Introduction
Colorectal cancer (CRC), a commonly diagnosed
cancer, was associated with a high incidence and mortality rates
worldwide in 2012. In addition, at present, its incidence and
mortality rates are increasing (1). In China, CRC was the fifth most
common cancer type, with high incidence and mortality rates in 2011
(2). Currently, treatment of
patients with relapsed and metastatic CRC is challenging (3). Therefore, there is an urgent
requirement for novel early diagnostic markers and therapeutic
targets for CRC treatment.
Accumulating evidence has demonstrated that
abnormally expressed long non-coding RNAs (lncRNAs) in cancer may
regulate the proliferation, apoptosis, metastasis and
differentiation of cancer cells by regulating epigenetic, splicing,
transcriptional or post-transcriptional processes and subsequently
tumorigenesis and tumor progression (4). In renal cell carcinoma, lncRNA
(lnc)FTX and lncTUG1 promote cell proliferation, while lncGAS-5
promotes cell apoptosis and inhibits cell proliferation. In
addition, certain lncRNAs, including HOTAIR, MALAT1 and linc00152,
perform multiple functions, including roles in the promotion of
cell proliferation, invasion, migration and metastasis (5). In hepatocellular carcinoma, lncRNAs
DANCR, Lalr1 and UFC1 are reported to regulate Wnt signaling to
promote or inhibit cancer cell invasion, migration and metastasis,
and lncRNAs UCA1 and lncSOX4 were demonstrated to promote cancer
cell proliferation (6). In
melanoma, lncRNAs Lime23, BANCR, ANRIL and UCA1 regulate cancer
cell proliferation and growth, and lncRNAs GAS5, PRC2 and PAUPAR
modulate cancer cell invasion, migration and metastasis (7). Previous studies have demonstrated
that various lncRNAs, including CCAT1, H19, FEZF-1-AS1, MALAT1 and
UCA1, enhance the proliferation, invasion and metastasis of CRC
cells (8).
LncTCF7 binds to the T-cell factor 7 (TCF7) gene and
recruits the Switch/sucrose non-fermentable (SWI/SNF) complex to
the TCF7 promoter to upregulate TCF7 expression and activate Wnt
signaling (9). TCF7 serves an
important role in T-cell development and multipotential
hematopoietic cell self-renewal by activating Wnt signaling
(10,11). TCF7 expression is reported to be
significantly higher in clear cell renal cell carcinoma tissue
compared with normal tissue (12),
while TCF7 silencing inhibited the survival of prostate cancer
cells and inhibited the development of androgen deprivation
resistance in prostate cancer (13,14).
TCF7 expression was reported to be increased in human osteosarcoma,
and microRNA-192 suppressed TCF7 expression to inhibit cell growth
and invasion, and induce apoptosis (15). Furthermore, TCF7 promoted
tumorigenesis following the ocular transplantation of embryonic
stem cell-derived retinal progenitor cells (11). These results from previous studies
indicate that lncTCF7 may promote the expression levels of
oncogenic TCF7 and may therefore be an oncogene. In addition,
previous studies have revealed that lncTCF7 promotes the
self-renewal of human cancer stem cells and enhances the
aggressiveness of human non-small cell lung cancer and
hepatocellular carcinoma cells (9,16,17).
However, the effect of lncTCF7 on CRC tumorigenesis and progression
remains unclear. In the current study, lncTCF7 expression in CRC
tissues was investigated, and the association between lncTCF7
expression and different clinicopathological features of patients
with CRC was determined. Furthermore, the effect of lncTCF7
silencing on the proliferation, cell cycle, apoptosis, migration
and invasion of CRC cells was investigated. The results
demonstrated that lncTCF7 predicted the progression, and promoted
the growth and metastasis, of CRC, indicating that lncTCF7 may be a
novel diagnostic marker and a potential therapeutic target for
CRC.
Materials and methods
Patients and tissue samples
In total, 112 patients with CRC were recruited from
Yantai City Hospital of Traditional Chinese Medicine (Yantai,
China) between March 2014 and December 2016. The sex and age
distribution of patients is presented in Table I. The CRC patients age ranged from
31–72 years with a median of 56 years. None of the recruited
patients with CRC received chemotherapy or radiation therapy prior
to surgery. Diagnosis of CRC and clinicopathological
characteristics of the patients were confirmed by two pathologists.
All participants signed informed consent forms and experiments were
approved by the Ethics Committee of the Yantai City Hospital of
Traditional Chinese Medicine. CRC tissues and adjacent normal
tissues (at least 2 cm away from the primary cancer site) were
obtained by performing radical resection and were immediately
stored at −80°C until use for total RNA extraction.
 | Table I.Association between
clinicopathological characteristics and lncRNA TCF7 expression in
patients with CRC. |
Table I.
Association between
clinicopathological characteristics and lncRNA TCF7 expression in
patients with CRC.
|
|
| lncRNA TCF7
expression |
|
|---|
|
|
|
|
|
|---|
| Clinicopathological
characteristics | All patients,
n=112 | Low group, n=56 | High group, n=56 | P-value |
|---|
| Sex |
|
|
| 0.705 |
| Male | 54 | 26 | 28 |
|
|
Female | 58 | 30 | 28 |
|
| Age, years |
|
|
| 0.674 |
|
<50 | 40 | 19 | 21 |
|
| ≥50 | 72 | 37 | 35 |
|
| Location |
|
|
| 0.693 |
|
Colon | 70 | 34 | 36 |
|
|
Rectum | 42 | 22 | 20 |
|
| Organization |
|
|
| 0.674 |
|
Adenocarcinoma | 48 | 22 | 26 |
|
| Mucinous
carcinoma | 64 | 34 | 30 |
|
| Tumor size, cm |
|
|
| 0.001a |
|
<5 | 73 | 45 | 28 |
|
| ≥5 | 39 | 11 | 28 |
|
| Differentiation |
|
|
| 0.005a |
| Poor | 29 | 8 | 21 |
|
| Moderate
+ high | 83 | 48 | 35 |
|
| Lymph node
metastasis |
|
|
| 0.010a |
| Yes | 23 | 6 | 17 |
|
| No | 89 | 50 | 39 |
|
| Depth of
invasion |
|
|
| 0.027a |
| Serosa
layer | 37 | 13 | 24 |
|
| Muscular
layer | 75 | 43 | 32 |
|
| TNM grade |
|
|
| 0.004a |
| I+II | 65 | 40 | 25 |
|
|
III+IV | 47 | 16 | 31 |
|
RNA extraction and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
To perform RT-qPCR, total RNA was extracted from CRC
tissues and adjacent normal tissues, and CRC cell lines, using
TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.,
Waltham, MA, USA), according to the manufacturer's protocol.
Subsequently, mRNA was reverse transcribed into cDNA at 37°C for 15
min and at 85°C for 5 sec using a PrimeScript RT Reagent kit with
gDNA Eraser (Takara Biotechnology Co., Ltd., Dalian, China),
according to the manufacturer's protocol. LncTCF7 expression was
determined by performing qPCR with SYBR Premix Ex Taq (Takara
Biotechnology Co., Ltd.) and a 7500 Real-Time PCR system (Applied
Biosystems; Thermo Fisher Scientific, Inc.). Thermocycling
conditions for qPCR was as follow: Initial denaturation at 95°C for
5 min, followed by 40 cycles of denaturation at 95°C for 10 sec and
combined annealing/extension at 60°C for 30 sec. Sequences of
primers used for performing qPCR were as follows: lncTCF7 forward,
5′-AGGAGTCCTTGGACCTGAGC-3′ and reverse,
5′-AGTGGCTGGCATATAACCAACA-3′; GAPDH forward,
5′-CCCATCACCATCTTCCAGGAG-3′ and reverse,
5′-GTTGTCATGGATGACCTTGGC-3′. GAPDH was used as an internal control
for normalizing lncTCF7 mRNA expression. Relative lncTCF7
expression was expressed as fold induction by using the
2−ΔΔCq method (18).
All samples were assessed in triplicate.
Cell culture and small interfering
(si)RNA transfection
SW-620 and HT29 CRC cell lines (American Type
Culture Collection, Manassas, VA, USA) were cultured in RPMI-1640
(Gibco; Thermo Fisher Scientific, Inc.) supplemented with 10% fetal
bovine serum (Gibco; Thermo Fisher Scientific, Inc.), 100 U/ml
penicillin and 100 µg/ml streptomycin at 37°C in a humidified
atmosphere of 5% CO2. LncTCF7 siRNA (si-lncTCF7-1,
5′-AGCCAACATTGTTGGTTAT-3′; and si-lncTCF7-2,
5′-CACCTAGGTGCTCACTGAA-3′) and negative control siRNA (si-NC,
5′-UUCUCCGAACGUGUCACGUTT-3′) were purchased from Guangzhou RiboBio
Co., Ltd., (Guangzhou, China). SW-620 and HT29 cells
(3×105) were transfected with the siRNAs (50 nM) using
Lipofectamine 2000 (Invitrogen; Thermo Fisher Scientific, Inc.),
according to the manufacturer's protocol. A total of 48 h following
transfection, lncTCF7 expression was determined by performing
RT-qPCR.
Proliferation assay
A cell proliferation assay was performed using Cell
Counting Kit (CCK)-8 assay kit reagent (Beyotime Institute of
Biotechnology, Haimen, China), according to the manufacturer's
protocol. At 24 h following transfection, Cells in si-negative
control (NC) and si-lncTCF7 groups were trypsinized and cultured in
96-well plates in triplicate at a density of 5×103 cells
per well at 37°C in a humidified atmosphere of 5% CO2.
Following culture for 0, 24, 48 and 72 h, 10 µl CCK-8 reagent was
added to each well and the cells were incubated for a further 4 h
at 37°C. Absorbance was measured at a wavelength of 450 nm using a
microplate reader.
Apoptosis and cell cycle assay
Cell apoptosis assay was performed using the Annexin
V-FITC apoptosis detection kit (Nanjing KeyGen Biotech Co., Ltd.,
Nanjing, China). For investigating apoptosis, SW-620 and HT29 cells
were digested with trypsin and were centrifuged at 200 × g for 5
min at 4°C 48 h after transfection. Subsequently, cell pellets were
resuspended in 500 µl binding buffer, which was followed by the
addition of 5 µl Annexin V-fluorescein isothiocyanate and 5 µl
propidium iodide (PI) to the cell suspension, and the cells were
incubated in the dark for 15 min at room temperature. Cell
apoptosis was detected by performing flow cytometry using a BD
FACSCalibur system and data was analyzed using the CellQuest
software version 5.1 (BD Biosciences, Franklin Lakes, NJ, USA). For
the cell cycle assay, SW-620 and HT29 cells were fixed in 500 µl
70% pre-cooled ethanol at 4°C overnight. An equal amount of PBS was
added twice for washing. A total of 100 µl RNase A was added at
37°C for 30 min, followed by the addition of 100 µl PI at 4°C in
the dark for 30 min. Cell cycle analysis was performed by flow
cytometry using a BD FACSCalibur system (BD Biosciences) and data
was analyzed using the ModFit software version 4.1 (Verity Software
House, Inc., Topsham, ME, USA). Each experiment was repeated three
times.
Cell migration and invasion
assays
Cell migration was determined using a wound healing
assay, according to a method described previously (19). SW-620 and HT29 cells transfected
with si-NC and si-lncTCF7 were seeded into 6-well plates
(5×105 cells/well), and after reaching 90% confluency, a
scratch was made in the plate with a 200 µl pipette tip. The
remaining cells were washed with RPMI-1640 medium and incubated
with 5% CO2 at 37°C for 48 h. An image of the migration
distance of cells was captured using a FSX100 Biological Image
system (Olympus Corporation, Tokyo, Japan) and the images were
analyzed using Image J version 6.0 (Media Cybernetics, Inc.,
Rockville, MD, USA). For invasion assays, the upper chamber had
been precoated with Matrigel (BD Biosciences) and incubated for 24
h. Then cell invasion was evaluated by performing a Transwell
assay, according to a method described previously (20,21).
Cells in six independent fields per well were imaged at
magnification, ×200 by using a microscope (Olympus Corporation),
and the number of migrating and invading cells were counted.
Statistical analysis
Statistical analysis was performed using SPSS
version 19.0 software (IBM Corp., Armonk, NY, USA). Continuous
variables were presented as the mean ± standard deviation.
Differences between CRC tissues and adjacent normal tissues were
analyzed using an independent t-test. Differences among si-NC,
si-lncTCF7-1 and si-lncTCF7-2 groups were evaluated using one-way
analysis of variance followed by least significant difference
post-hoc test. Differences between si-lncTCF7 and si-NC groups were
also analyzed using the independent t-test. A receiver operating
characteristic (ROC) curve was applied to evaluate the diagnostic
value (sensitivity and specificity). According to sensitivity and
specificity calculated by SPSS, the value of the Youden index
(sensitivity + specificity-1) was calculated, and the best
diagnostic cutoff value is the maximum value of the Youden index.
P<0.05 was considered to indicate a statistically significant
difference.
Results
LncTCF7 expression is increased in CRC
tissues
LncTCF7 expression in CRC tissues and adjacent
normal tissues was analyzed by performing RT-qPCR. Results of
RT-qPCR revealed that lncTCF7 expression (mean value) was
significantly increased in CRC tissues compared with adjacent
normal tissues (P<0.05; Fig.
1A). In addition, results of RT-qPCR demonstrated that 102
(102/112; 91.07%) patients with CRC exhibited higher lncTCF7
expression in CRC tissues compared with adjacent normal tissues
(Fig. 1B). To investigate the
association between the different clinicopathological features of
patients with CRC and lncTCF7 expression, the patients were divided
into the following two groups based on the comparison between
lncTCF7 expression and the median lncTCF7 expression value (6.59)
of all CRC tissues: Low expression group (lncTCF7 expression value,
<6.59) and high expression group (lncTCF7 expression value,
≥6.59). Results for lncTCF7 expression are presented in Table I. LncTCF7 expression was
significantly associated with tumor size, differentiation degree,
tumor-node-metastasis (TNM) grade, lymph node metastasis and depth
of invasion (P<0.05), but was not significantly associated with
sex, age, tumor location or organization (P>0.05). Furthermore,
lncTCF7 expression was increased in patients with large tumors
(size, >5 cm) compared with patients with small tumors (size,
<5 cm; Fig. 1C), in patients
exhibiting poor tumor differentiation compared with patients
exhibiting moderate + high tumor differentiation (Fig. 1D), in patients with lymph node
metastasis compared with patients without lymph node metastasis
(Fig. 1E), in patients with serosa
layer invasion compared with patients with muscular layer invasion
(Fig. 1F) and in patients with TNM
grade III + IV tumors compared with patients with TNM grade I + II
tumors (Fig. 1G). Receiver
operating characteristic (ROC) curve analysis was performed to
determine the sensitivity and specificity of lncTCF7 for diagnosing
CRC based on its expression. The ROC curves demonstrated
significant separation between the CRC tissues and adjacent normal
tissues, with an area under the curve of 0.885 (95% confidence
interval, 0.844–0.926; Fig. 1H).
According to sensitivity and specificity calculated by SPSS, the
maximum value of the Youden index was 0.616, and the corresponding
optimal lncTCF7 expression cutoff value was 4.48. At the optimal
expression cutoff value, the sensitivity and specificity of lncTCF7
for diagnosing CRC were 67.9 and 93.7%, respectively.
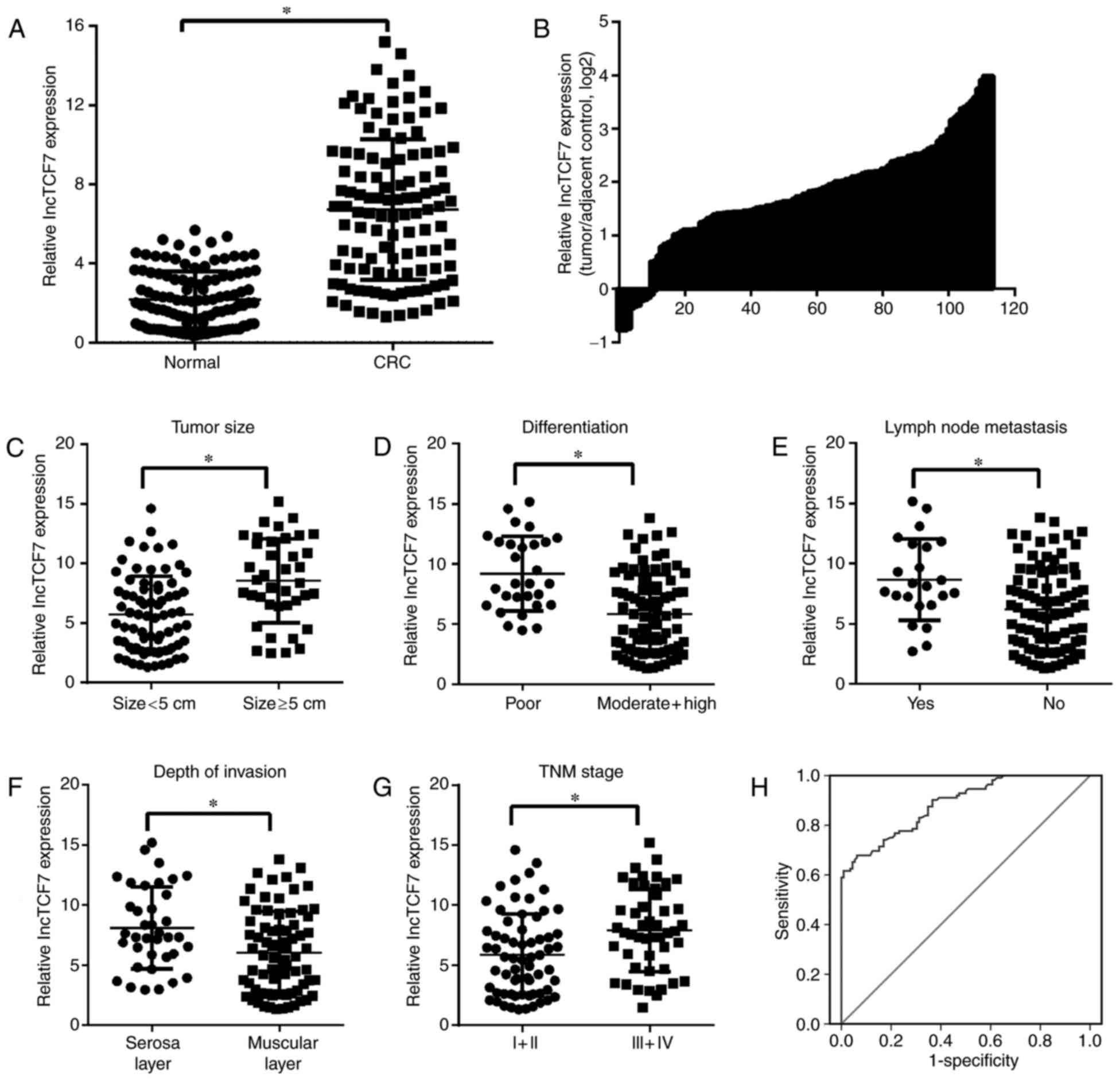 | Figure 1.LncTCF7 was overexpressed in CRC
tissues and its expression was associated with tumor size,
differentiation degree, TNM grade, lymph node metastasis, depth of
invasion and CRC diagnosis. (A) LncTCF7 expression in CRC tissues
and adjacent normal tissues was evaluated by performing reverse
transcription-quantitative polymerase chain reaction. (B)
Distribution plot of log2 values of relative expression ratio
(lncTCF7 expression in CRC tissues/lncTCF7 expression in adjacent
normal tissues) vs. the number of patients with CRC. LncTCF7
expression in CRC tissues of patients with (C) large- and
small-sized tumors, (D) poor and moderate + high tumor
differentiation, (E) with or without lymph node metastasis, (F)
muscular and serosa layer invasion and (G) TNM grades I + II and
III + IV tumors. Data are presented as the mean ± standard
deviation. (H) Receiver operating characteristic curve analysis for
assessing the diagnostic value of lncTCF7 expression in CRC.
*P<0.05, as indicated. LncTCF7, long non-coding RNA TCF7; CRC,
colorectal cancer; TNM, tumor-node-metastasis; lncTCF7, long
non-coding RNA TCF7; AUC, area under curve. |
LncTCF7 silencing suppresses
proliferation without affecting the apoptosis of CRC cells
To evaluate the effect of lncTCF7 expression on the
biological function of CRC, SW-620 and HT29 cells were transfected
with si-lncTCF7 and lncTCF7 expression was quantified by performing
RT-qPCR. Results of RT-qPCR revealed that lncTCF7 expression was
significantly inhibited in si-lncTCF7-1- and
si-lncTCF7-2-transfected cells, to a larger extent in
si-lncTCF7-1-transfected cells, compared with si-NC-transfected
cells (P<0.05; Fig. 2A). These
results indicated that si-lncTCF7-1 had a higher silencing
efficiency than si-lncTCF7-2; therefore, si-lncTCF7-1-transfected
cells were used for subsequent experiments. Results of the CCK-8
assay demonstrated that the proliferation of si-lncTCF7-transfected
SW-620 and HT29 cells at 24, 48 and 72 h after transfection was
significantly lower compared with si-NC-transfected SW-620 and HT29
cells at the same time points (P<0.05; Fig. 2B). Cell cycle analysis demonstrated
that lncTCF7 silencing induced a significant G1-phase arrest in
SW-620 and HT29 cells (P<0.05; Fig.
2C and D). However, results of apoptosis analysis indicated
that apoptosis was not significantly different between si-lncTCF7-
and si-NC-transfected SW-620 and HT29 cells (P>0.05; Fig. 2E and F).
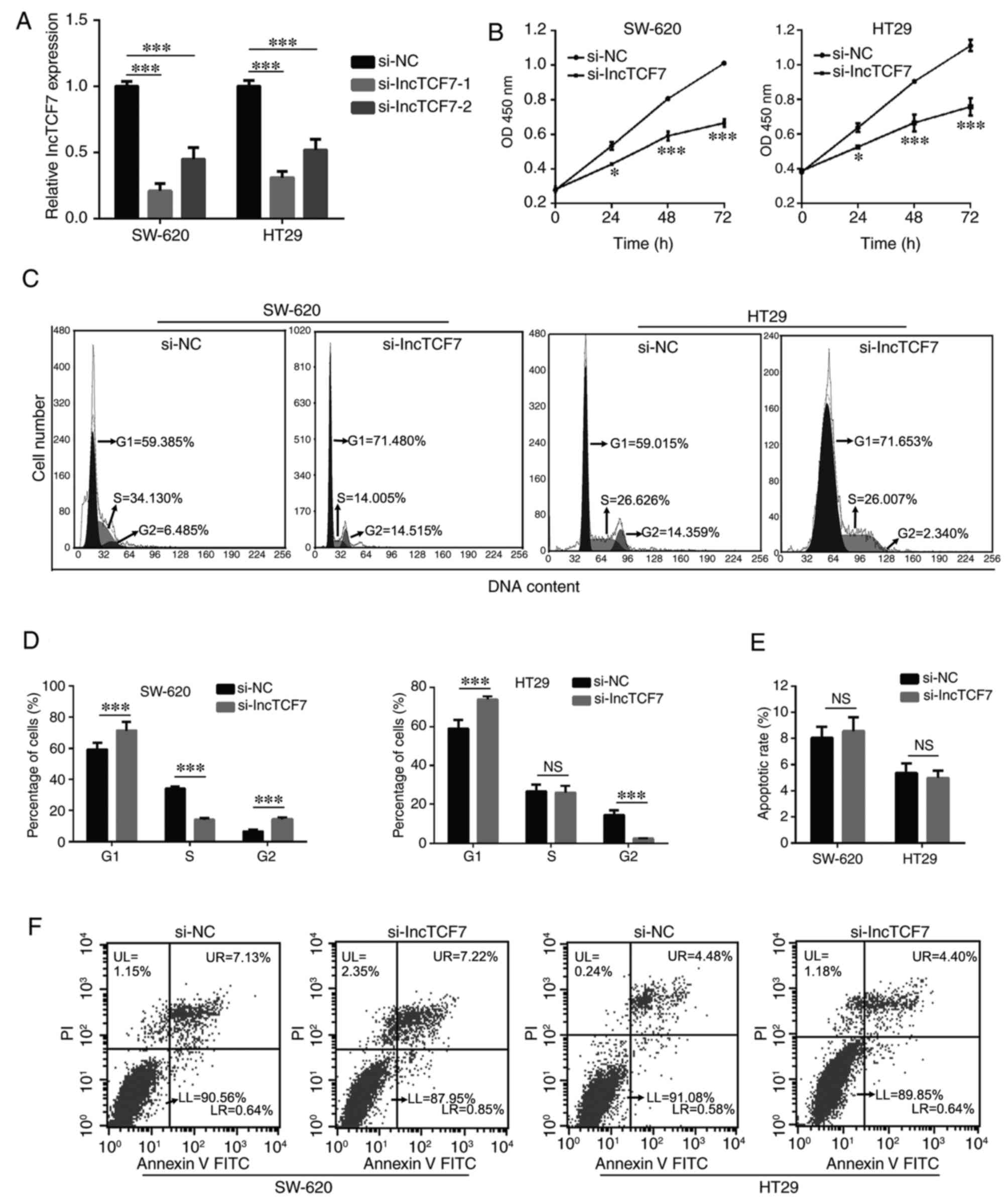 | Figure 2.Silencing of lncTCF7 inhibited the
proliferation and cell cycle without affecting the apoptosis of
SW-620 and HT29 cells. (A) LncTCF7 expression in si-NC- and
si-lncTCF7-1/2 transfected SW-620 and HT29 cells was detected by
performing reverse transcription-quantitative polymerase chain
reaction. (B) Proliferation of si-NC- and si-lncTCF7-transfected
SW-620 and HT29 cells was determined using a Cell Counting kit-8
assay. (C) Representative images of the cell cycle of SW-620 and
HT29 cells. (D) Cell cycle stages of si-NC- and
si-lncTCF7-transfected SW-620 and HT29 cells were investigated and
quantified by performing flow cytometry following staining with PI.
(E) Apoptosis of si-NC- and si-lncTCF7-transfected SW-620 and HT29
cells was determined by performing flow cytometry following
staining with Annexin V-FITC and PI. (F) Representative flow
cytometry plots of the apoptosis of SW-620 and HT29 cells. Cells in
upper right (UR) + lower right (LR) quadrants were considered as
apoptotic cells. Data are presented as the mean ± standard
deviation. For parts A and D, ***P<0.001, as indicated; for part
B, *P<0.05 and ***P<0.001 vs. si-NC group. LncTCF7, long
non-coding RNA TCF7; si, small interfering RNA; si-NC, si-negative
control; PI, propidium iodide; FITC, fluorescein isothiocyanate;
OD, optical density; NS, not significant. |
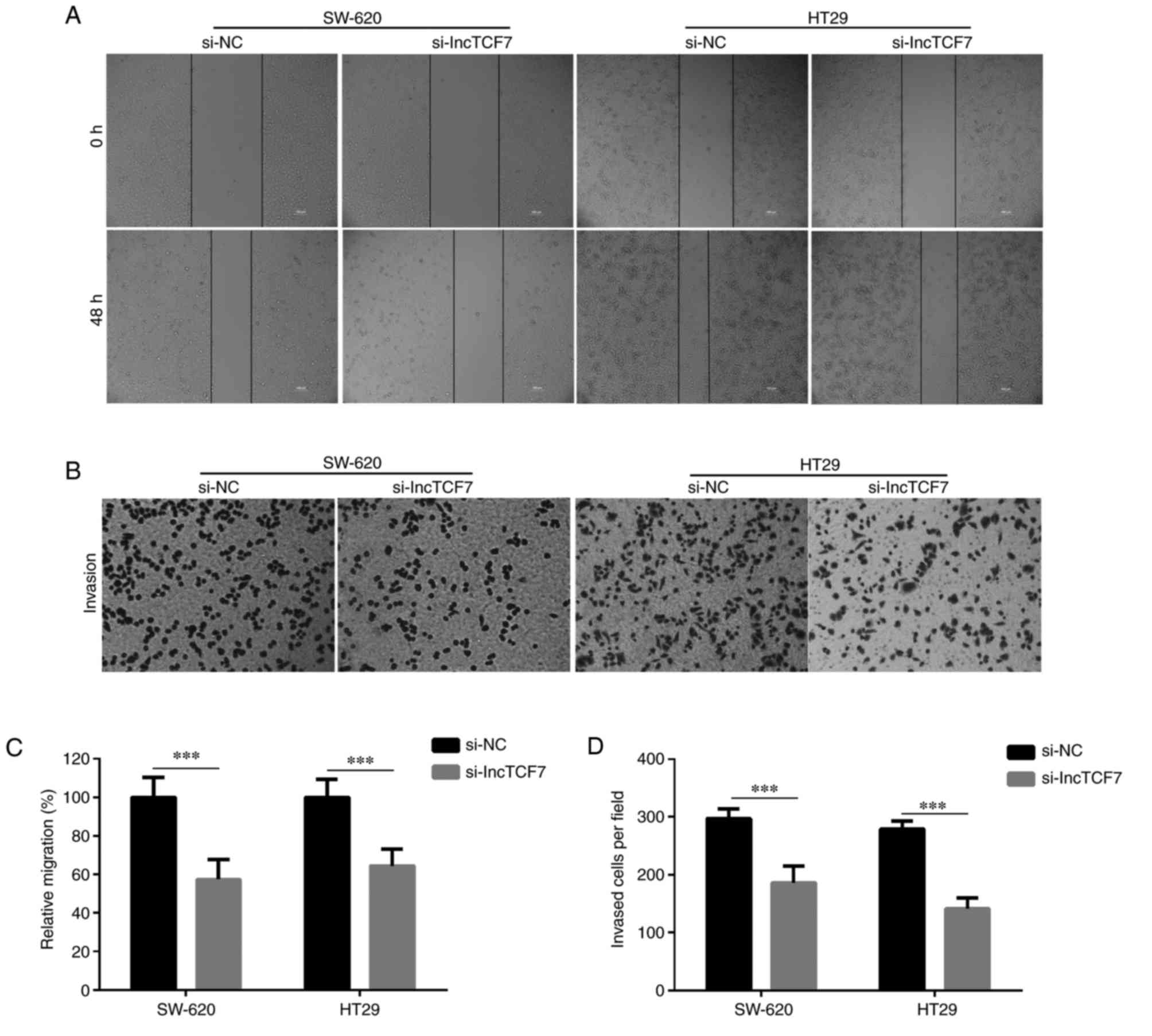
LncTCF7 silencing inhibits the
migration and invasion of CRC cells
The migration and invasion of SW-620 and HT29 cells
was determined by performing wound healing and Transwell assays,
respectively, after 48 h transfection. The migration and invasion
of cells was significantly lower among si-lncTCF7-transfected cells
compared with si-NC-transfected cells (P<0.05; Fig. 3).
Discussion
The present study investigated the expression of
lncTCF7 to assess its efficacy as a novel diagnostic marker of CRC.
The results revealed that lncTCF7 expression was significantly
higher in CRC tissues compared with adjacent normal tissues. In
addition, lncTCF7 expression was significantly associated with
tumor size, differentiation degree, TNM grade, lymph node
metastasis and invasion depth; however, it was not significantly
associated with sex, age, location and organization, indicating
that lncTCF7 may be associated with tumorigenesis and tumor
progression. These results are similar to those of previous
studies, which reported that lncTCF7 was overexpressed in liver
cancer and non-small cell lung cancer tissues (9,16).
Results of ROC analysis performed in the present study indicated
that lncTCF7 may be an independent diagnostic marker of CRC.
In addition, the effect of lncTCF7 on CRC
tumorigenesis and progression was investigated to determine its
potential as a therapeutic target for CRC treatment. The results
demonstrated that silencing of lncTCF7 inhibited the proliferation,
migration and invasion, but did not significantly affect the
apoptosis, of CRC cells. Additionally, cell cycle progression is an
essential requirement for cell growth. In the current study,
lncTCF7 silencing inhibited SW-620 and HT29 cell cycle transition
from G1 to S phase, which indicated that lncTCF7 silencing may
inhibit SW-620 and HT29 proliferation by suppressing cell cycle
progression. Proliferation and cell cycle dysfunction serve an
important role in CRC development, and cell migration and invasion
serve a central role in CRC metastasis (22–24).
The results of the present study also indicated that lncTCF7
silencing inhibited CRC development and metastasis, indicating that
lncTCF7 may function as an oncogene to promote CRC development and
metastasis. The results of the present study are consistent with
those of previous studies. Wang et al (9) demonstrated that lncTCF7 promoted the
self-renewal and propagation of liver cancer stem cells by
recruiting the Switch/sucrose non-fermentable complex to the TCF7
promoter to upregulate TCF7 expression and to activate Wnt
signaling. Furthermore, Wu and Wang (16) demonstrated that lncTCF7 increased
the self-renewal and metastasis of human non-small cell lung cancer
cells by promoting Slug and epithelial cell adhesion molecule
expression. In another study, Wu et al (17) revealed that lncTCF7 increased the
aggressiveness of hepatocellular carcinoma cells by activating
epithelial-mesenchymal transition signaling.
However, there are several limitations to the data
generated in the current study. Firstly, the overall survival of
lncTCF7 on CRC prognosis was not analyzed due to the short
follow-up time, and the overall survival of lncTCF7 on CRC
prognosis may be investigated in further research. In addition, the
effect of lncTCF7 on the animal models of CRC may be the focus of
further research. Finally, further studies are required to
determine the mechanism underlying lncTCF7-induced regulation of
CRC tumorigenesis and progression. In conclusion, the results of
the current indicated that lncTCF7 may be a potential biomarker and
therapeutic target for CRC treatment.
References
|
1
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chen W, Zheng R, Zeng H, Zhang S and He J:
Annual report on status of cancer in China, 2011. Chin J Cancer
Res. 27:2–12. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Fleshman JW and Smallwood N: Current
concepts in rectal cancer. Clin Colon Rectal Surg. 28:5–11. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rao AKDM, Rajkumar T and Mani S:
Perspectives of long non-coding RNAs in cancer. Mol Biol Rep.
44:203–218. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Li M, Wang Y, Cheng L, Niu W, Zhao G, Raju
JK, Huo J, Wu B, Yin B, Song Y and Bu R: Long non-coding RNAs in
renal cell carcinoma: A systematic review and clinical
implications. Oncotarget. 8:48424–48435. 2017.PubMed/NCBI
|
|
6
|
Klingenberg M, Matsuda A, Diederichs S and
Patel T: Non-coding RNA in hepatocellular carcinoma: Mechanisms,
biomarkers and therapeutic targets. J Hepatol. 67:603–618. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Richtig G, Ehall B, Richtig E,
Aigelsreiter A, Gutschner T and Pichler M: Function and clinical
implications of long non-coding RNAs in melanoma. Int J Mol Sci.
18:pii: E715. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Yang Y, Zhao L, Lei L, Lau WB, Lau B, Yang
Q, Le X, Yang H, Wang C, Luo Z, et al: LncRNAs: The bridge linking
RNA and colorectal cancer. Oncotarget. 8:12517–12532.
2017.PubMed/NCBI
|
|
9
|
Wang Y, He L, Du Y, Zhu P, Huang G, Luo J,
Yan X, Ye B, Li C, Xia P, et al: The long noncoding RNA lncTCF7
promotes self-renewal of human liver cancer stem cells through
activation of Wnt signaling. Cell Stem Cell. 16:413–425. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Weber BN, Chi AW, Chavez A, Yashiro-Ohtani
Y, Yang Q, Shestova O and Bhandoola A: A critical role for TCF-1 in
T-lineage specification and differentiation. Nature. 476:63–68.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Cui L, Guan Y, Qu Z, Zhang J, Liao B, Ma
B, Qian J, Li D, Li W, Xu GT and Jin Y: WNT signaling determines
tumorigenicity and function of ESC-derived retinal progenitors. J
Clin Invest. 123:1647–1661. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
12
|
Nikuseva-Martic T, Serman L, Zeljko M,
Vidas Z, Gašparov S, Zeljko HM, Kosović M and Pećina-Šlaus N:
Expression of secreted frizzled-related protein 1 and 3, T-cell
factor 1 and lymphoid enhancer factor 1 in clear cell renal cell
carcinoma. Pathol Oncol Res. 19:545–551. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Siu MK, Chen WY, Tsai HY, Chen HY, Yin JJ,
Chen CL, Tsai YC and Liu YN: TCF7 is suppressed by the androgen
receptor via microRNA-1-mediated downregulation and is involved in
the development of resistance to androgen deprivation in prostate
cancer. Prostate Cancer Prostatic Dis. 20:172–178. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Chen WY, Liu SY, Chang YS, Yin JJ, Yeh HL,
Mouhieddine TH, Hadadeh O, Abou-Kheir W and Liu YN: MicroRNA-34a
regulates WNT/TCF7 signaling and inhibits bone metastasis in
Ras-activated prostate cancer. Oncotarget. 6:441–457.
2015.PubMed/NCBI
|
|
15
|
Wang Y, Zhang S, Xu Y, Zhang Y, Guan H, Li
X, Li Y and Wang Y: Upregulation of miR-192 inhibits cell growth
and invasion and induces cell apoptosis by targeting TCF7 in human
osteosarcoma. Tumour Biol. 37:15211–15220. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wu J and Wang D: Long noncoding RNA TCF7
promotes invasiveness and self-renewal of human non-small cell lung
cancer cells. Hum Cell. 30:23–29. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wu J, Zhang J, Shen B, Yin K, Xu J, Gao W
and Zhang L: Long noncoding RNA lncTCF7, induced by IL-6/STAT3
transactivation, promotes hepatocellular carcinoma aggressiveness
through epithelial-mesenchymal transition. J Exp Clin Cancer Res.
34:1162015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li C, Liang G, Yang S, Sui J, Yao W, Shen
X, Zhang Y, Peng H, Hong W, Xu S, et al: Dysregulated lncRNA-UCA1
contributes to the progression of gastric cancer through regulation
of the PI3K-Akt-mTOR signaling pathway. Oncotarget. 8:93476–93491.
2017.PubMed/NCBI
|
|
20
|
Xu C, Sun L, Jiang C, Zhou H, Gu L, Liu Y
and Xu Q: SPP1, analyzed by bioinformatics methods, promotes the
metastasis in colorectal cancer by activating EMT pathway. Biomed
Pharmacother. 91:1167–1177. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Pan Y, Jiao G, Wang C, Yang J and Yang W:
MicroRNA-421 inhibits breast cancer metastasis by targeting
metastasis associated 1. Biomed Pharmacother. 83:1398–1406. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Romagnoli P, Filipponi F, Bandettini L and
Brugnola D: Increase of mitotic activity in the colonic mucosa of
patients with colorectal cancer. Dis Colon Rectum. 27:305–308.
1984. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Paganelli GM, Biasco G, Santucci R, Brandi
G, Lalli AA, Miglioli M and Barbara L: Rectal cell proliferation
and colorectal cancer risk level in patients with nonfamilial
adenomatous polyps of the large bowel. Cancer. 68:2451–2454. 1991.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Yao M, Lam EC, Kelly CR, Zhou W and Wolfe
MM: Cyclooxygenase-2 selective inhibition with NS-398 suppresses
proliferation and invasiveness and delays liver metastasis in
colorectal cancer. Br J Cancer. 90:712–719. 2004. View Article : Google Scholar : PubMed/NCBI
|