Introduction
Gastric cancer (GC) is one of the most common
malignancies with ~1 million new cases reported globally every year
according to the GLOBOCAN (2002) and Cancer Incidence in Five
Continents databases (1). The
mortality rate of GC is the second highest amongst all malignant
tumors (2). A good prognosis in
patients with GC requires a timely diagnosis and is also associated
with different pathological characteristics, genetic background and
the treatment method used (3,4). Due to
developments in cellular and molecular biology, understanding of
the pathogenesis of GC has gradually increased over the past 20
years, but the overall survival rate of patients remains unchanged
(5). Tumor related molecules,
signaling pathways, proteases and their inhibitors are all involved
in the process of tumor development (6,7).
Therefore, the molecular analysis of these processes has important
significance in the development of therapeutics and the prognosis
of GC in clinical practice (8,9).
Transforming growth factor β1 (TGFβ1) is a type of
polypeptide cytokine with multiple functions in humans (10). Almost all cells in the body can
produce TGFβ1 and its receptors, including epithelial, endothelial,
hematopoietic, nerve and connective tissue cells (11). Assoian et al successfully
extracted TGFβ1 from human platelets for the first time in 1983
(8,12). TGFβ1 has since been reported to play
an important role in the regulation of cellular proliferation
(12). TGFβ receptors (TGFβR) are
high affinity binding proteins of TGFβ1 located on the cell
membrane (13). These receptors have
been categorized into 3 isoforms according to electrophoretic
mobility; TGFβR1, TGFβR2 and TGFβR3 (14). By binding to TGFβR, TGFβ1 exerts a
wide range of biological effects (14). Previous studies have focused on the
relationship of TGFβ1 and TGFβRs with cancer (14,15).
TGFβ1 demonstrates diverse functions in tumors, such as the
inhibition of cell proliferation, differentiation and apoptosis in
the early stages of tumor development (14). In advanced stage cancer, TGFβ
promotes angiogenesis, induction of extracellular matrix
production, invasion and metastasis (16,17).
TGFβ1 and TGFβR are important members of the TGFβ/SMAD signaling
pathway, which is involved in the regulation of cell proliferation
and differentiation. The TGFβ/SMAD pathway is one of the most
frequently altered signaling pathways in tumors, including GC
(18–20).
The online Kaplan-Meier plotter (K-M plotter) is
capable of assessing the effect of any gene or gene combination on
survival in breast, ovarian, lung and gastric cancer, using patient
samples on gene chips or RNA-seq data (21). To date, the K-M plotter has been used
to identify and validate a number of genes in these cancer types
(22–27). The K-M plotter database contains the
prognostic and mRNA mapping information of 876 patients with GC
(21). In the present study, the K-M
plotter was used to determine the prognostic value of mRNA
expression of TGFβ1 and its receptors in patients with GC, and the
effects of TGFβ1 were validated in GC cell lines.
Materials and methods
Prognostic analyses of patients with
GC
Using the K-M plotter (kmplot.com/analysis/) the
association between the mRNA expression of TGFβ1 and its receptors,
and overall survival (OS) time was analyzed. Using the K-M plotter
online software, gene expression, relapse free and OS time data can
be downloaded from the Gene Expression Omnibus (Affymetrix
microarrays only), the European Genome-Phenome Archive and The
Cancer Genome Atlas databases
(kmplot.com/analysis/index.php?p=service&cancer=gastric).
Clinical data were collected from 876 patients with GC, including
sex, perforation history, Tumor Node Metastasis (TNM) stage
(28), Lauren classification
(29), HER2 status, pathological
grade and treatment method. The mRNA expression levels of TGFβ1 and
its receptors were entered into the database, and Kaplan-Meier
survival curves were generated for the OS time of patients with GC.
The patients were split into low- and high-expression groups
according to the expression levels of TGFβ1 and its receptors with
auto select best cutoff. The log rank P-value and the hazard ratio
(HR) with a 95% confidence interval (CI) was calculated.
Cell culture and transfection
The AGS and MKN45 human gastric cancer cell lines
were purchased from the Cell Bank of the Chinese Academy of
Sciences. Cells were cultured in DMEM supplemented with 10% fetal
bovine serum, 100 U/ml penicillin and 0.1 mg/ml streptomycin, and
incubated in a 5% CO2 incubator at 37°C for 48 h. Cells
in the exponential growth phase were harvested and transfected with
TGFβ1, TGFβR1, TGFβR2- or TGFβR3-specific siRNA (3 µg) using
Lipofectamine® 2000 (Thermo Fisher Scientific, Inc)
according to the manufacturer's protocol. The cells were incubated
for 48 h prior to further experimentation. Following siRNAs were
used (Ruibo; ribobio.com/): TGFβ1, 5′-GCCCATCTAGGTTATTTCCGTGG-3′;
TGFβR1, 5′-AGGGTACTACGTTGAAAGACTTA-3′; TGFβR2,
5′-ACGATAATGTTTGGTAGTATTCA-3′; TGFβR3,
5′-AACTTAAGATAGCAAGAAATATC-3′; negative control siRNA (a scrambled
siRNA control, siC) 5′-UUCUCCGAACGUGUCACGUTT-3′. Untreated AGS and
MKN45 cells were used as the blank control, and cells treated with
the scrambled siRNA control were used as the negative control.
Cell Counting Kit-8 (CCK-8) assay
After transfection, cells (1×103
cells/well) were seeded into a 96-well plate, cultured at 37°C in a
5% CO2 incubator. The proliferation of cells was
measured every 24 h. Fresh DMEM containing 10 µl CCK-8 solution
(Beijing Solarbio Science & Technology Co., Ltd.) was added to
each well to detect cell proliferation according to the
manufacturer's protocol. After incubation for 2 h at 37°C, cell
proliferation was determined by measuring the optical density (OD)
value at a wavelength of 450 nm. The CCK-8 assay was performed in
triplicate.
Transwell migration and invasion
assays
Following transfection, cells (1×103
cells/well) in serum-free medium were seeded into the upper
chambers of transwell inserts, while medium supplemented with 10%
FBS was added into the lower chambers. After incubation for 48 h,
the cells that had migrated into the lower chamber were fixed with
4% paraformaldehyde for 15 min and stained with 0.1% crystal violet
for 5 min. Images were captured using a light microscope at ×100
magnification. For the invasion assay, the upper chambers were
coated with Matrigel prior at 37°C for 4 h to the addition of the
cells.
Reverse transcription fluorescence
quantitative PCR (RTq-PCR)
An Ultrapure RNA kit (CWBio) was used of the
extraction of total RNA form AGS and MKN45 cells after the
transfection for 24 h. A HiFiScript cDNA Synthesis kit (CWBio) was
used for reverse transcription. The following thermocycling
conditions were used for reverse transcription: Incubation at 42°C
for 15 min and at 85°C for 5 min. Then, qPCR was performed using
MagicSYBR Mixture (CWBio). The following primers was used: TGFβ1
forward, 5′-CCCCTACATTTGGAGCCTGG-3′ and reverse,
5′-GCACGATCATGTTGGACAGC-3′; TGFβR1 forward,
5′-ACCGCACTGTCATTCACCAT-3′ and reverse, 5′-CTGAGCCAGAACCTGACGTT-3′;
TGFβR2 forward, 5′-GCTCTGGTGCTCTGGGAAAT-3′ and reverse,
5′-CCAGCACTCAGTCAACGTCT-3′; TGFβR3 forward,
5′-GCCCTGATGAGCTCCTGTTT-3′ and reverse, 5′-GGCACAGCCTGACAAAACAG-3′;
β-actin forward, 5′-CCCGAGCCGTGTTTCCT-3′ and reverse,
5′-GTCCCAGTTGGTGACGATGC-3′. The following thermocycling conditions
were used for qPCR: Initial denaturation at 95°C for 30 sec; 95°C
for 5 sec, 60°C for 30 sec, with a total of 40 cycles. The relative
expression levels of genes were analyzed using 2−ΔΔCq
method (30).
Western blotting
Total protein was extracted from the AGS and MKN45
cells after the transfection for 48 h using a RIPA lysis buffer
(Beijing Solarbio Science & Technology Co., Ltd.). Protein
determination was detected using a BCA Protein Assay kit (CWBio). A
total of 20 µg protein of each group was loaded on a 10% gel,
resolved using SDS-PAGE and subsequently transferred to a PVDF
membrane. The membrane was blocked with 5% non-fat milk at room
temperature for 1 h. The protein was incubated with primary
antibodies for at 4°C overnight and secondary antibodies at room
temperature for 1 h. The following antibodies were used in this
research: Anti-TGFβ1 antibody (1:500; ab92486; Abcam), anti-TGFβR1
antibody (1:500; ab31013; Abcam), anti-TGFβR2 antibody (1:500;
ab186838; Abcam), anti-TGFβR3 antibody (1:200; ab97459; Abcam) and
goat anti-rabbit secondary antibody (1:5000; ab6721; Abcam). An
Enhanced ECL Chemiluminescent Substrate kit (Shanghai Maokang;
maokangbio.com/index.action) was used for visualization. Protein
level was analyzed using ImageJ version 1.41 (National Institutes
of Health).
Statistical analysis
SPSS 20.0 (IBM Corp.) was used for the statistical
analysis. All data in the present study are presented as the mean ±
SD. The data were analyzed from three separate experiments.
Statistical significance was determined using one-way ANOVA
followed by the Bonferroni's post-hoc test. P<0.05 was
considered to indicate a statistically significant difference.
Results
Low expression of TGFβ1 and its
receptors is associated with improved prognosis in patients with
GC
The prognostic values of the mRNA expression of
TGFβ1 and its receptors was determined using the online K-M plotter
tool. The Affymetrix IDs of TGFβ1, TGFβR1, TGFβR2 and TGFβR3 are
203084_at, 206943_at, 207334_s_at and 204731_at respectively.
Survival curves were generated for all patients with GC (n=876),
patients with intestinal type GC (n=320) and patients with diffuse
type GC (n=241). In 876 cases, only the above patients have clear
pathological classification information, therefore only these
patient data were analyzed.
Firstly, the prognostic value of TGFβ1 mRNA
expression was determined (Fig. 1).
Low mRNA expression levels of TGFβ1 was associated with higher OS
time and therefore, improved prognosis in patients with GC (HR,
1.53; 95% CI, 1.24–1.90; P<0.0001; Fig. 1A). Low TGFβ1 mRNA expression was also
observed to be associated with a higher OS time in patients with
intestinal type GC (HR, 1.55; 95% CI, 1.04–2.30; P=0.028; Fig. 1B), and patients with diffuse type GC
(HR, 2.09; 95% CI, 1.35–3.26; P=0.00081; Fig. 1C).
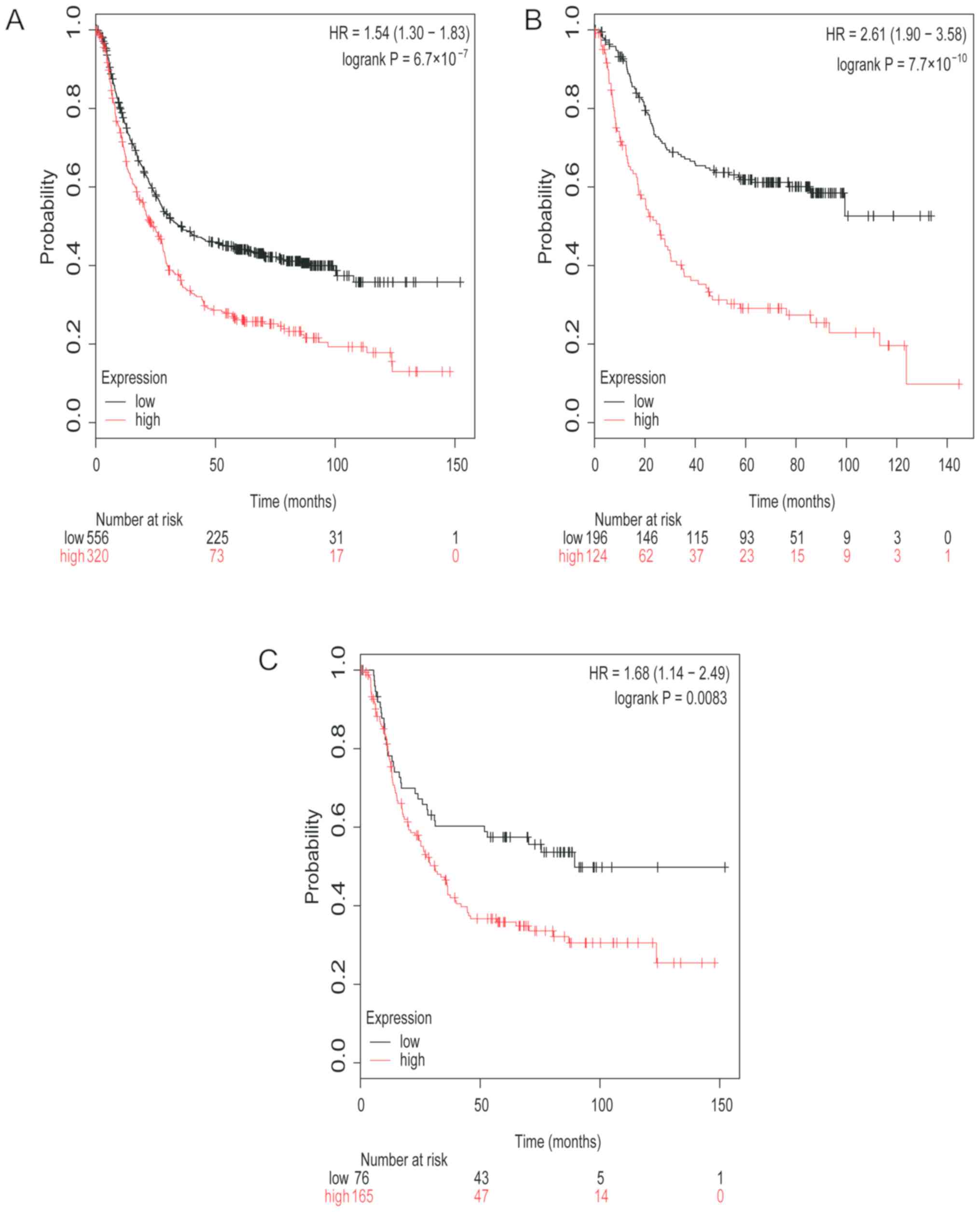
Next, the prognostic value of TGFβR1 mRNA expression
was analyzed. Low TGFβR1 mRNA expression in patients with GC was
associated with higher OS time (HR, 1.54; 95% CI, 1.30–1.83;
P<0.0001; Fig. 2A). Low TGFβR1
mRNA expression was also found to be associated with higher OS time
in patients with intestinal type GC (HR, 2.61; 95% CI, 1.90–3.58;
P<0.0001; Fig. 2B) and patients
with diffuse type GC (HR, 1.68; 95% CI, 1.14–2.49; P=0.0083;
Fig. 2C).
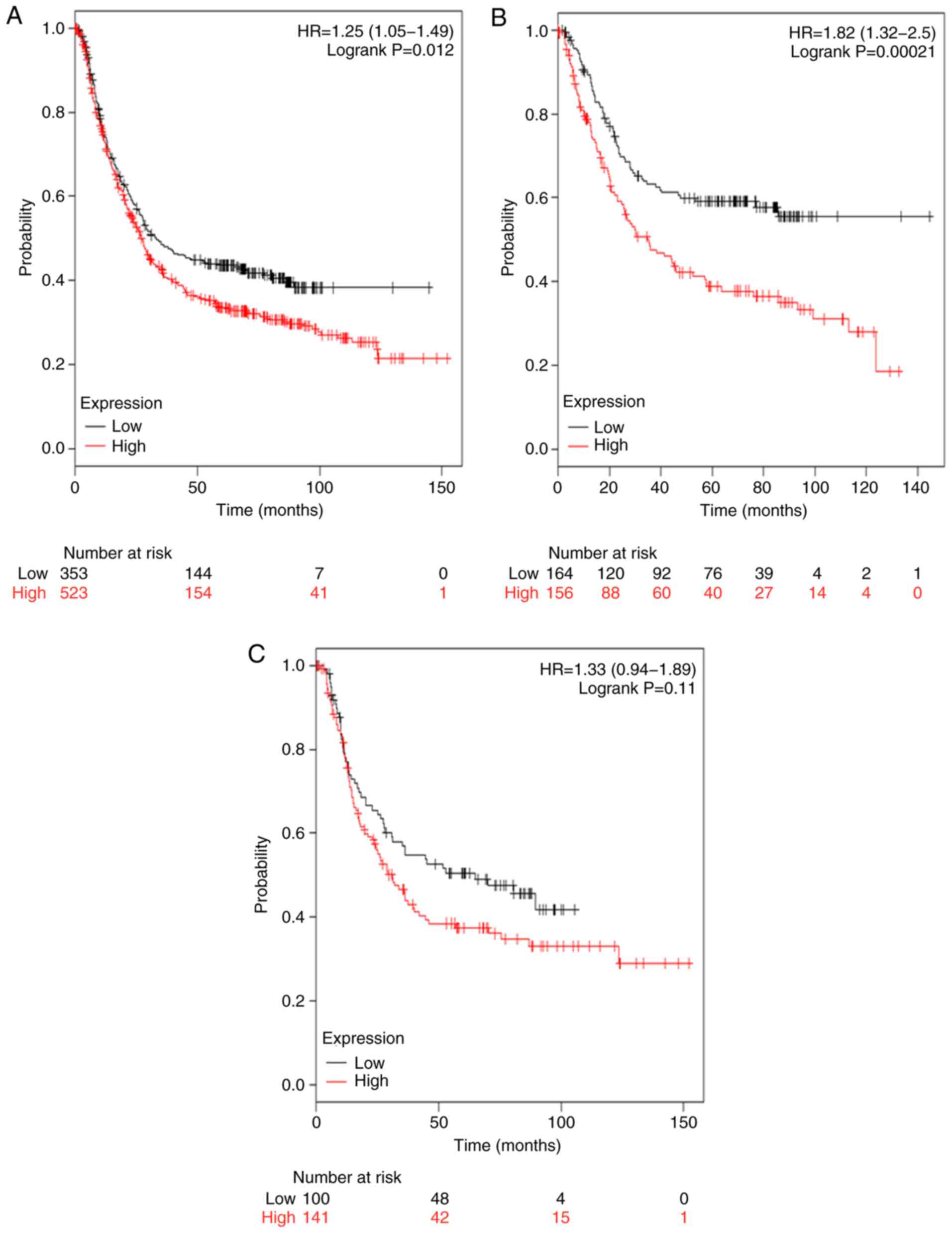
The survival curves associated with TGFβR2 mRNA
expression are represented in Fig.
3. Low expression levels of TGFβR2 mRNA were associated with an
improved prognosis in patients with GC (HR, 1.25; 95% CI,
1.05–1.49; P=0.012; Fig. 3A) and in
patients with intestinal type GC (HR=1.82; 95% CI, 1.32–2.50;
P=0.012; Fig. 3B). TGFβR2 was also
associated with a modest improvement in the prognosis of patients
with diffuse type GC; however, this increase was not statistically
significant (HR, 1.33; 95% CI, 0.94–1.89; P=0.11; Fig. 3C).
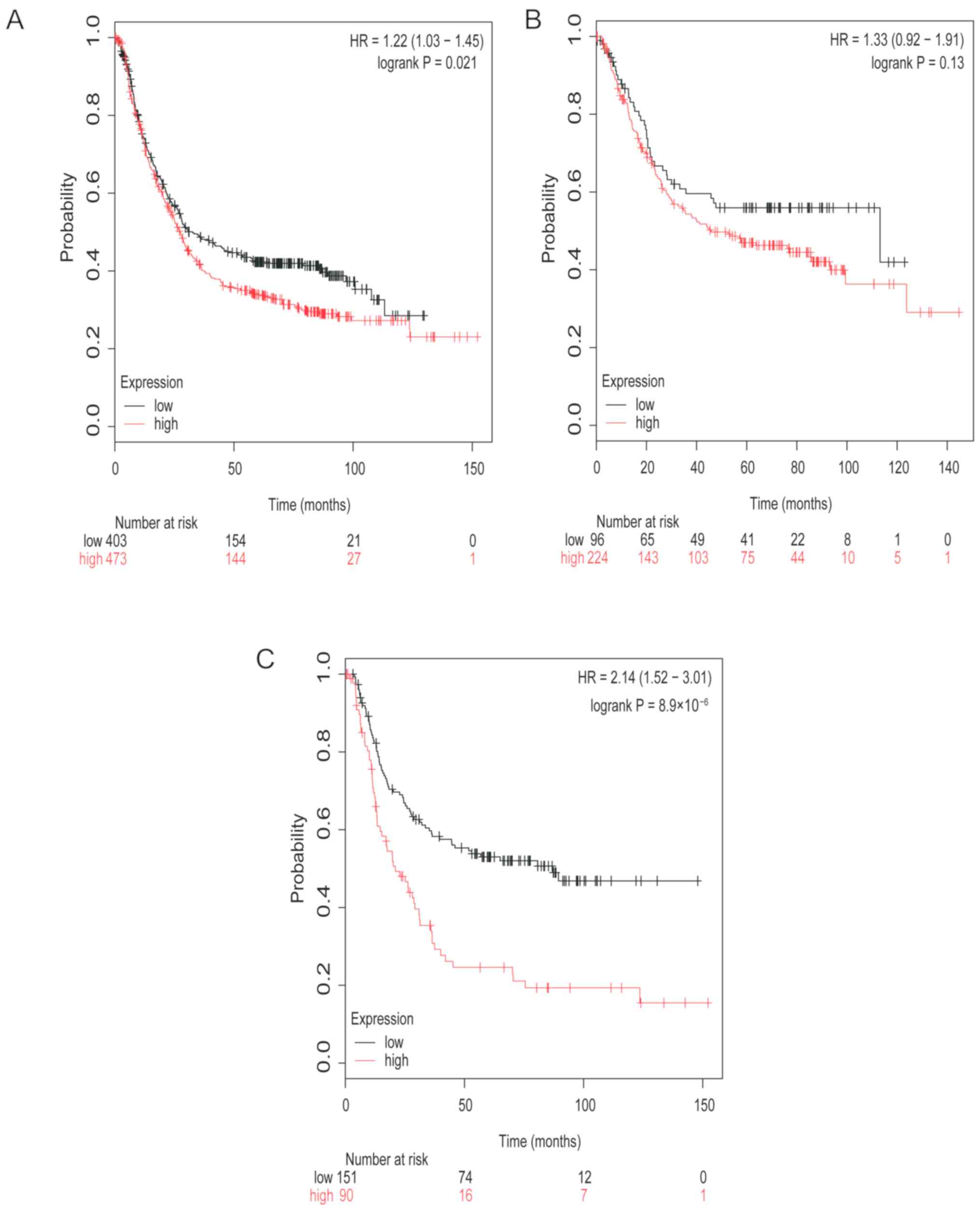
The survival curves of TGFβR3 mRNA expression for
all patient groups investigated are represented in Fig. 4. Low mRNA expression level of TGFβR3
was associated with improved prognosis in patients with GC (HR,
1.22; 95% CI, 1.03–1.45; P=0.021; Fig.
4A). Low mRNA expression of TGFβR3 was also associated with
improved prognosis in patients with diffuse type GC (HR, 2.14; 95%
CI, 1.52–3.01; P<0.0001; Fig.
4C). TGFβR3 was also associated with a modest improvement in
the prognosis of patients with intestinal type GC; however, this
increase was not statistically significant (HR, 1.33; 95% CI,
0.92–1.91; P=0.13; Fig. 4B).
According to the results of the present study, low mRNA expression
levels of TGFβ1, TGFβR1, TGFβR2 and TGFβR3 were all associated with
a higher OS time in patients with GC.
Furthermore, the association between TGFβ signaling
and prognosis in patients with GC with different
clinicopathological features, including clinical stages (Table I), HER2 status (Table II), pathological grades (Table III) and different treatment methods
(Table IV) was analyzed. As
presented in Table I, low TGFβ1 mRNA
expression was associated with an improved prognosis at clinical
stages 2 of GC (HR, 2.61; 95% CI, 1.16–5.86; P=0.016). Low mRNA
expression of TGFβR1 was associated with a better prognosis at
clinical stages 2 (HR, 3.39; 95% CI, 1.86–6.61; P<0.0001;
Table I) and 3 (HR, 1.9; 95% CI,
1.42–2.55; P<0.0001; Table I) in
patients with GC. Low mRNA expression of TGFβR2 was also associated
with a more favorable prognosis at clinical stages 1 (HR, 9.1; 95%
CI, 1.19–69.50; P=0.0099; Table I),
2 (HR, 2.32; 95% CI, 1.27–4.25; P= 0.0051; Table I) and 4 (HR, 1.76; 95% CI, 1.13–2.67;
P= 0.012; Table I). Low mRNA
expression of TGFβR3 was also found to be associated with better
prognosis in clinical stages 2 (HR, 2.79; 95% CI, 1.53–5.08;
P=0.00048; Table I), 3 (HR, 1.36;
95% CI, 1.02–1.8; P=0.035; Table I)
and 4 (HR, 2; 95% CI, 1.33-3; P=0.00063; Table I) patients with GC.
 | Table I.Association between mRNA expression
of TGFβ1 and its receptors and clinical stage in patients with
gastric cancer. |
Table I.
Association between mRNA expression
of TGFβ1 and its receptors and clinical stage in patients with
gastric cancer.
| Gene | Clinical
stages | Cases, n | HR | 95% CI | P-value |
|---|
| TGFβ1 | 1 | 69 | 3.92 | 0.89–17.33 | 0.052 |
|
| 2 | 145 | 2.61 | 1.16–5.86 | 0.016a |
|
| 3 | 319 | 0.77 | 0.58–1.03 | 0.074 |
|
| 4 | 152 | 0.84 | 0.57–1.24 | 0.380 |
| TGFβR1 | 1 | 69 | 1.95 | 0.61–6.19 | 0.250 |
|
| 2 | 145 | 3.39 | 1.86–6.61 |
<0.001c |
|
| 3 | 319 | 1.90 | 1.42–2.55 |
<0.001c |
|
| 4 | 152 | 1.39 | 0.94–2.07 | 0.100 |
| TGFβR2 | 1 | 69 | 9.10 | 1.19–69.51 | 0.010a |
|
| 2 | 145 | 2.32 | 1.27–4.25 | 0.005b |
|
| 3 | 319 | 1.28 | 0.96–1.71 | 0.090 |
|
| 4 | 152 | 1.76 | 1.13–2.67 | 0.012a |
| TGFβR3 | 1 | 69 | 1.63 | 0.60–4.41 | 0.330 |
|
| 2 | 145 | 2.79 | 1.53–5.08 |
<0.001c |
|
| 3 | 319 | 1.36 | 1.02–1.80 | 0.035a |
|
| 4 | 152 | 2.00 | 1.33–3.00 |
<0.001c |
 | Table II.Association between mRNA expression
of TGFβ1 and its receptors and HER 2 status of patients with
gastric cancer. |
Table II.
Association between mRNA expression
of TGFβ1 and its receptors and HER 2 status of patients with
gastric cancer.
| Gene | HER status | Cases, n | HR | 95% CI | P-value |
|---|
| TGFβ1 | – | 532 | 1.66 | 1.27–2.15 |
<0.001c |
|
| + | 344 | 1.26 | 0.96–1.65 | 0.090 |
| TGFβR1 | – | 532 | 1.67 | 1.33–2.09 |
<0.001c |
|
| + | 344 | 1.48 | 1.14–1.92 | 0.003b |
| TGFβR2 | – | 532 | 1.33 | 1.05–1.67 | 0.016a |
|
| + | 344 | 0.76 | 0.57–1.03 | 0.072 |
| TGFβR3 | – | 532 | 1.48 | 1.16–1.88 | 0.001 |
|
| + | 344 | 1.32 | 1.01–1.72 | 0.041a |
 | Table III.Association between mRNA expression
of TGFβ1 and its receptors and pathological grades of patients with
gastric cancer. |
Table III.
Association between mRNA expression
of TGFβ1 and its receptors and pathological grades of patients with
gastric cancer.
| Gene | Pathological
grades | Cases, n | HR | 95% CI | P-value |
|---|
| TGFβ1 | I | 166 | 0.60 | 0.37–0.99 | 0.042a |
|
| II | 67 | 0.57 | 0.30–1.09 | 0.087 |
|
| III | 32 | 0.74 | 0.30–1.78 | 0.500 |
| TGFβR1 | I | 166 | 1.52 | 0.93–2.50 | 0.094 |
|
| II | 67 | 2.62 | 1.32–5.20 | 0.004b |
|
| III | 32 | 2.23 | 0.68–7.90 | 0.170 |
| TGFβR2 | I | 166 | 0.57 | 0.37–0.89 | 0.012a |
|
| II | 67 | 2.16 | 0.90–5.19 | 0.078 |
|
| III | 32 | 0.67 | 0.27–1.63 | 0.370 |
| TGFβR3 | I | 166 | 1.20 | 0.79–1.83 | 0.390 |
|
| II | 67 | 0.29 | 0.08–1.03 | 0.043a |
|
| III | 32 | 4.48 | 1.04–19.34 | 0.028a |
 | Table IV.Association between mRNA expression
of TGFβ1 and its receptors and different treatment methods of
patients with gastric cancer. |
Table IV.
Association between mRNA expression
of TGFβ1 and its receptors and different treatment methods of
patients with gastric cancer.
| Gene | Treatment: | Cases, n | HR | 95% CI | P-value |
|---|
| TGFβ1 | Surgery only | 393 | 2.19 | 1.47–3.25 |
<0.001c |
|
| 5 FU based
adjuvant | 158 | 0.84 | 0.58–1.22 |
<0.001c |
|
| Other adjuvant | 80 | 2.72 | 1.12–6.58 | 0.021a |
| TGFβR1 | Surgery only | 393 | 1.53 | 1.14–2.04 | 0.004b |
|
| 5 FU based
adjuvant | 158 | 0.66 | 0.47–0.94 | 0.020a |
|
| Other adjuvant | 80 | 1.69 | 0.70–4.09 | 0.240 |
| TGFβR2 | Surgery only | 393 | 1.37 | 1.03–1.84 | 0.031a |
|
| 5 FU based
adjuvant | 158 | 0.42 | 0.29–0.61 |
<0.001c |
|
| Other adjuvant | 80 | 1.66 | 0.69–4.00 | 0.260 |
| TGFβR3 | Surgery only | 393 | 1.52 | 1.12–2.07 | 0.008b |
|
| 5 FU based
adjuvant | 158 | 0.60 | 0.40–0.89 | 0.010a |
|
| Other adjuvant | 80 | 3.27 | 1.26–8.52 | 0.010a |
Low mRNA expression levels of TGFβ1 (HR, 1.66; 95%
CI, 1.27–2.15; P=0.00014; Table II)
and TGFβR2 (HR, 1.33; 95% CI, 1.05–1.67; P=0.016; Table II) were associated with a better
prognosis in HER2− patients with GC. Low mRNA expression
of TGFβR1 [(HER2−: HR, 1.67; 95% CI; 1.33–2.09;
P<0.0001; Table II)
(HER2+: HR, 1.48; 95% CI, 1.14–1.92; P=0.0028; Table II)] and TGFβR3 [(HER2−:
HR, 1.48; 95% CI, 1.16–1.88; P=0.0013; Τable II) (HER2+:
HR,1.32; 95% CI, 1.01–1.72; P=0.041, Table II)] were associated with a better
prognosis in both HER2− and HER2+ patients
with GC.
Low mRNA expression of TGFβ1 (HR, 0.60; 95% CI,
0.37–0.99; P=0.042; Table III) and
TGFβR2 (HR, 0.57; 95% CI, 0.37–0.89; P=0.012; Table III) was associated with higher OS
time in grade I patients with GC. Additionally, TGFβR1 low mRNA
expression was associated with higher OS time in grade II patients
with GC (HR, 2.62; 95% CI, 1.32–5.20; P=0.0041; Table III). Low expression of TGFβR3 was
associated with higher OS time in pathological grades II (HR, 0.29;
95% CI, 0.08–1.03; P=0.043; Table
III) and III (HR, 4.48; 95% CI, 1.04–19.34; P=0.028; Table III) patients with GC.
Finally, as represented in Table IV, low mRNA expression of TGFβ1 was
associated with higher OS times in patients with GC who had been
treated with surgery alone (HR, 2.19; 95% CI, 1.47–3.25;
P<0.0001). Concurrently, low mRNA expression levels of TGFβR1
were associated with higher OS times in patients with GC with the
same method of treatment (HR, 1.53; 95% CI, 1.14–2.04; P=0.004;
Table IV) and patients with GC who
had fluorouracil (5-FU)-based adjuvant treatment (HR, 0.66; 95% CI,
0.47–0.94; P=0.02; Table IV). Low
mRNA expression of TGFβR2 was associated with higher OS times in
patients with GC who had received surgery alone (HR, 1.37; 95% CI,
1.03–1.84; P=0.031; Table IV) and
patients with GC who had received 5-FU-based adjuvant treatment
(HR, 0.42; 95% CI, 0.29–0.61; P<0.0001; Table IV). Low mRNA expression of TGFβR3
was associated with higher OS times in patients with GC who were
surgically treated (HR, 1.52; 95% CI, 1.12–2.07; P=0.0075; Table IV) and patients with GC who were
treated with a 5-FU-based adjuvant (HR, 0.60; 95% CI 0.40–0.89;
P=0.01; Table IV). The low
expression levels of TGFβ1 (HR, 2.72; 95% CI, 1.12–6.58; P=0.021)
and TGFβR3 (HR, 3.27; 95% CI, 1.26–8.52; P=0.01) (Table IV) were associated with higher OS
times in patients receiving other adjuvant treatments.
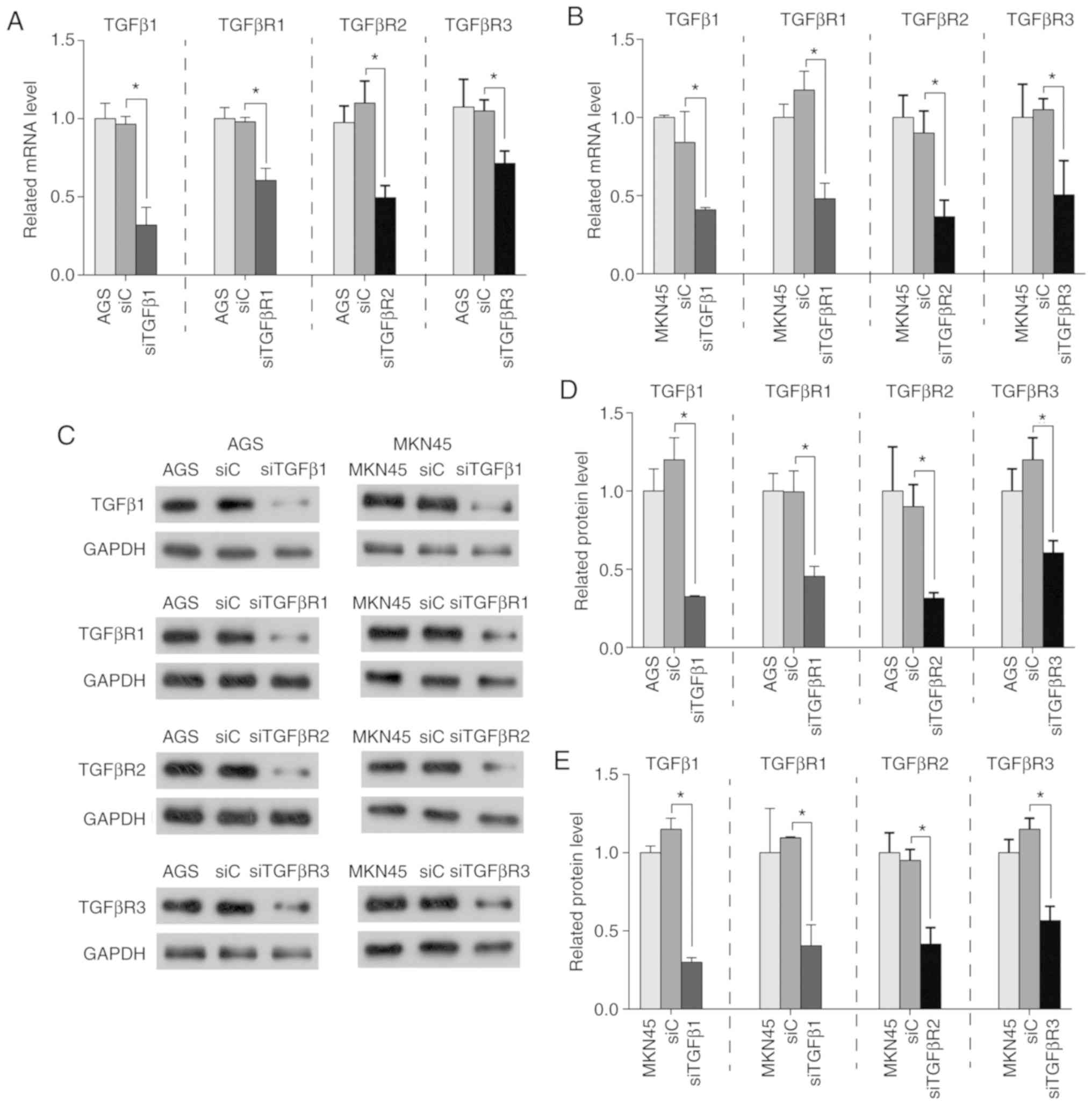
Knockdown of TGFβ1 and its receptors
inhibits the proliferation of human GC cells
Since a high mRNA expression level of TGFβ1 and its
receptors is predicative of a poor prognosis in patients with GC,
their direct effects on GC cells were subsequently investigated. In
order to evaluate the role of TGFβ1 and its receptors in AGS and
MKN45 cells, specific siRNAs were transfected into cells and
expression was quantified by RT-qPCR and western blotting. As
presented in Fig. 5, the expression
of TGFβ1 and its receptors was significantly suppressed in
transfected GC cells. The proliferation of AGS and MKN45 cells was
then determined using a CCK8 assay. Based on these results, it was
determined that the knockdown of TGFβ1 and its receptors (with the
exception of TGFβR3) inhibited the proliferation of GC cells
(Fig. 6A and B).
Knockdown of TGFβ1 and its receptors
inhibits the migration and invasion of human GC cells
Next, transwell assays were performed to explore the
effects of TGFβ1 and its receptors on the migration and invasion of
GC cells (Fig. 6C). With the
exception of TGFβR3, TGFβ1 and its receptors significantly
inhibited the migration of AGS and MKN45 cells (Fig. 6C-E). Moreover, the results of the
transwell assay for cell invasion demonstrated that except for
TGFβ1, knockdown of TGFβ1 and its receptors suppressed cell
invasiveness (Fig. 6C-E).
Cumulatively, the data confirm that knockdown of TGFβ1, TGββR1 and
TGFβR2 inhibit the progression of human GC.
Discussion
The TGFβ superfamily is a large class of cytokines
that perform various biological activities. This superfamily is
mainly comprised of TGFβ, activin and bone morphogenetic protein.
These molecules are important in the regulation of cell growth,
adhesion, migration, differentiation and apoptosis. In mammals,
three subtypes of TGFβ have been discovered: TGFβ1; TGFβ2; and
TGFβ3 (31). TGFβ1 is the most
commonly expressed form of TGFβ in human tissues, and plays an
important role in the regulation of cell growth, apoptosis,
differentiation and the maintenance of normal immune homeostasis
(32–34). TGFβ signaling is a double-edged sword
in the process of tumor formation and development (35). In tumor formation, the TGFβ signaling
pathway regulates downstream target genes, such as p21 cyclin
dependent kinase (CDKN)1A and p15CDKN2B, to arrest cells in the
G1 phase of the cell cycle, and inhibit the
proliferation of tumor cells (35).
In tumor progression, TGFβ can promote invasion and metastasis
through a variety of mechanisms, including immune suppression or
escape, angiogenesis and by increasing the interaction between
tumor cells and the extracellular matrix (35).
In previous years, numerous studies have
demonstrated that TGFβ1 is associated with tumor occurrence and
development, and is highly expressed in a variety of malignant
tumor types, including prostate, breast gastric and colorectal
cancer (36,37). Docea et al (38) noticed that the highest level of TGFβ
was exhibited in GC compared with normal tissue and the expression
of TGFβ progressively increased in the epithelium-intestinal
metaplasia-dysplasia-carcinoma sequence. In intestinal variants,
TGFβ immunoreactivity was significantly associated with the degree
of tumor differentiation and proliferative activity (38). According to another report, TGFβ1
mRNA levels were higher in tumor cells and were positively
associated with Smad2 and Smad7 mRNA levels (39). Serum TGFβ1 levels have been
demonstrated to be significantly higher in patients at both early
and advanced cancer stages, compared with controls (39). TGFβ1 is closely linked to the
initiation of the epithelial-mesenchymal transition (EMT) in the
development and progression of carcinomas (40,41). In
GC cells, TGFβ1 can induce the mRNA and protein expression of
Krüppel-like factor 8 expression (42). It can also contribute to EMT via the
downregulation of E-cadherin, and the upregulation of vimentin
expression (43–45). TGFβ1 can interact with a variety of
tumor-related genes and proteins in TGFβ-induced EMT in GC, such as
SAM-domain and SH3-domain containing 1, microRNA-21 and Grainy head
like 2 (43–45). In the present study, low mRNA
expression of TGFβ1 was associated with an improved prognosis in
patients with GC, including the intestinal and diffuse subtypes of
GC. In addition, TGFβ1 can be associated with patient prognosis in
GC, based on certain clinical features, including HER2 status,
pathological grade I and different treatment methods. These results
suggest that TGFβ1 has potential as a new prognostic indicator of
GC, including the intestinal and diffuse types.
The TGFβR includes three subtypes: TGFβR1, TGFβR2
and TGFβR3. TGFβR1 and 2 are categorized as type I transmembrane
glycoproteins with serine/threonine kinase activity and
collectively participate in the TGFβ/Smad signaling pathway.
Initially, TGFβ binds to TGFβR2, and then activates TGFβR3 through
phosphorylation. Together they form the TGFβR1-TGFβ1-TGFβR2
heterotetramer for transduction of cell signaling. TGFβR3 can
enhance the binding of ligand to TGFβR1 and 2, functioning as an
accessory receptor (14). Wild-type
TGFβR2 expression in GC cell lines can result in reduced
proliferation compared with control cells (46). A case-control study was performed to
evaluate the possible association of polymorphisms in TGF-β
receptors with susceptibility to developing GC (47).
Polymorphisms of TGFβR1 and 2 may be associated with
the risk of GC in the population of North China (48,49).
However, TGFβR3 has not yet been studied in the context of GC. In
the present study, it was revealed that low mRNA expression of
TGFβR1, TGFβR2 and TGFβR3 was associated with a more favorable
prognosis in patients with GC. While TGFβR2 was associated with OS
time in patients with intestinal type GC, this was not observed for
patients with diffuse type GC. Pak et al (50) determined that the expression of TβR2
was higher in patients with intestinal type GC compared with those
with diffuse type GC. In addition, TGFβR was also associated with
prognosis based on different clinical features in the
aforementioned study (50). While
TGFβ1 blockade has been proposed as an anti-cancer therapy, it is
imperative to understand the best method of administration, and
which specific pathological features will be most improved by this
therapy (51).
Finally, the present study investigated the specific
roles of TGFβ1 and its receptors on GC cells. The results of the
present study demonstrated that knockdown of TGFβ1, TGFβR1 and
TGFβR2 could significantly suppress the proliferation, migration
and invasion of human GC cells. These results are consistent with
previous studies (48,52–54).
While TGFβR2 has been widely studied in different types of cancer
(54,55), studies investigating TGFβR1 and
TGFβR3 are limited. The current study confirms the inhibitory
effect of TGFβR1 on the proliferation of GC cells, suggesting the
involvement of TGFβ1 and TGFβR2, but also TGFβR1 in the progression
of GC. However, the downstream targets and regulatory mechanism of
TGFβR1 remain unclear, and cells cultured in vitro cannot
precisely simulate the tumor microenvironment. The results of the
present study need to be verified by further in vivo
experiments.
In conclusion, the present study showed that TGFβ1
and its receptors were all associated with the prognosis of
patients with GC. Consistently, low mRNA expression levels of TGFβ1
and TGFβR indicated a better OS time. Furthermore, knockdown of
TGFβ1, TGFβR1 and TGFβR2 inhibited cell proliferation in GC. This
suggests that TGFβ1 and TGFβR play important roles in the
development of GC and may be investigated as therapeutic targets.
These findings provide novel insights and approaches for the
treatment of GC.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
FL and HW mainly performed the experiments and
analyzed the data. FL performed the online analysis and wrote the
paper. MZ carried out the experiment design and manuscript
drafting. All authors have read and approved the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Kamangar F, Dores GM and Anderson WF:
Patterns of cancer incidence, mortality, and prevalence across five
continents: Defining priorities to reduce cancer disparities in
different geographic regions of the world. J Clin Oncol.
24:2137–2150. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Alberts SR, Cervantes A and van de Velde
CJ: Gastric cancer: Epidemiology, pathology and treatment. Ann
Oncol. 14 (Suppl 2):ii31–ii36. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Choi YY, Noh SH and Cheong JH: Evolution
of gastric cancer treatment: From the golden age of surgery to an
era of precision medicine. Yonsei Med J. 56:1177–1185. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chang HM, Lee SW, Nomura E and Tanigawa N:
Laparoscopic versus open gastrectomy for gastric cancer patients
with COPD. J Surg Oncol. 100:456–458. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Foidart JM and Muschel RJ: Proteases and
Their Inhibitors in Cancer Metastasis. Kluwer Academic Publicers;
The Netherlands: pp. 225–252. 2002
|
|
7
|
Mesri M, Wall NR, Li J, Kim RW and Altieri
DC: Cancer gene therapy using survivin mutant adenovirus. J Clin
Invest. 108:981–990. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liang L, Fang JY and Xu J: Gastric cancer
and gene copy number variation: Emerging cancer drivers for
targeted therapy. Oncogene. 35:1475–1482. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lin Y, Wu Z, Guo W and Li J: Gene
mutations in gastric cancer: A review of recent next-generation
sequencing studies. Tumour Biol. 36:7385–7394. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kajdaniuk D, Marek B, Borgielmarek H and
Koskudła B: Transforming growth factor β1 (TGFβ1) in physiology and
pathology. Endokrynol Pol. 64:384–396. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Massagué J: TGF-beta signal transduction.
Annu Rev Biochem. 67:753–791. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Assoian RK, Komoriya A, Meyers CA, Miller
DM and Sporn MB: Transforming growth factor-beta in human
platelets. Identification of a major storage site, purification,
and characterization. J Biol Chem. 258:7155–7160. 1983.PubMed/NCBI
|
|
13
|
Sam R, Wanna L, Gudehithlu KP, Garber SL,
Dunea G, Arruda JA and Singh AK: Glomerular epithelial cells
transform to myofibroblasts: Early but not late removal of
TGF-beta1 reverses transformation. Transl Res. 148:142–148. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ikushima H and Miyazono K: TGFbeta
signalling: A complex web in cancer progression. Nat Rev Cancer.
10:415–424. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
15
|
Massagué J: TGFbeta in cancer. Cell.
134:215–230. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhuang J, Lu Q, Shen B, Huang X, Shen L,
Zheng X, Huang R, Yan J and Guo H: TGFβ1 secreted by
cancer-associated fibroblasts induces epithelial-mesenchymal
transition of bladder cancer cells through lncRNA-ZEB2NAT. Sci Rep.
5:119242015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Neuzillet C, Tijeras-Raballand A, Cohen R,
Cros J, Faivre S, Raymond E and de Gramont A: Targeting the TGFβ
pathway for cancer therapy. Pharmacol Ther. 147:22–31. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Busch S, Acar A, Magnusson Y, Gregersson
P, Rydén L and Landberg G: TGF-beta receptor type-2 expression in
cancer-associated fibroblasts regulates breast cancer cell growth
and survival and is a prognostic marker in pre-menopausal breast
cancer. Oncogene. 34:27–38. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ji Q, Liu X, Han Z, Zhou L, Sui H, Yan L,
Jiang H, Ren J, Cai J and Li Q: Resveratrol suppresses
epithelial-to-mesenchymal transition in colorectal cancer through
TGF-β1/Smads signaling pathway mediated Snail/E-cadherin
expression. BMC Cancer. 15:972015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ma ZL, Hou PP, Li YL, Wang DT, Yuan TW,
Wei JL, Zhao BT, Lou JT, Zhao XT, Jin Y and Jin YX: MicroRNA-34a
inhibits the proliferation and promotes the apoptosis of non-small
cell lung cancer H1299 cell line by targeting TGFβR2. Tumour Biol.
36:2481–2490. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Szász AM, Lánczky A, Nagy Á, Förster S,
Hark K, Green JE, Boussioutas A, Busuttil R, Szabó A and Győrffy B:
Cross-validation of survival associated biomarkers in gastric
cancer using transcriptomic data of 1,065 patients. Oncotarget.
7:49322–49333. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wu X, Liu W, Tang D, Xiao H, Wu Z, Chen C,
Yao X, Liu F and Li G: Prognostic values of four Notch receptor
mRNA expression in gastric cancer. Sci Rep. 6:280442016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhang S, Zhen W, Liu W, Lei R, Shan J, Li
L and Wang X: Distinct prognostic values of S100 mRNA expression in
breast cancer. Sci Rep. 7:397862017. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wu S, Xue W, Huang X, Yu X, Luo M, Huang
Y, Liu Y, Bi Z, Qiu X and Bai S: Distinct prognostic values of
ALDH1 isoenzymes in breast cancer. Tumor Biol. 36:2421–2426. 2015.
View Article : Google Scholar
|
|
25
|
Zhou X, Teng L and Wang M: Distinct
prognostic values of four-Notch-receptor mRNA expression in ovarian
cancer. Tumor Biol. 37:6979–6985. 2016. View Article : Google Scholar
|
|
26
|
Ivanova L, Zandberga E, Siliņa K, Kalniņa
Z, Ābols A, Endzeliņš E, Vendina I, Romanchikova N, Hegmane A,
Trapencieris P, et al: Prognostic relevance of carbonic anhydrase
IX expression is distinct in various subtypes of breast cancer and
its silencing suppresses self-renewal capacity of breast cancer
cells. Cancer Chemother Pharmacol. 75:235–246. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Győrffy B, Surowiak P, Budczies J and
Lánczky A: Online survival analysis software to assess the
prognostic value of biomarkers using transcriptomic data in
non-small-cell lung cancer. PLoS One. 8:e822412013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sun Z, Wang ZN, Zhu Z, Xu YY, Xu Y, Huang
BJ, Zhu GL and Xu HM: Evaluation of the seventh edition of American
Joint Committee on Cancer TNM staging system for gastric cancer:
Results from a Chinese monoinstitutional study. Ann Surg Oncol.
19:1918–1927. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ma J, Shen H, Kapesa L and Zeng S: Lauren
classification and individualized chemotherapy in gastric cancer.
Oncol Lett. 11:2959–2964. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Memon MA, Anway MD, Covert TR, Uzumcu M
and Skinner MK: Transforming growth factor beta (TGFbeta1, TGFbeta2
and TGFbeta3) null-mutant phenotypes in embryonic gonadal
development. Mol Cell Endocrinol. 294:70–80. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Meng XM, Tang PM, Li J and Lan HY:
TGF-β/Smad signaling in renal fibrosis. Front Physiol. 6:822015.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Okamura T, Morita K, Iwasaki Y, Inoue M,
Komai T, Fujio K and Yamamoto K: Role of TGF-β3 in the regulation
of immune responses. Clin Exp Rheumatol. 33 (Suppl 92):S63–S69.
2015.PubMed/NCBI
|
|
34
|
Potter RM, Huynh RT, Volper BD, Arthur KA,
D'Lugos AC, Sørensen MA, Magnusson SP, Dickinson JM, Hale TM and
Carroll CC: Impact of TGF-β inhibition during acute exercise on
Achilles tendon extracellular matrix. Am J Physiol Regul Integr
Comp Physiol. 312:R157–R164. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Akhurst RJ and Derynck R: TGF-beta
signaling in cancer-a double-edged sword. Trends in Cell Biol.
11:S44–S51. 2001. View Article : Google Scholar
|
|
36
|
Costanza B, Umelo IA, Bellier J,
Castronovo V and Turtoi A: Stromal modulators of TGF-β in cancer. J
Clin Med. 6(pii): E72017. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Pang MF, Georgoudaki AM, Lambut L,
Johansson J, Tabor V, Hagikura K, Jin Y, Jansson M, Alexander JS,
Nelson CM, et al: TGF-β1-induced EMT promotes targeted migration of
breast cancer cells through the lymphatic system by the activation
of CCR7/CCL21-mediated chemotaxis. Oncogene. 35:748–760. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Docea AO, Mitruţ P, Grigore D, Pirici D,
Călina DC and Gofiţă E: Immunohistochemical expression of TGF beta
(TGF-β), TGF beta receptor 1 (TGFBR1), and Ki67 in intestinal
variant of gastric adenocarcinomas. Rom J Morphol Embryol. 53
(Suppl 3):683–692. 2012.PubMed/NCBI
|
|
39
|
Ma GF, Miao Q, Zeng XQ, Luo TC, Ma LL, Liu
YM, Lian JJ, Gao H and Chen SY: Transforming growth factor-β1 and
-β2 in gastric precancer and cancer and roles in tumor-cell
interactions with peripheral blood mononuclear cells in vitro. PLoS
One. 8:e542492013. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
David CJ, Huang YH, Chen M, Su J, Zou Y,
Bardeesy N, Iacobuzio-Donahue CA and Massagué J: TGF-β tumor
suppression through a lethal EMT. Cell. 164:1015–1030. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Lamouille S, Xu J and Derynck R: Molecular
mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell
Biol. 15:178–196. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zhang H, Liu L, Wang Y, Zhao G, Xie R, Liu
C, Xiao X, Wu K, Nie Y, Zhang H and Fan D: KLF8 involves in
TGF-beta-induced EMT and promotes invasion and migration in gastric
cancer cells. J Cancer Res Clin Oncol. 139:1033–1042. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Xiang J, Fu X, Ran W and Wang Z: Grhl2
reduces invasion and migration through inhibition of TGFβ-induced
EMT in gastric cancer. Oncogenesis. 6:e2842017. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Li C, Song L, Zhang Z, Bai XX, Cui MF and
Ma LJ: MicroRNA-21 promotes TGF-β1-induced epithelial-mesenchymal
transition in gastric cancer through up-regulating PTEN expression.
Oncotarget. 7:66989–67003. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zong W, Yu C, Wang P and Dong L:
Overexpression of SASH1 inhibits TGF-β1-induced EMT in gastric
cancer cells. Oncol Res. 24:17–23. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Chang J, Park K, Bang YJ, Kim WS, Kim D
and Kim SJ: Expression of transforming growth factor beta type II
receptor reduces tumorigenicity in human gastric cancer cells.
Cancer Res. 57:2856–2859. 1997.PubMed/NCBI
|
|
47
|
Guo W, Dong Z, Guo Y, Chen Z, Yang Z and
Kuang G: Association of polymorphisms in transforming growth
factor-β receptors with susceptibility to gastric cardia
adenocarcinoma. Mol Biol Rep. 39:4301–4309. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Chen J, Miao L, Jin G, Ren C, Ke Q, Qian
Y, Dong M, Li H, Zhang Q, Ding Y, et al: TGFBR1 tagging SNPs and
gastric cancer susceptibility: A two-stage case-control study in
Chinese population. Mol Carcinog. 53:109–116. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Xu L, Zeng Z, Chen B, Wu X, Yu J, Xue L,
Tian L, Wang Y, Chen M, Sung JJ and Hu P: Association between the
TGFB1 −509C/T and TGFBR2 −875A/G polymorphisms and gastric cancer:
A case-control study. Oncol Lett. 2:371–377. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Pak KH, Dong HK, Kim H, Lee DH and Cheong
JH: Differences in TGF-β1 signaling and clinicopathologic
characteristics of histologic subtypes of gastric cancer. BMC
Cancer. 16:602016. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Suzuki E, Kapoor V, Kaiser LR and Albelda
SM: Soluble type II TGF-β receptor augments or inhibits murine
malignant mesothelioma tumor growth depending on when it is
administered. Cancer Res. 64:53362004.
|
|
52
|
Chen ZL, Qin L, Peng XB, Hu Y and Liu B:
INHBA gene silencing inhibits gastric cancer cell migration and
invasion by impeding activation of the TGF-β signaling pathway. J
Cell Physiol. 234:18065–18074. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Zhou H, Wang K, Hu Z and Wen J: TGF-β1
alters microRNA profile in human gastric cancer cells. Chin J
Cancer Res. 25:102–111. 2013.PubMed/NCBI
|
|
54
|
Duan J, Zhang H, Qu Y, Deng T, Huang D,
Liu R, Zhang L, Bai M, Zhou L, Ying G and Ba Y: Onco-miR-130
promotes cell proliferation and migration by targeting TGFβR2 in
gastric cancer. Oncotarget. 7:44522–44533. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Jin G, Deng Y, Miao R, Hu Z, Zhou Y, Tan
Y, Wang J, Hua Z, Ding W, Wang L, et al: TGFB1 and TGFBR2
functional polymorphisms and risk of esophageal squamous cell
carcinoma: A case-control analysis in a Chinese population. J
Cancer Res Clin Oncol. 134:345–351. 2008. View Article : Google Scholar : PubMed/NCBI
|