Introduction
Hepatocellular carcinoma (HCC) is one of the most
common types of cancer worldwide (1), and is currently the third leading
cause of cancer-associated death (2). The majority of patients with HCC are
not suitable candidates for surgery as they are diagnosed at an
advanced stage. Chemotherapy with cytotoxic drugs, including
anthracyclines, fluoropyrimidines and platinum complexes, serve a
significant role in the management of terminal HCC. However,
patients with HCC often do not respond to chemotherapy due to the
development of multidrug resistance (MDR). Therefore, research into
the development of a safe and effective MDR reversal agent is
urgently required.
Nucleophosmin (NPM) is a major nucleolar
phosphoprotein that has been implicated in multiple cellular
functions, including ribosomal protein assembly and transport
(3,4), centrosome duplication (5–7),
molecular chaperone activity to prevent protein aggregation
(8,9) and regulating the activity of the
tumor suppressors p53 (10–12)
and p14ARF (13–15).
Previous studies have demonstrated that the level of NPM expression
is markedly increased when cells are committed to mitogenesis
(16,17). In addition, excessive NPM
expression has been linked to cellular transformation and
oncogenesis (18). NPM
overexpression is often observed in human cancers, including those
of the stomach (19), colon
(20), bladder (21), prostate (22), thyroid (23), ovary (24), myeloid and lymphoid cells (25). It has been demonstrated that NPM
overexpression in bladder cancer is independently associated with
recurrence and progression to more advanced stages, which suggests
that overexpression of NPM may be an important prognostic indicator
for cancer recurrence (21). These
findings suggest that NPM may be involved in the regulation of
cellular growth in normal and neoplastic cells. Thus, it may have
potential as a clinical indicator in cancer patients (21). However, it remains unknown whether
NPM may regulate cellular growth in MDR HCC cell lines.
One of the most important and extensively studied
mechanisms of MDR in cancer cells is the efflux mechanism, which is
based on P-glycoprotein (P-gp) function (26,27).
P-gp is a 170 kDa plasma membrane glycoprotein encoded by the human
multidrug resistance gene 1 (MDR1) gene, which functions as an
adenosine triphosphate (ATP)-binding cassette transporter (26). P-gp is a drug efflux pump that
removes a number of chemotherapeutic drugs from MDR cancer cells
(27). In addition to producing
drug resistance at a cellular level, P-gp has also been
demonstrated to alter the pharmacokinetics of numerous drugs and
has been correlated with poor bioavailability (28–30).
Therefore, P-gp inhibition may lead to the reversal of MDR during
treatment with chemotherapeutic agents, and may lead to successful
chemotherapy results in patients with MDR tumors (31). However, the association between
P-gp and NPM in MDR HCC is currently unknown.
In the present study, the authors hypothesized that
downregulated expression of NPM may increase the uptake and
retention of chemotherapeutic agents via the inhibition of MDR1
expression and altered expression of P-gp in MDR HCC cells.
Therefore, the aim of the present study was to investigate the
cellular mechanisms of NPM-mediated reversal of MDR in HCC cells,
which may re-sensitize the MDR HCC cells to chemotherapy. This
novel strategy used the downregulated expression of NPM as a
targeted tool in combination with chemotherapeutic agents for
optimal therapeutic efficacy.
Materials and methods
Cell culture
The human HCC cell lines, HepG2 and SMMC7721, were
purchased from the Institute of Biochemistry and Cell Biology
(Shanghai Institutes for Biological Science, Chinese Academy of
Sciences, Shanghai, China). HepG2 was cultured in Dulbecco's
modified Eagle's medium (DMEM; Hyclone; GE Healthcare Life
Sciences, Logan, UT, USA) and SMMC7721 was cultured in RPMI-1640
(Hyclone; GE Healthcare Life Sciences). The media were supplemented
with 10% fetal bovine serum (FBS; Hyclone; GE Healthcare Life
Sciences).
Multidrug resistance human HCC cell lines, HepG2/ADM
and SMMC7721/ADM, were developed by the Department of General
Surgery, Shanxi Dayi Hospital, Taiyuan, China). HepG2 and SMMC7721
cells were plated in a 6-well plate at a concentration of
2×106 in 2 ml of medium. To develop the HepG2/ADM and
SMMC7721/ADM cells lines, ADM (Shanghai Pharmaceuticals Holding Co.
Ltd, Shanghai, China) was added respectively to HepG2 and SMMC7721
cells at increasing concentrations from 0.01 to 0.2 mg/l over 10
months. MDR was maintained by culturing the cells in the presence
of 0.2 mg/l ADM. MDR HCC cells were termed HepG2/ADM and
SMMC7721/ADM.
Cell viability assay
HepG2, HepG2/ADM, SMMC7721 and SMMC7721/ADM cells
were plated into 96-well plates at the density of 1×104 cells/ml
medium. When the cells were 80% confluent, they were cultured in
the presence of ADM, diamminedichloroplatinum (DDP), fluorouracil
(5-Fu), vincristine sulfate (VCR) or etoposide (VP-16) for 48 h at
37°C in an incubator containing 5% CO2. The cells were
respectively treated with 0, 0.1, 1, 10, 20, 30 mg/l ADM, DDP, VCR
and 0, 1, 10, 20,30, 40 mg/l 5-Fu and VP-16 in the presence of 10%
serum medium. DDP, 5-Fu, VCR and VP-16 were purchased from
Sigma-Aldrich (Merck KGaA, Darmstadt, Germany). In addition, cells
were cultured in the presence of the NPM inhibitor, NSC348884
(Sigma-Aldrich; Merck Millipore, Darmstadt, Germany). Cell lines
HepG2/ADM+NSC348884 and SMMC7721/ADM+NSC348884 were cultured in
DMEM or RPMI-1640 containing 10% FBS and 0.2 mg/l ADM, together
with 1, 2, 3, 4, 5 or 6 µmol/l NSC348884. Cell proliferation was
determined using a cell counting kit-8 assay (CCK-8; Dojindo
Molecular Technologies, Inc., Kumamoto, Japan). A total of 100 ul
cell suspension was added into one well of a 96-well culture plate,
and 10 ul CCK-8 was then added into the well to measure cell
proliferation following medication, and absorbance was measured at
a wavelength of 540 nm on a plate reader (PerkinElmer Wallac 1420
Victor2, Waltham, MA, USA). Data were expressed as the percentage
of the survival of control, calculated from the absorbance and
corrected for background. The half maximal inhibitory concentration
(IC50) was estimated by the dose of drug that resulted
in 50% decrease in cell viability.
Flow cytometric analysis of cell cycle
distribution
Cultured HepG2/ADM and SMMC7721/ADM cells and their
parental cells were collected via trypsinization, washed with
ice-cold PBS, centrifuged at 500 × g for 5 min at 4°C, washed twice
with ice-cold PBS and fixed in 70% ethanol for 2 h at 4°C. Samples
were rehydrated with PBS and the cells were incubated for 30 min at
room temperature with a propidium iodide staining solution in PBS
containing 0.2 mg/ml propidium iodide, 0.2 mg/ml DNAse-free RNAse A
(Roche Diagnostics, Basel, Switzerland), and 0.1% Triton X-100.
Using red propidium DNA fluorescence, 20,000 events were acquired
with an Epics@ XL Beckman Coulter FACS machine (Beckman Coulter
Inc., Brea, CA, USA) for each sample and the percentage of cells in
G0/G1, S and G2/M phases of the cell cycle was calculated using the
System II™ software (Beckman Coulter Inc.) (32).
Western blot analysis
The cells were lysed at 4°C in a lysis buffer (Cell
Signaling Technology Inc., Danvers, MA, USA). The cell lysates were
centrifuged at 21,000 × g for 15 min at 4°C. The protein
concentration in the supernatant was detected using a BCA kit. Then
proteins from tissue homogenate were loaded on sodium dodecyl
sulfate-polyacrylamide gel (12% SDS-PAGE), transferred onto a
polyvinylidene membrane, blocked with bovine serum albumin, and
then incubated using the primary antibodies anti-NPM (catalog no.
3542; 1:1,000; Cell Signaling Technology Inc.), anti-MDR-1 (catalog
no. 13342; 1:1,000; Cell Signaling Technology Inc.), anti-P-gp
(catalog no. A10436R; 1:500; Beijing Solarbio Science &
Technology Co., Ltd, Beijing, China) and anti-β-actin (catalog no.
A10938R; 1:1,000; Beijing Solarbio Science & Technology Co.,
Ltd) at 4°C, overnight. Membranes were washed three times and then
incubated with horseradish peroxide -conjugated secondary antibody
(catalog no. 7074S; 1:2,000; Cell Signaling Technology Inc.) for 40
min at room temperature. Specific antibody binding was detected
using electrochemiluminescence (Chemi Doc XRS+ Imaging system,
Bio-Rad Laboratories, Inc. Hercules, CA, USA). The abundance of
western blot signaling was determined using the image analysis
software (Chemi Doc XRS+ Imaging system, Bio-Rad Laboratories,
Inc.). Western blot analysis was carried out as described
previously (33).
Reverse-transcription-quantitative
polymerase chain reaction (RT-qPCR) analysis
RT-qPCR analysis was performed as described
previously (33). Cells were
plated in a 6-well plate at a concentration of 5×106 in 2 ml of
growth medium. Total RNA was extracted using TRIzol®
(Takara Bio, Inc., Otsu, Japan) according to the manufacturer's
protocol. Two micrograms of total RNA was reverse-transcribed into
first-strand cDNA using Mx3005P. The following primers were used:
NPM forward, 5′-GCAGTCGACGACACCAACATGGAAGATTCGATGGAC-3′ and
reverse, 5′-CGCGTTAACAAGAGACTTCCTCCACTG-3′; MDR1 forward,
5′-GGGGTACCCCAGTCTCTACG-3′ and reverse,
5′-CAAGCTTGTCCGACCTGAAGAG-3′; β-actin forward,
5′-TAAAGGGCATCCTGGGCTACACT-3′ and reverse,
5′-TTACTCCTTGGAGGCCATGTAGG-3′. PCR was performed for 35 cycles,
each cycle was comprised of a denaturation step at 94°C for 45 sec,
annealing at 50°C for 45 sec and extension at 72°C for 45 sec,
prior to a final extension step at 72°C for 10 min. As a control,
the housekeeping gene β-actin was amplified and quantified.
Relative quantification of target gene expression was conducted
using the 2-ΔΔCq method (34). RT-qPCR analysis was repeated >3
times.
Statistical analysis
All of the data were processed using the statistical
software SPSS version 17.0 (SPSS Inc., Chicago, IL, USA). Samples
were analyzed in triplicate, and three independent experiments were
performed. Data are expressed as the mean ± standard deviation, and
differences between two groups were analyzed with the Student's
t-test. P<0.05 was considered to indicate a statistically
significant difference.
Results
Determination of MDR in HepG2/ADM and
SMMC7721/ADM cells
ADM is a chemotherapeutic agent that is used for the
primary treatment of tumors, including HCC (35). In the present study, ADM was
applied to two HCC cell lines to generate MDR HepG2/ADM and
SMMC7721/ADM cells. MDR HCC cell lines were generated over the
course of 10 months. The IC50 values of different
anticancer drugs in HepG2/ADM and SMMC7721/ADM cells were
significantly higher when compared with that of their parental
cells (Table I), and the CCK-8
assay revealed that HepG2/ADM and SMMC7721/ADM were not only
resistant to ADM but also to multiple anticancer drugs, including
DDP, 5-Fu, VCR and VP-16 (Table
I). These results suggested that acquired MDR of HepG2/ADM and
SMMC7721/ADM was successfully established.
 | Table I.Determination of the IC50
values of different anticancer drugs in multidrug-resistant
hepatocellular carcinoma cells. |
Table I.
Determination of the IC50
values of different anticancer drugs in multidrug-resistant
hepatocellular carcinoma cells.
| Anticancer
drug | HepG2 (mg/l) | HepG2/ADM
(mg/l) | HepG2/ADM+NSC348884
(mg/l) | SMMC7721
(mg/l) | SMMC7721/ADM
(mg/l) |
SMMC7721/ADM+NSC348884 (mg/l) |
|---|
| ADM | 0.24±0.07 |
14.45±1.41aa |
1.52±0.28b | 1.58±0.22 |
21.04±1.67c |
8.65±0.62d |
| DDP | 1.31±0.18 |
5.17±0.29a |
2.83±0.19b | 3.5±0.17 |
8.59±0.33cc |
5.12±0.31d |
| 5-Fu | 8.54±0.16 |
34.46±1.39a |
11.69±0.81bb | 6.66±0.26 |
15.97±1.03c |
9.84±0.12dd |
| VCR | 0.48±0.03 |
16.49±1.02aa |
7.82±0.11b | 0.32±0.02 |
12.51±0.6cc |
6.74±0.1dd |
| VP-16 | 2.53±0.14 |
26.38±0.96aa |
17.07±1.24bb | 3.86±0.25 |
28.86±1.76c |
19.1±1.64d |
NPM protein and mRNA levels increased
in HepG2/ADM and SMMC7721/ADM cells when compared with their
parental cells
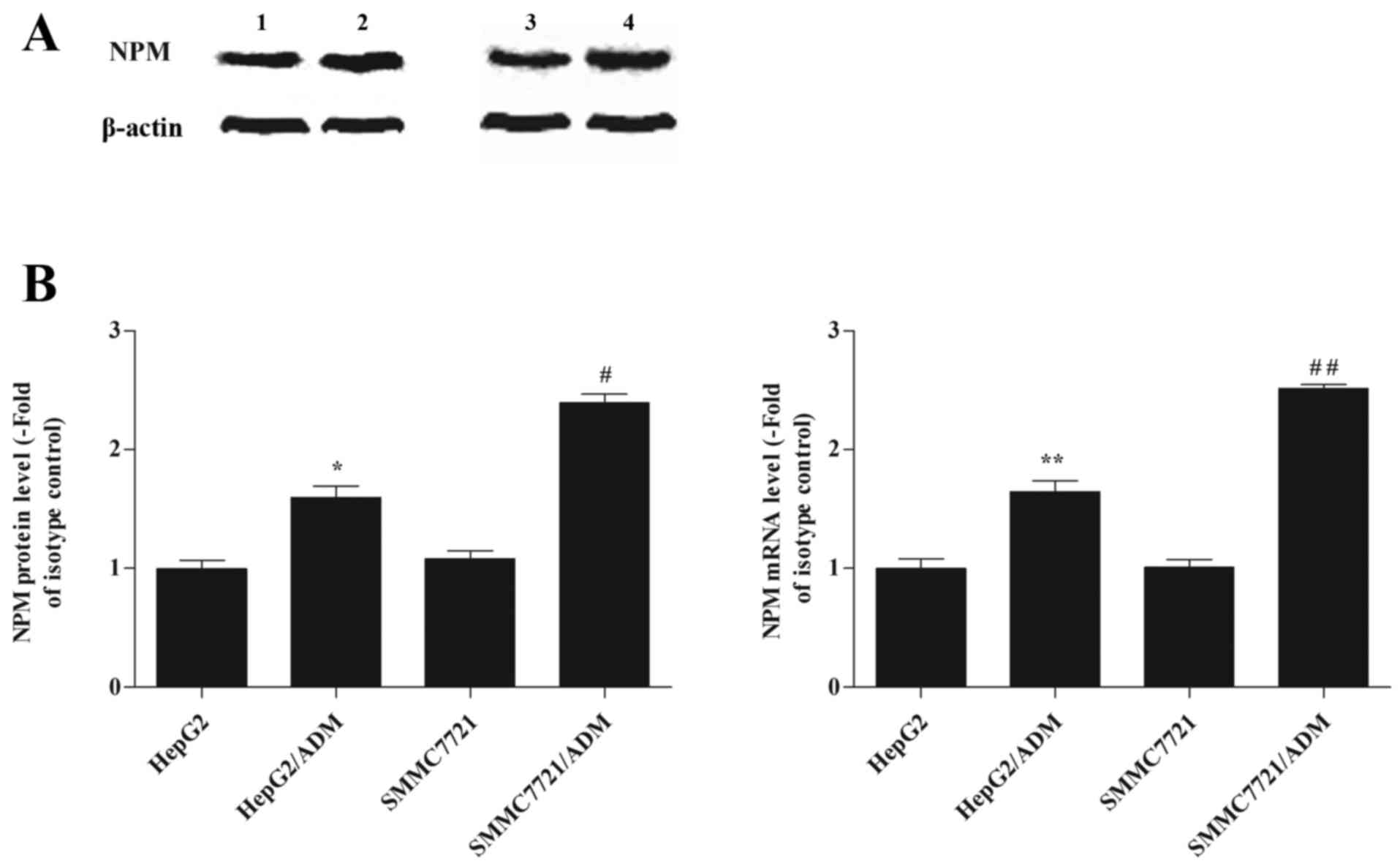
As shown in Fig.
1A, NPM protein levels were significantly higher in the
HepG2/ADM and SMMC7721/ADM cells when compared with their
respective parental cells (1.63±0.18 vs. 0.99±0.25, P<0.05;
2.39±0.19 vs. 1.74±0.09, P<0.05). RT-qPCR analysis demonstrated
that NPM mRNA levels in the HepG2/ADM group were significantly
higher when compared with that of the HepG2 group (1.64±0.23 vs.
1.01±0.2, P<0.01), and the levels in the SMMC7721/ADM group were
significantly higher than that of the SMMC7721 group (2.51±0.08 vs.
1.63±0.07, P<0.01; Fig. 1B).
The results suggested that expression of NPM was upregulated in
HepG2/ADM and SMMC7721/ADM cells when compared to their respective
parental cells, and that these alterations occurred at the
transcriptional level.
MDR-1 protein and mRNA levels
increased in HepG2/ADM and SMMC7721/ADM cells when compared to
their parental cells
Western blotting and RT-qPCR analyses were used to
determine the level of MDR expression in the two cell lines. MDR-1
protein and mRNA levels were significantly increased in the
HepG2/ADM and SMMC7721/ADM cells when compared to their respective
parental cells (MDR-1 protein, HepG2/ADM vs. HepG2, P<0.05;
MDR-1 protein, SMMC7721/ADM vs. SMMC7721, P<0.01; MDR-1 mRNA,
HepG2/ADM vs. HepG2, P<0.01; MDR-1 mRNA, SMMC7721/ADM vs.
SMMC7721, P<0.01; Fig. 2).
MDR-1 protein expression in each lane was normalized to β-actin
expression.
Cell cycle phase distribution was
significantly altered in HepG2/ADM and SMMC7721/ADM cells when
compared to their parental cells
Cell cycle distribution was determined by flow
cytometry analysis to examine differences between MDR HepG2/ADM and
SMMC7721/ADM cells and their respective parental cells. The
percentage of HepG2/ADM cells in the G2/M-phase and
SMMC7721/ADM cells in the S-phase was significantly increased
(G2/M-phase, HepG2/ADM vs. HepG2, P<0.01;
G2/M-phase, SMMC7721/ADM vs. SMMC7721, P<0.05;
S-phase, HepG2/ADM vs. HepG2, P<0.01; S-phase, SMMC7721/ADM vs.
SMMC7721, P<0.05; Table II),
when compared with the parental cells. In addition, the percentage
of HepG2/ADM and SMMC7721/ADM cells were significantly decreased at
the G0/G1 phase (HepG2/ADM vs. HepG2,
P<0.05; SMMC7721/ADM vs. SMMC7721, P<0.01; Table II) when compared with the parental
cells.
 | Table II.Cell cycle distribution of parental
and multidrug-resistant hepatocellular carcinoma cells. |
Table II.
Cell cycle distribution of parental
and multidrug-resistant hepatocellular carcinoma cells.
| Cells |
G0/G1 | S |
G2/M |
|---|
| HepG2 | 66.69±2.26 | 18.27±0.53 | 14.98±0.73 |
| HepG2/ADM |
59.97±1.37a |
12.67±0.29aa |
27.32±1.14aa |
|
HepG2/ADM+NSC348884 |
63.48±1.83cc |
16.38±0.79c |
20.12±1.59c |
| SMMC7721 | 72.25±1.41 | 17.48±0.39 | 6.2±0.64 |
| SMMC7721/ADM |
62.88±1.32bb |
33.32±1.41b |
3.61±0.65b |
|
SMMC7721/ADM+NSC348884 |
68.21±1.04dd |
26.34±1.06dd |
5.43±0.34d |
NSC348884 downregulates NPM
levels
It has been previously reported that NSC348884 is a
specific inhibitor of NPM (36).
NSC348884 was used in the present study to determine whether
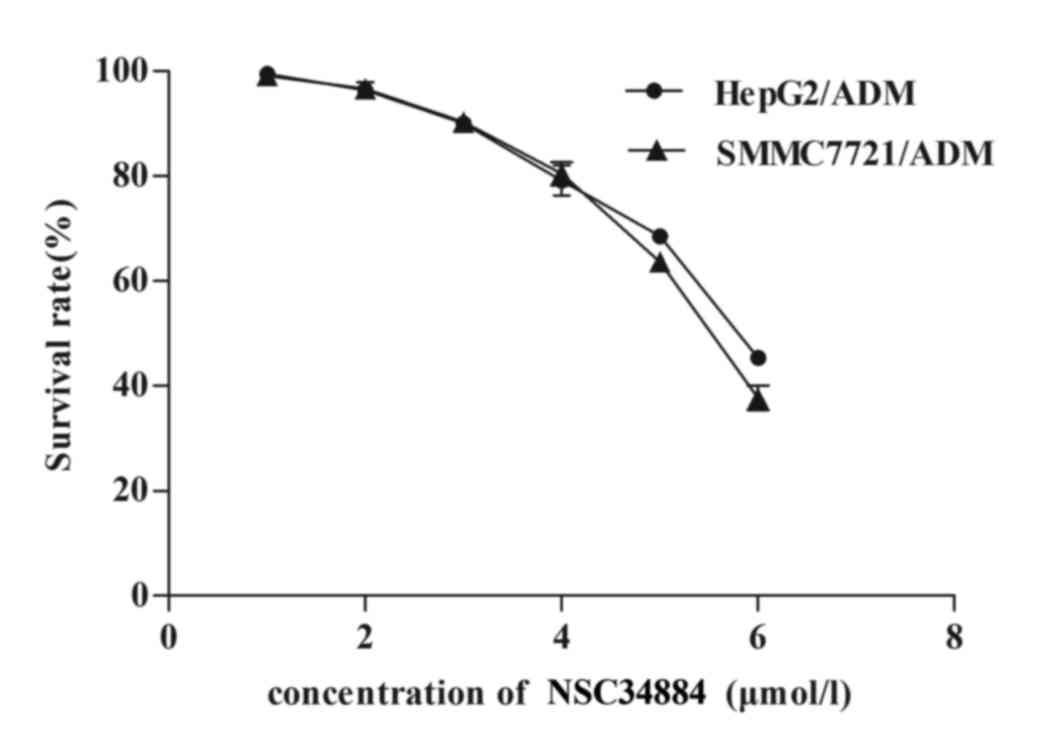
downregulation of NPM reverses the MDR of HCC cell lines. MDR HCC
cells were exposed to a variety of concentrations (1, 2, 3, 4, 5 or
6 µmol/l) of NSC348884. When cultured with ≤3 µmol/l NSC348884,
HepG2/ADM and SMMC7721/ADM cells did not exhibit significant
toxicity (Fig. 3). However, when
cultured with >3 µmol/l NSC348884, the cell survival rate of
HepG2/ADM and SMMC7721/ADM cells markedly decreased (Fig. 3). As shown in Fig. 4, pretreatment of HepG2/ADM and
SMMC7721/ADM cells with NSC348884, significantly decreased NPM
protein and mRNA expression when compared to that of the parental
cells (NPM protein, HepG2/ADM+NSC348884 vs. HepG2/ADM, P<0.05;
NPM protein, SMMC7721/ADM+NSC348884 vs. SMMC7721/ADM, P<0.01;
NPM mRNA, HepG2/ADM+NSC348884 vs. HepG2/ADM, P<0.01; NPM mRNA,
SMMC7721/ADM+NSC348884 vs. SMMC7721/ADM, P<0.01; Fig. 4).
NSC348884 reversed MDR in HepG2/ADM
and SMMC7721/ADM cells
As demonstrated in Table I, HepG2/ADM and SMMC7721/ADM were
resistant to ADM, as well as DDP, 5-Fu, VCR and VP16 anticancer
drugs. The IC50 values were 5.17±0.29 and 34.46±1.39
mg/l in HepG2/ADM cells treated with DDP and 5-Fu, respectively,
and 8.59±0.33 and 15.97±1.03 mg/l in SMMC7721/ADM treated with DDP
and 5-Fu, respectively (Table I).
HepG2 and SMMC7721 cells were more sensitive to these drugs, with
IC50 values of 1.31±0.18 and 8.54±0.16 mg/l in HepG2
cells treated with DDP and 5-Fu, respectively, and 3.5±0.17 and
6.66±0.26 mg/l in SMMC7721 cells treated with DDP and 5-Fu,
respectively. Pretreatment of HepG2/ADM and SMMC7721/ADM cells with
3 µmol/l NSC348884 was associated with increased sensitivity to
these agents. The IC50 values were 2.83±0.19 and
11.69±0.81 mg/l in HepG2/ADM+NSC348884 cells treated with DDP and
5-Fu, respectively, and 5.12±0.31 and 9.84±0.12 mg/l in
SMMC7721/ADM+NSC348884 cells treated with DDP and 5-Fu,
respectively (Table I). In
addition, pretreatment of HepG2/ADM and SMMC7721/ADM cells with 3
µmol/l NSC348884, was associated with a significant decrease in
MDR-1 protein and mRNA levels in the HepG2/ADM+NSC348884 and
SMMC7721/ADM+NSC348884 cells when compared with the HepG2/ADM and
SMMC7721/ADM cells (MDR-1 protein, HepG2/ADM+NSC348884 vs.
HepG2/ADM, P<0.05; MDR-1 protein, SMMC7721/ADM+NSC348884 vs.
SMMC7721/ADM, P<0.01; MDR-1 mRNA, HepG2/ADM+NSC348884 vs.
HepG2/ADM, P<0.01; MDR-1 mRNA, SMMC7721/ADM+NSC348884 vs.
SMMC7721/ADM, P<0.01; Fig. 5).
The quantity of product in each lane was normalized to β-actin
expression. Alterations in the cell cycle distribution of HepG2/ADM
and SMMC7721/ADM cells were significantly reversed following
treatment with NSC348884 (Table
II). The percentage of HepG2/ADM cells in G2/M-phase
and SMMC7721/ADM cells in S-phase was significantly increased, when
compared with the parental cells. In addition, the percentage of
HepG2/ADM and SMMC7721/ADM cells were significantly decreased at
the G0/G1 phase when compared with the
parental cells. These results suggest that NSC348884 may reverse
the MDR of HepG2/ADM and SMMC7721/ADM cells.
The effect of NPM on P-gp
expression
In order to investigate the effect of NPM on P-gp
expression, western blot analysis was performed (Fig. 6). It was revealed that P-gp
expression was significantly higher in HepG2/ADM and SMMC7721/ADM
cells when compared with the parental cells (P<0.01 and
P<0.01, respectively; Fig. 6).
By contrast, when HepG2/ADM and SMMC7721/ADM cells were pretreated
with NSC348884, P-gp expression was significantly reduced
(P<0.01 and P<0.01, respectively; Fig. 6).
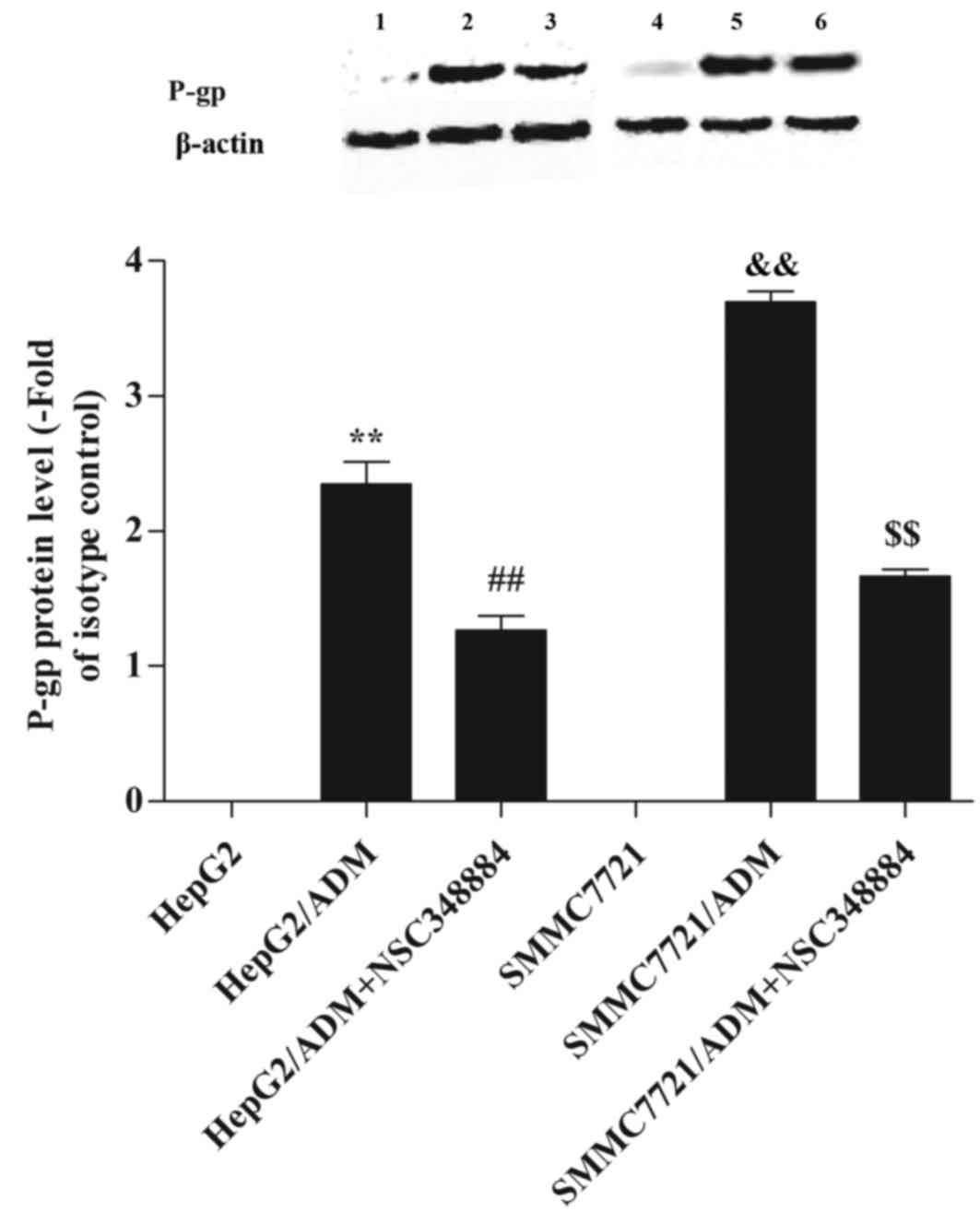 | Figure 6.Effect of nucleophosmin on P-gp
expression. A high level of P-gp expression was detected in
HepG2/ADM and SMMC7721/ADM cells. However, when HepG2/ADM and
SMMC7721/ADM cells were pretreated with NSC348884, the P-gp level
was significantly decreased. Lane 1, HepG2; lane 2, HepG2/ADM; lane
3, HepG2/ADM+NSC348884; lane 4, SMMC7721; lane 5, SMMC7721/ADM;
lane 6, SMMC7721/ADM+NSC348884. **P<0.01 vs. HepG2;
##P<0.01 vs. HepG2/ADM;
&&P<0.01 vs. SMMC7721 and
$$P<0.01 vs. SMMC7721/ADM. ADM, Adriamycin; P-gp,
P-glycoprotein. |
Discussion
MDR is characterized by the development of
anticancer drug resistance, which may lead to the development of
resistance to other pharmacokinetic and structurally unrelated
drugs (37,38). For a number of years, MDR has been
a major issue for scientists and clinicians in the treatment of
cancer, however an effective solution has remained elusive.
Due to the difficulties encountered in the reversal
of MDR, alternative methods to overcome MDR in cancer cells are
continuously being investigated. Recently, NPM has received
significant interest due to its association with ADM-resistant
cells (39). NPM, also known as
B23, NO38 or Numatrin, is a 38-kDa estrogen-regulated nucleolar
phosphoprotein that shuttles between the nucleus and cytoplasm
(40). NPM function has been
implicated in a number of cellular processes, including ribosome
shuttling between precursor proteins in the cytoplasm and nucleus,
nuclear protein chaperone activity, the maintenance of genomic
stability and the indirect regulation of growth and proliferation
(39,41,42).
NPM overexpression has frequently been associated with tumor
progression, and may be a marker for some cancers, including
gastric, ovarian, prostate (42)
and Ewing's sarcoma (43). A
previous study reported that NPM was highly expressed in human MDR
gastric cancer cell lines (44),
radiotherapy-resistant HeLa cells (45,46)
and in the MCF-7 breast cancer cell line where upregulated NPM
expression enhances interferon regulating factor 1 mediated
estrogen-resistance (47).
However, the effect of NPM downregulation on the reversal of MDR in
HCC, as well as the underlying molecular mechanisms involved,
remain unknown. Therefore, the present study was designed to
investigate the effect of NPM downregulation on MDR, and the
molecular mechanisms involved in this process. The results
demonstrated that NPM expression was significantly increased in MDR
HCC cell lines when compared to that of their parental cells.
To identify the mechanisms involved in the
development of MDR in HepG2/ADM and SMMC7721/ADM cells, the cell
cycle distribution was analyzed by flow cytometry. The percentage
of MDR HepG2/ADM and SMMC7721/ADM cells was markedly decreased in
the G0/G1 phase and increased in the S and
G2/M phases when compared to their parental cells. This
may have been responsible for the reduced cell proliferation
ability (date not shown). In addition, delayed cell cycle
progression may facilitate the removal of specific cytotoxic agents
from the cell, thus leading to MDR in the cells.
An improved understanding of the possible molecular
mechanisms and signaling pathways involved in MDR is important to
overcome MDR and improve chemotherapeutic efficacy in patients with
HCC. Multiple hypotheses have been proposed regarding the
mechanisms underlying the development of MDR, including the
involvement of P-gp, which is encoded by the MDR1 gene (26). Previous studies have demonstrated
that P-gp relies on the actin cytoskeleton for its localization in
lipid rafts on the cell membrane, thereby influencing drug influx
and possibly counteracting uptake (48,49).
The action of P-gp as a drug efflux pump for therapies including
ADM, docetaxel, paclitaxel and daunorubicin (50), has led to the development of
chemosensitizing agents including cyclosporine, verapamil and
quinine, which competitively or noncompetitively inhibit this
protein (51). The expression of
P-gp is increased in drug-resistant tumors of the colon, kidney and
adrenal gland, as well as in some tumors that have acquired MDR
following chemotherapy (52).
Excessive P-gp has been demonstrated to bind and transport
anticancer drugs through ATP-dependent anticancer drug efflux
pumps, leading to an increased efflux of the anticancer agent from
the cancer cells, and a lower intracellular concentration (26,31,53).
The results of the present study demonstrated that P-gp expression
was increased in MDR HepG2/ADM and SMMC7721/ADM cells when compared
with their parental cells, indicating that MDR of HepG2/ADM and
SMMC7721/ADM cells may be attributed to the overexpression of
P-gp.
In order to further explore the role of NPM in the
HCC MDR cell lines, NSC348884, a specific inhibitor of NPM, was
applied to investigate whether downregulation of NPM may reverse
MDR in HCC cell lines. Previous studies have demonstrated that
application of NSC348884 suppressed the proliferation of prostate,
colon, breast, lung and lymphoma tumor cells, thereby enhancing ADM
sensitivity (36). Following
NSC348884 treatment of MDR HCC cells in the present study, cellular
resistance to anticancer drugs was reversed, and corresponding
alterations in the cell cycle distributions were observed. Further
experiments suggested that NSC348884 may reverse MDR, via
inhibition of P-gp function.
NSC348884 significantly reversed HCC MDR in the
present study. The results implied that NSC348884 may be effective
in reversing MDR in vitro. In addition, RT-qPCR and western
blot analysis revealed that the expression of P-gp at the mRNA and
protein level were decreased. Reduced expression of P-gp at the
transcriptional and translational levels has been proposed to be
one of the mechanisms for certain modulators or agents to reverse
the MDR phenotype (54).
In conclusion, the results of the present study have
provided evidence demonstrating that NPM protein and mRNA levels
were increased in HepG2/ADM and SMMC7721/ADM cells when compared to
that of their parental cells. In addition, treatment of cells with
a specific inhibitor of NPM (NSC348884) was able to reverse the MDR
of HepG2/ADM and SMMC7721/ADM cells, potentially via the
downregulation of P-gp expression. The results suggest that NPM may
be involved in MDR of HCC. It is a novel MDR reversal agent and may
be a potential adjuvant agent for tumor chemotherapy. However,
further research is required to optimize NPM exposure, and to
determine the mechanisms underlying how downregulation of NPM leads
to enhanced sensitivity of MDR HCC cells to anticancer drugs.
Acknowledgements
The present study was supported by grants from the
Nature Scientific Foundation of Shanxi Province (grant no.
2011021035-3) and the Scientific Foundation of Shanxi Provincial
Health Department (grant no. 200810).
Glossary
Abbreviations
Abbreviations:
|
MDR
|
multidrug resistance
|
|
MDR1
|
multidrug resistance gene 1
|
|
P-gp
|
P-glycoprotein
|
|
HCC
|
hepatocellular carcinoma
|
|
NPM
|
nucleophosmin
|
References
|
1
|
Schlageter M, Terracciano LM, D'Angelo S
and Sorrentino P: Histopathology of hepatocellular carcinoma. World
J Gastroenterol. 20:15955–15964. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Gu W, Fang FF, Li B, Cheng BB and Ling CQ:
Characterization and resistance mechanisms of A 5-fluorouracil
resistance hepatocellular carcinoma cell line. Asian Pac J Cancer
Prev. 13:4807–4814. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Verheggen C, Almouzni G and
Hernandez-Verdun D: The ribosomal RNA processing machinery is
recruited to the nucleolar domain before RNA polymerase I during
Xenopus laevis development. J Cell Biol. 149:293–306. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Huang N, Negi S, Szebeni A and Olson MO:
Protein NPM3 interacts with the multifunctional nucleolar protein
B23/nucleophosmin and inhibits ribosome biogenesis. J Biol Chem.
280:5496–5502. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Okuda M, Horn HF, Tarapore P, Tokuyama Y,
Smulian AG, Chan PK, Knudsen ES, Hofmann IA, Snyder JD, Bove KE and
Fukasawa K: Nucleophosmin/B23 is a target of CDK2/cyclin E in
centrosome duplication. Cell. 103:127–140. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Okuda M: The role of nucleophosmin in
centrosome duplication. Oncogene. 21:6170–6174. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Grisendi S, Bernardi R, Rossi M, Cheng K,
Khandker L, Manova K and Pandolfi PP: Role of nucleophosmin in
embryonic development and tumorigenesis. Nature. 437:147–153. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hingorani K, Szebeni A and Olson MO:
Mapping the functional domains of nucleolar protein B23. J Biol
Chem. 275:24451–24457. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Szebeni A, Hingorani K, Negi S and Olson
MO: Role of protein kinase CK2 phosphorylation in the molecular
chaperone activity of nucleolar protein b23. J Biol Chem.
278:9107–9115. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Colombo E, Marine JC, Danovi D, Falini B
and Pelicci PG: Nucleophosmin regulates the stability and
transcriptional activity of p53. Nat Cell Biol. 4:529–533. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li J, Zhang X, Sejas DP, Bagby GC and Pang
Q: Hypoxia-induced nucleophosmin protects cell death through
inhibition of p53. J Biol Chem. 279:41275–41279. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Maiguel DA, Jones L, Chakravarty D, Yang C
and Carrier F: Nucleophosmin sets a threshold for p53 response to
UV radiation. Mol Cell Biol. 24:3703–3711. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Itahana K, Bhat KP, Jin A, Itahana Y,
Hawke D, Kobayashi R and Zhang Y: Tumor suppressor ARF degrades
B23, a nucleolar protein involved in ribosome biogenesis and cell
proliferation. Mol Cell. 12:1151–1164. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bertwistle D, Sugimoto M and Sherr CJ:
Physical and functional interactions of the Arf tumor suppressor
protein with nucleophosmin/B23. Mol Cell Biol. 24:985–996. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Brady SN, Yu Y, Maggi LB Jr and Weber JD:
ARF impedes NPM/B23 shuttling in an Mdm2-sensitive tumor suppressor
pathway. Mol Cell Biol. 24:9327–9338. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Feuerstein N and Mond JJ: ‘Numatrin,’ a
nuclear matrix protein associated with induction of proliferation
in B lymphocytes. J Biol Chem. 262:11389–11397. 1987.PubMed/NCBI
|
|
17
|
Feuerstein N, Spiegel S and Mond JJ: The
nuclear matrix protein, numatrin (B23), is associated with growth
factor-induced mitogenesis in Swiss 3T3 fibroblasts and with T
lymphocyte proliferation stimulated by lectins and anti-T cell
antigen receptor antibody. J Cell Biol. 107:1629–1642. 1988.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Pulford K, Morris SW and Mason DY:
Anaplastic lymphoma kinase proteins and malignancy. Curr Opin
Hematol. 8:231–236. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Tanaka M, Sasaki H, Kino I, Sugimura T and
Terada M: Genes preferentially expressed in embryo stomach are
predominantly expressed in gastric cancer. Cancer Res.
52:3372–3377. 1992.PubMed/NCBI
|
|
20
|
Nozawa Y, Van Belzen N, van der Made AC,
Dinjens WN and Bosman FT: Expression of nucleophosmin/B23 in normal
and neoplastic colorectal mucosa. J Pathol. 178:48–52. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Tsui KH, Cheng AJ, Chang Pe, Pan TL and
Yung BY: Association of nucleophosmin/B23 mRNA expression with
clinical outcome in patients with bladder carcinoma. Urology.
64:839–844. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Subong EN, Shue MJ, Epstein JI, Briggman
JV, Chan PK and Partin AW: Monoclonal antibody to prostate cancer
nuclear matrix protein (PRO:4-216) recognizes nucleophosmin/B23.
Prostate. 39:298–304. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Onda M, Emi M, Yoshida A, Miyamoto S,
Akaishi J, Asaka S, Mizutani K, Shimizu K, Naqahama M, Ito K, et
al: Comprehensive gene expression profiling of anaplastic thyroid
cancer with cDNA microarray of 25 344 genes. Endocr Relat Cancer.
11:843–854. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang Y: The ARF-B23 connection:
implications for growth control and cancer treatment. Cell Cycle.
3:259–262. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Schnittger S, Schoch C, Kern W, Mecucci C,
Tschulik C, Martelli MF, Haferlach T, Hiddemann W and Falini B:
Nucleophosmin gene mutations are predictors of favorable prognosis
in acute myelogenous leukemia with a normal karyotype. Blood.
106:3733–3739. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Gottesman MM, Fojo T and Bates SE:
Multidrug resistance in cancer: Role of ATP-dependent transports.
Nat Rev Cancer. 2:48–58. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ambudkar SV, Kimchi-Sarfaty C, Sauna ZE
and Gottesman MM: P-glycoprotein: From genomics to mechanism.
Oncogene. 22:7468–7485. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Glavinas H, Krajcsi P, Cserepes J and
Sarkadi B: The role of ABC transporters in drug resistance,
metabolism and toxicity. Curr Drug Deliv. 1:27–42. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Varma MV, Ashokraj Y, Dey CS and
Panchagnula R: P-glycoprotein inhibitors and their screening: A
perspective from bioavailability enhancement. Pharmacol Res.
48:347–359. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Johnson WW: P-glycoprotein-mediated efflux
as a major factor in the variance of absorption and distribution of
drugs: Modulation of chemotherapy resistance. Methods Find Exp Clin
Pharmacol. 24:501–514. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Fojo T and Bates S: Strategies for
reversing drug resistance. Oncogene. 22:7512–7523. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Nguyen T, Zhang XD and Hersey P: Relative
resistance of fresh isolates of melanoma to tumor necrosis
factor-related apoptosis-inducing ligand (TRAIL)-induced apoptosis.
Clin Cancer Res. 7:966s–973s. 2001.PubMed/NCBI
|
|
33
|
Li X, Zhao H, Wu Y, Zhang S, Zhao X, Zhang
Y, Wang J, Wang J and Liu H: Up-regulation of hypoxia-inducible
factor-1α enhanced the cardioprotective effects of ischemic
postconditioning in hyperlipidemic rats. Acta Biochim Biophys Sin
(Shanghai). 46:112–118. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Minotti G, Menna P, Salvatorelli E, Cairo
G and Gianni L: Anthracyclines: Molecular advances and
pharmacologic developments in antitumor activity and
cardiotoxicity. Pharmacol Rev. 56:185–229. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Qi W, Shakalya K, Stejskal A, Goldman A,
Beeck S, Cooke L and Mahadevan D: NSC348884, a nucleophosmin
inhibitor disrupts oligomer formation and induces apoptosis in
human cancer cells. Oncogene. 2:4210–4220. 2008. View Article : Google Scholar
|
|
37
|
Pérez-Tomás R: Multidrug resistance:
Retrospect and prospects in anti-cancer drug treatment. Curr Med
Chem. 13:1859–1876. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Daniel C, Bell C, Burton C, Harguindey S,
Reshkin SJ and Rauch C: The role of proton dynamics in the
development and maintenance of multidrug resistance in cancer.
Biochim Biophys Acta. 1832:606–617. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Yung BY: Oncogenic role of
nucleophosmin/B23. Chang Gung Med J. 30:285–293. 2007.PubMed/NCBI
|
|
40
|
Lam L, Aktary Z, Bishay M, Werkman C, Kuo
CY, Heacock M, Srivastava N, Mackey JR and Pasdar M: Regulation of
subcellular distribution and oncogenic potential of nucleophosmin
by plakoglobin. Oncogenesis. 1:e42012. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Skaar TC, Prasad SC, Sharareh S, Lippman
ME, Brunner N and Clarke R: Two-dimensional gel electrophoresis
analyses identify nucleophosmin as an estrogen regulated protein
associated with acquired estrogen-independence in human breast
cancer cells. J Steroid Biochem Mol Biol. 67:391–402. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Grisendi S, Mecucci C, Falini B and
Pandolfi PP: Nucleophosmin and cancer. Nat Rav Cancer. 6:493–505.
2006. View Article : Google Scholar
|
|
43
|
Kikuta K, Tochigi N, Shimoda T, Yabe H,
Morioka H, Toyama Y, Hosono A, Beppu Y, Kawai A, Hirohashi S and
Kondo T: Nucleophosmin as a candidate prognostic biomarker of
Ewing's sarcoma revealed by proteomics. Clin Cancer Res.
15:2885–2894. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Yang YX, Hu HD, Zhang DZ and Ren H:
Identification of proteins responsible for the development of
adriamycin resistance in human gastric cancer cells using
comparative proteomics analysis. J Biochem Mol Biol. 40:853–860.
2007.PubMed/NCBI
|
|
45
|
Wu MH, Chang JH, Chou CC and Yung BY:
Involvement of nucleophosmin-B23 in the response of HeLa cells to
UV irradiation. Int J Cancer. 97:297–305. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Wu MH, Chang JH and Yung BY: Resistance to
UV-induced cell-killing in nucleophosmin/B23 over-expressed NIH3T3
fibroblasts: Enhancement of DNA repair and up-regulation of PCNA in
association with nucleophosmin/B23 over-expression. Carcinogenesis.
23:93–100. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Clarke R, Liu MC, Bouker KB, Gu Z, Lee RY,
Zhu Y, Skaar TC, Gomez B, O'Brien K, Wang Y, et al: Antiestrogen
resistance in breast cancer and the role of estrogen receptor
signaling. Oncogene. 22:7316–7339. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zhang L, Xiao R, Xiong J, Leng J, Ehtisham
A, Hu Y, Ding Q, Xu H, Liu S, Wang J, et al: Activated ERM Protein
plays a critical role in drug resistance of MOLT4 cells induced by
CCL25. PLoS One. 8:e523842013. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Meszaros P, Hummel I, Klappe K, Draghiciu
O, Hoekstra D and Kok JW: The fuction of the ATP-bnding cassette
(ABC) transporter ABCB1 is not susceptible to actin disruption.
Biochim Biophys Acta. 1828:340–351. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Avendaño C and Menéndez JC: Inhibitors of
multidrug resistance to antitumor agents (MDR). Curr Med Chem.
9:159–193. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Leonard GD, Fojo T and Bates SE: The role
of ABC transporters in clinical practice. Oncologist. 8:411–424.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Tsuruo T: Molecular cancer therapeutics:
Recent progress and targets in drug resistance. Intern Med.
42:237–243. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Ozben T: Mechanism and strategies to
overcome multiple drug resistance in cancer. FEBS Lett.
580:2903–2909. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Hu YP, Pourquer P, Doignon F, Crouzet M
and Robert J: Effects of modulators of multidrug resistance on the
expression of the MDR1 gene in human KB cells in culture.
Anticancer Drugs. 7:738–744. 1996. View Article : Google Scholar : PubMed/NCBI
|