Introduction
Breast cancer is the most prevalent cancer and the
leading cause of cancer mortality in females worldwide, as well as
in Thailand. Angiogenesis is the formation of new blood vessels and
has been involved in the initiation and aggressiveness of breast
cancer (1–3). The most important key modulator in
this complex process is vascular endothelial growth factor
(VEGF). VEGF plays a role in breast cancer (4) and the VEGF pathway is targeted
in the treatment of breast cancer (5).
Human VEGF is localized on chromosome 6p21.3
and organized as eight exons, separated by seven introns (6,7).
Several polymorphisms in the promoter and 5′ untranslated region of
(UTR) of VEGF have been identified (8,9).
Awata et al previously identified seven polymorphisms in the
promoter region as well as 5′ and 3′UTR of VEGF in a
Japanese population. Serum VEGF levels have also been found
to be significantly higher in healthy subjects with the −634C/C
genotype (10). However, in
vitro experiments using lipopolysaccharide-stimulated
peripheral blood mononuclear cells demonstrated that −634 G/G
correlates with elevated VEGF production (9). In non-small cell lung cancer, a low
VEGF expression in cancer tissue was significantly
associated with the presence of the −2578C/C, −634G/G and −1154A/A
and GA alleles in the VEGF promoter (11). The association between VEGF
polymorphisms and breast cancer has been previously investigated
(12). Based on these
observations, we hypothesized that polymorphisms in the VEGF
promoter and 5′UTR contribute to varied levels of VEGF
expression, subsequently leading to susceptibility to aggressive
breast cancer. To investigate this hypothesis, the association
between VEGF polymorphisms and breast cancer susceptibility
and aggressiveness was investigated, as well as mRNA expression in
breast cancer tissue.
Materials and methods
Study population
The study population was recruited from the Division
of Head, Neck and Breast Surgery (Siriraj Hospital, Bangkok,
Thailand) between 2000 and 2003. Patients with histopathologically
confirmed breast carcinoma were included in this prospective study
and newly diagnosed breast cancer patients were included in the
case group. Patients with histories of other types of cancer were
excluded. Healthy individuals and patients who attended the
hospital due to benign conditions with no previous diagnosis of
cancer were included in the control group and frequency matched to
the breast carcinoma cases with regard to age (±5 years). Informed
consent was obtained from all subjects. This study was approved by
the ethics committee of the Siriraj Hospital, Mahidol University,
Bangkok, Thailand.
Genotyping of VEGF polymorphisms
VEGF −634G/C and −1154G/A were genotyped by
the allele refractory mutation system-PCR. PCR was performed as
summarized in Table I. VEGF
−1498C/T and −2578C/A were genotyped by the PCR-restriction
fragment length polymorphisms. Representative PCR products were
sequenced to validate the assay.
 | Table IPCR primer pairs and conditions for
VEGF genotyping. |
Table I
PCR primer pairs and conditions for
VEGF genotyping.
| Polymorphisms | Primers | Product size, bp | PCR conditions (T,
MgCl2)a | Alleles | Restriction
enzyme | DNA fragment sizes,
bp |
|---|
| −634G/C | F:
CATTGATCCGGGTTTTATCCC | 282 | 65–60b, 1 | | | |
| G:
CACTCACTTTGTCCCTGTAG | | | G | - | - |
| C:
CACTCACTTTGTCCCTGTAC | | | C | | - |
| Inc F:
AGATGGTCCCTCACCTTCCT | 352 | | | | |
| Inc R:
GTCTACCCTCCTGAGCTTGC | | | | | |
| −1154G/A | F:
GTCCGCACGTAACCTCACTT | 220 | 62, 1.5 | | | |
| G:
GACAGGCGAGCTTCAGCACC | | | G | - | - |
| A:
GACAGGCGAGCTTCAGCACT | | | A | - | |
| Inc F:
AGATGGTCCCTCACCTTCCT | 352 | | | | |
| Inc R:
GTCTACCCTCCTGAGCTTGC | | | | | |
| −1498C/T | F:
TGTGCGTGTGGGGTTGAGCG | 175 | 60, 1.5 | T | BstUI | 175 |
| R:
TACGTGCGGACAGGGCCTGA | | | C | | 155, 20 |
| −2578C/A | F:
ATTGCTGCATTCCCATTCTC | 251 | 60, 2.5 | C | BstYI | 251 |
| R:
CCCTTTTCCTCCAACTCTC | 268 | | A | | 180, 88 |
Assessment of VEGF mRNA expression
levels
The correlation between VEGF polymorphisms
and expression levels was determined in breast tissue. VEGF
mRNA expression was assayed by semi-quantitative RT-PCR as
described previously (13). A
primer pair that amplified β-actin was employed to check RNA
integrity and used as an internal control. Primer sequences, PCR
conditions and the number of PCR cycles are presented in Table II. To account for variability
between gels, an RT-PCR product from the MDA-MB-231 cell line was
electrophoresed as a control on each gel. PCR product intensity was
analyzed using GeneTools® software (Syngene, Cambridge,
UK). mRNA levels were calculated as the ratio of tissue sample to
corresponding β-actin and then corrected as a ratio to the
MDA-MB-231 sample on the same scan. Each RNA sample was assayed in
duplicate and in two separate settings.
 | Table IIPCR primer pairs and conditions for
semi-quantitative RT-PCR of VEGF and β-actin
mRNA. |
Table II
PCR primer pairs and conditions for
semi-quantitative RT-PCR of VEGF and β-actin
mRNA.
| Gene | Primers | Product size,
bp | PCR conditions (T,
MgCl2)a | PCR cycles |
|---|
| VEGF | F:
CTCACCAAGGCCAGCACATAGG | 159 | 55, 2.5 | 32 |
| R:
ATCTGGTTCCCGAAACCCTGAG | 291 | | |
| | 363 | | |
| | 414 | | |
| β-actin | F:
TCGACAACGGCTCCGGCAT | 239 | 50, 2.5 | 26 |
| R:
AAGGTGTGGTGCCAGATTTTC | | | |
Statistical analysis
Distribution of VEGF allele frequencies and
genotypes among the case and control groups was analyzed using the
χ2 test. Odds ratios (ORs) and 95% confidence intervals
(CIs), obtained from unconditional logistic regression, were used
to measure the strength of the association between VEGF
polymorphisms and susceptibility and aggressiveness of breast
cancer. Individual haplotypes were determined using the PHASE
program available at http://www.stat.washington.edu/stephens/phase.html
(14). The end-point of overall
survival (OS) analysis was breast cancer-associated mortality. The
disease-free survival (DFS) analysis end-point was cancer
recurrent/metastasis or breast cancer-associated mortality. DFS and
OS time was calculated as the time from diagnosis to the end point
of the study, censoring at the date of last contact or non-cancer
mortality. The survival curves were determined using a Kaplan-Meier
product-limit method. Statistical significance between the survival
curves was assessed using the log-rank test. Multivariate analysis
was performed to evaluate the effect of prognostic factors on OS,
using the Cox proportional hazards model. P<0.05 was considered
to indicate a statistically significant difference. mRNA levels
were calculated as the ratio of tissue sample to corresponding
β-actin and corrected as a ratio to the MDA-MB-231 cell line
on the same scan. Each RNA sample was assayed in duplicate and in
two separate settings.
Results
Correlation between VEGF genotypes and
breast cancer susceptibility
Genotyping was performed on 483 breast cancer
patients and 524 controls. −634C allele distribution was
significantly higher in breast cancer patients compared with
control subjects (33.23 vs. 23.71%; P<0.001). VEGF
−634G/C and −634C/C genotype distributions were significantly
higher in breast cancer patients (GC, 41.20 and CC, 12.63 vs. GC,
25.06 and CC, 11.18%; P<0.001). Allele and genotype frequency
distributions of other loci were not found to be different. The
mean age of the control group was 48.56±14.45 years (SEM). The mean
age of the breast cancer patients was 50.8±11.326 years (SEM). The
mean age of the breast cancer and control groups were statistically
different [OR, −2.238; 95% CI, (−5.20)-(−3.957); P=0.011], thus,
ORs and 95% CIs calculated by logistic regression were adjusted for
age. Individuals with the −634G/C genotype had an increased risk of
breast cancer when compared with the −634G/G genotype (OR, 2.544;
95% CI, 1.852–3.496; P<0.001). Homozygous CC had an increased
risk when compared with −634G/G (OR, 1.600; 95% CI, 1.030–2.485;
P=0.036; Table III). VEGF
polymorphisms in other loci did not demonstrate any increased risk
for breast cancer.
 | Table IIICorrelation between VEGF
genotype and breast cancer susceptibility. |
Table III
Correlation between VEGF
genotype and breast cancer susceptibility.
| VEGF
genotype polymorphisms | Controls, n
(%) | Cases, n (%) | OR (95% CI),
P-value |
|---|
| −634 |
| GG | 234 (65.91) | 223 (46.17) | 1 (ref.) |
| GC | 81 (22.82) | 199 (41.20) | 2.544
(1.852–3.496), 0.001 |
| CC | 40 (11.27) | 61 (12.63) | 1.600
(1.030–2.485), 0.036 |
| −1154 |
| GG | 279 (69.23) | 318 (65.84) | 1 (ref.) |
| GA | 118 (29.28) | 149 (30.85) | 1.096
(0.819–1.467), 0.538 |
| AA | 6 (1.49) | 16 (3.31) | 2.320
(0.819–6.014), 0.084 |
| −1498 |
| TT | 215 (52.31) | 243 (50.31) | 1 (ref.) |
| CT | 172 (41.85) | 214 (44.31) | 1.097
(0.835–1.441), 0.507 |
| CC | 24 (5.84) | 26 (5.38) | 1.002
(0.557–1.802), 0.995 |
| −2578 |
| CC | 214 (51.69) | 240 (49.69) | 1 (ref.) |
| AC | 173 (41.79) | 213 (44.10) | 1.091
(0.830–1.435), 0.530 |
| AA | 27 (6.52) | 30 (6.21) | 1.016
(0.584–1.767), 0.957 |
Correlation between VEGF genotypes and
clinicopathological parameters
Table IV shows
known clinicopathological parameters and demographic factors of the
breast cancer patients. Numerous patients received surgery
(mastectomy in 385 patients and wide excision in 91 patients). The
patients with invasive carcinoma who underwent wide excision
received radiotherapy. VEGF −634G/C genotype was associated
with tumor size >20 mm (OR, 1.638; 95% CI, 1.103–2.434;
P=0.015), perineural invasion (OR, 2.261; 95% CI, 1.217–4.202;
P=0.010) and stage II–IV at diagnosis (OR, 1.915; 95% CI,
1.255–2.944; P=0.003). Separate analysis of invasive ductal
carcinoma revealed a marked association with tumor size >20 mm
(OR, 1.722; 95% CI, 1.097–2.703; P=0.018), perineural invasion (OR,
2.36; 95% CI, 1.227–4.539; P=0.010) and stage II–IV at diagnosis
(OR, 2.078; 95% CI, 1.237–3.490; P=0.006). The VEGF −1498C/C
genotype correlated with decreased risk of lymphovascular invasion
(LVI; OR, 0.308; 95% CI, 0.102–0.927; P=0.036).
 | Table IVClinicopathological parameters and
demographic factors of breast cancer. |
Table IV
Clinicopathological parameters and
demographic factors of breast cancer.
|
Characteristics | Breast cancer
patients, n (%) |
|---|
| Age at diagnosis,
years |
| ≤40 | 68 (14.08) |
| 40–49 | 178 (36.85) |
| 50–59 | 137 (28.36) |
| >60 | 100 (20.70) |
| Tumor type |
| Ductal carcinoma
in situ | 36 (7.45) |
| Invasive ductal
carcinoma | 396 (81.99) |
| Invasive lobular
carcinoma | 12 (2.48) |
| Others | 39 (8.07) |
| Tumor size, mm |
| In
situ | 30 (6.21) |
| ≤20 | 159 (32.92) |
| >20–50 | 250 (51.76) |
| >50 | 44 (9.11) |
| Axillary nodal
metastasis |
| No | 271 (56.11) |
| Yes | 205 (42.44) |
| Unknown | 7 (1.45) |
| Distant
metastasis |
| No | 466 (96.48) |
| Yes | 17 (3.52) |
| Stage at
diagnosis |
| 0 | 28 (5.80) |
| I | 115 (23.81) |
| II | 214 (44.31) |
| III | 109 (22.57) |
| IV | 17 (3.52) |
| Histological
grading |
|
Well-differentiated | 36 (7.45) |
| Moderately
differentiated | 230 (47.62) |
| Poorly
differentiated | 146 (30.23) |
| Unknown | 71 (14.70) |
| ER |
| Negative | 191 (39.54) |
| Positive | 261 (54.04) |
| Unknown | 31 (6.42) |
| PR |
| Negative | 236 (48.86) |
| Positive | 210 (43.48) |
| Unknown | 37 (7.66) |
| Surgery |
| Yes | 476 (98.55) |
| No | 3 (0.62) |
| Unknown | 4 (0.83) |
| Chemotherapy |
| Yes | 308 (63.77) |
| No | 166 (34.37) |
| Unknown | 9 (1.86) |
| Radiotherapy |
| Yes | 198 (40.99) |
| No | 272 (56.31) |
| Unknown | 13 (2.69) |
Haplotype analysis
The −2578C/−1498T/−1154G/−634G haplotype was the
most common haplotype in the two groups (frequency, 0.3778 and
0.4739, respectively). Permutation testing revealed a significant
difference between haplotype frequencies in the breast cancer and
control groups (P=0.01). CTGG and CTGC haplotype copy number
distributions were significantly different between the groups.
Bearing 1 or 2 copies of the CTGG haplotype had a protective effect
against breast cancer (OR, 0.55; 95% CI, 0.42–0.73; P<0.001). By
contrast, bearing 1 or 2 copies of the CTGC haplotype was found to
have an increased risk for breast cancer (OR, 1.81; 95% CI,
1.39–2.35; P<0.001). Patients with 1 or 2 copies of the CTGC
haplotype significantly correlated with a tumor size >20 mm,
(OR, 1.60; 95% CI, 1.09–2.35; P=0.0126), presence of perinural
invasion (PNI; OR, 1.84; 95% CI, 0.97–3.52; P=0.046) and stage
II–IV (OR, 1.73; 95% CI, 1.14–2.61; P=0.0062). Patients with 1 or 2
copies of ACAG haplotype exhibited a reduced risk for LVI and
poorly differentiated cell types (OR, 0.74; 95% CI, 0.48–1.15;
P=0.1656 and OR, 0.74; 95% CI, 0.46–1.17; P=0.1768,
respectively).
Survival analysis
The median follow-up time was 65 months (range,
1.25–136.7 months). Among 446 patients, there were 37 mortalities
during the study period, 30 of which were breast cancer-related.
Locoregional recurrence was observed in 29 patients. Distant
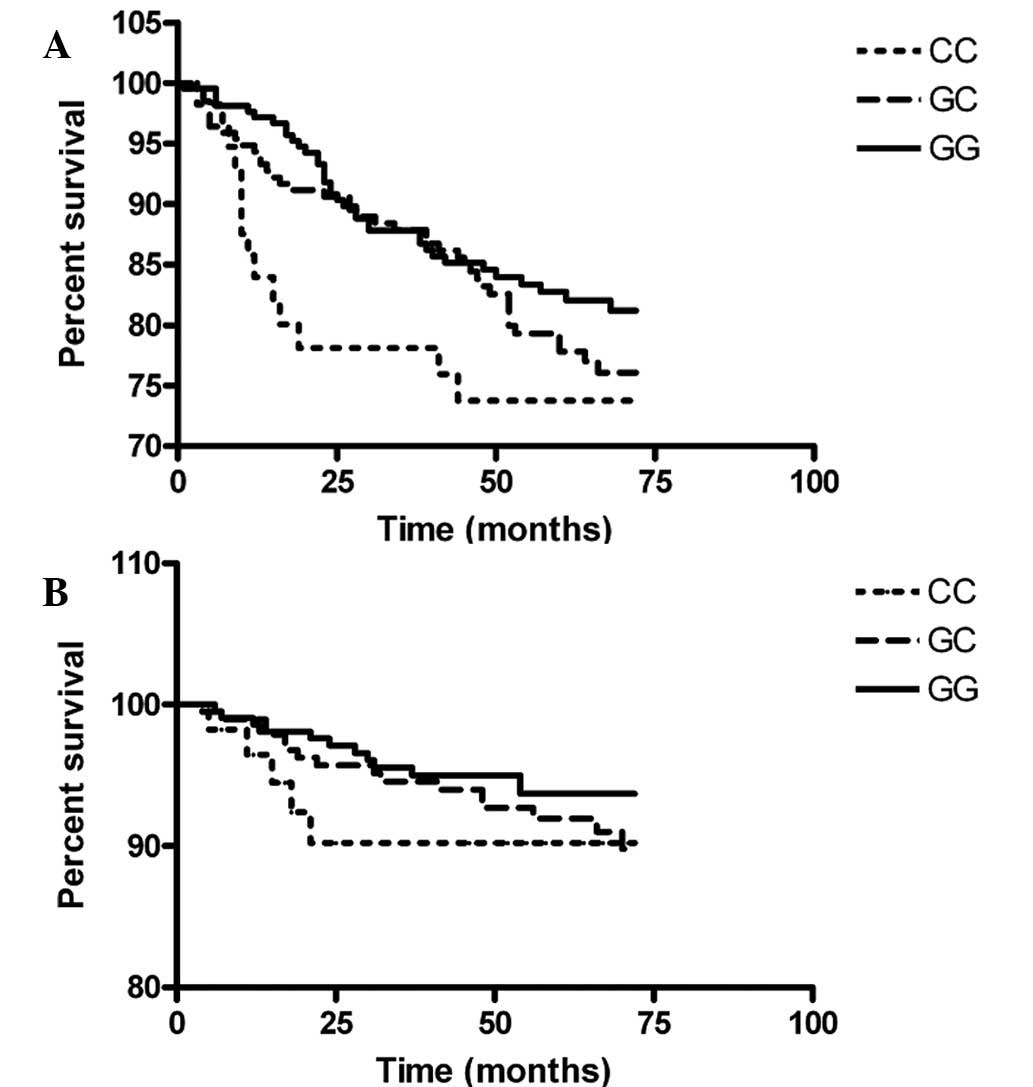
metastasis occurred in 86 patients. The Kaplan-Meier survival
curves of various VEGF genotypes are shown in Fig. 1. Patients with the −634CC genotype
(5 mortalities of 57 patients) succumbed to breast cancer in the
first 21 months following breast cancer diagnosis. Univariate
analysis between clinicopathological parameters, VEGF
genotypes and corresponding 5-year survival are provided in
Table V. Age >50 years
correlated with lower 5-year OS. Larger tumor size, LVI, PNI,
estrogen receptor- and progesterone receptor-negative, regional
lymph-node positive, distant metastasis and advanced staging were
the major prognostic factors for OS and DFS. No statistically
significant correlation was found between VEGF genotypes and
5-year survival in the univariate analysis. Factors with a P-value
≤0.2 were included in the Cox regression analysis and poor
differentiation, presence of PNI, PR-negative, axillary nodal
metastasis and having the −634C/C and −1154G/A genotypes were found
to significantly correlate with increased hazard ratio (HR) for
DFS. HRs were 3.050 (95% CI, 1.354–6.871; P=0.007) and 2.452 (95%
CI, 1.384–4.343; P=0.002) for the −634C/C and −1154G/A genotypes,
respectively.
 | Table VDFS and OS by clinicopathological
parameters and VEGF polymorphisms. |
Table V
DFS and OS by clinicopathological
parameters and VEGF polymorphisms.
| DFS | OS |
|---|
|
|
|
|---|
| Parameters | Cases, n | Event | 5-year survival,
% | P-value | Cases, n | Event | 5-year survival,
% | P-value |
|---|
| Age at diagnosis,
years |
| ≤50 | 250 | 50 | 79.70 | 0.863 | 253 | 12 | 95.02 | 0.047 |
| >50 | 197 | 40 | 78.46 | | 205 | 19 | 90.89 | |
| Tumor size, mm |
| In situ and
≤20 | 177 | 12 | 93.13 | <0.001 | 180 | 4 | 97.66 | 0.001 |
| >20 | 274 | 78 | 70.03 | | 285 | 27 | 90.18 | |
| Histological
grading |
| Well-/moderately
differentiated | 245 | 44 | 81.48 | 0.014 | 253 | 15 | 94.93 | 0.158 |
| Poorly
differentiated | 134 | 38 | 70.23 | | 137 | 13 | 88.45 | |
| LVI |
| Absent | 277 | 39 | 84.80 | <0.001 | 284 | 15 | 95.22 | 0.047 |
| Present | 147 | 45 | 69.31 | | 154 | 15 | 89.15 | |
| PNI |
| Absent | 332 | 51 | 83.79 | <0.001 | 343 | 15 | 95.80 | 0.003 |
| Present | 49 | 20 | 55.11 | | 52 | 7 | 85.92 | |
| ER |
| Positive | 244 | 42 | 82.24 | 0.032 | 254 | 10 | 96.36 | 0.003 |
| Negative | 179 | 44 | 74.48 | | 183 | 20 | 88.19 | |
| PR |
| Positive | 196 | 25 | 87.31 | <0.001 | 204 | 7 | 97.39 | 0.007 |
| Negative | 222 | 59 | 72.06 | | 228 | 22 | 89.38 | |
| Regional nodal
metastasis |
| No | 258 | 25 | 89.24 | <0.001 | 262 | 7 | 97.41 | <0.001 |
| Yes | 187 | 64 | 65.05 | | 197 | 23 | 87.48 | |
| Distant
metastasis |
| No | N/A | N/A | N/A | N/A | 449 | 28 | 93.72 | 0.007 |
| Yes | N/A | N/A | N/A | N/A | 16 | 3 | 74.29 | |
| Staging |
| 0–I | 134 | 8 | 94.06 | <0.001 | 133 | 1 | 99.25 | 0.001 |
| II–IV | 312 | 82 | 72.73 | | 324 | 30 | 90.55 | |
| 634G/C |
| GG | 209 | 35 | 82.87 | 0.088 | 215 | 12 | 94.18 | 0.490 |
| GC | 187 | 40 | 77.54 | | 193 | 14 | 92.98 | |
| CC | 55 | 15 | 71.14 | | 57 | 5 | 90.29 | |
| −1154G/A |
| GG | 297 | 52 | 81.23 | 0.166 | 304 | 18 | 93.99 | 0.627 |
| GA | 138 | 35 | 74.45 | | 145 | 12 | 91.83 | |
| AA | 16 | 3 | 81.25 | | 16 | 1 | 92.31 | |
| −1498C/T |
| TT | 225 | 38 | 82.33 | 0.224 | 231 | 13 | 93.72 | 0.542 |
| CT | 201 | 48 | 74.66 | | 208 | 17 | 92.52 | |
| CC | 25 | 4 | 87.50 | | 26 | 1 | 94.74 | |
| −2578C/A |
| CC | 223 | 39 | 81.82 | 0.207 | 229 | 13 | 93.67 | 0.162 |
| AC | 199 | 47 | 74.82 | | 206 | 18 | 91.73 | |
| AA | 29 | 4 | 88.82 | | 30 | 0 | 100.00 | |
VEGF mRNA expression in breast cancer
tissue
VEGF mRNA expression was evaluated in 124
breast cancer tissues. Characteristics of breast cancer patients
are provided in Table VI.
Expression ranged between 0 and 3.27 with a median of 1.10.
Patients with the VEGF −634C/C genotype had significantly
higher VEGF mRNA in breast cancer tissue compared with those
with the −634G/G or −634G/C genotype (Fig. 2). Patients with heterozygous
−1154G/A, −1498C/T and −2578A/C exhibited lower VEGF mRNA
when compared with homozygous −1154G/G, −1498T/T and −2578C/C. Due
to the presence of outliers and a small number of patients with the
homozygous −1154A/A, −1498C/C and −2578A/A genotypes, VEGF
mRNA expression in these groups appeared to be high. Following
exclusion of the outlier, VEGF mRNA expression was decreased
in patients with homozygous −1154A/A, −1498C/C and −2578A/A (data
not shown).
 | Table VICharacteristics of 124 breast cancer
patients included in VEGF mRNA evaluation. |
Table VI
Characteristics of 124 breast cancer
patients included in VEGF mRNA evaluation.
|
Characteristics | Patients, n
(%) |
|---|
| Age at diagnosis,
years |
| ≤50 | 66 (53.23) |
| >50 | 58 (47.77) |
| Tumor type |
| Ductal carcinoma
in situ | 2 (1.61) |
| Invasive ductal
carcinoma | 108 (87.10) |
| Invasive lobular
carcinoma | 4 (3.23) |
| Others | 9 (7.26) |
| Tumor size, mm |
| In
situ | 2 (1.61) |
| ≤20 | 28 (22.58) |
| >20–50 | 68 (54.84) |
| >50 | 26 (20.97) |
| Axillary nodal
metastasis |
| No | 59 (47.58) |
| Yes | 65 (52.42) |
| Distant
metastasis |
| No | 117 (94.35) |
| Yes | 7 (5.65) |
| Stage at
diagnosis |
| 0 | 2 (1.61) |
| I | 20 (16.13) |
| II | 61 (49.19) |
| III | 34 (27.42) |
| IV | 7 (5.65) |
| Histological
grading |
| Well
differentiated | 3 (2.61) |
| Moderately
differentiated | 69 (60.00) |
| Poorly
differentiated | 43 (37.39) |
| LVI |
| Absent | 68 (57.14) |
| Present | 51 (42.86) |
| PNI |
| Absent | 93 (83.78) |
| Present | 18 (16.22) |
| ER |
| Negative | 60 (48.78) |
| Positive | 63 (51.22) |
| PR |
| Negative | 70 (56.91) |
| Positive | 53 (43.09) |
Correlation between VEGF mRNA levels and
clinicopathological parameters
Breast cancer patients were classified into low and
high expression groups using a median value of 1.10. Patient
distribution in each group and clinicopathological parameters are
provided in Table VII. Elevated
VEGF expression correlated with a tumor size >20 mm (OR,
2.476; 95% CI, 1.047–5.858; P=0.039), axillary nodal metastasis
(OR, 2.288; 95% CI, 1.110–4.713; P=0.025) and presence of LVI (OR,
2.406; 95% CI, 1.142–5.070; P=0.021).
 | Table VIICorrelation between VEGF mRNA
expression and clinicopathological parameters. |
Table VII
Correlation between VEGF mRNA
expression and clinicopathological parameters.
| VEGF mRNA
expression | | |
|---|
|
| | |
|---|
|
Characteristics | Low | High | OR (95% CI) | P-value |
|---|
| Age, years | | | | |
| ≤50 | 36 (58.06) | 30 (48.39) | | |
| >50 | 26 (41.94) | 32 (51.61) | 1.477
(0.727–3.001) | 0.281 |
| Tumor size, mm | | | | |
| ≤20 | 20 (32.26) | 10 (16.13) | | |
| >20 | 42 (67.74) | 52 (83.87) | 2.476
(1.047–5.858) | 0.039 |
| Axillary nodal
metastasis | | | | |
| No | 36 (58.06) | 23 (37.70) | | |
| Yes | 26 (41.94) | 38 (62.30) | 2.288
(1.110–4.713) | 0.025 |
| Distant
metastasis | | | | |
| No | 60 (96.77) | 56 (91.80) | | |
| Yes | 2 (3.23) | 5 (8.20) | 2.679
(0.499–14.369) | 0.250 |
| Staging | | | | |
| 0–II | 46 (74.19) | 37 (59.68) | | |
| III–IV | 16 (25.81) | 25 (40.32) | 1.943
(0.906–4.163) | 0.088 |
| Histological
grading | | | | |
| Well-/moderately
differentiated | 38 (65.52) | 34 (59.65) | | |
| Poorly
differentiated | 20 (34.48) | 23 (40.35) | 1.285
(0.603–2.740) | 0.516 |
| LVI | | | | |
| Absent | 40 (67.80) | 28 (46.67) | | |
| Present | 19 (32.20) | 32 (53.33) | 2.406
(1.142–5.070) | 0.021 |
| PNI | | | | |
| Absent | 48 (87.27) | 45 (80.36) | | |
| Present | 7 (12.73) | 11 (19.64) | 1.676
(0.598–4.701) | 0.326 |
| ER | | | | |
| Positive | 36 (58.06) | 27 (44.26) | | |
| Negative | 26 (41.94) | 34 (55.74) | 0.574
(0.281–1.171) | 0.127 |
| PR | | | | |
| Positive | 31 (50.00) | 22 (36.07) | | |
| Negative | 31 (50.00) | 39 (63.93) | 0.564
(0.274–1.161) | 0.120 |
| Hormone
receptor | | | | |
| Negative | 39 (62.90) | 30 (49.18) | | |
| Positive | 23 (37.10) | 31 (50.82) | 0.571
(0.278–1.172) | 0.126 |
Discussion
In the current study, it was observed that
alteration of nucleotides from G to C at −634, resulted in an
increased risk of breast cancer. However, previous studies have not
reported this correlation in breast cancer (15–20).
In the present study, the −634G/C genotype was significantly
associated with more aggressive features. Due to a limited number
of patients with the −634C/C genotype, the difference was not
observed to be statistically significant. This was consistent with
a previous study by Balasubramanian et al and Jin et
al which reported that an alteration of G to C at this position
was associated with a larger tumor size and high grade breast
cancer (16,18). By contrast, Langsenlehner et
al observed a significant correlation between the −634G/C and
−634C/C genotypes and smaller tumor size (19). Survival analysis revealed a
significant correlation between the −634C/C genotype and lower DFS.
However, OS of the patients with different −634G/C genotypes was
similar, which may be due to the relatively short term follow up of
this study. By contrast, the survival analysis of 1,455 Chinese
breast cancer patients revealed that patients with the −634G/G
genotype had a lower OS compared with those with the −634C/C
genotype, however, this polymorphism was not found to correlate
with DFS (21). The variance in
the demographic results of the cancer population may contribute to
discrepancies observed. In the present study, 26.09% of the
patients were diagnosed as stage III and IV and 63.77% of the
patients received chemotherapy, while in a study by Lu et
al, only 11.34% of the patients were diagnosed as stage III and
IV and a large number of patients received chemotherapy (93.95%)
(21).
In the present study, no correlation was found
between −1154G/A polymorphisms and breast cancer risk, consistent
with previous studies in Caucasian populations (18,20,22).
No correlation between breast cancer aggressiveness and −1154G/A
polymorphisms was observed, consistent with a study by Jin et
al(18). Breast cancer
susceptibility or aggressiveness was not associated with −1498C/T
polymorphisms, consistent with previous large case-control studies
in Asian and Caucasian populations (15,16).
There was no association between −2578A/C polymorphisms and breast
cancer susceptibility/aggressiveness as previously observed in
Caucasian populations by Jin et al and Langsenlehner et
al(18,19). By contrast, two additional studies
in Caucasian populations revealed conflicting results. Schneider
et al reported an association between the −2578A/A genotype
and breast cancer risk while Jacobs et al reported that
−2578C was associated with an increased risk of invasive breast
cancer (17,20). However, the latter study stated the
importance of LD of −2578A/C and −1154G/A. Thus, the association of
the two polymorphisms and breast cancer risk may not be
individually demonstrated.
Haplotype analysis revealed that the −2578C/−1498T/
−1154G/−634G haplotypes had a protective effect against breast
cancer. Patients with the −2578C/−1498T/−1154G/−634C haplotype had
an increased risk of breast cancer and were associated with a tumor
size >20 mm, stage II–IV and PNI. In a Swedish population,
haplotypes −2578C/−634C were significantly associated with a large
tumor size and higher grade. Having 2 copy numbers of haplotypes
−2578A/−634G was associated with lower tumor grade (18). Jacobs et al reported that
the −2578A/−1154A/−634G haplotypes correlated with a reduced risk
of breast cancer in an American population (17). These observations and the
observations of the present study indicate that haplotypes bearing
the−634C allele contribute to increased breast cancer risk and
aggressiveness. Non-replication of genetic association results is
common in genetic epidemiology. In addition, the polymorphisms on
the promoter region and 5′UTR were in LD. Interpretation of
haplotype analysis revealed that the alleles included in the
haplotypes may be associated with other functional polymorphisms
that were not assessed. Comparison of haplotype analysis may be
complicated due to variation in alleles in the haplotype, different
software used to generate the haplotype and determining haplotype
frequency.
The VEGF −634CC genotype correlated with
elevated levels of mRNA expression. Elevated VEGF mRNA
expression correlated with tumor size >20 mm, lymph node
involvement and presence of LVI. These observations are consistent
with a previous study employing the RT-PCR technique. Gomez-Esquer
et al demonstrated a correlation between VEGF mRNA
expression higher than the 25th percentile and more aggressive
features in 103 breast cancer patients (23).
Incorporation of bevacizumab, a humanized monoclonal
antibody that targets VEGF in chemotherapy, is a rapidly
evolving area in the treatment of breast cancer. An association
study of VEGF polymorphisms in 180 advanced breast cancer
patients treated with paclitaxel alone or with bevacizumab and 183
untreated patients, revealed that the VEGF −2578A/A genotype
was associated with an improved median OS in the combination arm
when compared with AC combined with the CC genotype. The
VEGF −1154A/A genotype also demonstrated an improved median
OS when compared with GG combined with the GA genotype in the
combination arm (24). These
observations indicate that selection methods to identify the
patients suitable for anti-VEGF therapy must be established.
−634G/C polymorphisms may identify populations at risk and predict
the outcome of breast cancer. It is possible that genotyping of
VEGF −634 polymorphisms in breast cancer patients may be
used to select appropriate patients for anti-angiogenesis
treatment.
Acknowledgements
This study was supported by the Faculty of Medicine
Siriraj Hospital, Mahidol University.
References
|
1
|
Guinebretière JM, Lê Monique G, Gavoille
A, Bahi J and Contesso G: Angiogenesis and risk of breast cancer in
women with fibrocystic disease. J Natl Cancer Inst. 86:635–636.
1994.PubMed/NCBI
|
|
2
|
Weidner N, Folkman J, Pozza F, et al:
Tumor angiogenesis: a new significant and independent prognostic
indicator in early-stage breast carcinoma. J Natl Cancer Inst.
84:1875–1887. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Weidner N, Semple JP, Welch WR and Folkman
J: Tumor angiogenesis and metastasis - correlation in invasive
breast carcinoma. N Engl J Med. 324:1–8. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Gasparini G: Prognostic value of vascular
endothelial growth factor in breast cancer. Oncologist. 5(Suppl 1):
S37–S44. 2000. View Article : Google Scholar
|
|
5
|
Chan A, Miles DW and Pivot X: Bevacizumab
in combination with taxanes for the first-line treatment of
metastatic breast cancer. Ann Oncol. 21:2305–2315. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Vincenti V, Cassano C, Rocchi M and
Persico G: Assignment of the vascular endothelial growth factor
gene to human chromosome 6p21.3. Circulation. 93:1493–1495. 1996.
View Article : Google Scholar
|
|
7
|
Tischer E, Mitchell R, Hartman T, Silva M,
Gospodarowicz D, Fiddes JC and Abraham JA: The human gene for
vascular endothelial growth factor. Multiple protein forms are
encoded through alternative exon splicing. J Biol Chem.
266:11947–11954. 1991.
|
|
8
|
Brogan IJ, Khan N, Isaac K, Hutchinson JA,
Pravica V and Hutchinson IV: Novel polymorphisms in the promoter
and 5′ UTR regions of the human vascular endothelial growth factor
gene. Hum Immunol. 60:1245–1249. 1999.
|
|
9
|
Watson CJ, Webb NJ, Bottomley MJ and
Brenchley PE: Identification of polymorphisms within the vascular
endothelial growth factor (VEGF) gene: correlation with variation
in VEGF protein production. Cytokine. 12:1232–1235. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Awata T, Inoue K, Kurihara S, et al: A
common polymorphism in the 5′-untranslated region of the VEGF gene
is associated with diabetic retinopathy in type 2 diabetes.
Diabetes. 51:1635–1639. 2002.
|
|
11
|
Koukourakis MI, Papazoglou D,
Giatromanolaki A, Bougioukas G, Maltezos E and Sivridis E: VEGF
gene sequence variation defines VEGF gene expression status and
angiogenic activity in non-small cell lung cancer. Lung Cancer.
46:293–298. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sa-Nguanraksa D and O-Charoenrat P: The
role of vascular endothelial growth factor a polymorphisms in
breast cancer. Int J Mol Sci. 13:14845–14864. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
O-charoenrat P, Rhys-Evans P, Modjtahedi H
and Eccles SA: Vascular endothelial growth factor family members
are differentially regulated by c-erbB signaling in head and neck
squamous carcinoma cells. Clin Exp Metastasis. 18:155–161. 2000.
View Article : Google Scholar
|
|
14
|
Stephens M, Smith NJ and Donnelly P: A new
statistical method for haplotype reconstruction from population
data. Am J Hum Genet. 68:978–989. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Kataoka N, Cai Q, Wen W, et al:
Population-based case-control study of VEGF gene polymorphisms and
breast cancer risk among Chinese women. Cancer Epidemiol Biomarkers
Prev. 15:1148–1152. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Balasubramanian SP, Cox A, Cross SS,
Higham SE, Brown NJ and Reed MW: Influence of VEGF-A gene variation
and protein levels in breast cancer susceptibility and severity.
Int J Cancer. 121:1009–1016. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Jacobs EJ, Feigelson HS, Bain EB, et al:
Polymorphisms in the vascular endothelial growth factor gene and
breast cancer in the Cancer Prevention Study II cohort. Breast
Cancer Res. 8:R222006. View
Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jin Q, Hemminki K, Enquist K, et al:
Vascular endothelial growth factor polymorphisms in relation to
breast cancer development and prognosis. Clin Cancer Res.
11:3647–3653. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Langsenlehner U, Wolf G, Langsenlehner T,
et al: Genetic polymorphisms in the vascular endothelial growth
factor gene and breast cancer risk. The Austrian ‘tumor of breast
tissue: incidence, genetics and environmental risk factors’ study.
Breast Cancer Res Treat. 109:297–304. 2008.PubMed/NCBI
|
|
20
|
Schneider BP, Radovich M, Sledge GW, et
al: Association of polymorphisms of angiogenesis genes with breast
cancer. Breast Cancer Res Treat. 111:157–163. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lu H, Shu XO, Cui Y, et al: Association of
genetic polymorphisms in the VEGF gene with breast cancer survival.
Cancer Res. 65:5015–5019. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Smith KC, Bateman AC, Fussell HM and
Howell WM: Cytokine gene polymorphisms and breast cancer
susceptibility and prognosis. Eur J Immunogenet. 31:167–173. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Gómez-Esquer F, Agudo D, Martinez-Arribas
F, Nuñez-Villar MJ and Schneider J: mRNA expression of the
angiogenesis markers VEGF and CD105 (endoglin) in human breast
cancer. Anticancer Res. 24:1581–1585. 2004.PubMed/NCBI
|
|
24
|
Schneider BP, Wang M, Radovich M, et al;
ECOG 2100. Association of vascular endothelial growth factor and
vascular endothelial growth factor receptor-2 genetic polymorphisms
with outcome in a trial of paclitaxel compared with paclitaxel plus
bevacizumab in advanced breast cancer: ECOG 2100. J Clin Oncol.
26:4672–4678. 2008. View Article : Google Scholar
|