Introduction
Hepatocellular carcinoma (HCC) is the fifth most
common malignancy and the second leading cause of cancer-related
deaths worldwide (1). Potentially
curative treatment strategies such as tumor resection, liver
transplantation, or local ablation are only limited to HCC at early
stage (2). Patients with advanced
stage HCC show poor responses to traditional chemotherapy and
sorafenib is the only approved systemic therapy for advanced HCC,
however, the response rate is quite low (3,4).
Therefore, new therapeutic strategies for HCC treatment are
urgently needed.
1-benzyl-3-(5′-hydroxymethyl-2′-furyl) indazole
(YC-1) is a synthetic compound that exhibits various potent
biological and pathological activities, including antiplatelet
activity, soluble guanylyl cyclase activity, suppression of
hypoxia-induced factor-1α (HIF-1α), and anti-cancer activity
(5). A previous report showed that
YC-1 exerts antitumor effects in HCC through suppressing HIF-1α and
inducing S cell cycle arrest or G0-G1 cell cycle arrest (6,7). YC-1 also
enhanced chemosensitivity in HCC via inhibition of signal
transducer and activator of transcription (STAT3) activity
(8). Our previous study also
suggested that YC-1 enhanced the antitumor activity of sorafenib
through STAT3 in HCC (9). These
studies suggest that YC-1 acts as an HIF-1α inhibitor or sensitizer
to enhance the effect of chemotherapeutics, however, the precise
mechanisms underlying the antitumor effects of YC-1 against HCC
have not been fully elucidated.
ATPase inhibitory factor 1 (IF1) is encoded by the
ATPIF1 gene that is located in chromosomes 1 and 4 of human
and mouse genomes, respectively (10). IF1 is implicated in the control of
both mitochondrial bioenergetics and structure by regulating the
activity and oligomerization of the F1Fo-ATPsynthase (hereinafter
referred to as ATP synthase) (11–16). The
function of IF1 as an ATP synthase inhibitor is regulated by matrix
pH under conditions of mitochondrial de-energization and by the
phosphorylation of S39 under several physiological situations, such
as progression through the cell cycle, hypoxia, and rapid changes
in metabolic demand (11). A previous
study demonstrated that overexpression of HIF-1α by hypoxia or
CoCl2 treatment augmented IF1 protein levels in a rat
hepatic epithelial cell line (17).
The pathway may be suitable to the mechanism of IF1 in HCC. Some
studies showed that IF is upregulated in many human carcinomas and
the level of IF1 expression in HCC, gliomas, and gastric cancers
correlates with aggressiveness and invasiveness of tumors and poor
prognosis in patients (14,18). Whether IF1 is involved in the
antitumor effect of YC-1 against HCC remains unclear.
In the present study, we examined the function of
IF1 in HCC cells and the potential role or involvement of IF1 in
the antitumor effects of YC-1 in the HCCLM3 and Huh7 HCC cell lines
in vitro.
Materials and methods
Cell lines and cell culture
Human HCC cell line Huh7 was obtained from the
American Type Culture Collection (Manassas, VA, USA). The human HCC
cell line HCCLM3 was obtained from the China Center for Type
Culture Collection and Cell Bank of the Chinese Academy of Sciences
(Shanghai, China). Cells were maintained in high-glucose Dulbecco's
modified Eagle's medium (DMEM) supplemented with 10% fetal bovine
serum (FBS), 100 U/ml penicillin and 100 µg/ml streptomycin (Thermo
Fisher Scientific, Inc., Waltham, MA, USA) in a humidified
atmosphere of 5% CO2 at 37°C. Cell lines were
immediately expanded and frozen down such that cell lines could be
restarted every 3 months from a frozen vial of the same batch of
cells. No further authentication was done. All cell lines were
routinely tested to rule out mycoplasma contamination. YC-1 was
obtained from Sigma-Aldrich (Merck KGaA, Darmstadt, Germany).
Establishment of stable knockdown or
overexpression cell lines
Lentiviral vectors encoding the human IF1 gene
(NM_016311.4) and shRNA-IF1 (CACCATGAAGAAGAAATCGTT) were
constructed by Beijing LiKeli BioTECH Co. Ltd. (Beijing, China) and
designated as LV-IF1 and LV-shRNA-IF1, respectively. The empty
vectors (LV-Ctrl or LV-shRNA-Ctrl) were used as a negative control.
The lentiviral vectors were transfected into HCC cells with a
multiplicity of infection of 20 to 30 in the presence of polybrene
(2 µg/ml). At 48 h after transfection, transfected cells were
selected for 2 weeks with 2 µg/ml puromycin (Sigma-Aldrich; Merck
KGaA). Pooled populations of knockdown cells and overexpression
cells, which were obtained 2 weeks after drug selection without
subcloning, were used in in vitro experiments.
Proliferation assay
Cell proliferation was analyzed using the
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
assay. Briefly, HCC cells were seeded in 96-well plates at a
density of 3×103 cells/well and treated with various
concentrations of YC-1 for 24, 48, or 72 h. MTT solution was added
to each well at a final concentration of 0.5 mg/ml and cells were
incubated for 4 h. Formazan crystals resulting from MTT reduction
were then dissolved by the addition of 150 µl dimethyl sulfoxide
per well. The absorbance was measured at 570 nm using an automated
ELISA plate reader.
Colony formation assay
Six-well dishes were seeded with 1×103
cells and cells were cultured for 24 h. The cells were then
incubated in the presence of various concentrations of YC-1 for 24
h in complete medium, washed with media, and allowed to grow in
complete medium for 2 weeks. The obtained colonies were washed with
PBS, fixed in 4% paraformaldehyde for 20 min at room temperature,
washed with PBS, and then stained with crystal violet. The stained
colonies were then counted.
Invasion assay
Cell invasion assays were performed using a modified
Boyden chamber (Costar; Corning Inc., Corning, NY, USA) that was
precoated with Matrigel. HCCLM3 or Huh7 cells (2×104
cells per well) and 10 µM YC-1 were added into the upper chamber,
and 600 µl DMEM with 10% FBS were added into the lower chamber. The
chambers were incubated for 24 h. After removing the filter inserts
and the cells on the upper side of the filter, the migrated cells
on the lower chamber were stained with crystal violet for 20 min,
washed with PBS, and photographed under an inverted fluorescence
microscope (Olympus IX51) equipped with an Olympus Qcolor 3 digital
camera (both Olympus Corp., Tokyo, Japan). Migration was assessed
by counting the number of stained cells from 5 random fields at
magnification, ×10.
Western blot analysis
Equivalent amounts of whole cell extracts were
subjected to SDS-PAGE and transferred to nitrocellulose membranes.
The membranes were blocked with 5% non-fat milk for 2 h and then
incubated with primary antibodies overnight at 4°C, followed by
incubation with the appropriate HRP-conjugated secondary antibody
for 1.5 h at room temperature. Immunoreactivity was detected with
SuperSignal West Pico substrate (Thermo Fisher Scientific, Inc.)
according to the manufacturer's instructions. Primary antibodies
anti-IF1, anti-E-cadherin, anti-phosphorylated (p-)STAT3, and
anti-β-actin were obtained from CST (Danvers, MA, USA). Horseradish
peroxidase (HRP)-labeled anti-mouse and anti-rabbit secondary
antibodies were from Santa Cruz Biotechnology, Inc. (Dallas, TX,
USA). All other antibodies were purchased from Abcam (Cambridge,
MA, USA).
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total mRNA was extracted using the TRIzol reagent
(Invitrogen; Thermo Fisher Scientific, Inc.) and RT was performed
using an RT-PCR kit (TransGen Biotech Co., Ltd., Beijing, China).
qPCR experiments were conducted on an DNA Engine Opticon System (MJ
Research Inc.; Bio-Rad Laboratories, Inc.) using SYBR-Green PCR
Master Mix kit in triplicate specific primers. The sequences of
primers to determine the expression of the target gene were listed
as follows: IF1 forward, 5′-GGGCCTTCGGAAAGAGAG-3′ and reverse,
5′-TTCAAAGCTGCCAGTTGTTC-3′; and glyceraldehydes 3-phosphate
dehydrogenase (GAPDH) forward, 5′-CGGAGTCAACGGATTTGGTCGTAT-3′ and
reverse, 5′-AGCCTTCTCCATGGTGGTGAAGAC-3′. The PCR thermocycling
conditions consisted of 5 min at 95°C followed by 40 cycles of
denaturation for 30 sec at 95°C, annealing for 30 sec at 56°C and a
primer extension for 30 sec at 72°C. Relative gene expression
levels were quantified using the 2−∆∆Cq method (19).
Statistical analysis
All values are expressed as the mean ± standard
error of the mean. The data were analyzed using Student's t-test or
the analysis of variance test with Tukey's post hoc test. P<0.05
was considered to indicate a statistically significant difference.
GraphPad Prism 5 (GraphPad Software Inc., San Diego, CA, USA) was
used for these analyses.
Results
IF1 regulates the proliferation and
invasion of HCC cells
To investigate the function of IF1 in HCC, we first
examined the expression of IF1 in HCCLM3 and Huh7 cell lines.
Western blot analysis revealed that the expression of IF1 was
higher in HCCLM3 cells compared with Huh7 cells (Fig. 1A). To examine the effects of altered
IF1 expression, we transfected a lentiviral expression vector that
expresses IF1 in Huh7 cells (Huh7-LV-IF1) or shRNA for IF1
knockdown in HCCLM3 cells (HCCLM3-shRNA-IF1) and generated stable
cell lines, together with the appropriate controls (Huh7-LV-Ctrl
and HCCLM3-shRNA-Ctrl) and RT-qPCR was used to confirm the results
(Fig. 1B).
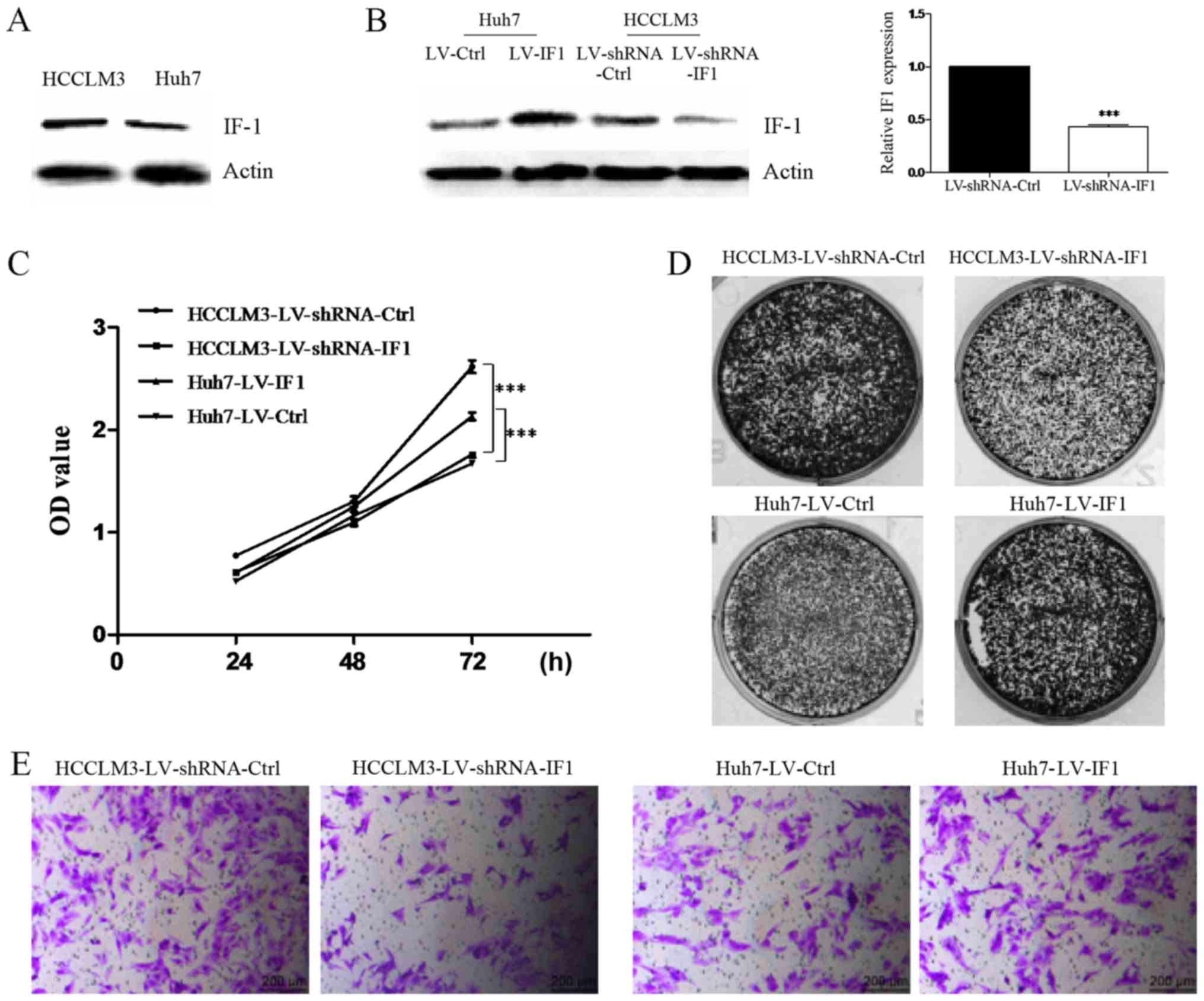 | Figure 1.IF1 regulates proliferation and
invasion of HCC cells. (A) Western blot analysis of IF1 in Huh7 and
HCCLM3 cells. Actin was used as loading control. (B) Lentiviral
overexpression of IF1 in Huh7 cells (Huh7-LV-IF1 or Huh7-LV-Ctrl)
or IF1 shRNA in HCCLM3 cells (HCCLM3-shRNA-IF1 or
HCCLM3-shRNA-Ctrl). Western blot analysis and RT-qPCR were
performed for the indicated proteins and genes. ***P<0.001 vs.
LV-shRNA-Ctrl. (C) MTT assays, (D) colony formation assays and (E)
invasion assays of HCC cells with IF1 overexpression or IF1 shRNA
knockdown compared with the appropriate controls (scale bars, 200
µm). ***P<0.001, as indicated. IF1, inhibitory factor 1; HCC,
hepatocellular carcinoma; shRNA, short hairpin RNA; Ctrl, control;
OD, optical density; MTT,
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide. |
We next performed a series of assays to clarify the
function of IF1 in HCC cells. We found that knockdown of IF1 could
significantly inhibit the proliferation, colony formation and
invasion activities of HCCLM3 cells compared with controls
(Fig. 1C-E). In contrast,
overexpression of IF1 in Huh7 cells resulted in increased
proliferation, colony formation and invasion (Fig. 1C-E). Together these results indicated
a role for IF1 in regulating proliferation and invasion of HCC
cells.
Overexpression of IF1 reduced the
inhibitory effects of YC-1 in Huh7 cells
We next examined whether IF1 could impact the
effects of YC-1 on HCC cells using Huh7-LV-IF1 and Huh7-LV-Ctrl
cells. Consistent with previous studies on YC-1 anti-cancer
effects, we found that YC-1 treatment reduced the proliferation,
colony formation and invasion activities of Huh7 cells (Fig. 2). However, Huh7 cells with
overexpression of IF1 showed reduced sensitivity to the negative
effects of YC-1 on proliferation and attenuated the YC-1-induced
inhibition of invasion (Fig. 2).
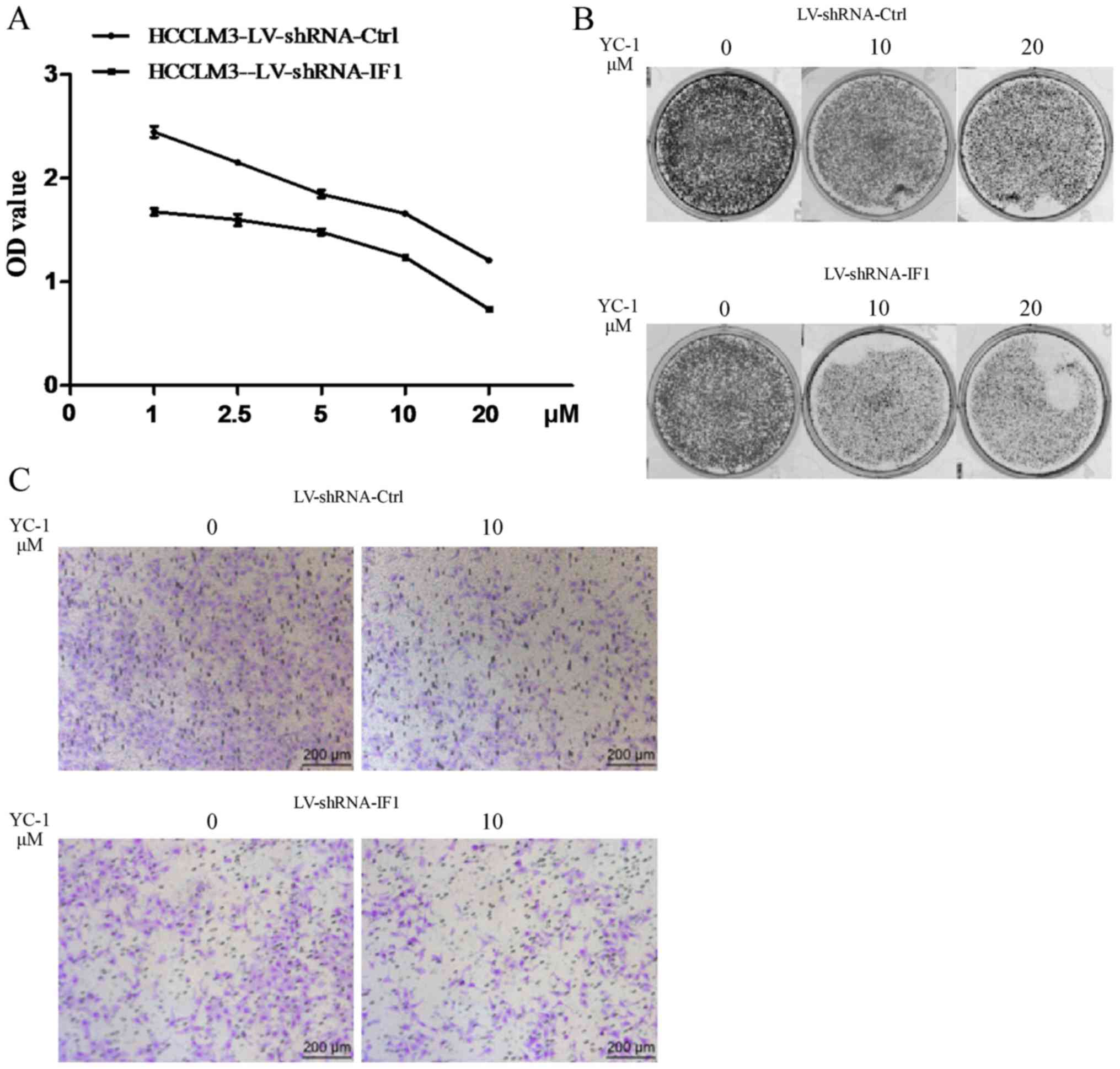
Knockdown of IF1 elevated the
sensitivity of HCCLM3 cells to YC-1
We next evaluated the effect of IF1 knockdown on the
sensitivity of HCC cells to YC-1. We found that HCCLM3 cells with
knockdown of IF1 showed elevated sensitivity to the inhibitory
effects of YC-1 on cell proliferation, colony formation and
invasion activities compared with cells transfected with the shRNA
control (Fig. 3).
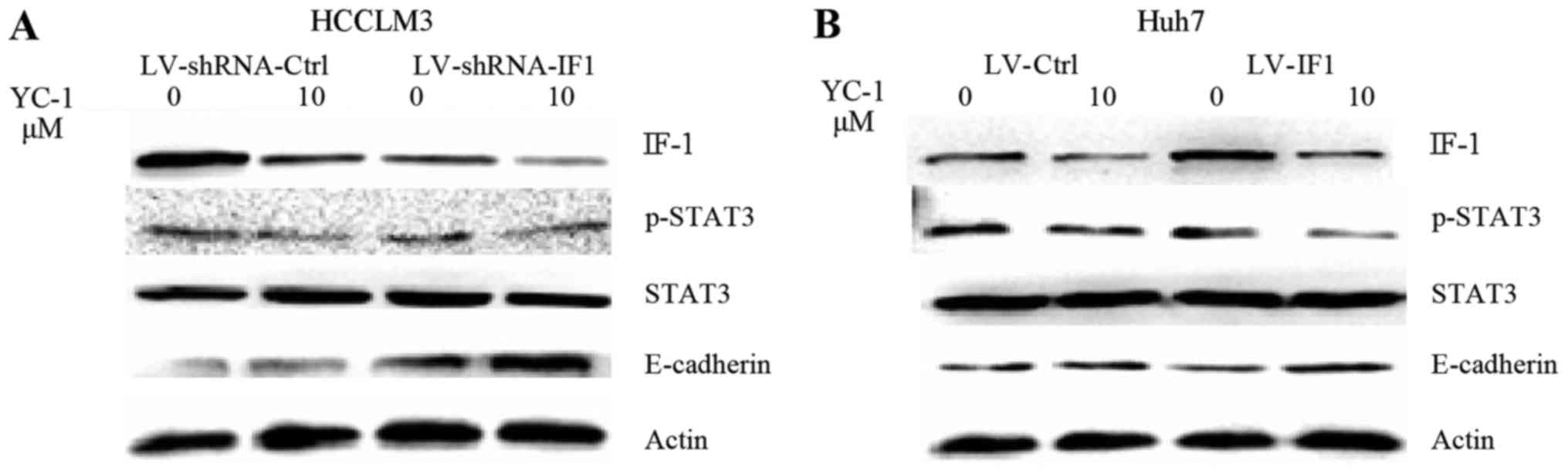
STAT3 activation is involved in the
effect of IF1 on YC-1 treatment in HCC cells
To investigate the underlying mechanism of the role
of IF1 in the effect of YC-1 on HCC cells, we examined a panel of
several molecular factors by western blot analysis. YC-1 treatment
in HCCLM3 and Huh7 cells decreased the expression of p-STAT3 and
IF1 and increased the expression of E-cadherin (Fig. 4). IF1 knockdown significantly reduced
the levels of p-STAT3 and increased the expression of E-cadherin,
while IF1 overexpression increased the expression of p-STAT3 and
decreased the expression of E-cadherin (Fig. 4).
Discussion
In the present study, we provided evidence that IF1
plays an important role in the antitumor effects of YC-1 in HCC.
Our results demonstrated that IF1 was involved in the
proliferation, colony formation and invasion activities of HCC
cells. Overexpression of IF1 reversed the inhibitory effects of
YC-1 in Huh7 cells, and knockdown of IF1 elevated the sensitivity
of HCCLM3 cells to YC-1. STAT3 signaling pathways may also be
involved in the process. Our results suggest that inhibition of IF1
could improve the antitumor effects of YC-1 against HCC.
IF1 is an inhibitor of the mitochondrial H (+)-ATP
synthase, and several studies have suggested that IF1 is involved
in tumor progression. IF1 was shown to be an independent prognostic
factor in non-small cell lung cancer (13). In the liver, IF1 could downregulate
oxidative phosphorylation and induce a tumor-promoting metabolic
state (20). Another study showed
that IF1 was a prognostic marker in gastric cancer and that IF1
contributes to proliferation and invasion of human gastric cancer
cells (14). Silencing of IF1 in
bladder cancer inhibited cell growth via cell cycle arrest
(15). A study in HCC showed that a
reciprocal activation between IF1 and NF-κB drives angiogenesis and
metastasis (16). Analysis of IF1
expression in tumors in breast and colon cancer patient cohorts
revealed its relevance as a predictive marker for clinical outcome
(21). Another report showed that
upregulation of IF1 in human tumors mediated the metabolic shift of
cancer cells to a Warburg phenotype (22). Our results also showed that knockdown
of IF1 inhibited the proliferation, colony formation and invasion
activities of HCC cells, while IF1 overexpression had the opposite
effect.
YC-1 functions to inhibit HIF-1α and suppress tumor
progression (23), and the direct
inhibiting growth of tumor is limit. Indeed, resistance to YC-1
necessitates the use of increased therapeutic doses and can result
in increased adverse side effects. A previous study on HCC cells
showed that YC-1 exhibited an antiproliferative effect and arrests
the cell cycle in G0-G1, however, the IC50 value was
about 46 µM (24). Notably, our
previous study demonstrated that YC-1 enhanced the antitumor
activity of sorafenib through inhibition of STAT3 in HCC (9). Another report showed that YC-1 could
potentiate the apoptotic effect of licochalcone A on human
epithelial ovarian carcinoma cells via activation of death receptor
and mitochondrial pathways (25).
These studies suggest YC-1 could be also used as a sensitizer to
enhance the effect of chemotherapeutics. Our current results
demonstrated that increased IF1 expression reduced the inhibitory
effects of YC-1 and knockdown of IF1 enhanced the growth and
invasion suppressive effects of YC-1 in HCC cells. These findings
indicate that IF1 may be involved in the effects of YC-1 on HCC
cells and suggest that a combination of silencing IF1 and YC-1 may
be a useful strategy for HCC treatment.
STAT3 is a central mediator of cancer metastasis and
a target of anticancer drugs for blocking tumor metastasis. STAT3
is activated by its binding to various ligands, such as
interleukin-6 (IL-6), interferon, and IL-10 (26–29).
Numerous studies have revealed that the JAK/STAT signaling pathway
is involved in several aspects of tumorigenesis, including
proliferation, apoptosis, angiogenesis, and metastasis (30,31).
Activated STAT3 has been reported in various human cancers, and
elevated level of activated STAT3 is associated with poor
prognosis. Our results showed that overexpression of IF1 could
induce the levels of p-STAT3, while IF1 knockdown reduced p-STAT3
levels. Another study showed that YC-1 could inhibit STAT3 activity
by enhancing the polyubiquitination of p-STAT3 (705) and
overexpression of STAT3 reversed YC-1-induced cell death. YC-1 may
also suppress the expression of p-STAT3 through inhibiting SHP-1
activity. Together this suggests that STAT3 may function as a
bridge in the interaction between IF1 and YC-1.
In conclusion, our results demonstrate that
inhibition of IF1 improves the antitumor effects of YC-1 in HCC
cells. This finding supports the clinical development of combining
YC-1 and an IF1 inhibitor for the treatment of HCC.
Acknowledgements
Not applicable.
Funding
The present study was supported by the National
Natural Science Foundation of China (grant nos. 81572957 and
81502650).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
WS conceived and supervised the study, and helped to
draft the manuscript. JG, SW and SK participated in the design of
the study. XD and JK performed the experiments and drafted the
manuscript. SD, WX, YD and CY analyzed the data. All authors read
and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that there are no competing
interests.
References
|
1
|
Forner A, Llovet JM and Bruix J:
Hepatocellular carcinoma. Lancet. 379:1245–1255. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Xu XL, Liu XD, Liang M and Luo BM:
Radiofrequency ablation versus hepatic resection for small
hepatocellular carcinoma: Systematic review of randomized
controlled trials with meta-analysis and trial sequential analysis.
Radiology. 287:461–472. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Niu L, Liu L, Yang S, Ren J, Lai PBS and
Chen GG: New insights into sorafenib resistance in hepatocellular
carcinoma: Responsible mechanisms and promising strategies. Biochim
Biophys Acta. 1868:564–570. 2017.PubMed/NCBI
|
|
4
|
Bruix J, Cheng AL, Meinhardt G, Nakajima
K, De Sanctis Y and Llovet J: Prognostic factors and predictors of
sorafenib benefit in patients with hepatocellular carcinoma:
Analysis of two phase III studies. J Hepatol. 67:999–1008. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chun YS, Yeo EJ and Park JW: Versatile
pharmacological actions of YC-1: Anti-platelet to anticancer.
Cancer Lett. 207:1–7. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Yeo EJ, Ryu JH, Chun YS, Cho YS, Jang IJ,
Cho H, Kim J, Kim MS and Park JW: YC-1 induces S cell cycle arrest
and apoptosis by activating checkpoint kinases. Cancer Res.
66:6345–6352. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lau CK, Yang ZF, Lam CT, Tam KH, Poon RT
and Fan ST: Suppression of hypoxia inducible factor-1alpha
(HIF-1alpha) by YC-1 is dependent on murine double minute 2 (Mdm2).
Biochem Biophys Res Commun. 348:1443–1448. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lau CK, Yang ZF, Lam SP, Lam CT, Ngai P,
Tam KH, Poon RT and Fan ST: Inhibition of Stat3 activity by YC-1
enhances chemo-sensitivity in hepatocellular carcinoma. Cancer Biol
Ther. 6:1900–1907. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Kong J, Kong F, Gao J, Zhang Q, Dong S, Gu
F, Ke S, Pan B, Shen Q, Sun H, et al: YC-1 enhances the anti-tumor
activity of sorafenib through inhibition of signal transducer and
activator of transcription 3 (STAT3) in hepatocellular carcinoma.
Mol Cancer. 13:72014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
García-Bermúdez J and Cuezva JM: The
ATPase inhibitory factor 1 (IF1): A master regulator of energy
metabolism and of cell survival. Biochim Biophys Acta.
1857:1167–1182. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Garcia-Ledo L, Nuevo-Tapioles C,
Cuevas-Martin C, Martínez-Reyes I, Soldevilla B, González-Llorente
L and Cuezva JM: Overexpression of the ATPase inhibitory factor 1
favors a non-metastatic phenotype in breast cancer. Front Oncol.
7:692017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Faccenda D, Nakamura J, Gorini G, Dhoot
GK, Piacentini M, Yoshida M and Campanella M: Control of
mitochondrial remodeling by the ATPase inhibitory factor 1 unveils
a pro-survival relay via OPA1. Cell Rep. 18:1869–1883. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Gao YX, Chen L, Hu XG, Wu HB, Cui YH,
Zhang X, Wang Y, Liu XD and Bian XW: ATPase inhibitory factor 1
expression is an independent prognostic factor in non-small cell
lung cancer. Am J Cancer Res. 6:1141–1148. 2016.PubMed/NCBI
|
|
14
|
Yin T, Lu L, Xiong Z, Wei S and Cui D:
ATPase inhibitory factor 1 is a prognostic marker and contributes
to proliferation and invasion of human gastric cancer cells. Biomed
Pharmacother. 70:90–96. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wei S, Fukuhara H, Kawada C, Kurabayashi
A, Furihata M, Ogura S, Inoue K and Shuin T: Silencing of ATPase
inhibitory factor 1 inhibits cell growth via cell cycle arrest in
bladder cancer. Pathobiology. 82:224–232. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Song R, Song H, Liang Y, Yin D, Zhang H,
Zheng T, Wang J, Lu Z, Song X, Pei T, et al: Reciprocal activation
between ATPase inhibitory factor 1 and NF-κB drives hepatocellular
carcinoma angiogenesis and metastasis. Hepatology. 60:1659–1673.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Huang LJ, Chuang IC, Dong HP and Yang RC:
Hypoxia-inducible factor 1α regulates the expression of the
mitochondrial ATPase inhibitor protein (IF1) in rat liver. Shock.
36:90–96. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wu J, Shan Q, Li P, Wu Y, Xie J and Wang
X: ATPase inhibitory factor 1 is a potential prognostic marker for
the migration and invasion of glioma. Oncol Lett. 10:2075–2080.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Santacatterina F, Sánchez-Cenizo L,
Formentini L, Mobasher MA, Casas E, Rueda CB, Martínez-Reyes I,
Núñez de Arenas C, García-Bermúdez J, Zapata JM, et al:
Down-regulation of oxidative phosphorylation in the liver by
expression of the ATPase inhibitory factor 1 induces a
tumor-promoter metabolic state. Oncotarget. 7:490–508. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sánchez-Aragó M, Formentini L,
Martinez-Reyes I, García-Bermudez J, Santacatterina F,
Sánchez-Cenizo L, Willers IM, Aldea M, Nájera L, Juarránz A, et al:
Expression, regulation and clinical relevance of the ATPase
inhibitory factor 1 in human cancers. Oncogenesis. 2:e462013.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Sánchez-Cenizo L, Formentini L, Aldea M,
Ortega AD, García-Huerta P, Sánchez-Aragó M and Cuezva JM:
Up-regulation of the ATPase inhibitory factor 1 (IF1) of the
mitochondrial H+-ATP synthase in human tumors mediates the
metabolic shift of cancer cells to a Warburg phenotype. J Biol
Chem. 285:25308–25313. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kong J, Kong J, Pan B, Ke S, Dong S, Li X,
Zhou A, Zheng L and Sun WB: Insufficient radiofrequency ablation
promotes angiogenesis of residual hepatocellular carcinoma via
HIF-1α/VEGFA. PLoS One. 7:e372662012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wang SW, Pan SL, Guh JH, Chen HL, Huang
DM, Chang YL, Kuo SC, Lee FY and Teng CM: YC-1
[3-(5′-Hydroxymethyl-2′-furyl)-1-benzyl Indazole] exhibits a novel
antiproliferative effect and arrests the cell cycle in G0-G1 in
human hepatocellular carcinoma cells. J Pharmacol Exp Ther.
312:917–925. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Lee CS, Kwak SW, Kim YJ, Lee SA, Park ES,
Myung SC, Kim W, Lee MS and Lee JJ: Guanylate cyclase activator
YC-1 potentiates apoptotic effect of licochalcone A on human
epithelial ovarian carcinoma cells via activation of death receptor
and mitochondrial pathways. Eur J Pharmacol. 683:54–62. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Huynh J, Etemadi N, Hollande F, Ernst M
and Buchert M: The JAK/STAT3 axis: A comprehensive drug target for
solid malignancies. Semin Cancer Biol. 45:13–22. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Bixel K, Saini U, Kumar Bid H, Fowler J,
Riley M, Wanner R, Deepa Priya Dorayappan K, Rajendran S, Konishi
I, Matsumura N, et al: Targeting STAT3 by HO3867 induces apoptosis
in ovarian clear cell carcinoma. Int J Cancer. 141:1856–1866. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Khan MW, Saadalla A, Ewida AH, Al-Katranji
K, Al-Saoudi G, Giaccone ZT, Gounari F, Zhang M, Frank DA and
Khazaie K: The STAT3 inhibitor pyrimethamine displays anti-cancer
and immune stimulatory effects in murine models of breast cancer.
Cancer Immunol Immunother. 67:13–23. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zeng R, Tang Y, Zhou H, Liu Y, Huang J, Li
L, Liu W, Feng Y, Zhou Y, Chen T, et al: STAT3 mediates multidrug
resistance of Burkitt lymphoma cells by promoting antioxidant
feedback. Biochem Biophys Res Commun. 488:182–188. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Gao P, Niu N, Wei T, Tozawa H, Chen X,
Zhang C, Zhang J, Wada Y, Kapron CM and Liu J: The roles of signal
transducer and activator of transcription factor 3 in tumor
angiogenesis. Oncotarget. 8:69139–69161. 2017.PubMed/NCBI
|
|
31
|
Kim BH, Yi EH and Ye SK: Signal transducer
and activator of transcription 3 as a therapeutic target for cancer
and the tumor microenvironment. Arch Pharm Res. 39:1085–1099. 2016.
View Article : Google Scholar : PubMed/NCBI
|