Introduction
Moyamoya disease (MMD) is a cerebrovascular disease,
which was first described by Takeuchi and Shimizui in 1957
(1). It is characterized by
progressive stenosis or occlusion of the distal portions of the
internal carotid arteries, with compensatory development of an
aberrant vascular network at the base of the brain. The incidence
of MMD is higher in East Asian countries compared with the rest of
the world (2,3). Its initial clinical manifestations
include transient ischemic attack, infarction, hemorrhage,
headaches and epilepsy. Genetic factors are likely to be involved
in the development of MMD; epidemiological investigations have
revealed that 7–12% of patients in Japan have a familial occurrence
(4). In addition, studies have
revealed that non-coding RNAs, including micro(mi)RNAs and long
non-coding (lnc)RNAs exert an important regulatory role in this
disease (5,6). However, the etiology and pathogenesis
of MMD remain unclear.
With the rapid progress and wide use of next
generation sequencing technologies, several independent studies
have revealed the existence of transfer RNA (tRNA)-derived
fragments (tRFs) (7–9). These derivatives were originally
viewed as tRNA degradation products, but accumulating evidence
suggests that they serve roles in various biological processes,
including regulation of gene expression, cell proliferation and
tumor suppression (10–12). For example, a study on mouse sperm
indicated that 5′-tiRNAs (tRNA-derived stress-induced RNAs), a
subset of tRFs, may alter expression profiles and modify RNA, and
act as a paternal epigenetic factor regulating intergenerational
inheritance of certain metabolic diseases (13). Accumulation of 5′-tiRNAs due to
tRNA hypomethylation could specifically repress translation and
activate stress pathways, eventually leading to human
neurodevelopmental disorders (14). However, the potential function of
tRFs in MMD has yet to be elucidated.
In the present study, a prospective case-control
study was performed to investigate and compare the expression
profiles of tRFs in patients with MMD and healthy controls.
Furthermore, bioinformatics analyses were used to further
investigate the molecular basis of tRFs in MMD.
Materials and methods
Patient samples
The present study was conducted in accordance with
the Declaration of Helsinki, and was approved by the Research
Ethics Board of Beijing Tiantan Hospital, Capital Medical
University (Beijing, China). Written informed consent was obtained
from all individuals who were included in the study. Between March
2017 and April 2017, three adult patients who were diagnosed with
MMD and underwent neurosurgical treatment at the Department of
Neurosurgery, Beijing Tiantan Hospital were enrolled for the
RNA-sequencing experiment. Cerebral digital subtraction angiography
was performed in all patients. MMD was diagnosed according to the
criteria put forward by the Japanese Ministry of Health and Welfare
in 2012 (15). Patients diagnosed
with quasi-MMD, hypertension, hyperlipidemia or diabetes, and
patients taking certain drugs (including antibiotics, antiplatelet
drugs and antiepileptic drugs, among others) were excluded from the
study to prevent potential interference to the experimental
results. Overall, three male patients with MMD (age range, 30–51
years) were included in this study, which were compared with three
age-matched healthy male volunteers (age range, 25–29 years). The
duration from disease onset to hospital admission in patients with
MMD was >2 weeks. Due to ethical issues, blood vessels could not
be obtained from the patients; therefore, peripheral blood was used
as an alternative, which is consistent with other studies on MMD
(5,6). In the present study, all the blood
samples were collected prior to surgical treatment. To verify the
RNA sequencing results, five patients (three females, two males)
with MMD (age range, 30–42 years) were included in the study, and
were compared with five healthy male volunteers (age range, 26–32
years). Samples were collected between March 2017 and April 2017. A
total of 2.5 ml peripheral blood was collected from each
participant.
RNA extraction and small
RNA-sequencing
A total of 2.5 ml peripheral blood was collected
from each participant in K2EDTA-coated
Vacutainer® tubes (BD Biosciences, Franklin Lakes, NJ,
USA). Total RNA was extracted from the six blood samples using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc., Waltham, MA, USA), according to the manufacturer's protocol.
Subsequently, RNA was quantified using a NanoDrop™ ND-1000
spectrophotometer (NanoDrop; Thermo Fisher Scientific, Inc.,
Wilmington, DE, USA). RNA integrity was assessed by 1% agarose gel
electrophoresis. RNA samples were firstly pretreated using the
rtStar™ tRF&tiRNA Pretreatment kit (cat. no. AS-FS-005;
Arraystar Inc., Rockville, MD, USA) to remove any RNA
modifications, which may interfere with small RNA sequencing
library construction. The pretreated total RNA samples were
subjected to sequencing library preparation using a commercial kit
(NEB Next® Multiplex Small RNA Library Prep Set for
Illumina®; cat. no. 7580; New England BioLabs, Inc.,
Ipswich, MA, USA). The procedure was conducted according to the
manufacturer's protocol, which included 3′- and 5′-adapter
ligation, cDNA synthesis and library polymerase chain reaction
(PCR) amplification. The completed libraries were quantified on an
Agilent Bioanalyzer 2100 (Agilent Technologies, Inc., Santa Clara,
CA, USA), and then sequencing was performed using an Illumina
NextSeq 500 system (Illumina, Inc., San Diego, CA, USA), according
to the manufacturer's protocol.
Data analysis
Analysis of the sequence reads was performed using
the Solexa pipeline (Off-Line Base Caller software version 1.8,
Illumina, Inc.). The sequencing quality was examined using FastQC
software version 0.11.5 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/).
The trimmed reads were first compared to the genomic tRNA database
(http://gtrnadb.ucsc.edu/) using Novo Align
software version 2.07.11 (http://www.novocraft.com/main/index.php) to identify
known mature- and pre-tRNAs. The unidentified reads were further
aligned to several other small RNA databases (mRNA/ribosomal
RNA/small nuclear RNA/small nucleolar RNA/Piwi-interacting
RNA/miRNA), including GtRNA database (http://gtrnadb.ucsc.edu), tRFs database (http://genome.bioch.virginia.edu/trfdb/)
and Mintbase (https://cm.jefferson.edu/MINTbase/). The tRF
expression levels were measured and normalized to the number of
transcripts per million of total aligned tRNA reads. Expression
profiles between patients and healthy controls were compared by
calculating the fold change (i.e. the ratio of the group means) for
each tRF. Statistical significance was determined using Student's
t-test. In total, 38 tRFs with fold change ≥2.0 and P<0.05 were
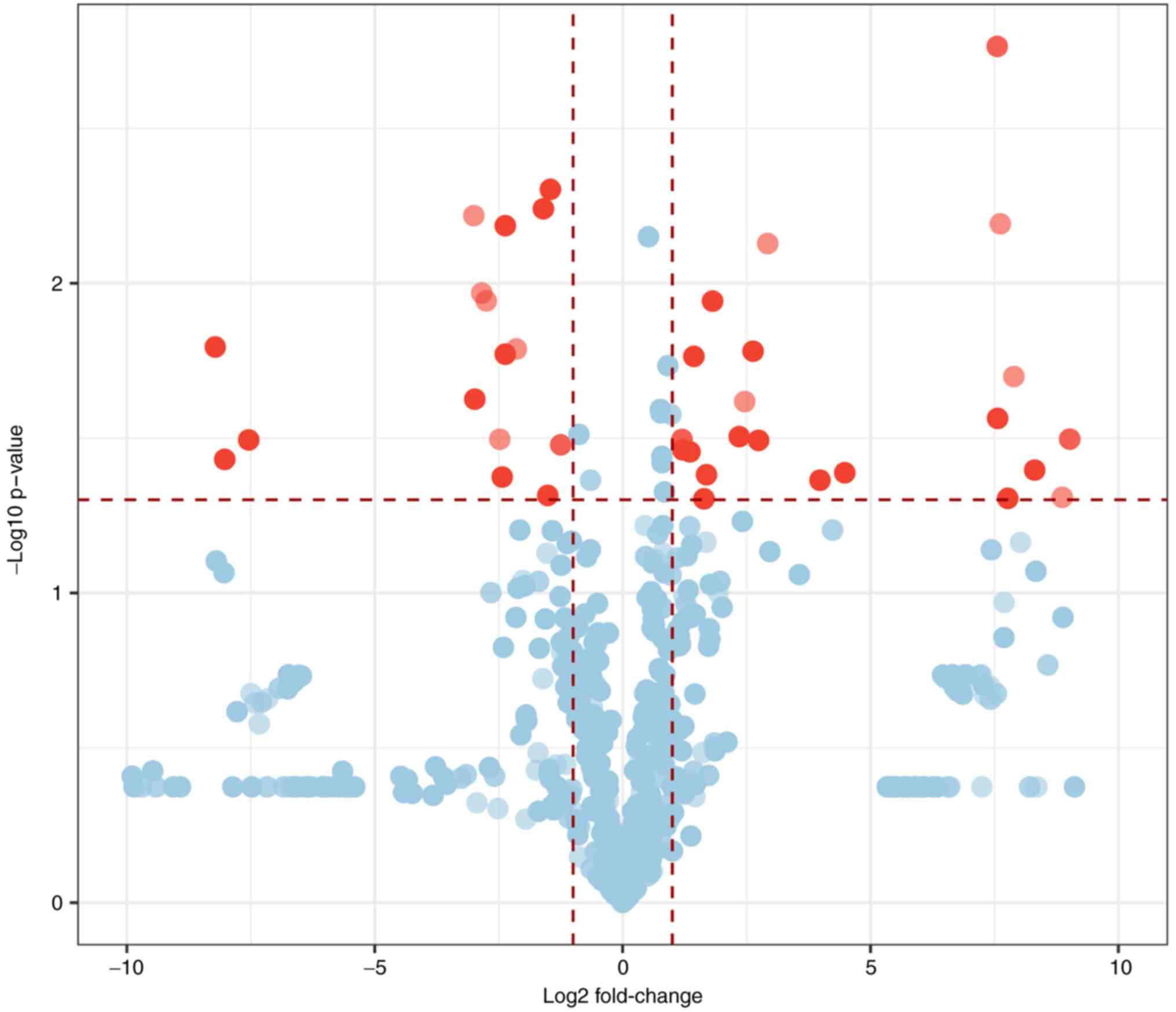
considered significantly differentially expressed. Volcano plot and
heat map were constructed to analyze the differentially expressed
tRFs between the two groups.
Validation using reverse
transcription-quantitative PCR (RT-qPCR)
RT-qPCR is considered the gold standard for gene
expression quantification (16,17).
To verify the RNA sequencing results, four tRFs were randomly
selected and their expression levels were detected in five patients
with MMD and five healthy controls (An independent cohort, n=5).
Total RNA was extracted from peripheral blood using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.). RNA samples were pretreated using the rtStar™ tRF&tiRNA
Pretreatment kit (cat. no. AS-FS-005; Arraystar Inc.) to remove RNA
modifications, and were then reverse transcribed to cDNA using the
rtStar™ First-Strand cDNA Synthesis kit (cat. no. AS-FS-003;
Arraystar Inc.) according to the manufacturer's protocol. qPCR was
performed using ViiA 7 Real-time PCR system, using Arraystar
SYBR® Green Real-time qPCR Master mix (cat. no.
AS-MR-005-25; Applied Biosystems; Thermo Fisher Scientific, Inc.)
with specific primers (Table I).
The thermocycling conditions were the following: Initial
denaturation at 95°C for 10 min, followed by 40 cycles of 95°C for
10 sec and 60°C for 60 sec. U6 was used as an internal control to
assess the efficiency of the reaction. RT-qPCR experiments were
performed in triplicate. Data were analyzed using the
2−ΔΔCq method (17); a
standard dilution curve was used to determine amplification
efficiency.
 | Table I.Primers for reverse
transcription-quantitative polymerase chain reaction. |
Table I.
Primers for reverse
transcription-quantitative polymerase chain reaction.
| Gene | Primers |
|---|
| U6 |
F:5′-GCTTCGGCAGCACATATACTAAAAT-3′ |
|
|
R:5′-CGCTTCACGAATTTGCGTGTCAT-3′ |
| AS-tDR-000586 |
F:5′-TCTCCCACATGGTCTAGCGGT-3′ |
|
|
R:5′-TGCTCTTCCGATCTGAAAACCA-3′ |
| AS-tDR-000924 | F:
5′-AGAGTTCTACAGTCCGACGATC-3′ |
|
|
R:5′-CCGATCTACTGAGCTATCCG-3′ |
| AS-tDR-007725 |
F:5′-CAGTCCGACGATCTCCCTGT-3′ |
|
|
R:5′-ATCTGAGAGCGCCGAATCC-3′ |
| AS-tDR-011363 |
F:5′-CAGTCCGACGATCCTGTCACG-3’ |
|
| R:
5′-TTCCGATCTGCCCCGGTCT-3′ |
Bioinformatics analysis
As with miRNAs, tRFs can interact with Argonaute
(AGO) proteins (18). Previous
studies have demonstrated that tRFs are able to target RNAs
similarly to miRNAs (19,20). In addition, it has been proposed
that tRF seed sequences have similar matching profiles to miRNAs
(21–23). Therefore, miRNA target-predicting
algorithms including miRanda (http://www.microrna.org/microrna/home.do; August 2010
release) and TargetScan (version 71; http://www.targetscan.org/vert_71/), were used to
identify the potential target genes of the selected tRFs, based on
the presence of binding sites in the 3′untranslated region.
Subsequently, the predicted target genes were inputted into the
Database for Annotation, Visualization and Integrated Discovery
(http://david.abcc.ncifcrf.gov/) using
Gene Ontology (GO; http://www.geneontology.org/) to analyze their
molecular functions. Kyoto Encyclopedia of Genes and Genomes (KEGG;
http://www.genome.ad.jp/kegg/) database
was used to identify significant pathways for these target genes.
DAVID (version 6.8; http://david.ncifcrf.gov/) software was used to
generate the pathway map.
Statistical analysis
Data are presented as the means ± standard
deviation. Data analysis was performed using R software version
3.3.1 (R Project for Statistical Computing, Vienna, Austria).
Differences between two groups were analyzed using the Student's
t-test. P<0.05 was considered to indicate a statistically
significant difference.
Results
Expression profiles of tRFs
The clinical characteristics of patients with MMD
are summarized in Table II,
including age, sex, initial presentation, disease duration, Suzuki
stage (24) and Modified Rankin
Score (25). All individuals were
of Han Chinese ethnicity. Differentially expressed tRFs in patients
with MMD vs. healthy controls were identified by RNA sequencing. A
total of 22 tRFs were upregulated and 16 tRFs were downregulated
(fold change ≥2.0; P<0.05). Notably, 26 tRFs were significantly
differentially expressed by at least 5-fold between the two groups
(fold change ≥5.0; P<0.05); with 15 upregulated and 11
downregulated tRFs. Among the differentially expressed tRFs,
5′-tiRNA derived from tRNAGlu(TTC) was the most
upregulated, whereas i-tRF from tRNAGln(CTG) and tRF-1
from pre-tRNAGln(CTG) were the most downregulated. More
upregulated tRFs were observed compared with downregulated tRFs.
The 38 upregulated and downregulated tRFs ranked by fold change are
summarized in Tables III and
IV. A volcano plot was
constructed to identify differentially expressed tRFs (Fig. 1). The heatmap revealed systematic
variations between the MMD and control groups in terms of tRF
expression (Fig. 2).
 | Table II.Clinical characteristics of patients
with moyamoya disease. |
Table II.
Clinical characteristics of patients
with moyamoya disease.
| Patient no. | Age (years) | Sex | Clinical
presentation | Duration from
disease onset to hospital admission (months) | Suzuki stage (I to
VI) | mRS |
|---|
| 1 | 51 | Male | Intracerebral
hemorrhage | 6 | R, III; L, III | 2 |
| 2 | 38 | Male | Limb weakness | 12 | R, II; L, II | 2 |
| 3 | 30 | Male | Intraventricular
hemorrhage | 6 | R, III; L, II | 3 |
 | Table III.Summary of the 22 tRFs upregulated in
patients with moyamoya disease vs. healthy controls. |
Table III.
Summary of the 22 tRFs upregulated in
patients with moyamoya disease vs. healthy controls.
| tRF ID | Mintbase ID | tRF sequence
(5′-3′) | Amino
acid-anticodon | Fold change | P-value |
|---|
| AS-tDR-000586 |
tRF-36-86J8WPMN1E8Y7ZD |
TCCCACATGGTCTAGCGGTTAGGATTCCTGGTTTTC | Glu-TTC | 518.143 | 0.031859 |
| AS-tDR-007710 | N/A |
GCCCGGATGATCCTCAGTGGTCTGGGGTGCAGGCTTC | SeC-TCA | 466.968 | 0.049086 |
| AS-tDR-011363 | N/A |
CTGTCACGCGGGAGACCGGGGC | Asp-GTC | 317.306 | 0.040083 |
| AS-tDR-005857 |
tRF-37-EH623K7SIR3DR2M |
ACGCGGGAGACCGGGGTTCGATTCCCCGACGGGGAGC | Asp-GTC | 237.194 | 0.019999 |
| AS-tDR-006177 | tRF-16-K1JKE1B |
CCACCCAGGGACGCCA | Asn-GTT | 217.935 | 0.049457 |
| AS-tDR-011429 | N/A |
TAGGGAGGTTATGATTAACT | Arg-TCG | 196.348 | 0.006424 |
| AS-tDR-000107 |
tRF-22-WEK879652 |
TCGATCCCGGGTTTCGGCACCA | Phe-GAA | 188.997 | 0.027295 |
| AS-tDR-004084 |
tRF-18-S8V0JUDR |
GTCGGTAGAGCATGAGAC | Lys-CTT | 187.974 | 0.001717 |
| AS-tDR-007725 | N/A |
TCCCTGTGGTCTAGTGGTTAGGATTCGGCGCTCTC | Glu-CTC | 22.298 | 0.040897 |
| AS-tDR-000547 |
tRF-36-87R8WP9N1EWJQ7D |
TCCCTGGTGGTCTAGTGGTTAGGATTCGGCGCTCTC | Glu-CTC | 15.778 | 0.043218 |
| AS-tDR-007304 | N/A |
GCATTGGTGGTTCAGTGGTAGAATTCTCGCCTT | Glu-TTC | 7.596 | 0.007439 |
| AS-tDR-005361 | tRF-17-9LON4V3 |
TGGTAGAATTCTCGCCT | Gly-CCC | 6.683 | 0.032141 |
| AS-tDR-006553 |
tRF-18-P6KP6HD2 |
GCCCACCCAGGGACGCCA | Asn-GTT | 6.191 | 0.016570 |
| AS-tDR-008080 | N/A |
GCCCGGATGATCCTCAGTGGTCTGGGGTGCAGGCT | SeC-TCA | 5.507 | 0.024088 |
| AS-tDR-004206 | tRF-18-1SS2PMX |
AGTCGGTAGAGCATCAGA | Lys-TTT | 5.097 | 0.031258 |
| AS-tDR-006922 | tRF-17-884U1D2 |
TCCGGCTCGAAGGACCA | Tyr-GTA | 3.513 | 0.011413 |
| AS-tDR-001611 |
tRF-22-8XF6RE98N |
TCCTAAGCCAGGGATTGTGGGT | Arg-CCT | 3.227 | 0.041503 |
| AS-tDR-000122 | tRF-17-K5KKOV2 |
CCCACCCAGGGACGCCA | Asn-GTT | 3.119 | 0.049625 |
| AS-tDR-001889 | tRF-16-941QKSD |
TGGTGAGTATCCCCGC | Asp-GTC | 2.707 | 0.017218 |
| AS-tDR-008258 | N/A |
TGGTTCAGTGGTAGAAT | Gly-GCC &
Gly-CCC | 2.561 | 0.035012 |
| AS-tDR-008237 | N/A |
CTAAGCCAGGGATTGTGGGC | Arg-CCT | 2.316 | 0.034470 |
| AS-tDR-004490 |
tRF-21-JOWU1M70E |
CATAATCTGAAGGTCGTGAGT | Met-CAT | 2.293 | 0.031923 |
 | Table IV.Summary of the 16 tRFs downregulated
in patients with moyamoya disease vs. healthy controls. |
Table IV.
Summary of the 16 tRFs downregulated
in patients with moyamoya disease vs. healthy controls.
| tRF ID | Mintbase ID | tRF sequence
(5′-3′) | Amino
acid-anticodon | Fold change | P-value |
|---|
| AS-tDR-007510 | N/A |
ACTCTGGACTCTGAATCT | Gln-CTG | 297.887 | 0.016055 |
| AS-tDR-000924 | tRF-16-PS5P4PE |
GCCCGGATAGCTCAGT | Lys-TTT | 260.969 | 0.037030 |
| AS-tDR-003801 | tRF-17-MMUKLYK |
CGGTCTAAGGCGCTGCG | Leu-CAG | 186.513 | 0.032042 |
| AS-tDR-011424 | N/A |
ATATCCAACCTTCGGCTATAGGG | Thr-CGT | 8.018 | 0.006046 |
| AS-tDR-011366 |
tRF-19-WB86N7HU |
TCGAATCCCACTTCTGACA | Leu-CAG &
Leu-CAA | 7.902 | 0.023668 |
| AS-tDR-005697 |
tRF-18-BS6PDFD2 |
AACCGGGCAGAAGCACCA | Val-CAC | 7.168 | 0.010754 |
| AS-tDR-000053 |
tRF-31-79MP9P9NH57SD |
GTTTCCGTAGTGTAGTGGTTATCACGTTCGC | Val-AAC | 6.730 | 0.011393 |
| AS-tDR-009236 | N/A |
GGCTCGAAGGACTTCGTCTGTAA | Tyr-GTA | 5.578 | 0.031881 |
| AS-tDR-009369 |
tRF-21-WD8YQ84VD |
TCGACTCCTGGCTGGCTCGCC | Arg-ACG | 5.389 | 0.042221 |
| AS-tDR-001449 |
tRF-32-79MP9P9NH57SJ |
GTTTCCGTAGTGTAGTGGTTATCACGTTCGCC | Val-AAC &
Val-CAC | 5.162 | 0.016935 |
| AS-tDR-001400 |
tRF-32-6978WPRLXN4VQ |
GGCTCGTTGGTCTAGGGGTATGATTCTCGCTT | Pro-CGG &
Pro-TGG & Pro-AGG | 5.159 | 0.006515 |
| AS-tDR-011405 | N/A |
TTGAGATACTGAGTAGTCTGGTG | Arg-TCG | 4.422 | 0.016291 |
| AS-tDR-000009 |
tRF-31-P4R8YP9LON4VD |
GCATGGGTGGTTCAGTGGTAGAATTCTCGCC | Gly-GCC | 3.029 | 0.005745 |
| AS-tDR-007276 | N/A |
GTTTCCGTAGTGTAGTGGTTATCACGTTCGCT | Val-CAC &
Val-AAC | 2.853 | 0.048408 |
| AS-tDR-009331 | N/A |
TCGACTCCTGGCTGGCTCG | Arg-ACG | 2.749 | 0.004973 |
| AS-tDR-001273 |
tRF-35-86V8WPMN1E8Y7Z |
TCCCATATGGTCTAGCGGTTAGGATTCCTGGTTTT | Glu-TTC | 2.377 | 0.033231 |
RT-qPCR validation of RNA sequencing
results
Four tRFs (AS-tDR-000586, AS-tDR-000924,
AS-tDR-007725 and AS-tDR-011363) were randomly selected for
validation of RNA sequencing results using RT-qPCR in MMD and
control samples (n=5). The expression profiles of these tRFs were
in agreement with the results obtained from RNA sequencing. There
were significant differences in tRF expression levels in patients
with MMD compared with healthy controls (P<0.05; Fig. 3). These findings indicated that the
RNA sequencing data were reliable enough to warrant further
analysis.
tRF target gene prediction
In order to investigate the function of these tRFs,
their putative target genes were predicted using miRanda and
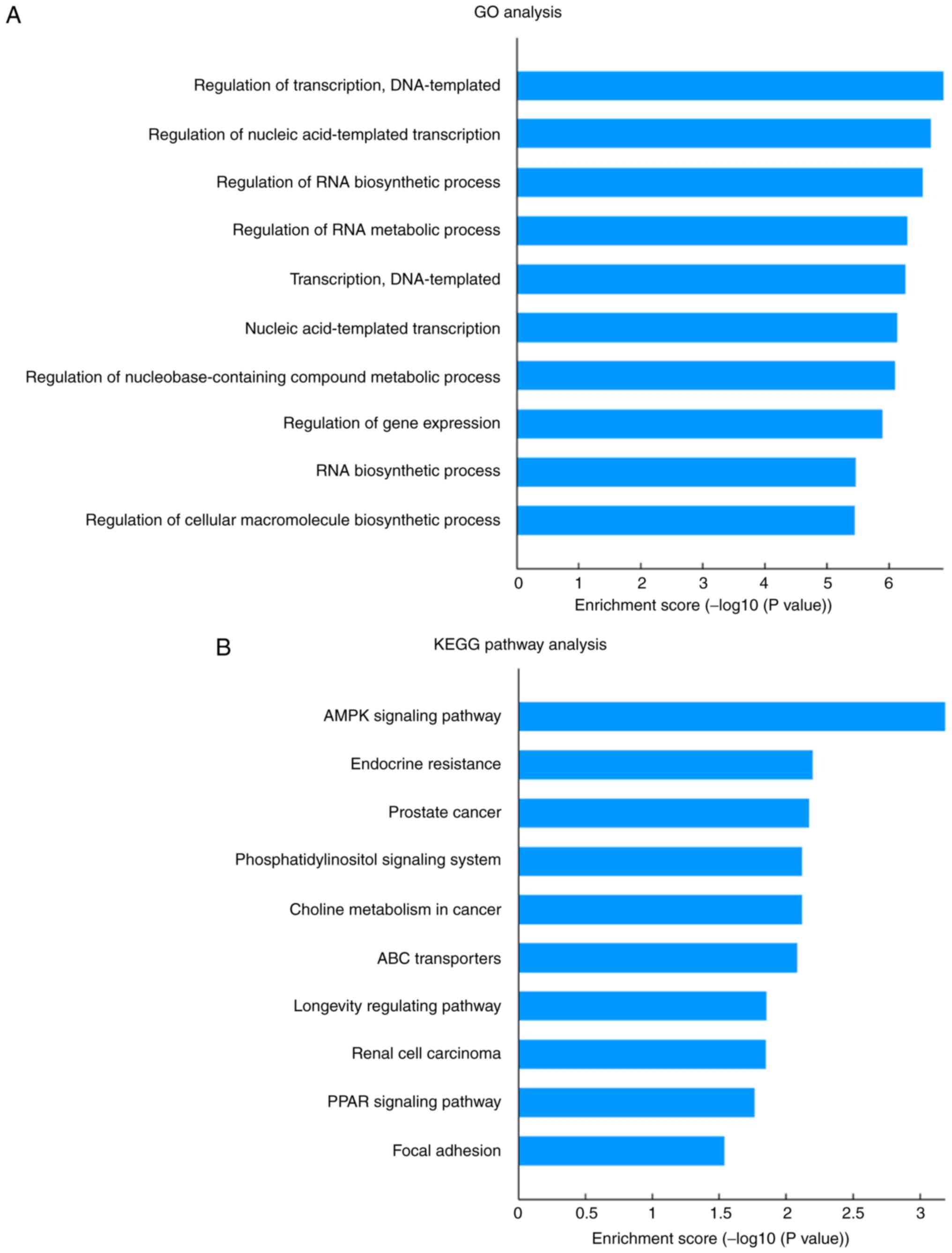
TargetScan. GO analysis of these predicted tRF targets revealed 220
significantly enriched GO terms for biological process. The top
five GO biological process terms were ‘regulation of transcription,
DNA-templated’, ‘regulation of nucleic acid-templated
transcription’, ‘regulation of RNA biosynthetic process’,
‘regulation of RNA metabolic process’ and ‘transcription,
DNA-templated’ (Fig. 4A). KEGG
pathway analysis revealed 15 significantly enriched pathways that
corresponded to the target genes (P<0.05; Fig. 4B). These pathways were mainly
angiogenesis- and metabolism-associated pathways, including ‘AMPK
signaling pathway’, ‘phosphatidylinositol signaling system’, ‘ABC
transporters’ and ‘endocrine resistance’. As shown in Fig. 5, a number of genes involved in the
AMP-activated protein kinase (AMPK) signaling pathway were targets
of tRFs.
Discussion
MMD is a complex disease with unknown
pathophysiology. Several factors, including genetic, acquired and
environmental factors, have been implicated in MMD (26). Previous studies have demonstrated
that familial MMD is inherited in an autosomal dominant manner with
incomplete penetrance; therefore, genetic factors may serve a
pivotal role in MMD development and progression (26,27).
Through the extensive development of high-throughput sequencing,
tRFs have become a highly studied topic in the field of biology and
medicine. Numerous studies have been conducted to explore their
biogenesis and biological functions (11,13,28,29);
however, to the best of our knowledge, there are no reports
demonstrating tRF expression profiles in patients with MMD, and the
association between tRFs and MMD. In the present study, the
expression of tRFs in patients with MMD and healthy controls were
explored; 38 tRFs were differentially expressed and maybe linked to
the development of the disease.
tRFs are a type of small non-coding RNA, which were
first identified in 1977 in the urine and serum of patients with
cancer (30,31). Due to their size, tRFs are broadly
classified into two groups. tiRNAs are 30–40 nucleotides (nt) long
and are generated by specific cleavage at the anticodon loop of
full-length tRNAs. Another class of tRFs is 18–22 nt long and
consists of four subtypes: tRF-1, tRF-3, tRF-5 and i-tRF. Although
the function of tRFs remains largely undefined, increasing evidence
suggests that tRFs can act in canonical miRNA pathways (19,22,32–34).
For example, Maute et al revealed that CU1276, a tRF-3
derived from tRNAGly(GCC), is abundantly expressed in
normal germinal center B cells and manifests the same functional
features as an miRNA (22). In
addition, it has been demonstrated that this particular tRF can
bind AGO proteins and post-transcriptionally repress replication
protein A1 in a sequence-specific miRNA-like manner, demonstrating
a clear regulatory role for tRFs. In 2014, Park et al
compared the expression levels of miRNAs in patients with advanced
breast cancer and healthy controls; it was revealed that miRNA-1280
is differentially expressed in patients and could reflect tumor
status, highlighting its potential as a clinical biomarker.
Subsequently, miRNA-1280 has been confirmed to be a tRF from
tRNALeu rather than a classical miRNA (34). In addition, tRFs have been reported
to have roles in cerebrovascular diseases. By using deep sequencing
to profile small RNA transcripts in rat brain tissues, Li et
al revealed that the expression of tRFs is significantly
enhanced following ischemia (28).
Two specific tRFs derived from tRNAVal and
tRNAGly could suppress angiogenesis by inhibiting the
function of vascular endothelial cells, thus suggesting that these
tRFs may have important roles in cerebral ischemic stroke.
It had previously been demonstrated that miRNAs and
lncRNAs are differentially expressed in patients with MMD, and both
are considered to be associated with MMD pathogenesis (5,6). In
the present study, tRF expression profiles in peripheral blood
samples were investigated, and it was revealed that tRF expression
levels in patients with MMD were altered compared with in healthy
controls. Amongst the differentially expressed tRFs, 5′-tiRNA
derived from tRNAGlu(TTC) was the most significantly
upregulated tRF, whereas i-tRF from tRNAGln(CTG) and
tRF-1 from pre-tRNAGln(CTG) were significantly
downregulated. The results suggested that these tRFs may have the
potential to serve as novel candidate biomarkers for MMD. In
addition, tRFs from tRNAGly and tRNAVal were
upregulated in patients with MMD, which was in accordance with a
previous study performed in rat ischemic brain (28). This may be due to progressive
stenosis or occlusion of the intracranial vessels in patients with
MMD leading to reduced blood supply to the brain and cerebral
ischemia. By using GO enrichment analysis, it was identified that
these tRFs were mainly involved in biological regulation. Notably,
it has previously been reported that tRFs can act as signaling
molecules and modulators invarious biological processes (9,35,36).
AMPK is a stress-activated protein kinase, which can
be activated by nutrient deprivation, including hypoxia/ischemia,
vigorous exercise and starvation (37,38).
The activation of AMPK appears to be crucial for angiogenesis
(38), the main process through
which new blood vessels are formed. Adiponectin, an AMPK activator,
can promote angiogenesis in response to tissue ischemia by
activating AMPK signaling in endothelial cells (39). Vascular endothelial growth factor
(VEGF) is an endothelial cell-specific mitogen, which has a
critical role in angiogenesis (39). Nagata et al identified that
suppression of AMPK signaling could inhibit endothelial cell
migration to VEGF and in vitro differentiation into
tube-like structures under hypoxic conditions, further suggesting
that AMPK signaling is essential for angiogenesis in response to
ischemic stress (37). Notably,
aberrant angiogenesis of blood vessel endothelial cells is
considered to be the pathological basis of the onset of MMD
(40). In the present study,
pathway enrichment analysis revealed the ‘AMPK signaling pathway’
as the most enriched pathway, which was composed of several tRF
target genes, indicating that tRFs may be involved in the
pathogenesis of MMD.
The phosphatidylinositol signaling system is another
affected pathway identified in the present study. Phosphoinositide
3-kinases (PI3Ks) are a conserved family of enzymes that
phosphorylate the 3′-OH of the inositol ring of
phosphatidylinositol (41). The
PI3K pathway has been reported to be closely correlated with
angiogenesis (41). Activation of
the PI3K/protein kinase B pathway can increase VEGF production. In
addition, this pathway can regulate the expression of other
angiogenic factors, including nitric oxide and angiopoietins
(41). The PI3K pathway also
serves a pivotal role in regulating the proliferation of vascular
smooth muscle cells (42).
Other pathways identified in the present study are
also closely associated with vascular diseases, including ‘ABC
transporters’, ‘ErbB signaling pathway’ and ‘Hippo signaling
pathway’. There are many signaling pathways that are associated
with MMD; therefore, it is difficult to verify each pathway
individually. Nevertheless, the results suggested that tRFs may be
involved in the development and progression of MMD, and provided
novel therapeutic strategies for clinical treatment of the disease.
However, one of the limitations of the present study was the small
sample size. Future studies with a larger sample size are required
to better understand the potential role of tRFs in the
pathophysiology of MMD. Additionally, the functions of tRFs in MMD
are based on bioinformatics prediction; therefore, further in
vitro and in vivo experiments are required to validate
the findings.
In conclusion, the expression profile of tRFs in
patients with MMD was revealed using small RNA sequencing, which
also identified tRFs that were expressed in an MMD-specific manner.
In addition, bioinformatics analyses were used to predict tRFs that
may be involved in the pathophysiology of MMD. The results of the
present study provided novel insight into the mechanisms underlying
MMD pathogenesis, and suggested tRFs that may serve as potential
therapeutic targets for the future treatment of MMD.
Acknowledgements
The authors would like to thank Dr Meng Wang
(Capital Institute of Pediatrics, Beijing, China) for proofreading
the manuscript.
Funding
The present study was supported by the National
Natural Science Foundation of China (grant no. 81371292) and the
‘13th Five-Year Plan’ National Science and Technology Supporting
Plan (grant no. 2015BAI12B04).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
CW conceived the study, reviewed the literature, and
wrote the initial and final drafts of the manuscript. MZ, JW, DZ
and SW collected, analyzed and interpreted the data. JZ designed
the present study, critically revised the manuscript and gave the
final approval for publication. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
This study was conducted in accordance with the
Declaration of Helsinki, and was approved by the Research Ethics
Board of Beijing Tiantan Hospital, Capital Medical University.
Written informed consent was obtained from all individuals who were
included in the study.
Patient consent for publication
All patients within this study provided consent for
the publication of their data.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Takeuchi K and Shimizu K: Hypoplasia of
the bilateral internal carotid arteries. Brain Nerve. 9:37–43.
1957.
|
|
2
|
Kim JS: Moyamoya disease: Epidemiology,
clinical features, and diagnosis. J Stroke. 18:2–11. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Acker G, Fekonja L and Vajkoczy P:
Surgical management of moyamoya disease. Stroke. 49:476–482. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kuriyama S, Kusaka Y, Fujimura M, Wakai K,
Tamakoshi A, Hashimoto S, Tsuji I, Inaba Y and Yoshimoto T:
Prevalence and clinicoepidemiological features of moyamoya disease
in Japan: Findings from a nationwide epidemiological survey.
Stroke. 39:42–47. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Dai D, Lu Q, Huang Q, Yang P, Hong B, Xu
Y, Zhao W, Liu J and Li Q: Serum miRNA signature in Moyamoya
disease. PLoS One. 9:e1023822014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Gao F, Yu L, Zhang D, Zhang Y, Wang R and
Zhao J: Long noncoding RNAs and their regulatory network: Potential
therapeutic targets for adult moyamoya disease. World Neurosurg.
93:111–119. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cole C, Sobala A, Lu C, Thatcher SR,
Bowman A, Brown JW, Green PJ, Barton GJ and Hutvagner G: Filtering
of deep sequencing data reveals the existence of abundant
Dicer-dependent small RNAs derived from tRNAs. RNA. 15:2147–2160.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kawaji H, Nakamura M, Takahashi Y,
Sandelin A, Katayama S, Fukuda S, Daub CO, Kai C, Kawai J, Yasuda
J, et al: Hidden layers of human small RNAs. BMC Genomics.
9:1572008. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lee YS, Shibata Y, Malhotra A and Dutta A:
A novel class of small RNAs: TRNA-derived RNA fragments (tRFs).
Genes Dev. 23:2639–2649. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Martens-Uzunova ES, Olvedy M and Jenster
G: Beyond microRNA-novel RNAs derived from small non-coding RNA and
their implication in cancer. Cancer Lett. 340:201–211. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kumar P, Kuscu C and Dutta A: Biogenesis
and function of transfer RNA-related fragments (tRFs). Trends
Biochem Sci. 41:679–689. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Anderson P and Ivanov P: TRNA fragments in
human health and disease. FEBS Lett. 588:4297–4304. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen Q, Yan M, Cao Z, Li X, Zhang Y, Shi
J, Feng GH, Peng H, Zhang X, Zhang Y, et al: Sperm tsRNAs
contribute to intergenerational inheritance of an acquired
metabolic disorder. Science. 351:397–400. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Blanco S, Dietmann S, Flores JV, Hussain
S, Kutter C, Humphreys P, Lukk M, Lombard P, Treps L, Popis M, et
al: Aberrant methylation of tRNAs links cellular stress to
neuro-developmental disorders. EMBO J. 33:2020–2039. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Research Committee on the Pathology and
Treatment of Spontaneous Occlusion of the Circle of Willis; Health
Labour Sciences Research Grant for Research on Measures for
Infractable Diseases, : Guidelines for diagnosis and treatment of
moyamoya disease (spontaneous occlusion of the circle of Willis).
Neurol Med Chir (Tokyo). 52:245–266. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Heid CA, Stevens J, Livak KJ and Williams
PM: Real time quantitative PCR. Genome Res. 6:986–994. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Shigematsu M and Kirino Y: tRNA-derived
short non-coding RNA as interacting partners of argonaute proteins.
Gene Regul Syst Bio. 9:27–33. 2015.PubMed/NCBI
|
|
19
|
Karaiskos S and Grigoriev A: Dynamics of
tRNA fragments and their targets in aging mammalian brain. F1000Res
5. ISCB Comm J. 27582016.
|
|
20
|
Haussecker D, Huang Y, Lau A, Parameswaran
P, Fire AZ and Kay MA: Human tRNA-derived small RNAs in the global
regulation of RNA silencing. RNA. 16:673–695. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kumar P, Anaya J, Mudunuri SB and Dutta A:
Meta-analysis of tRNA derived RNA fragments reveals that they are
evolutionarily conserved and associate with AGO proteins to
recognize specific RNA targets. BMC Biol. 12:782014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Maute RL, Schneider C, Sumazin P, Holmes
A, Califano A, Basso K and Dalla-Favera R: tRNA-derived microRNA
modulates proliferation and the DNA damage response and is
down-regulated in B cell lymphoma. Proc Natl Acad Sci USA.
110:1404–1409. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Miyoshi K, Miyoshi T and Siomi H: Many
ways to generate microRNA-like small RNAs: Non-canonical pathways
for microRNA production. Mol Genet Genomics. 284:95–103. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Suzuki J and Takaku A: Cerebrovascular
‘moyamoya’ disease. Disease showing abnormal net-like vessels in
base of brain. Arch Neurol. 20:288–299. 1969. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
van Swieten JC, Koudstaal PJ, Visser MC,
Schouten HJ and van Gijn J: Interobserver agreement for the
assessment of handicap in stroke patients. Stroke. 19:604–607.
1988. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Scott RM and Smith ER: Moyamoya disease
and moyamoya syndrome. N Engl J Med. 360:1226–1237. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Mineharu Y, Liu W, Inoue K, Matsuura N,
Inoue S, Takenaka K, Ikeda H, Houkin K, Takagi Y, Kikuta K, et al:
Autosomal dominant moyamoya disease maps to chromosome 17q25.3.
Neurology. 70:2357–2363. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Li Q, Hu B, Hu GW, Chen CY, Niu X, Liu J,
Zhou SM, Zhang CQ, Wang Y and Deng ZF: tRNA-derived small
non-coding RNAs in response to ischemia inhibit angiogenesis. Sci
Rep. 6:208502016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ivanov P: Emerging roles of tRNA-derived
fragments in viral infections: The case of respiratory syncytial
virus. Mol Ther. 23:1557–1558. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Borek E, Baliga BS, Gehrke CW, Kuo CW,
Belman S, Troll W and Waalkes TP: High turnover rate of transfer
RNA in tumor tissue. Cancer Res. 37:3362–3366. 1977.PubMed/NCBI
|
|
31
|
Speer J, Gehrke CW, Kuo KC, Waalkes TP and
Borek E: tRNA breakdown products as markers for cancer. Cancer.
44:2120–2123. 1979. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Karaiskos S, Naqvi AS, Swanson KE and
Grigoriev A: Age-driven modulation of tRNA-derived fragments in
Drosophila and their potential targets. Biol Direct. 10:512015.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Sobala A and Hutvagner G: Transfer
RNA-derived fragments: Origins, processing, and functions. Wiley
Interdiscip Rev RNA. 2:853–862. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Park IH, Kang JH, Lee KS, Nam S, Ro J and
Kim JH: Identification and clinical implications of circulating
microRNAs for estrogen receptor-positive breast cancer. Tumour
Biol. 35:12173–12180. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yeung ML, Bennasser Y, Watashi K, Le SY,
Houzet L and Jeang KT: Pyrosequencing of small non-coding RNAs in
HIV-1 infected cells: Evidence for the processing of a
viral-cellular double-stranded RNA hybrid. Nucleic Acids Res.
37:6575–6586. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Gebetsberger J and Polacek N: Slicing
tRNAs to boost functional ncRNA diversity. RNA Biol. 10:1798–1806.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Nagata D, Mogi M and Walsh K:
AMP-activated protein kinase (AMPK) signaling in endothelial cells
is essential for angiogenesis in response to hypoxic stress. J Biol
Chem. 278:31000–31006. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Fisslthaler B and Fleming I: Activation
and signaling by the AMP-activated protein kinase in endothelial
cells. Circ Res. 105:114–127. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Ouchi N, Shibata R and Walsh K:
AMP-activated protein kinase signaling stimulates VEGF expression
and angiogenesis in skeletal muscle. Circ Res. 96:838–846. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Chmelova J, Kolar Z, Prochazka V, Curik R,
Dvorackova J, Sirucek P, Kraft O and Hrbac T: Moyamoya disease is
associated with endothelial activity detected by anti-nestin
antibody. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub.
154:159–162. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Karar J and Maity A: PI3K/AKT/mTOR pathway
in angiogenesis. Front Mol Neurosci. 4:512011. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Morello F, Perino A and Hirsch E:
Phosphoinositide 3-kinase signalling in the vascular system.
Cardiovasc Res. 82:261–271. 2009. View Article : Google Scholar : PubMed/NCBI
|