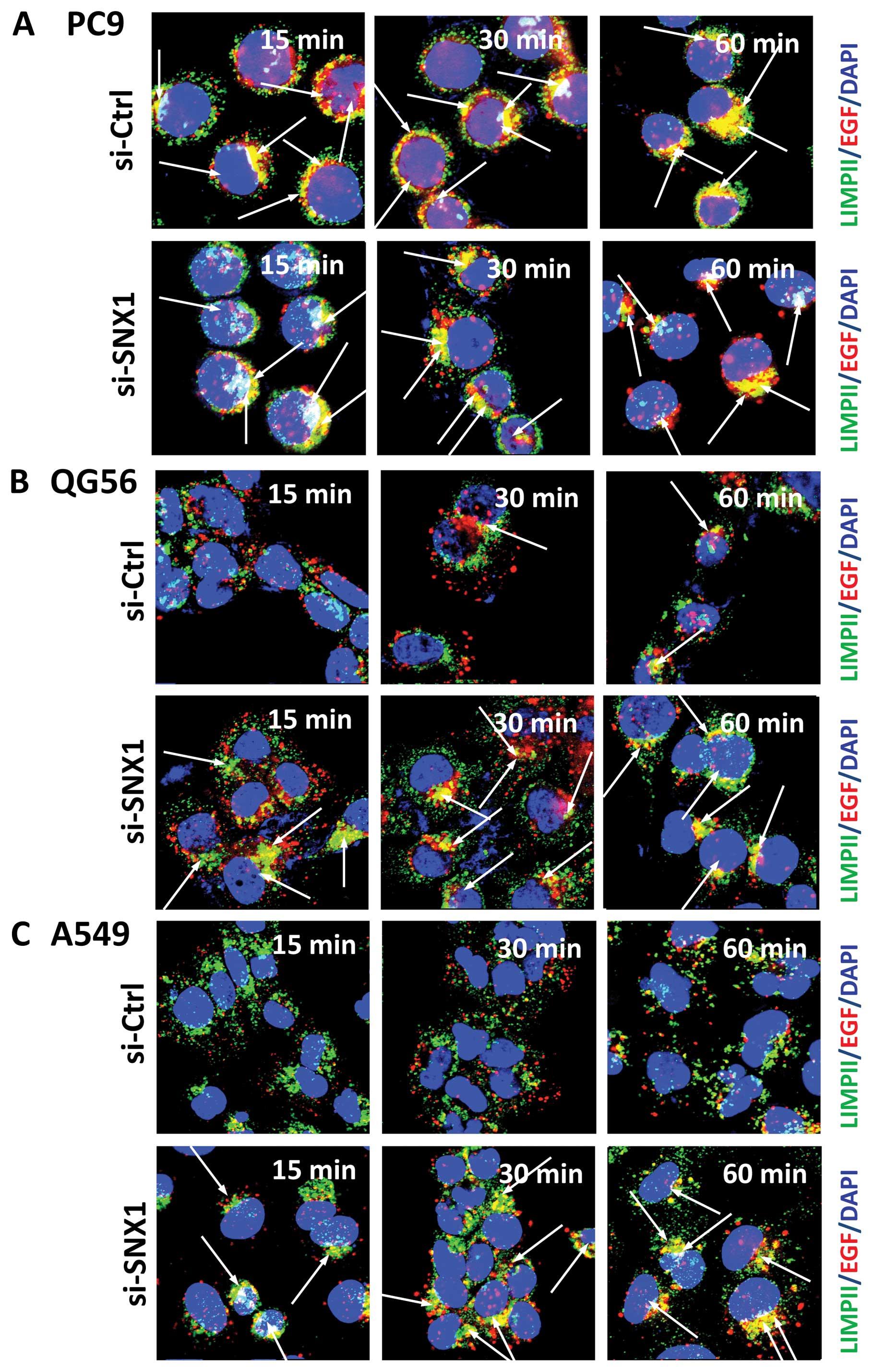
immunofluorescence microscopy. In gefitinib-sensitive PC9 cells
transfected with siRNA-SNX1, a rapid endocytosis of Texas red-EGF
was observed, and the distribution of small punctate vesicles that
stained positive for internalized EGF overlapped with the
LIMPII-positive late endosomes/lysosomes in the perinuclear region
(Fig. 3A). Moreover, a
considerable amount of the internalized Texas red-EGF co-localized
with the LIMPII-positive late endosomes/lysosomes after 30 min.
These observations are consistent with those for PC9 cells
transfected with siRNA-control, therefore, suggesting that
silencing of SNX1 does not have any inhibitory effect on EGFR
endocytosis.
On the other hand, in the gefitinib-resistant QG56
and A549 cells transfected with siRNA-control, the internalization
of Texas red-EGF was suppressed and the endocytosed Texas
red-EGF-positive staining was not co-localized with LIMPII-positive
vesicular structures even after 30 min internalization (Fig. 3B and C). Therefore, the
transfection of siRNA-control did not change the suppressive
internalization of the ligand-induced EGFR in the
gefitinib-resistant QG56 and A549 cell lines. In contrast, in the
gefitinib-resistant QG56 and A549 cells transfected with
siRNA-SNX1, we found an increase in the co-localization of Texas
red-EGF and LIMPII at 15 and 30 min after EGF stimulation.
Quantitative analysis was carried out to determine
the amounts of LIMPII-positive late endosomes/lysosomes marker
co-localized with the endocytosed Texas red-EGF after 30-min
internalization in PC9, QG56 and A549 cells transfected with
siRNA-control or siRNA-SNX1 (Fig.
3D). Our data confirm that silencing of SNX1 by siRNA rescues a
rapid endocytosis and trafficking of EGFR via the early/late
endocytic pathway in gefitinib-resistant cells. In contrast, an
aberration of EGFR endocytosis was noted through the early to late
endocytic pathway in the gefitinib-resistant cells transfected with
siRNA-control. These results demonstrate that depletion of SNX1 by
siRNA considerably stimulates ligand-induced EGFR endocytosis in
the gefitinib-resistant QG56 and A549 cells.
To further substantiate the effect of SNX1 silencing
on the ligand-induced phosphorylation of EGFR and the endocytosis
of pEGFR, the 3 NSCLC cell lines transfected with siRNA-control or
siRNA-SNX1 were stimulated with EGF for 60 min, and then each cell
type was double-stained for pEGFR and the endocytosed transferrin
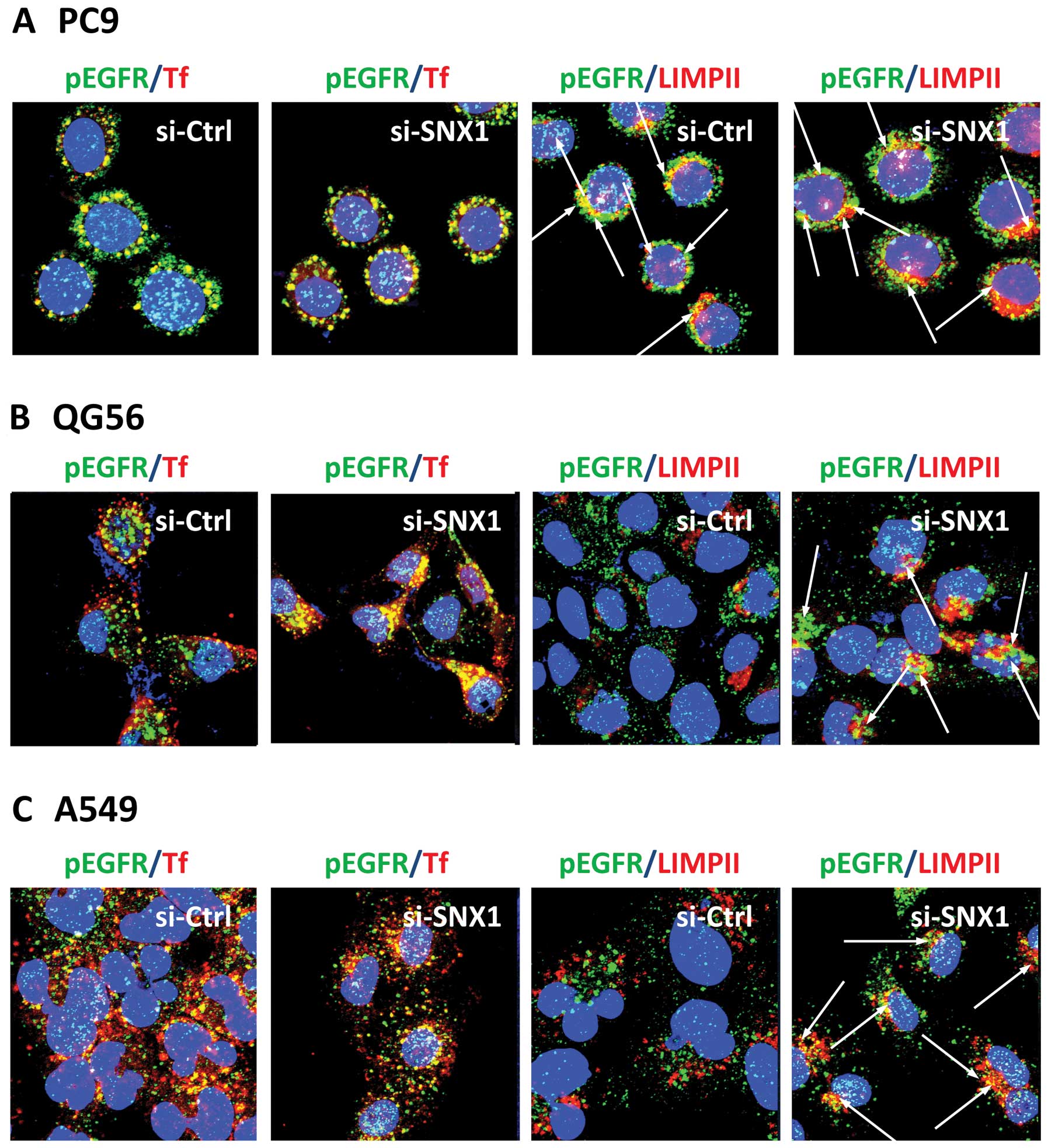
or for pEGFR and LIMPII. In the gefitinib-resistant cell lines QG56
and A549 transfected with siRNA-control, pEGFR remained
predominantly associated with the transferrin-positive early
endosomes, but no colocalization of pEGFR with LIMPII-positive
vesicles was seen in the cells (Fig.
4B and C). In contrast, we found a significant increase in
co-localized pEGFR and LIMPII in the cells transfected with
siRNA-SNX1 (Fig. 4B and C). These
results confirm that silencing of SNX1 stimulates ligand-induced
pEGFR endocytosis via the early/late endocytic pathway in a
gefitinib-resistant NSCLC cell line. On the other hand, in the
gefitinib-sensitive PC9 cells transfected with siRNA-control or
siRNA-SNX1, we observed an efficinet pEGFR endocytosis from early
endosomes to late endosomes. In addition, EGFR co-localized with
the LIMPII-positive vesicles as well as the transferrin-positive
early endosomes after EGF stimulation (Fig. 4A). These results indicate that
depletion of SNX1 by siRNA does not have any inhibitory effect on
ligand-induced pEGFR endocytosis in gefitinib-sensitive PC9 cells.
Quantitative analysis of LIMPII-positive late endosome/lysosome
marker (Fig. 4D) that co-localized
with the endocytosed pEGFR after 60 min of internalization further
confirmed the stimulating effect of SNX1 silencing on the
ligand-induced endocytic trafficking of pEGFR from early endosomes
to late endosomes in the gefitinib-resistant NSCLC cells.
Depletion of SNX1 stimulates the
ligand-induced degradation of EGFR and increases EGFR
phosphorylation in gefitinib-resistant NSCLC cell line
To analyze the EGF-stimulated degradation of EGFR or
pEGFR in the gefitinib-sensitive cell line PC9, the cells were
stimulated with EGF at 37°C for the indicated times, and then
analyzed by western blotting. The results revealed that the
degradation of EGFR proceeded efficiently in the PC9 cells
(Fig. 5A), and the amounts of EGFR
or pEGFR decreased by about 80% after 120 min (Fig. 5B and C). Further, we tested if the
lysosomal protease inhibitor bafilomycin A1 prevents the
EGF-induced accelerated degradation of EGFR and pEGFR in PC9 cells.
Cells were pretreated with bafilomycin A1 for 30 min, and then
stimulated with EGF for various periods, and the lysates were
analyzed by western blotting. The results showed that bafilomycin
A1 treatment completely blocked EGF-induced degradation of EGFR and
pEGFR, as indicated by the expression of these molecules remaining
in the cells following incubation (Fig. 5). However, no inhibitory effect was
seen when the cells were treated in the presence of MG132, a
proteasomal inhibitor (data not shown). The strong inhibition of
bafilomycin A1 on EGF-stimulated EGFR degradation was consistent
with previously reported data (29).
Quantitative analysis showed that EGFR and pEGFR
were degraded by more than 70% within 1 h of EGF stimulation, and
they gradually disappeared from the cells in the 120 min following
stimulation (Fig. 5B and C);
however, in the cells treated with bafilomycin A1, more than 94% of
the EGFR or pEGFR remained in the cells after 120 min of EGF
stimulation. Therefore, these results indicate that intracellular
degradation of EGFR or pEGFR proceeds efficiently via an
endosomal/lysosomal pathway in PC9 cells.
Next, to further examine the effect of the silencing
of SNX1 protein levels on the EGF-induced degradation of EGFR or
pEGFR in the gefitinib-resistant A549 cells, the cells transfected
with siRNA-control or siRNA-SNX1 were stimulated with EGF at 37°C
for the indicated times, and then, the cell lysates were analyzed
by western blotting. As shown in Fig.
6A and B, we found an accelerated EGF-dependent degradation of
EGFR in the siRNA-SNX1-transfected A549 cells, with degradation of
approximately 72% at 60 min and 85% at 120 min after EGF
stimulation. In contrast, the EGFR degradation was about 45% at 60
min and 65% at 120 min after EGF stimulation in the
siRNA-control-transfected cells. These results indicate that the
depletion of SNX1 by siRNA stimulates the EGF-dependent EGFR
downregulation via the early/late endocytic pathway in the
gefitinib-resistant cell line.
Furthermore, we found novel evidence that the
expression level of pEGFR was dramatically increased at 60 and 120
min after EGF stimulation in the siRNA-SNX1-transfected A549 cells
compared to the increase seen in siRNA-control-transfected A549
cells (Fig. 6A and C), and that
the induced expression level of pEGFR protein following EGF
stimulation was about 33-fold and 9.4-fold at 60 and 120 min,
respectively (Fig. 6A and C). In
contrast, in the siRNA-control-transfected A549 cells, the increase
of the pEGFR expression was only 7.5-fold and 4.3-fold at 60 and
120 min, respectively (Fig. 6A and
C). The increased pEGFR appeared to be rapidly degraded within
120 min in both the siRNA-control- and siRNA-SNX1-transfected cells
(Fig. 6C and D), indicating that
pEGFR is efficiently trafficked to the lysosomal degradation
pathway. It should be noted that the expression levels of pEGFR in
the siRNA-SNX1-transfected cells was 13.8-fold (at 60 min) and
6.7-fold (at 120 min) higher than that in the
siRNA-control-transfected cells after EGF stimulation (Fig. 6E). These results indicate that
depletion of SNX1 by siRNA efficiently increases endogenous
phosphorylation of EGFR via the endocytic pathway, implying that
SNX1 plays a suppressive role in the phosphorylation and
downregulation of EGFR via endocytic pathway in human lung cancer
cells.
Discussion
In the present study, we demonstrate for the first
time the intracellular regulatory function of SNX1 with regard to
EGF-induced endocytosis and downregulation of EGFR/pEGFR using
RNAi-mediated knockdown approaches via the early/late endocytic
pathway in gefitinib-sensitive and gefitinib-resistant NSCLC cell
lines. Using confocal immunofluorescence microscopy and qRT-PCR, we
verified a considerable reduction of endogenous SNX1 protein
expression and mRNA expression, respectively, in all 3 NSCLC cell
lines transfected with siRNA-SNX1. Moreover, we found that
endogenous expression of EGFR transcripts in the
gefitinib-resistant cell lines QG56 and A549 was significantly
higher than in the gefitinib-sensitive cell line PC9, and knockdown
of SNX1 in all 3 NSCLC cell lines considerably increased the
expression of the EGFR transcript. From these findings, we propose
that the expression of SNX1 protein might play a negative role in
the regulation of EGFR mRNA expression in these NSCLC cell
lines.
Most importantly, we provided evidence by using
confocal immunofluorescence microscopy that knockdown of endogenous
SNX1 by siRNA-SNX1 induced efficient endocytosis of ligand-induced
EGFR or pEGFR via the early/late endocytic pathway in the
gefitinib-resistant NSCLC cell lines QG56 and A549. We showed that
increased co-localization of Texas red-EGF and LIMPII after 15 min
internalization, while in the siRNA-control-tranfected
gefitinib-resistant cells, the internalization of EGFR was
suppressed and the endocytosed Texas red-EGF-positive staining did
not overlap with late endosome-positive vesicular structures even
after 30 min stimulation. These data suggest that knockdown of SNX1
stimulates ligand-induced EGFR endocytosis in the
gefitinib-resistant cells.
Inhibition of EGFR/pEGFR degradation in PC9 cells
was verified using bafilomycin A1, a lysosomal inhibitor. We found
that bafilomycin A1 treatment in PC9 cells completely blocked the
efficient EGF-induced degradation of EGFR/pEGFR, confirming that
the EGF-induced EGFR or pEGFR is trafficked to late
endosomes/lysosomes where extensive degradation for these proteins
takes place in the PC9 cells. We further showed, by using western
blot analysis, that depletion of SNX1 considerably stimulates the
ligand-induced downregulation of EGFR in the gefitinib-resistant
A549 cells. This result indicates that silencing of SNX1 stimulates
EGFR traffic out of early endosomes for targeting lysosomal
degradation pathway. Accordingly, we postulate that SNX1 might be a
negative regulator of ligand-induced EGFR endocytosis, followed by
downregulation via the early/late endocytic pathway.
It was also interesting to note that silencing of
SNX1 induced a dramatic increase in the expression of pEGFR in the
siRNA-SNX1-transfected A549 cells. Further, the observed increase
of pEGFR at 60 min in the siRNA-SNX1-transfected cells was
approximately 14-fold higher than that in the
siRNA-control-transfected cells. This marked increase in pEGFR
expression was not seen in the siRNA-control-transfected cells.
These results indicate that depletion of SNX1 protein by siRNA
considerably increases endogenous phosphorylation of EGFR via the
endocytic pathway. It is known that following EGF stimulation, the
phosphorylated EGFR is internalized by rapid clathrin-mediated
endocytosis, and the internalized EGFR is then sorted via the
endosomal-sorting complex required for transport (ESCRT)-dependent
pathway for targeting degradation or recycling pathway (30). In this context, our present
results, demonstrating the suppressive role of SNX1 on the
phosphorylation of EGFR via the endocytic pathway, suggest a
critical function for SNX1 in the maintenance of tightly regulated
EGFR-mediated signaling.
We recently reported the novel observation that in
the gefitinib-resistant NSCLC cell line, early endosomes labeled
with endocytosed Texas red-transferrin formed aggregated vesicular
structures distributed in the perinuclear region, and that
considerable amounts of cytosolic SNX1 were distributed in these
aggregated early endosomal vesicles (13,14).
Conversely, no such aggregation of SNX1-positive early endosomes
was observed in the gefitinib-sensitive NSCLC cell line. On the
basis of these data, we postulated that membrane trafficking of
pEGFR from early endosomes to late endosomes might be significantly
impaired in the gefitinib-resistant NSCLC A549 and QG56 cells.
Therefore, we assume that abrogation of certain SNX1 trafficking
machinery could cause this perturbation of EGFR endocytosis, which
might lead to the acquisition of gefitinib-resistance in NSCLC cell
lines.
It was originally reported that SNX1 interacts with
EGFR, and the overexpression of SNX1 enhanced EGF-dependent EGFR
degradation in lysosomes, thereby demonstrating that SNX1 is a
likely mediator in the intracellular sorting of EGFR for targeting
the lysosomes for degradation (15). In this context, our findings
regarding the suppressive role of SNX1 appear to be in contrast to
previous findings (15,20). Further studies to investigate the
role of SNX1 on EGFR activation and signaling in NSCLC cell lines
will be required.
Abbreviations:
|
EGFR
|
epidermal growth factor receptor
|
|
pEGFR
|
phosphorylated epidermal growth factor
receptor
|
|
SNX1
|
sorting nexin 1
|
|
NSCLC
|
non-small cell lung cancer
|
|
LIMPII
|
lysosomal integral membrane protein
II
|
Acknowledgements
This work was supported by JSPS
KAKENHI Grant numbers 23390372, 23659734 and 23592202.
References
|
1
|
Ullrich A and Schlessinger J: Signal
transduction by receptors with tyrosine kinase activity. Cell.
61:203–212. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Carpenter G: The EGF receptor: a nexus for
trafficking and signaling. Bioessays. 22:697–707. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Yarden Y: The EGFR family and its ligands
in human cancer signaling mechanisms and therapeutic opportunities.
Eur J Cancer. 37:3–8. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Schlessinger J: Common and distinct
elements in cellular signaling via EGF and FGF receptors. Science.
306:1506–1507. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Mendelsohn J and Baserga J: The EGF
receptor family as targets for cancer therapy. Oncogene.
19:6550–6565. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
De Bono JS and Rowinsky EK: The ErbB
receptor family: a therapeutic target for cancer. Trends Mol Med.
8:19–26. 2002.PubMed/NCBI
|
|
7
|
Woodburn JR: The epidermal growth factor
receptor and its inhibition in cancer therapy. Pharmacol Ther.
82:241–250. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Baselga J and Averbuch SD: ZD1839
(‘Iressa’) as an anticancer agent. Drugs. 60(Suppl 1): S33–S42.
2000.
|
|
9
|
Arteaga CL and Johnson DH: Tyrosine kinase
inhibitors-ZD1839 (Iressa). Curr Opin Oncol. 13:491–498. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Barker AJ, Gibson KH, Grundy W, Godfrey
AA, Barlow JJ, Healy MP, Woodburn JR, Ashton SE, Curry BJ, Scarlett
L, Henthorn L and Richards L: Studies leading to the identification
of ZD1839 (IRESSA): an orally active, selective epidermal growth
factor receptor tyrosine kinase inhibitor targeted to the treatment
of cancer. Bioorg Med Chem Lett. 11:1911–1914. 2001. View Article : Google Scholar
|
|
11
|
Ono M, Hirata A, Kometani T, Miyagawa M,
Ueda S, Kinoshita H, Fujii T and Kuwano M: Sensitivity to gefitinib
(Iressa, ZD1839) in non-small cell lung cancer cell lines
correlates with dependence on the EGF receptor/extracellular
signal-regulated kinase 1/2 and EGF receptor/Akt pathway for
proliferation. Mol Cancer Ther. 3:465–472. 2004.
|
|
12
|
Nishimura Y, Bereczky B and Ono M: The
EGFR inhibitor gefitinib suppresses ligand-stimulated endocytosis
of EGFR via the early/late endocytic pathway in non-small cell lung
cancer cell lines. Histochem Cell Biol. 127:541–553. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Nishimura Y, Yoshioka K, Bereczky B and
Itoh K: Evidence for efficient phosphorylation of EGFR and rapid
endocytosis of phosphorylated EGFR via the early/late endocytic
pathway in a gefitinib-sensitive non-small cell lung cancer cell
line. Mol Cancer. 7:1–13. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Nishimura Y, Yoshioka K, Takiguchi S,
Bereczky B, Nakabeppu Y and Itoh K: A role for SNX1 in the
regulation of EGF-dependent phosphorylated EGFR endocytosis via the
early/late endocytic pathway in a gefitinib-sensitive human lung
cancer cells. Curr Signal Transduct Ther. 6:383–395. 2011.
View Article : Google Scholar
|
|
15
|
Kurten RC, Cadena DL and Gill GN: Enhanced
degradation of EGF receptors by a sorting nexin, SNX1. Science.
272:1008–1010. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Worby CA and Dixon JE: Sorting out the
cellular function of sorting nexins. Nat Rev Mol Cell Biol.
3:919–931. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
17
|
Horazdovsky BF, Davies BA, Seaman MN,
McLaughlin SA, Yoon S and Emr SD: A sorting nexin-1 homologue,
Vps5p, forms a complex with Vps17p and is required for recycling
the vacuolar protein-sorting receptor. Mol Biol Cell. 8:1529–1541.
1997. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Nothwehr SF and Hindes AE: The yeast
VPS5/GRD2 gene encodes a sorting nexin-1-like protein required for
localizing membrane proteins to the late Golgi. J Cell Sci.
110:1063–1072. 1997.PubMed/NCBI
|
|
19
|
Seaman MN, McCaffery JM and Emr SD: A
membrane coat complex essential for endosome-to-Golgi retrograde
transport in yeast. J Cell Biol. 142:665–681. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhong Q, Lasar CS, Tronchere H, Sato T,
Meerloo T, Yeo M, Songyang Z, Emr SD and Gill GN: Endosomal
localization and function of sorting nexin 1. Proc Natl Acad Sci
USA. 99:6767–6772. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Carlton J, Bujny M, Peter BJ, Oorschot VM,
Rutherford A, Mellor H, Klumperman J, McMahon HT and Cullen PJ:
Sorting nexin-1 mediates tubular endosome-to-TGN transport through
coincidence sensing of high-curvature membranes and
3-phosphoinositides. Curr Biol. 14:1791–1800. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Gullapalli A, Garrett TA, Paing MM,
Griffin CT, Yang Y and Trejo J: A role for sorting nexin 2 in
epidermal growth factor receptor down-regulation: evidence for
distinct functions of sorting nexin 1 and 2 in protein trafficking.
Mol Biol Cell. 15:2143–2155. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Okazaki I, Himeno M, Ezaki J, Ishikawa T
and Kato K: Purification and characterization of an 85 kDa
sialoglycoprotein in rat liver. J Biochem. 111:763–769.
1992.PubMed/NCBI
|
|
24
|
Nishimura Y, Yoshioka K, Bernard O, Himeno
M and Itoh K: LIM kinase 1: evidence for a role in the regulation
of intracellular vesicle trafficking of lysosomes and endosomes in
human breast cancer cells. Eur J Cell Biol. 34:189–213.
2004.PubMed/NCBI
|
|
25
|
Nishimura Y, Yoshioka K, Bernard O,
Bereczky B and Itoh K: A role of LIM kinase 1/cofilin pathway in
regulating endocytic trafficking of EGF receptor in human breast
cancer cells. Histochem Cell Biol. 126:627–638. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Nishimura Y, Bereczky B, Yoshioka K,
Taniguchi S and Itoh K: A novel role of Rho-kinase in the
regulation of ligand-induced phosphorylated EGFR endocytosis via
the early/late endocytic pathway in human fibrosarcoma cells. J Mol
Histol. 42:427–442. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kornfeld S and Mellman I: The biogenesis
of lysosomes. Ann Rev Cell Biol. 5:483–525. 1989. View Article : Google Scholar
|
|
28
|
Sandoval IV, Arredondo JJ, Alcalde J,
Gonzalez-Noriega A, Vandekerckhove J, Jimenez MA and Rico M: The
residues Leu (Ile) 475-Ile (Leu) 476, contained in the extended
carboxyl cytoplasmic tail, are critical for targeting of the
resident lysosomal membrane protein LIMPII to lysosomes. J Biol
Chem. 269:6622–6631. 1994.
|
|
29
|
Skarpen E, Johannessen LE, Bjerk K,
Fasteng H, Guren TK, Lindeman B, Thoresen GH, Christoffersen T,
Stang E, Huitfeldt HS and Madshus IH: Endocytosed epidermal growth
Factor (EGF) receptors contribute to the EGF-mediated growth arrest
in A431 cells by inducing a sustained increase in p21/CIP1. Exp
Cell Res. 243:161–172. 1998. View Article : Google Scholar
|
|
30
|
Sorkin A and von Zastrow M: Endocytosis
and signalling: intertwining molecular networks. Nat Rev Mol Cell
Biol. 10:609–622. 2009. View
Article : Google Scholar : PubMed/NCBI
|