Tumor initiation and progression were traditionally
described as a consequence of genetic variations; breakthroughs in
epigenetic mechanisms provided more evidence to show that
epigenetic changes have critical roles in tumor development.
Targeting epigenetics appears to be a promising therapy for cancer
treatment. Several targeting epigenetic drugs for cancer patients
have already obtained approval by the Food and Drug Administration
(1).
As epigenetic markers, post-translational
modification of histone, involving methylation, acetylation,
phosphorylation as well as ubiquitination, SUMOylation, adenosine
diphosphate-ribosylation, deimination and proline isomerization,
are critical determinants for tumor initiation and progression,
which could be passed to daughter cells. A number of previous
studies have demonstrated that histone modifications are important
for the recruitment or activity of downstream effectors. As
aberrant histone modifications are associated with divergent
reactions, the present review focuses on histone methylation.
Histone methylation was first described in 1964 by Murray (2), and frequently occurs at lysine and
arginine residues at the N-terminals of H3 and H4. Histone
methylation can engage in either gene activation or silencing
depending on specific sites, which may result in the promotion of
tumor development (3–5). The lysine residues can be either mono-,
di- or tri-methylated, and only the arginine residues can be mono-
or di-methylated, and this can have divergent effects on gene
transcription (6–9).
As essential tools to ensure accomplished
methylation, histone methyltransferases (HMTs) transfer methyl
groups from S-adenosyl methionine to the lysine and arginine
residues, which further affect gene transcription, chromatin
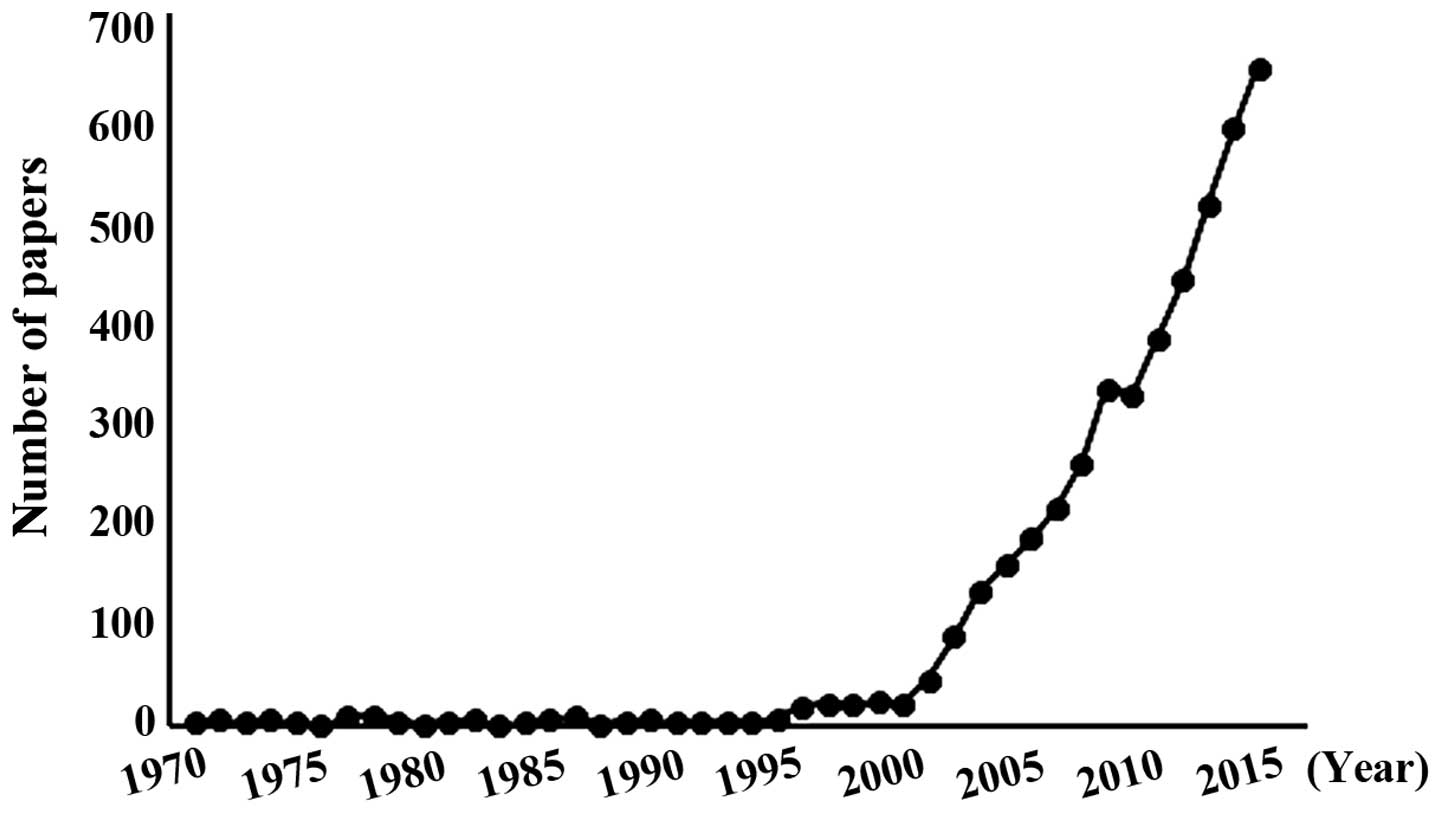
compaction and effector proteins binding (8,10,11). To date, histone methyltransferases has
received much attention (Fig. 1), and
~47% of previous studies regarding HMTs were associated with tumor
development. Currently, 51 SET domain lysine HMTs, 1 non-SET domain
lysine HMT (DOT1L) and 9 arginine HMTs have been identified, and
the majority of these are associated with cancer development
(11). Misregulation of HMTs shifts
the balance of transcription and leads to changes in cell fate,
resulting in tumor formation. The following are summaries of the
current knowledge of certain histone lysine methyltransferases
(HKMTs) and their key sites, whilst exploring the relevance of HMTs
and cancer development.
The HKMT family contains an evolutionarily conserved
SET domain, which is defined as a 130-amino acid sequence carrying
the two most-conserved sequence motifs ELXF/YDY and NHS/CXXPN. It
catalyzes the site- and state-specific methylation of different
lysine residues with relatively high substrate specificities. There
have been 52 family members identified, including 51 SET domain
lysine HMTs and 1 non-SET domain lysine HMT, known as DOT1L
(12).
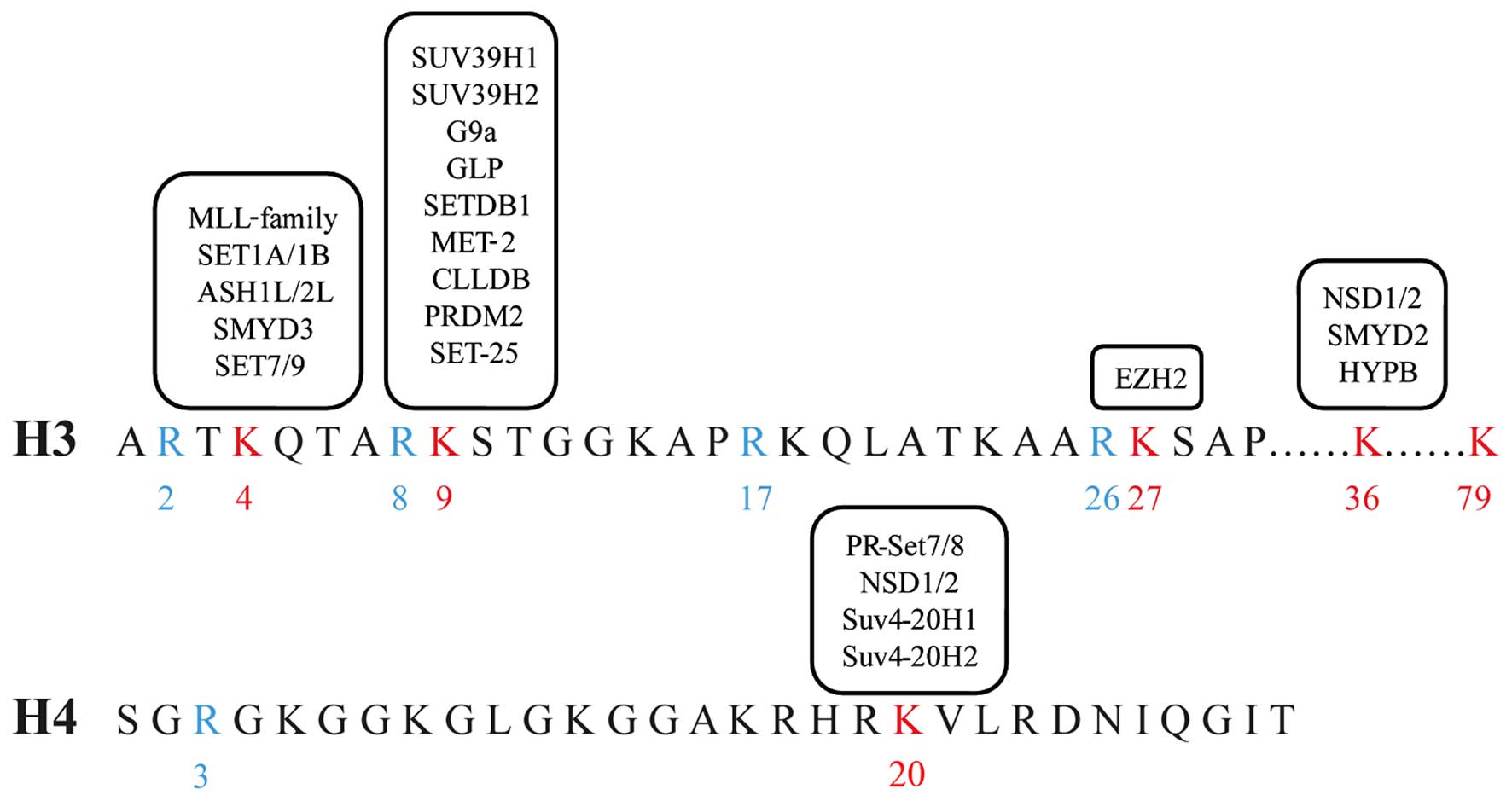
Studies have verified that histone lysines 4, 9, 27,
36 and 79 of histone H3 (H3K4, H3K9, H3K27, H3K36 and H3K79), and
20 of histone H4 (H4K20) may be methylated (Fig. 2). Methylation can take three forms:
Mono-(me1), di-(me2) and tri-(me3) methylation. Each histone
methylation has a function in regulating transcription and
chromatin conformation. H3K4me2/3, H3K9me1, H3K27me1, H3K36me3,
H3K79me3 and H4K20me1 are generally enriched in active transcribed
regions, while H3K9me2/3, H3K27me2/3 and H4K20me3 are associated
with gene repression (13).
Methylation of histone H3K9 is a well-conserved
epigenetic marker for transcriptional silencing (14). Histone H3K9 methylation and DNA
methylation can work together on the establishment and maintenance
of heterochromatin (15). The
methyltransferases responsible for histone H3K9 methylation are
able to catalyze different substrates and lead to various results
(16). These include Clr4/SUV39H1
(17), SUV39H2 (18), G9a (19),
GLP/Eu-HMTase1 (20), ESET/SETDB1
(21), Riz1/PRDM2 (22) and CLLD8/KMT1F (23).
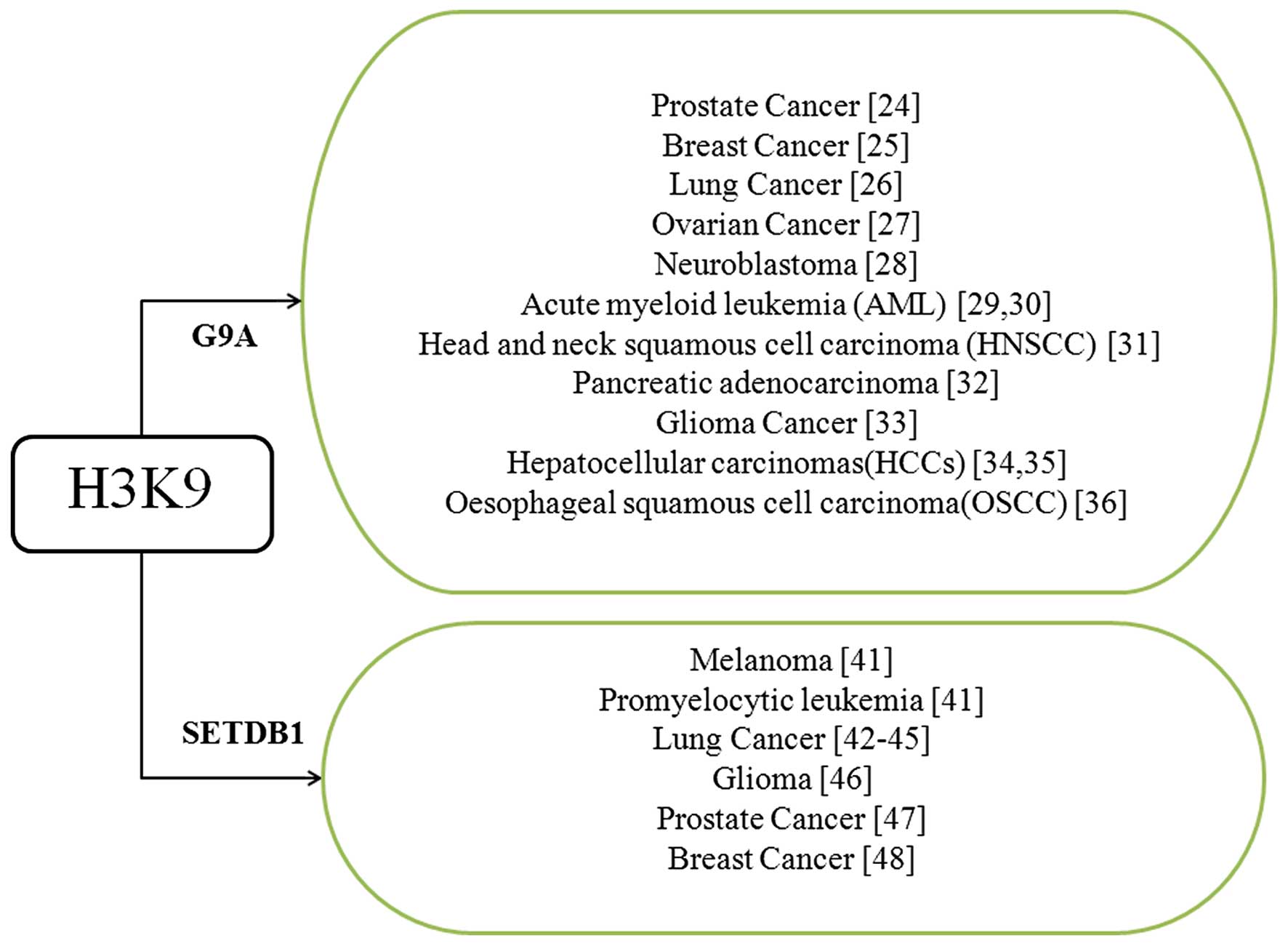
Among these HKMTs, G9a is critical for gene
silencing and embryo development. Currently, aberrant regulation of
G9a has been identified in a number of cancers (Fig. 3), and is involved in the control of
cancer metabolism by maintaining the serine-glycine biosynthetic
pathway (24). In PC3 prostate cancer
cells, knockdown of G9a significantly inhibits cell growth and
induces cellular senescence, and higher G9a expression is
associated with poorer prognosis in cancer patients (25). In addition, knockdown of G9a promotes
E-cadherin expression in claudin-low breast cancer (CLBC), and
inhibited cell migration and invasion in CLBC and lung cancer
(26,27). In ovarian cancer, higher G9a expression
predicts a greater mortality of patients (28). In neuroblastoma, our previous study
reported the importance of G9a in regulating the autophagy
signaling pathway, knockdown of G9a inhibited cell growth and
proliferation, and the activation of autophagy occurred (29). In acute myeloid squamous cell carcinoma
and pancreatic adenocarcinoma, inhibition of G9a induced
autophagy-related cell death (30,31). In
glioma cancer, previous studies have identified that G9a-dependent
H3K9me2 repressed cluster of differentiation 133 and Sox2
expression and in leukemia, and loss of G9a markedly delayed tumor
progression and repressed ATRA-mediated leukemia cell
differentiation (32,33). Additionally, in head and neck cancer,
G9a inhibited stem cell self-renewal (34). G9a was also upregulated in
hepatocellular carcinoma (HCC) tissues, and cooperated with the
H3K9 methylation effector protein CDYLb, which is involved in HCC
development (35,36). In oesophageal squamous cell carcinoma,
G9a may serve as an effective prognostic factor and be used as a
biomarker (37).
Methylation of H3K4, as an epigenetic phenomenon
conserved from yeast to human, is extremely important for
transcriptional initiation. It recruits proteins for
transactivation, and has reverse functionally to H3K9 methylation.
As a common marker of activated genes, H3K4 tri-methylation
provides an epigenetic signature of active enhancers (67). H3K4 methyltransferases include the
mixed lineage leukemia (MLL) family (68), SET1A/B (69), absent, small or homeotic disks 1-like
(ASH1L) (70), ASH2L (71), SET and MYND domain-containing protein 3
(SMYD3) (72), SET7/9 (73), and SMYD1 (74). However, in mammals, ≥6 H3K4
methyltransferases, including Set1A and Set1B and MLLs 1–4, exhibit
histone methyltransferase activity (68).
As cardiac- and muscle-specific histone
methyltransferases, except for the regulation of early heart
development, the oncogenic role of SMYD3 has been identified in
different types of cancer. In colorectal cancer, HCC and esophageal
squamous cell carcinoma, knockdown of SMYD3 impairs cell
proliferation (75–77). Similarly to CRC and HCC, in breast
cancer, silencing of SMYD3 also inhibits cell growth, and SMYD3
promotes breast carcinogenesis by directly regulating the
protooncogene WNT10B (78). In
cervical carcinoma cells and prostate cancer, reduction of SMYD3
expression by doxycycline or small hairpin RNA is able to
significantly inhibit cell proliferation, colony formation and
migration/invasion activity (79,80).
H3K36 methylation is an indicator of transcriptional
elongation, and H3K36 methyltransferases contain nuclear receptor
binding SET domain-containing protein 1 (NSD1), NSD2, NSD3
(81), SMYD2 and SETD2 (82). These are involved in diverse biological
processes, including alternative splicing and transcriptional
repression, as well as DNA repair and recombination (83).
The NSD family is known to be involved in multiple
types of cancer, and knockdown of NSD members would suppress cell
proliferation and tumor growth. NSD1 specifically mediates methyl
transfer onto H3K36 and H4K20. In prostate cancer, NSD1 can enhance
androgen receptor transactivation and is associated with prostate
tumorigenesis (84,85). In neuroblastoma, overexpression of NSD1
induces tumor suppressor-like features, such as reduced colony
formation density and inhibited cell growth (86). NSD1 has been reported in numerous types
of cancer, including multiple myeloma (87), acute myeloid leukemia (88,89), lung
cancer (90,91) and ganglioglioma (92). SETD2 is a novel tumor suppressor, which
is responsible for H3K36me3 reduction, further resulting in tumor
growth inhibition. Mutated SETD2 has been frequently identified in
human leukemia, thymic carcinoma (93), renal cell carcinoma (82,94),
non-small cell lung cancer (95) and
pediatric glioma (96).
Methylation of H4K20 has been implicated in multiple
biological processes, such as gene transcriptional regulation, cell
cycle control, development and genomic integrity maintenance
(102,103). Mono- and di-methylated H4K20 have
been attributed to DNA replication, DNA damage repair and chromatin
compaction. Lack of H4K20me1 results in chromosome condensation in
the interphase nucleus. Tri-methylation of H4K20 is required for
silencing heterochromatic regions (104).
HMTs have become more important in epigenetics and
cancer in recent years. There appear to be numerous connections
between SET-domain proteins and cancer. HKMTs are important in
regulating gene transcription, which may lead to various human
malignancies. Among these key sites, H3K9 and H3K27 occupy the
majority of the active proteins. Although the functions of HMTs
have been explored extensively, the downstream and pathological
mechanisms remain to be elucidated.
The present review summarizes the current
understanding of HMTs, provides a platform for exploring potential
therapy targets through histone modifications, and provides
insights into a potential role of aberrant histone modifications in
various human malignancies. The aforementioned methylation
accumulated the complexity of histone modifications, which provides
new insights of these functions in different patterns and their
involvement in additional diseases.
Previous studies have provided knowledge about
epigenetic heredity, a process whereby genetic information can be
preserved through modifications to chromatin without altering DNA
nucleotide sequences. Future studies may be able to uncover the
molecular mechanisms of histone modifiers, and further studies
could perform screening of downstream genes by chromatin
immunoprecipitation assays and microarrays to identify the specific
target genes and their roles in cancer therapy. This may improve
clinical outcomes or predict treatment outcomes for cancer
patients.
The present study was supported by the National
Basic Research Program of China (grant nos. 2012cb114603 and
2362015xk09), the National Natural Science Foundation of China
(grant nos. 31501100 and 81201551), the Research Fund for the
Doctoral Program of Higher Education of China (grant no.
20130182110003), the Fundamental Research Funds for the Central
Universities (grant no. XDJK2013B020) and the Fundamental Research
Funds for the Central University (grant no. XDJK2015C129).
|
1
|
Klener P: Epigenetic cancer drugs and
their role in anticancer therapy. Vnitr Lek. 59:463–465. 2013.(In
Czech).PubMed/NCBI
|
|
2
|
Murray K: The occurrence of E-N-methyl
lysine in histones. Biochemistry. 3:10–15. 1964. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Yun M, Wu J, Workman JL and Li B: Readers
of histone modifications. Cell Res. 21:564–578. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Biancotto C, Frigè G and Minucci S:
Histone modification therapy of cancer. Adv Genet. 70:341–386.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kouzarides T: Chromatin modifications and
their function. Cell. 128:693–705. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lee JC, Kang SU, Jeon Y, Park JW, You JS,
Ha SW, Bae N, Lubec G, Kwon SH, Lee JS, et al: Protein
L-isoaspartyl methyltransferase regulates p53 activity. Nat Commun.
3:9272012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Verma M and Srivastava S: Epigenetics in
cancer: Implications for early detection and prevention. Lancet
Oncol. 3:755–763. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Chaib H, Prébet T, Vey N and Collette Y:
Histone methyltransferases: A new class of therapeutic targets in
cancer treatment? Med Sci (Paris). 27:725–732. 2011.In French.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wang X and Zhu WG: Advances in histone
methyltransferases and histone demethylases. Ai Zheng.
27:1018–1025. 2008.(In Chinese). PubMed/NCBI
|
|
10
|
Collazo E, Couture JF, Bulfer S and
Trievel RC: A coupled fluorescent assay for histone
methyltransferases. Anal Biochem. 342:86–92. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Campagna-Slater V, Mok MW, Nguyen KT,
Feher M, Najmanovich R and Schapira M: Structural chemistry of the
histone methyltransferases cofactor binding site. J Chem Inf Model.
51:612–623. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Qian C and Zhou MM: SET domain protein
lysine methyltransferases: Structure, specificity and catalysis.
Cell Mol Life Sci. 63:2755–2763. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Vermeulen M, Eberl HC, Matarese F, Marks
H, Denissov S, Butter F, Lee KK, Olsen JV, Hyman AA, Stunnenberg
HG, et al: Quantitative interaction proteomics and genome-wide
profiling of epigenetic histone marks and their readers. Cell.
142:967–980. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Peterson CL and Laniel MA: Histones and
histone modifications. Curr Biol. 14:R546–R551. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Stancheva I: Caught in conspiracy:
Cooperation between DNA methylation and histone H3K9 methylation in
the establishment and maintenance of heterochromatin. Biochem Cell
Biol. 83:385–395. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Towbin BD, González-Aguilera C, Sack R,
Gaidatzis D, Kalck V, Meister P, Askjaer P and Gasser SM: Step-wise
methylation of histone H3K9 positions heterochromatin at the
nuclear periphery. Cell. 150:934–947. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Krouwels IM, Wiesmeijer K, Abraham TE,
Molenaar C, Verwoerd NP, Tanke HJ and Dirks RW: A glue for
heterochromatin maintenance: Stable SUV39H1 binding to
heterochromatin is reinforced by the SET domain. J Cell Biol.
170:537–549. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
O'Carroll D, Scherthan H, Peters AH,
Opravil S, Haynes AR, Laible G, Rea S, Schmid M, Lebersorger A,
Jerratsch M, et al: Isolation and characterization of Suv39h2, a
second histone H3 methyltransferase gene that displays
testis-specific expression. Mol Cell Biol. 20:9423–9433. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Tachibana M, Sugimoto K, Fukushima T and
Shinkai Y: Set domain-containing protein, G9a, is a novel
lysine-preferring mammalian histone methyltransferase with
hyperactivity and specific selectivity to lysines 9 and 27 of
histone H3. J Biol Chem. 276:25309–25317. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tachibana M, Matsumura Y, Fukuda M, Kimura
H and Shinkai Y: G9a/GLP complexes independently mediate H3K9 and
DNA methylation to silence transcription. EMBO J. 27:2681–2690.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Li H, Rauch T, Chen ZX, Szabó PE, Riggs AD
and Pfeifer GP: The histone methyltransferase SETDB1 and the DNA
methyltransferase DNMT3A interact directly and localize to
promoters silenced in cancer cells. J Biol Chem. 281:19489–19500.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Steele-Perkins G, Fang W, Yang XH, Van
Gele M, Carling T, Gu J, Buyse IM, Fletcher JA, Liu J, Bronson R,
et al: Tumor formation and inactivation of RIZ1, an Rb-binding
member of a nuclear protein-methyltransferase superfamily. Genes
Dev. 15:2250–2262. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Falandry C, Fourel G, Galy V, Ristriani T,
Horard B, Bensimon E, Salles G, Gilson E and Magdinier F:
CLLD8/KMT1F is a lysine methyltransferase that is important for
chromosome segregation. J Biol Chem. 285:20234–20241. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ding J, Li T, Wang X, Zhao E, Choi JH,
Yang L, Zha Y, Dong Z, Huang S, Asara JM, et al: The histone H3
methyltransferase G9A epigenetically activates the serine-glycine
synthesis pathway to sustain cancer cell survival and
proliferation. Cell Metab. 18:896–907. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kondo Y, Shen L, Ahmed S, Boumber Y,
Sekido Y, Haddad BR and Issa JP: Downregulation of histone H3
lysine 9 methyltransferase G9a induces centrosome disruption and
chromosome instability in cancer cells. PLoS One. 3:e20372008.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Dong C, Wu Y, Yao J, Wang Y, Yu Y,
Rychahou PG, Evers BM and Zhou BP: G9a interacts with Snail and is
critical for Snail-mediated E-cadherin repression in human breast
cancer. J Clin Invest. 122:1469–1486. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Chen MW, Hua KT, Kao HJ, Chi CC, Wei LH,
Johansson G, Shiah SG, Chen PS, Jeng YM, Cheng TY, et al: H3K9
histone methyltransferase G9a promotes lung cancer invasion and
metastasis by silencing the cell adhesion molecule Ep-CAM. Cancer
Res. 70:7830–7840. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Hua KT, Wang MY, Chen MW, Wei LH, Chen CK,
Ko CH, Jeng YM, Sung PL, Jan YH, Hsiao M, et al: The H3K9
methyltransferase G9a is a marker of aggressive ovarian cancer that
promotes peritoneal metastasis. Mol Cancer. 13:1892014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ke XX, Zhang D, Zhu S, Xia Q, Xiang Z and
Cui H: Inhibition of H3K9 methyltransferase G9a repressed cell
proliferation and induced autophagy in neuroblastoma cells. PLoS
One. 9:e1069622014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li KC, Hua KT, Lin YS, Su CY, Ko JY, Hsiao
M, Kuo ML and Tan CT: Inhibition of G9a induces DUSP4-dependent
autophagic cell death in head and neck squamous cell carcinoma. Mol
Cancer. 13:1722014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yuan Y, Tang AJ, Castoreno AB, Kuo SY,
Wang Q, Kuballa P, Xavier R, Shamji AF, Schreiber SL and Wagner BK:
Gossypol and an HMT G9a inhibitor act in synergy to induce cell
death in pancreatic cancer cells. Cell Death Dis. 4:e6902013.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Son HJ, Kim JY, Hahn Y and Seo SB:
Negative regulation of JAK2 by H3K9 methyltransferase G9a in
leukemia. Mol Cell Biol. 32:3681–3694. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Lehnertz B, Pabst C, Su L, Miller M, Liu
F, Yi L, Zhang R, Krosl J, Yung E, Kirschner J, et al: The
methyltransferase G9a regulates HoxA9-dependent transcription in
AML. Genes Dev. 28:317–327. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Tao H, Li H, Su Y, Feng D, Wang X, Zhang
C, Ma H and Hu Q: Histone methyltransferase G9a and H3K9
dimethylation inhibit the self-renewal of glioma cancer stem cells.
Mol Cell Biochem. 394:23–30. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Hung SY, Lin HH, Yeh KT and Chang JG:
Histone-modifying genes as biomarkers in hepatocellular carcinoma.
Int J Clin Exp Pathol. 7:2496–2507. 2014.PubMed/NCBI
|
|
36
|
Wu H, Zhang H, Wang P, Mao Z, Feng L, Wang
Y, Liu C, Xia Q, Li B, Zhao H, et al: Short-Form CDYLb but not
long-form CDYLa functions cooperatively with histone
methyltransferase G9a in hepatocellular carcinomas. Genes
Chromosomes Cancer. 52:644–655. 2013.PubMed/NCBI
|
|
37
|
Zhong X, Chen X, Guan X, Zhang H, Ma Y,
Zhang S, Wang E, Zhang L and Han Y: Overexpression of G9a and MCM7
in oesophageal squamous cell carcinoma is associated with poor
prognosis. Histopathology. 66:192–200. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Schultz DC, Ayyanathan K, Negorev D, Maul
GG and Rauscher FJ III: SETDB1: A novel KAP-1-associated histoneH3,
lysine 9-specific methyltransferase that contributes to
HP1-mediated silencing of euchromatic genes by KRAB zinc-finger
proteins. Genes Dev. 16:919–932. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Harte PJ, Wu W, Carrasquillo MM and Matera
AG: Assignment of a novel bifurcated SET domain gene, SETDB1, to
human chromosome band 1q21 by in situ hybridization and radiation
hybrids. Cytogenet Cell Genet. 84:83–86. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Frietze S, O'Geen H, Blahnik KR, Jin VX
and Farnham PJ: ZNF274 recruits the histone methyltransferase
SETDB1 to the 3 ends of ZNF genes. PLoS One. 5:e150822010.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Cho S, Park JS and Kang YK: Dual functions
of histone-lysine N-methyltransferase Setdb1 protein at
promyelocytic leukemia-nuclear body (PML-NB): Maintaining PML-NB
structure and regulating the expression of its associated genes. J
Biol Chem. 286:41115–41124. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Ceol CJ, Houvras Y, Jane-Valbuena J,
Bilodeau S, Orlando DA, Battisti V, Fritsch L, Lin WM, Hollmann TJ,
Ferré F, et al: The histone methyltransferase SETDB1 is recurrently
amplified in melanoma and accelerates its onset. Nature.
471:513–517. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Lee JK and Kim KC: DZNep, inhibitor of
S-adenosylhomocysteine hydrolase, down-regulates expression of
SETDB1 H3K9me3 HMTase in human lung cancer cells. Biochem Biophys
Res Commun. 438:647–652. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Rodriguez-Paredes M, de Paz Martinez A,
Simó-Riudalbas L, Sayols S, Moutinho C, Moran S, Villanueva A,
Vázquez-Cedeira M, Lazo PA, Carneiro F, et al: Gene amplification
of the histone methyltransferase SETDB1 contributes to human lung
tumorigenesis. Oncogene. 33:2807–2813. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Sun QY, Ding LW, Xiao JF, Chien W, Lim SL,
Hattori N, Goodglick L, Chia D, Mah V, Alavi M, et al: SETDB1
accelerates tumourigenesis by regulating the WNT signalling
pathway. J Pathol. 235:559–570. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Wu PC, Lu JW, Yang JY, Lin IH, Ou DL, Lin
YH, Chou KH, Huang WF, Wang WP, Huang YL, et al: H3K9 histone
methyltransferase, KMT1E/SETDB1, cooperates with the SMAD2/3
pathway to suppress lung cancer metastasis. Cancer Res.
74:7333–7343. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Spyropoulou A, Gargalionis A, Dalagiorgou
G, Adamopoulos C, Papavassiliou KA, Lea RW, Piperi C and
Papavassiliou AG: Role of histone lysine methyltransferases SUV39H1
and SETDB1 in gliomagenesis: Modulation of cell proliferation,
migration, and colony formation. Neuromolecular Med. 16:70–82.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Sun Y, Wei M, Ren SC, Chen R, Xu WD, Wang
FB, Lu J, Shen J, Yu YW, Hou JG, et al: Histone methyltransferase
SETDB1 is required for prostate cancer cell proliferation,
migration and invasion. Asian J Androl. 16:319–324. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zhang H, Cai K, Wang J, Wang X, Cheng K,
Shi F, Jiang L, Zhang Y and Dou J: MiR-7, inhibited indirectly by
lincRNA HOTAIR, directly inhibits SETDB1 and reverses the EMT of
breast cancer stem cells by downregulating the STAT3 pathway. Stem
Cells. 32:2858–2868. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Kouzarides T: Histone methylation in
transcriptional control. Curr Opin Genet Dev. 12:198–209. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Jansen MP, Reijm EA, Sieuwerts AM,
Ruigrok-Ritstier K, Look MP, Rodríguez-González FG, Heine AA,
Martens JW, Sleijfer S, Foekens JA, et al: High miR-26a and low
CDC2 levels associate with decreased EZH2 expression and with
favorable outcome on tamoxifen in metastatic breast cancer. Breast
Cancer Res Treat. 133:937–947. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Raaphorst FM, Meijer CJ, Fieret E,
Blokzijl T, Mommers E, Buerger H, Packeisen J, Sewalt RA, Otte AP
and van Diest PJ: Poorly differentiated breast carcinoma is
associated with increased expression of the human polycomb group
EZH2 gene. Neoplasia. 5:481–488. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Mu Z, Li H, Fernandez SV, Alpaugh KR,
Zhang R and Cristofanilli M: EZH2 knockdown suppresses the growth
and invasion of human inflammatory breast cancer cells. J Exp Clin
Cancer Res. 32:702013. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Zeidler M, Varambally S, Cao Q, Chinnaiyan
AM, Ferguson DO, Merajver SD and Kleer CG: The Polycomb group
protein EZH2 impairs DNA repair in breast epithelial cells.
Neoplasia. 7:1011–1019. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Hoffmann MJ, Engers R, Florl AR, Otte AP,
Muller M and Schulz WA: Expression changes in EZH2, but not in
BMI-1, SIRT1, DNMT1 or DNMT3B are associated with DNA methylation
changes in prostate cancer. Cancer Biol Ther. 6:1403–1412. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Varambally S, Dhanasekaran SM, Zhou M,
Barrette TR, Kumar-Sinha C, Sanda MG, Ghosh D, Pienta KJ, Sewalt
RG, Otte AP, et al: The polycomb group protein EZH2 is involved in
progression of prostate cancer. Nature. 419:624–629. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Bryant RJ, Cross NA, Eaton CL, Hamdy FC
and Cunliffe VT: EZH2 promotes proliferation and invasiveness of
prostate cancer cells. Prostate. 67:547–556. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Kim SH, Joshi K, Ezhilarasan R, Myers TR,
Siu J, Gu C, Nakano-Okuno M, Taylor D, Minata M, Sulman EP, et al:
EZH2 protects glioma stem cells from radiation-induced cell death
in a MELK/FOXM1-dependent manner. Stem Cell Rep. 4:226–238. 2015.
View Article : Google Scholar
|
|
59
|
Zhang W, Lv S, Liu J, Zang Z, Yin J, An N,
Yang H and Song Y: PCI-24781 down-regulates EZH2 expression and
then promotes glioma apoptosis by suppressing the PIK3K/Akt/mTOR
pathway. Genet Mol Biol. 37:716–724. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Xu ZQ, Zhang L, Gao BS, Wan YG, Zhang XH,
Chen B, Wang YT, Sun N and Fu YW: EZH2 promotes tumor progression
by increasing VEGF expression in clear cell renal cell carcinoma.
Clin Transl Oncol. 17:41–49. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Xia H, Zhang W, Li Y, Guo N and Yu C: EZH2
silencing with RNA interference induces G2/M arrest in
human lung cancer cells in vitro. BioMed Res Int. 2014:3487282014.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Guo SQ and Zhang YZ: Overexpression of
enhancer of zests homolog 2 in lymphoma. Chin Med J (Engl).
125:3735–3739. 2012.PubMed/NCBI
|
|
63
|
Fujii S, Ito K, Ito Y and Ochiai A:
Enhancer of zeste homologue 2 (EZH2) down-regulates RUNX3 by
increasing histone H3 methylation. J Biol Chem. 283:17324–17332.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Zhang P, Yang X, Ma X, Ingram DR, Lazar
AJ, Torres KE and Pollock RE: Antitumor effects of pharmacological
EZH2 inhibition on malignant peripheral nerve sheath tumor through
the miR-30a and KPNB1 pathway. Mol Cancer. 14:552015. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Dubuc AM, Remke M, Korshunov A, Northcott
PA, Zhan SH, Mendez-Lago M, Kool M, Jones DT, Unterberger A,
Morrissy AS, et al: Aberrant patterns of H3K4 and H3K27 histone
lysine methylation occur across subgroups in medulloblastoma. Acta
Neuropathol. 125:373–384. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Wan J, Zhan J, Li S, Ma J, Xu W, Liu C,
Xue X, Xie Y, Fang W, Chin YE, et al: PCAF-primed EZH2 acetylation
regulates its stability and promotes lung adenocarcinoma
progression. Nucleic Acids Res. 43:3591–3604. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Pekowska A, Benoukraf T, Zacarias-Cabeza
J, Belhocine M, Koch F, Holota H, Imbert J, Andrau JC, Ferrier P
and Spicuglia S: H3K4 tri-methylation provides an epigenetic
signature of active enhancers. EMBO J. 30:4198–4210. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Wang P, Lin C, Smith ER, Guo H, Sanderson
BW, Wu M, Gogol M, Alexander T, Seidel C, Wiedemann LM, et al:
Global analysis of H3K4 methylation defines MLL family member
targets and points to a role for MLL1-mediated H3K4 methylation in
the regulation of transcriptional initiation by RNA polymerase II.
Mol Cell Biol. 29:6074–6085. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Wu M, Wang PF, Lee JS, Martin-Brown S,
Florens L, Washburn M and Shilatifard A: Molecular regulation of
H3K4 trimethylation by Wdr82, a component of human Set1/COMPASS.
Mol Cell Biol. 28:7337–7344. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Gregory GD, Vakoc CR, Rozovskaia T, Zheng
X, Patel S, Nakamura T, Canaani E and Blobel GA: Mammalian ASH1L is
a histone methyltransferase that occupies the transcribed region of
active genes. Mol Cell Biol. 27:8466–8479. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Stoller JZ, Huang L, Tan CC, Huang F, Zhou
DD, Yang J, Gelb BD and Epstein JA: Ash2l interacts with Tbx1 and
is required during early embryogenesis. Exp Biol Med (Maywood).
235:569–576. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Hamamoto R, Furukawa Y, Morita M, Iimura
Y, Silva FP, Li M, Yagyu R and Nakamura Y: SMYD3 encodes a histone
methyltransferase involved in the proliferation of cancer cells.
Nat Cell Biol. 6:731–740. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Tao Y, Neppl RL, Huang ZP, Chen J, Tang
RH, Cao R, Zhang Y, Jin SW and Wang DZ: The histone
methyltransferase Set7/9 promotes myoblast differentiation and
myofibril assembly. J Cell Biol. 194:551–565. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Sirinupong N, Brunzelle J, Ye J, Pirzada
A, Nico L and Yang Z: Crystal structure of cardiac-specific histone
methyltransferase SmyD1 reveals unusual active site architecture. J
Biol Chem. 285:40635–40644. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Peserico A, Germani A, Sanese P, Barbosa
AJ, di Virgilio V, Fittipaldi R, Fabini E, Bertucci C, Varchi G,
Moyer MP, et al: A SMYD3 small-molecule inhibitor impairing cancer
cell growth. J Cell Physiol. 230:2447–2460. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Xu JY, Chen LB, Xu JY, Yang Z, Wei HY and
Xu RH: Inhibition of SMYD3 gene expression by RNA interference
induces apoptosis in human hepatocellular carcinoma cell line
HepG2. Ai Zheng. 25:526–532. 2006.(In Chinese). PubMed/NCBI
|
|
77
|
Dong SW, Zhang H, Wang BL, Sun P, Wang YG
and Zhang P: Effect of the downregulation of SMYD3 expression by
RNAi on RIZ1 expression and proliferation of esophageal squamous
cell carcinoma. Oncol Rep. 32:1064–1070. 2014.PubMed/NCBI
|
|
78
|
Hamamoto R, Silva FP, Tsuge M, Nishidate
T, Katagiri T, Nakamura Y and Furukawa Y: Enhanced SMYD3 expression
is essential for the growth of breast cancer cells. Cancer Sci.
97:113–118. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Wang SZ, Luo XG, Shen J, Zou JN, Lu YH and
Xi T: Knockdown of SMYD3 by RNA interference inhibits cervical
carcinoma cell growth and invasion in vitro. BMB Rep. 41:294–299.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Liu C, Wang C, Wang K, Liu L, Shen Q, Yan
K, Sun X, Chen J, Liu J, Ren H, et al: SMYD3 as an oncogenic driver
in prostate cancer by stimulation of androgen receptor
transcription. J Natl Cancer Inst. 105:1719–1728. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Morishita M and di Luccio E: Cancers and
the NSD family of histone lysine methyltransferases. Biochim
Biophys Acta. 1816:158–163. 2011.PubMed/NCBI
|
|
82
|
Duns G, van den Berg E, van Duivenbode I,
Osinga J, Hollema H, Hofstra RM and Kok K: Histone
methyltransferase gene SETD2 is a novel tumor suppressor gene in
clear cell renal cell carcinoma. Cancer Res. 70:4287–4291. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Maltby VE, Martin BJ, Schulze JM, Johnson
I, Hentrich T, Sharma A, Kobor MS and Howe L: Histone H3 lysine 36
methylation targets the Isw1b remodeling complex to chromatin. Mol
Cell Biol. 32:3479–3485. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Sampson ER, Yeh SY, Chang HC, Tsai MY,
Wang X, Ting HJ and Chang C: Identification and characterization of
androgen receptor associated coregulators in prostate cancer cells.
J Biol Regul Homeost Agents. 15:123–129. 2001.PubMed/NCBI
|
|
85
|
Bianco-Miotto T, Chiam K, Buchanan G,
Jindal S, Day TK, Thomas M, Pickering MA, O'Loughlin MA, Ryan NK,
Raymond WA, et al: Global levels of specific histone modifications
and an epigenetic gene signature predict prostate cancer
progression and development. Cancer Epidemiol Biomarkers Prev.
19:2611–2622. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Berdasco M, Ropero S, Setien F, Fraga MF,
Lapunzina P, Losson R, Alaminos M, Cheung NK, Rahman N and Esteller
M: Epigenetic inactivation of the Sotos overgrowth syndrome gene
histone methyltransferase NSD1 in human neuroblastoma and glioma.
Proc Natl Acad Sci USA. 106:21830–21835. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Zhao F, Chen Y, Zeng L, Li R, Zeng R, Wen
L, Liu Y and Zhang C: Role of triptolide in cell proliferation,
cell cycle arrest, apoptosis and histone methylation in multiple
myeloma U266 cells. Eur J Pharmacol. 646:1–11. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Thanasopoulou A, Tzankov A and Schwaller
J: Potent co-operation between the NUP98-NSD1 fusion and the
FLT3-ITD mutation in acute myeloid leukemia induction.
Haematologica. 99:1465–1471. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Ostronoff F, Othus M, Gerbing RB, Loken
MR, Raimondi SC, Hirsch BA, Lange BJ, Petersdorf S, Radich J,
Appelbaum FR, et al: NUP98/NSD1 and FLT3/ITD coexpression is more
prevalent in younger AML patients and leads to induction failure: A
COG and SWOG report. Blood. 124:2400–2407. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Job B, Bernheim A, Beau-Faller M,
Camilleri-Broët S, Girard P, Hofman P, Mazières J, Toujani S,
Lacroix L, Laffaire J, et al: LG Investigators: Genomic aberrations
in lung adenocarcinoma in never smokers. PLoS One. 5:e151452010.
View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Seiwert TY, Zuo Z, Keck MK, Khattri A,
Pedamallu CS, Stricker T, Brown C, Pugh TJ, Stojanov P, Cho J, et
al: Integrative and comparative genomic analysis of HPV-positive
and HPV-negative head and neck squamous cell carcinomas. Clin
Cancer Res. 21:632–641. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Deardorff MA, Maisenbacher M and Zackai
EH: Ganglioglioma in a Sotos syndrome patient with an NSD1
deletion. Am J Med Genet A. 130A:393–394. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Wang Y, Thomas A, Lau C, Rajan A, Zhu Y,
Killian JK, Petrini I, Pham T, Morrow B, Zhong X, et al: Mutations
of epigenetic regulatory genes are common in thymic carcinomas. Sci
Rep. 4:73362014. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Gossage L, Murtaza M, Slatter AF,
Lichtenstein CP, Warren A, Haynes B, Marass F, Roberts I, Shanahan
SJ, Claas A, et al: Clinical and pathological impact of VHL, PBRM1,
BAP1, SETD2, KDM6A, and JARID1c in clear cell renal cell carcinoma.
Genes Chromosomes Cancer. 53:38–51. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Hao C, Wang L, Peng S, Cao M, Li H, Hu J,
Huang X, Liu W, Zhang H, Wu S, et al: Gene mutations in primary
tumors and corresponding patient-derived xenografts derived from
non-small cell lung cancer. Cancer Lett. 357:179–185. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Huether R, Dong L, Chen X, Wu G, Parker M,
Wei L, Ma J, Edmonson MN, Hedlund EK, Rusch MC, et al: The
landscape of somatic mutations in epigenetic regulators across
1,000 paediatric cancer genomes. Nat Commun. 5:36302014. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Zhang Q, Xue P, Li H, Bao Y, Wu L, Chang
S, Niu B, Yang F and Zhang T: Histone modification mapping in human
brain reveals aberrant expression of histone H3 lysine 79
dimethylation in neural tube defects. Neurobiol Dis. 54:404–413.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Martin C and Zhang Y: The diverse
functions of histone lysine methylation. Nat Rev Mol Cell Biol.
6:838–849. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Kim W, Choi M and Kim JE: The histone
methyltransferase Dot1/DOT1L as a critical regulator of the cell
cycle. Cell Cycle. 13:726–738. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Chang MJ, Wu H, Achille NJ, Reisenauer MR,
Chou CW, Zeleznik-Le NJ, Hemenway CS and Zhang W: Histone H3 lysine
79 methyltransferase Dot1 is required for immortalization by MLL
oncogenes. Cancer Res. 70:10234–10242. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Kim W, Kim R, Park G, Park JW and Kim JE:
Deficiency of H3K79 histone methyltransferase Dot1-like protein
(DOT1L) inhibits cell proliferation. J Biol Chem. 287:5588–5599.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Oda H, Okamoto I, Murphy N, Chu J, Price
SM, Shen MM, Torres-Padilla ME, Heard E and Reinberg D:
Monomethylation of histone H4-lysine 20 is involved in chromosome
structure and stability and is essential for mouse development. Mol
Cell Biol. 29:2278–2295. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Qin Y, Ouyang H, Liu J and Xie Y: Proteome
identification of proteins interacting with histone
methyltransferase SET8. Acta Biochim Biophys Sin (Shanghai). 2013.
View Article : Google Scholar
|
|
104
|
Jørgensen S, Schotta G and Sørensen CS:
Histone H4 lysine 20 methylation: Key player in epigenetic
regulation of genomic integrity. Nucleic Acids Res. 41:2797–2806.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Wu S and Rice JC: A new regulator of the
cell cycle: The PR-Set7 histone methyltransferase. Cell Cycle.
10:68–72. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Morishita M and di Luccio E: Structural
insights into the regulation and the recognition of histone marks
by the SET domain of NSD1. Biochem Biophys Res Commun. 412:214–219.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Yang P, Guo L, Duan ZJ, Tepper CG, Xue L,
Chen X, Kung HJ, Gao AC, Zou JX and Chen HW: Histone
methyltransferase NSD2/MMSET mediates constitutive NF-κB signaling
for cancer cell proliferation, survival, and tumor growth via a
feed-forward loop. Mol Cell Biol. 32:3121–3131. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Beck DB, Oda H, Shen SS and Reinberg D:
PR-Set7 and H4K20me1: At the crossroads of genome integrity, cell
cycle, chromosome condensation, and transcription. Genes Dev.
26:325–337. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Yokoyama Y, Matsumoto A, Hieda M, Shinchi
Y, Ogihara E, Hamada M, Nishioka Y, Kimura H, Yoshidome K,
Tsujimoto M, et al: Loss of histone H4K20 trimethylation predicts
poor prognosis in breast cancer and is associated with invasive
activity. Breast Cancer Res. 16:R662014. View Article : Google Scholar : PubMed/NCBI
|