Introduction
Hepatocellular carcinoma (HCC), as one of the most
common malignancies worldwide (the fifth most frequent neoplastic
disease in men and seventh in women), represents a global health
concern. HCC accounts for >80% of all primary liver cancers,
with a strong male predominance (2–4 times more frequent in men
compared with women). Globally, HCC accounts for 4.6% of all
cancers and has a mortality rate of 94%, leading to approximately
one million deaths annually. Over the last 4 decades, the incidence
of HCC has been on the increase in the developed world. For
example, in the United States, the incidence has doubled since the
1970s and the mortality rate from HCC has increased by 41% over
this time period (1–6).
The vast majority of the burden of HCC is
concentrated in the developing world, accounting for 84% of the
total worldwide incidence and 83% of total deaths. HCC follows a
rather distinct geographic pattern and >80% of HCC cases
worldwide occur in sub-Saharan Africa and Eastern Asia, with
incidence rates of >20/100,000. Southern European countries,
including Spain, Italy and Greece, report incidence levels of
10.0–20.0/100,000, whereas North America, South America, Northern
Europe and Oceania report incidence levels of <5.0/100,000
individuals. This distribution and incidence disparity is very
similar to the global distribution pattern of hepatitis B virus
(HBV) and hepatitis C virus (HCV) infection: The HCC rate is
clearly highest in regions endemic for HBV and HCV. Even variations
in the age-, gender- and race-specific rates of HCC in various
geographical regions are suggested to be associated with the
different prevalence rates of hepatitis viruses in these regions.
For example, HCC is rarely encountered before the age of 40 years,
with the exception of the regions where HBV infection is
hyperendemic. In rather higher-risk populations, such as the
Chinese, the mean age range for a diagnosis of HCC is 55–59 years,
but it is 63–65 years in Europe and North America (7,8). Turkey is
considered to be an HBV-endemic country, with a carrier rate of
5–10%, whereas the incidence rate is 1.5% for HCV. The Turkish
Ministry of Health has reported the incidence of HCC to be
0.83/100,000 in 2003, which has remained approximately the same
between 2000 and 2003. According to the Turkiye Hepatitis
Prevalence's 2010 data, the rate of HCV carriage is 0.95% in Turkey
(9–11).
Despite the advances in the understanding of the
molecular pathogenesis of HCC, as well as its associated diagnostic
techniques and novel therapies, including targeted therapies, HCC
remains a dismal diagnosis. Unresectable HCC is an aggressive
neoplasm; in the case of intermediate disease, the median survival
is 16–20 months, whereas it is only 6 months for advanced-stage
untreated patients. Several factors, including high tumor
multiplicity rate, extent of vascular invasion and coexistent
cirrhosis, are HCC characteristics that contribute to these
unsatisfactory outcomes. Furthermore, the late detection of HCC is
a factor contributing to the poor outcome of HCC patients, as over
two-thirds of patients are diagnosed at advanced stages of the
disease. However, in the subpopulation of patients who are
diagnosed at an early disease stage and who receive potentially
curative treatment, such as liver transplantation, surgical
resection and tumor ablation, a 5-year survival rate of 40–70% may
be expected (1–6).
Similar to the majority of other cancers, HCC
results from a combination of environmental factors, such as
infection with HBV, and specific genetic alterations, although the
precise molecular pathogenesis of HCC remains unknown. Ras proteins
are a family of small guanosine triphosphate-regulated molecular
switches, which convey signals from the cell membrane to the
nucleus, and activate several molecular pathways involved in
proliferation, transformation and malignant progression. The Ras
family of proteins includes H-Ras, N-Ras and K-Ras (12–14). The
transcription factor Fos is the main nuclear target of the
Ras/Raf/mitogen-activated protein kinase kinase
(MEK)/mitogen-activated protein kinase (MAPK) pathway. Fos forms a
heterodimer with Jun, which yields the active AP1 complex that is
involved in tumor proliferation (13).
A number of single-point mutations of the Ras gene
have led to the constitutive activation of a Ras protein with
impaired GTPase activity, which results in the continuous
stimulation of cellular proliferation. The frequency of these gene
mutations is tumor type-dependent: They are occasionally observed
in breast, ovarian, esophageal, prostate, and gastric cancers, but
they are ubiquitous in pancreatic adenocarcinomas and present in
half of colon and thyroid cancers. In total, ~30% of all human
tumors harbor a Ras mutation, which occurs most often in the K-Ras
gene (13). K-Ras mutations have been
identified in 7% of human liver cancers, whereas H-Ras and N-Ras
mutations are also observed at lower rates. The activation of Ras
signaling has been universally observed in human HCC samples, and
Ras activation has been shown to lead to hepatocellular
proliferation and transformation (12,13).
The epidermal growth factor receptors (EGFRs) are
reported to be frequently expressed in human HCC, most likely
contributing to hepatocellular carcinogenesis and tumor
aggressiveness. Thus, targeting the EGFR function has been
suggested as a promising strategy in the treatment of advanced HCC.
Cetuximab is a monoclonal antibody against EGFR and is approved for
the treatment of K-Ras wild-type advanced colorectal and head and
neck cancer patients. Cetuximab has also been used in the treatment
of advanced HCC patients, with promising although conflicting
results. As it has been suggested by several studies on the role of
K-Ras mutations in the prediction of response to cetuximab in colon
cancer, it is possible that the K-Ras mutation status may also
predict the response to cetuximab in HCC patients (2,3,5,6,15).
Considering the significance of the prediction of
response to therapy in HCC patients who are considered for
cetuximab therapy, we designed the present pilot study to estimate
the K-Ras mutation rate in Turkish HCC patients.
Patients and methods
Patients and sampling
A total of 73 HCC patients from 6 comprehensive
cancer centers in 4 metropolitan cities of Turkey were enrolled in
this study. This study was approved by the Ethics Committee of
Marmara University, Faculty of Medicine (Istanbul, Turkey). Each
center committed to participate with approximately 10 random
patients, whose formalin-fixed paraffin-embedded (FFPE) tumor
tissues were available for K-Ras genotyping. Hematoxylin and
eosin-stained slides were prepared from FFPE tumor tissue blocks
and re-evaluated by an expert pathologist (B.S.) for confirmation
of the diagnosis and evaluation of the adequacy of the tumor cell
content of the blocks for further molecular analysis. A total of 58
patients whose FFPE tumor tissue blocks were found to be suitable
for molecular analysis were included in this study.
K-Ras mutation analysis
The most important K-Ras mutations in exon 2 were
assessed. FFPE tumor tissues of patients were used for K-Ras
genotyping. Genomic tumor DNA was extracted from 6-µm unstained
FFPE sections. Tumor tissue was manually scraped from
deparaffinized unstained slides into microcentrifuge tubes.
Following manual microdissection of tumor tissue under a light
microscope, DNA was isolated with a QIAamp® DNA FFPE Tissue kit
(Qiagen Inc., Valencia, CA, USA) according to the manufacturer's
instructions. Appropriate controls were included in each assay. Two
methods were applied based on the availability of adequate amounts
of tumor DNA. In the first method, the samples were processed using
TheraScreen® KRAS RGQ PCR kit (catalogue no. 870001; Qiagen
Manchester, Ltd., Manchester, UK) according to the manufacturer's
instructions. The genomic DNA was further used to detect 7 somatic
mutations (35G>A; 35G>C; 35G>T; 34G>A; 34G>C;
34G>T and 38G>A) in codons 12 and 13 in exon 2 of the K-Ras
oncogene by quantitative polymerase chain reaction (PCR) using a
LightCycler® 480 II instrument (Roche, Penzberg, Germany) with both
Scorpions and Amplification Refractory Mutation System
Technologies. In the second method, the genomic DNA was amplified
by PCR using primers specific for K-Ras exon 2 with GML® SeqFinder
Sequencing System's KRAS kit (catalogue no. 106025.v.3.0; GML
Corporation, Wallerau, Switzerland). The PCR conditions were as
follows: 10 sec at 95°C, 40 cycles of 30 sec at 95°C, 1 min at
59°C, 1 min at 72°C and 7 min at 72°C using GeneAmp® PCR System
9700 (Applied Biosystems, Foster City, CA, USA). The amplified PCR
products were then cleaned up using ExoSAP-IT® (GML Corporation).
Sequencing of the amplified fragments was performed using a
sequencing PCR reaction consisting of 25 cycles of 10 sec at 94°C,
5 sec at 50°C and 4 min at 60°C using the BigDye® Terminator v3.1
Cycle Sequencing kit (catalogue no. 4336917; Applied Biosystems)
according to the manufacturer's instructions. These samples were
processed using the ABI 310 Automated Sequencer (Applied
Biosystems). The identified DNA sequence alterations were confirmed
by sequencing both DNA strands in two independent experiments with
forward and reverse primers.
Results
Geographic distribution of enrolled
patients
In total, 58 patients were randomly selected from 6
comprehensive cancer centers in Turkey. Tumor samples from the
following cancer centers were found to be suitable for mutation
analysis: Marmara University, Kartal Training and Research
Hospital, Ege University and Bilim University.
Prevalence of K-Ras mutations in HCC
patients
The total prevalence of the investigated K-Ras
mutations based on the different detection methods is presented in
Table I. Of the 58 samples initially
retrieved for this study, 40 had adequate tumor tissue for mutation
analysis. The majority of these samples (33/40; 82.5%) had no
mutations at exon 2 (wild-type). The observed mutations were only
detected in the samples processed via the direct sequencing
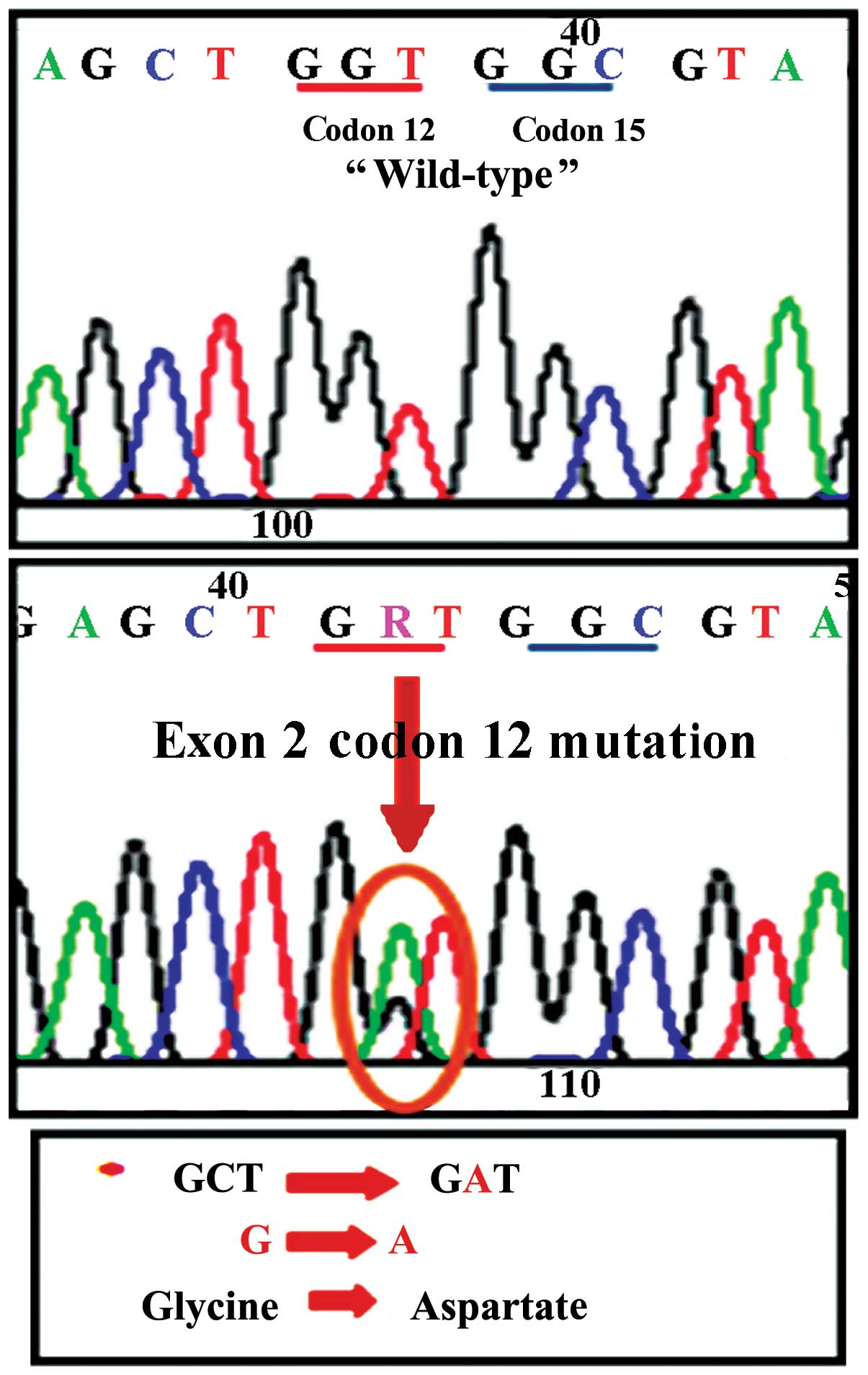
technique. A representative image of a mutation detected by the
direct sequencing analysis (replacement of glycine by aspartate in
codon 12 of exon 2 of the K-Ras gene) is shown in Fig. 1. The distribution of the detected
mutations based on the contributing centers is described in
Table II.
 | Table I.Distribution of K-Ras mutant and
wild-type tumors based on the mutation analysis method. |
Table I.
Distribution of K-Ras mutant and
wild-type tumors based on the mutation analysis method.
| Methods | Wild-type | Mutant | Total |
|---|
| TheraScreen | 22 | 0 | 22 |
| Direct
sequencing | 11 | 7 | 18 |
 | Table II.Distribution of the K-Ras wild-type
and mutant tumor samples based on the clinical sites of the
study. |
Table II.
Distribution of the K-Ras wild-type
and mutant tumor samples based on the clinical sites of the
study.
| Center | No. | TheraScreen exon 2
wild-type | TheraScreen exon 2
mutated | Direct sequencing
exon 2 wild-type | Direct sequencing
exon 2 mutated | Inadequate
tissue | Wild-type, % |
|---|
| Marmara
University | 7 | 5 | - | 1 | - | 1 | 85 |
| Kartal Training and
Research Hospital | 13 | 8 | - | 1 | 2 | 2 | 69 |
| Ege University | 13 | 5 | - | 6 | 2 | - | 84 |
| Bilim University | 10 | 4 | - | 3 | 3 | - | 70 |
| Total | 43 | 22 | - | 11 | 7 | 3 | 77 |
In our study, TheraScreen did not identify any
mutations, while direct sequencing identified 7 mutations in the
investigated patient population. Due to technical considerations
(the starting amount of DNA), we were unable to perform direct
sequencing on all samples. A number of previous studies
demonstrated the high sensitivity of TheraScreen in identifying
K-Ras mutations (approaching 100%) (16–18); thus,
we considered that the results of two mutation analysis methods are
valid and may be pooled together.
Discussion
HCC is a difficult disease to treat, particularly
when presenting at an advanced or non-resectable stage.
Chemotherapy with traditional cytotoxic agents (doxorubicin,
cisplatin, or 5-fluorouracil) is associated with a low response
rate (<10%) and offers little overall survival benefit. The most
common comorbidities, particularly cirrhosis and liver failure,
lead to poor tolerance of chemotherapy in these patients, as well
as an unpredictable clinical course, considering that these drugs
are mainly metabolized by the liver. Other potential treatment
options, such as interferon therapy, anti-androgens, or tamoxifen,
have been proven to be essentially ineffective. Recent advances in
the identification of molecules involved in hepatocarcinogenesis
has led to the identification of possible therapeutic targets for
the targeted therapy of HCC, such as growth factors and
neoangiogenesis factors, as well as their receptors, namely
tyrosine kinase intracellular enzymatic pathways and intracellular
signal transmission molecules. The focus of targeted therapy for
HCC has been largely on tyrosine kinase inhibitors and monoclonal
antibodies. The most widely investigated pathways, which are
involved in the process of enzymatic activation of growth- and
proliferation-promoting intracellular signals, include the
Ras/Raf/MEK/MAPK pathway, the phosphoinositol 3-kinase pathway and
the Wnt/catenin pathway (2,5). Thus, studies that assist in selecting
patients to receive novel treatments, such as cetuximab, are
pivotal, as such therapies may prove curative for these patients.
Cetuximab is considered to be a promising agent for advanced-stage
HCC patients, as nearly all HCC cells exhibit increased expression
of EGFR. The results of the few small-sized studies may be
summarized as follows: Cetuximab may be effective in the control of
advanced HCC, prolonging patient survival; however, combination
regimens of cetuximab with oxaliplatin and gemcitabine are
particularly effective. The advantage of cetuximab monotherapy in
the treatment of advanced HCC is that it exhibits a tolerable
toxicity profile, although combination therapy also has an
acceptable toxicity profile (2,3,5,6,15,19).
In our pilot study, we found that ~80% of HCC cancer
patients carry the wild-type K-Ras gene. This is similar to the
findings of previous studies, which reported that the K-Ras
mutation is an uncommon or even rare event in HCC. This is in
contrast to the frequent presence of K-Ras mutations in chemical
models of HCC in animals (1,19–24).
Considering the small sample size of the previous and present
studies, the true prevalence of K-Ras mutations in HCC has yet to
be determined. The discrepancy between various studies may be
attributed to the involved etiological factors, as well as to the
geographic origins of the patient populations (20–24). The
majority of K-Ras genes are located on exon 2 and mutations in
codons 12, 13, 146 and 154 are the most frequent; >80% of
mutations occur in codons 12 and 13 (25–27). Thus,
we may hypothesize that our results comprehensively represent K-Ras
mutations in the investigated group of patients. Additionally, we
acknowledge that the clinical characteristics of our patients may
be beneficial; however, these data were not added to this pilot
study. Our aim was to design the present pilot study to estimate
the K-Ras mutation rate using novel technologies in Turkish HCC
patients.
The majority of the previous studies on the role of
K-Ras mutations in response to cetuximab therapy have been
performed on colon and head and neck cancer patients. Activating
K-Ras gene mutations are detected in ~15–30% of non-small-cell lung
cancer patients and 40–45% of colorectal cancer patients. Several
studies have demonstrated that K-Ras mutations negatively predict
the lack of response to EGFR tyrosine kinase inhibitors in
non-small-cell lung cancer patients and to cetuximab or panitumumab
in colorectal cancer patients. Thus, the general consensus is that
patients with K-Ras mutations do not benefit from cetuximab
treatment. Patients with wild-type K-Ras, in contrast to the mutant
type, may even benefit from cetuximab dose escalation and the
overall survival time is twice as long in K-Ras wild-type patients
treated with cetuximab compared with patients harboring K-Ras
mutations (26–32). In addition to extensive studies on the
role of the K-Ras mutation status in determining the response to
cetuximab in colon cancer patients, the same phenomenon has been
observed in several other types of cancer, including advanced
rectal cancer, gastric cancer, inoperable biliary tract cancer and
squamous cell carcinoma of the skin (33–36).
In conclusion, our results demonstrated that, in a
population of HCC patients referred to 6 major comprehensive cancer
centers in Turkey, K-Ras mutations are a rather uncommon finding.
Thus, it may be feasible to initiate in these centers clinical
trials using cetuximab for the treatment of unresectable or
advanced-stage HCC patients.
References
|
1
|
Taketomi A, Shirabe K, Muto J, Yoshiya S,
Motomura T, Mano Y, Ikegami T, Yoshizumi T, Sugio K and Maehara Y:
A rare point mutation in the Ras oncogene in hepatocellular
carcinoma. Surg Today. 43:289–292. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Rossi L, Zoratto F, Papa A, Iodice F,
Minozzi M, Frati L and Tomao S: Current approach in the treatment
of hepatocellular carcinoma. World J Gastrointest Oncol. 2:348–359.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Poggi G, Montagna B, Melchiorre F,
Quaretti P, Delmonte A, Riccardi A, Tagliaferri B, Sottotetti F, Di
Cesare P, Stella MG, et al: Hepatic intra-arterial cetuximab in
combination with 5-fluorouracil and cisplatin as salvage treatment
for sorafenib-refractory hepatocellular carcinoma. Anticancer Res.
31:3927–3933. 2011.PubMed/NCBI
|
|
4
|
Dhanasekaran R, Limaye A and Cabrera R:
Hepatocellular carcinoma: Current trends in worldwide epidemiology,
risk factors, diagnosis, and therapeutics. Hepat Med. 4:19–37.
2012.PubMed/NCBI
|
|
5
|
Corey KE and Pratt DS: Current status of
therapy for hepatocellular carcinoma. Therap Adv Gastroenterol.
2:45–57. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhu AX, Stuart K, Blaszkowsky LS,
Muzikansky A, Reitberg DP, Clark JW, Enzinger PC, Bhargava P,
Meyerhardt JA, Horgan K, et al: Phase 2 study of cetuximab in
patients with advanced hepatocellular carcinoma. Cancer.
110:581–589. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Mittal S and El-Serag HB: Epidemiology of
hepatocellular carcinoma: Consider the population. J Clin
Gastroenterol. 47(Suppl): S2–S6. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
El-Serag HB: Epidemiology of viral
hepatitis and hepatocellular carcinoma. Gastroenterology.
142:1264–1273.e1. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Alacacioglu A, Somali I, Simsek I,
Astarcioglu I, Ozkan M, Camci C, Alkis N, Karaoglu A, Tarhan O,
Unek T, et al: Epidemiology and survival of hepatocellular
carcinoma in Turkey: Outcome of multicenter study. Jpn J Clin
Oncol. 38:683–688. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Dogan E, Yalcin S, Koca D and Olmez A:
Clinicopathological characteristics of hepatocellular carcinoma in
Turkey. Asian Pac J Cancer Prev. 13:2985–2990. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Can A, Dogan E, Bayoglu IV, Tatli AM,
Besiroglu M, Kocer M, Dulger AC, Uyeturk U, Kivrak D, Orakci Z, et
al: Multicenter epidemiologic study on hepatocellular carcinoma in
Turkey. Asian Pac J Cancer Prev. 15:2923–2927. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ye H, Zhang C, Wang BJ, Tan XH, Zhang WP,
Teng Y and Yang X: Synergistic function of Kras mutation and HBx in
initiation and progression of hepatocellular carcinoma in mice.
Oncogene. 33:5133–5138. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Adjei AA: Blocking oncogenic Ras signaling
for cancer therapy. J Natl Cancer Inst. 93:1062–1074. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kim JH, Kim HY, Lee YK, Yoon YS, Xu WG,
Yoon JK, Choi SE, Ko YG, Kim MJ, Lee SJ, et al: Involvement of
mitophagy in oncogenic K-Ras-induced transformation: Overcoming a
cellular energy deficit from glucose deficiency. Autophagy.
7:1187–1198. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Asnacios A, Fartoux L, Romano O, Tesmoingt
C, Louafi SS, Mansoubakht T, Artru P, Poynard T, Rosmorduc O,
Hebbar M, et al: Gemcitabine plus oxaliplatin (GEMOX) combined with
cetuximab in patients with progressive advanced stage
hepatocellular carcinoma: Results of a multicenter phase 2 study.
Cancer. 112:2733–2739. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Vallée A, Le Loupp AG and Denis MG:
Efficiency of the TheraScreen® RGQ PCR kit for the detection of
EGFR mutations in non-small cell lung carcinomas. Clin Chim Acta.
429:8–11. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Adams JA, Post KM, Bilbo SA, Wang X, Sen
JD, Cornwell AJ, Malek AJ and Cheng L: Performance evaluation
comparison of 3 commercially available PCR-based KRAS mutation
testing platforms. Appl Immunohistochem Mol Morphol. 22:231–235.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chang SC, Denne J, Zhao L, Horak C, Green
G, Khambata-Ford S, Bray C, Celik I, Van Cutsem E and Harbison C:
Comparison of KRAS genotype: TheraScreen assay vs. LNA-mediated
qPCR clamping assay. Clin Colorectal Cancer. 12:195–203.e2. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sanoff HK, Bernard S, Goldberg RM, Morse
MA, Garcia R, Woods L, Moore DT and O'Neil BH: Phase II study of
capecitabine, oxaliplatin and cetuximab for advanced hepatocellular
carcinoma. Gastrointest Cancer Res. 4:78–83. 2011.PubMed/NCBI
|
|
20
|
Colombino M, Sperlongano P, Izzo F,
Tatangelo F, Botti G, Lombardi A, Accardo M, Tarantino L, Sordelli
I, Agresti M, et al: BRAF and PIK3CA genes are somatically mutated
in hepatocellular carcinoma among patients from South Italy. Cell
Death Dis. 3:e2592012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lord PG, Hardaker KJ, Loughlin JM, Marsden
AM and Orton TC: Point mutation analysis of ras genes in
spontaneous and chemically induced C57Bl/10J mouse liver tumours.
Carcinogenesis. 13:1383–1387. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Weihrauch M, Benick M, Lehner G, Wittekind
M, Bader M, Wrbitzk R and Tannapfel A: High prevalence of K-ras-2
mutations in hepatocellular carcinomas in workers exposed to vinyl
chloride. Int Arch Occup Environ Health. 74:405–410. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Weihrauch M, Benicke M, Lehnert G,
Wittekind C, Wrbitzky R and Tannapfel A: Frequent k-ras-2 mutations
and p16(INK4A)methylation in hepatocellular carcinomas in workers
exposed to vinyl chloride. Br J Cancer. 84:982–989. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Bai F, Nakanishi Y, Takayama K, Pei XH,
Inoue K, Harada T, Izumi M and Hara N: Codon 64 of K-ras gene
mutation pattern in hepatocellular carcinomas induced by bleomycin
and 1-nitropyrene in A/J mice. Teratog Carcinog Mutagen. 23(Suppl
1): 161–170. 2003. View Article : Google Scholar
|
|
25
|
Ciardiello F and Tortora G: EGFR
antagonists in cancer treatment. N Engl J Med. 358:1160–1174. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yarom N and Jonker DJ: The role of the
epidermal growth factor receptor in the mechanism and treatment of
colorectal cancer. Discov Med. 11:95–105. 2011.PubMed/NCBI
|
|
27
|
Douillard JY, Oliner KS, Siena S,
Tabernero J, Burkes R, Barugel M, Humblet Y, Bodoky G, Cunningham
D, Jassem J, et al: Panitumumab-FOLFOX4 treatment and RAS mutations
in colorectal cancer. N Engl J Med. 369:1023–1034. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Garassino MC, Borgonovo K, Rossi A,
Mancuso A, Martelli O, Tinazzi A, Di Cosimo S, La Verde N, Sburlati
P, Bianchi C, et al: Biological and clinical features in predicting
efficacy of epidermal growth factor receptor tyrosine kinase
inhibitors: A systematic review and meta-analysis. Anticancer Res.
29:2691–2701. 2009.PubMed/NCBI
|
|
29
|
Yen LC, Uen YH, Wu DC, Lu CY, Yu FJ, Wu
IC, Lin SR and Wang JY: Activating KRAS mutations and
overexpression of epidermal growth factor receptor as independent
predictors in metastatic colorectal cancer patients treated with
cetuximab. Ann Surg. 251:254–260. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Folprecht G, Gruenberger T, Bechstein W,
Raab HR, Weitz J, Lordick F, Hartmann JT, Stoehlmacher-Williams J,
Lang H, Trarbach T, et al: Survival of patients with initially
unresectable colorectal liver metastases treated with
FOLFOX/cetuximab or FOLFIRI/cetuximab in a multidisciplinary
concept (CELIM study). Ann Oncol. 25:1018–1025. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Ji JH, Park SH, Lee J, Kim TW, Hong YS,
Kim KP, Kim SY, Baek JY, Kang HJ, Shin SJ, et al: Prospective phase
II study of neoadjuvant FOLFOX6 plus cetuximab in patients with
colorectal cancer and unresectable liver-only metastasis. Cancer
Chemother Pharmacol. 72:223–230. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Jiang Y, Kimchi ET, Staveley-O'Carroll KF,
Cheng H and Ajani JA: Assessment of K-ras mutation: A step toward
personalized medicine for patients with colorectal cancer. Cancer.
115:3609–3617. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Rubovszky G, Láng I, Ganofszky E, Horváth
Z, Juhos E, Nagy T, Szabó E, Szentirmay Z, Budai B and Hitre E:
Cetuximab, gemcitabine and capecitabine in patients with inoperable
biliary tract cancer: A phase 2 study. Eur J Cancer. 49:3806–3812.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Shi M, Shi H, Ji J, Cai Q, Chen X, Yu Y,
Liu B, Zhu Z and Zhang J: Cetuximab inhibits gastric cancer growth
in vivo, independent of KRAS status. Curr Cancer Drug Targets.
14:217–224. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Sun PL, Li B and Ye QF: Effect of
neoadjuvant cetuximab, capecitabine and radiotherapy for locally
advanced rectal cancer: Results of a phase II study. Int J
Colorectal Dis. 27:1325–1332. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Uribe P and Gonzalez S: Epidermal growth
factor receptor (EGFR) and squamous cell carcinoma of the skin:
Molecular bases for EGFR-targeted therapy. Pathol Res Pract.
207:337–342. 2011. View Article : Google Scholar : PubMed/NCBI
|