Introduction
Gastric cancer is the third most frequently
diagnosed cancer type, and has been the leading cause of
cancer-related death in less-developed countries (1). Although advances in therapeutic
strategies such as surgery and systemic chemotherapy, have improved
the clinical outcomes for gastric cancer patients, the prognosis of
these patients remains poor because of frequent cancer recurrence
(2). Nevertheless, due to the
encouraging development in newly targeted therapies, such a
disappointing clinical condition has been improving. Trastuzumab,
the first targeted antibody against human epidermal growth factor
receptor 2 (HER2), has been approved for the treatment of patients
with HER2-positive metastatic gastric cancer (3). However, even with trastuzumab
resistance, gastric cancer almost inevitably progresses, as the
tumors become resistant to trastuzumab after an initial period of
clinical benefits (4). Mechanisms
leading to trastuzumab resistance in breast cancer, such as
cross-talk between HER2 and other intracellular kinase receptors,
have been described recently (5),
however, the underlying mechanisms of trastuzumab resistance in
gastric cancer remain largely unknown.
Cancer cells are able to activate alternative
survival pathways yielding drug resistance in response to
chemotherapy, and thereby leads to the chemotherapeutic treatment
failure (6). Since drug resistance
is a common cause limiting the efficacy of cancer treatment,
several mechanisms are elucidated to be responsible for the
resistance development, such as increased rates of drug efflux,
apoptosis resistance, and microRNAs (miRNAs) mediated
overexpression of many drug resistance-related genes (7). Microarray technology, a
high-throughput platform that analyzes gene expression, in
combination with bioinformatics analysis has been widely used as a
promising tool to acquire gene signature during tumorigenesis or
drug resistance, and identify prognostic biomarkers in cancer
patients (8–10). Abnormal expression patterns of drug
resistance-related genes commonly play important roles in drug
resistance (11), thus, exploring
and identifying the critical drug resistance-related genes based on
microarray analysis would have a significant impact.
Therefore, in the present study, we sought to
identify the potential genes that promote trastuzumab resistance in
gastric cancer through retrieving microarray data from public
databases and comprehensive bioinformatics analysis.
Materials and methods
Microarray data
The gene expression profiles of GSE26899, GSE77346,
GSE54129, and GSE65801 were obtained from the Gene Expression
Omnibus (GEO; www.ncbi.nlm.nih.gov/geo). In detail, GSE26899 dataset
is consisted of 96 clinical gastric tumor tissues and 12 adjacent
normal tissues; GSE77346 dataset is consisted of 1
trastuzumab-sensitive cell line and 4 trastuzumab-resistant cell
lines (12); GSE54129 includes 111
human gastric cancer tissues and 21 non-cancerous tissues; GSE65801
contains 32 gastric cancer tissues and 32 paired non-cancerous
tissues (13).
Processing of microarray data
The raw microarray data files of the datasets
downloaded from the GEO website were subsequently analyzed via
using the GEO2R (www.ncbi.nlm.nih.gov/geo/geo2r/), an online tool
comparing two or more groups of samples in the same experimental
setting (14). False Discovery
Rate (FDR) of P-value adjusted (adj. P) to 0.05 and |logFC|>1
were set as the cut-off criteria.
Functional and pathway enrichment
analyses
Gene ontology (GO) analysis is a commonly used
approach for functional studies with three ontologies including
biological process, molecular function, and cellular component
(15), while Kyoto Encyclopedia of
Genes and Genomes (KEGG) is a knowledge base for the systematic
study of gene functions (16). To
study the functional annotations of differentially expressed genes
(DEGs), we next employed Database for Annotation, Visualization and
Integrated Discovery (DAVID, david.abcc.ncifcrf.gov/,) to process the GO and KEGG
analyses of DEGs identified in gastric cancer samples. P<0.05
was set as the threshold.
Protein-protein interaction (PPI)
network construction and module analysis
The Search Tool for the Retrieval of Interacting
Genes (STRING), an online database (string-db.org)
designed to evaluate PPI information, covers 9,643,763 proteins
from more than 2,000 organisms, which was used to construct the
PPI. To evaluate the interactive associations of DEGs identified
from GSE26899, we mapped these DEGs to the STRING (version 10.5)
database. Confidence score >0.4 was selected as significant. PPI
networks were constructed by STRING and visualized by Cytoscape.
Subsequently, the plug-in Molecular Complex Detection (MCODE) was
employed to screen the modules of PPI networks in Cytoscape with
the threshold set as follows: MCODE scores >10.
Survival analysis of collagen type IV
α1 chain (COL4A1)
To evaluate the association between COL4A1
level and its clinical outcomes, Kaplan-Meier plotter (KM plotter;
www.kmplot.com), an online survival analysis
tool, was performed. KM plotter is capable of assessing the effect
of 54,675 genes on overall survival via using 10,188 cancer samples
including 4,142 breast, 1,648 ovarian, 2,437 lung, and 1,065
gastric cancer patients (17).
Patients with gastric cancer were separated into high- and
low-expression groups according to the level of COL4A1, and the
overall survival was then analyzed. The hazard ratio (HR) with 95%
confidence intervals and log rank P-value were calculated.
Analysis of COL4A1 by geneMANIA and
coremine
GeneMANIA, an online tool (www.genemania.org/), can be used to generate
hypotheses of gene function, analyze gene lists, and prioritize
genes for functional assays (18).
After selecting Homo sapiens from the nine optional organisms,
COL4A1 was entered into the search bar and the results were
then collected. Annotation of biological processes involving
COL4A1 was performed by consulting the Coremine Medical
online database (www.coremine.com/medical/).
Prediction of miRNAs
To predict the miRNAs targeting the mRNA of
COL4A1, miRWalk (version 2.0, zmf.umm.uni-heidelberg.de/apps/zmf/mirwalk2/), an
online platform supplying information about predicted and
experimentally validated miRNA-target interactions, was then
employed (19). Herein, nine
prediction programs (miRWalk, miRanda, miRDB, miRNAMap, Pictar2,
PITA, RNA22, RNAhybrid and Targetscan) were selected. These
predicted miRNAs were then overlapped by at least seven programs,
and selected for further analysis. Pathway enrichment analysis of
these miRNAs was performed by using the DIANA-mirPath web server
(snf-515788.vm.okeanos.grnet.gr/index.php?r=mirpath) (20).
Statistical analysis
SPSS 22.0 software (IBM Corp., Armonk, NY, USA) was
used to analyze data. Two tailed Student's t-test was used to
compare the two groups. P<0.05 was considered to indicate a
statistically significant difference.
Results
Module acquisition based on the DEGs
identified in gastric cancer tissues
Drug resistance-related genes identified from in
vitro drug-induced resistant models may simply represent
transcriptional changes, thus, complementary agents targeting these
genes are usually failure to translate into clinical practice
(21). Because drug resistance
acquisition can arise before the malignant transformation stage
(22), the genes playing important
roles in tumorigenesis and drug resistance is thereby more likely
to be critical for resistance occurrence. Therefore, we first
identified 509 DEGs in gastric cancer tissues from the GSE26899
dataset using a 2-fold-change and adj. P<0.05 as the threshold
cutoff. Among these DEGs, the expression of these 172 genes was
significantly upregulated, while that of 337 genes was
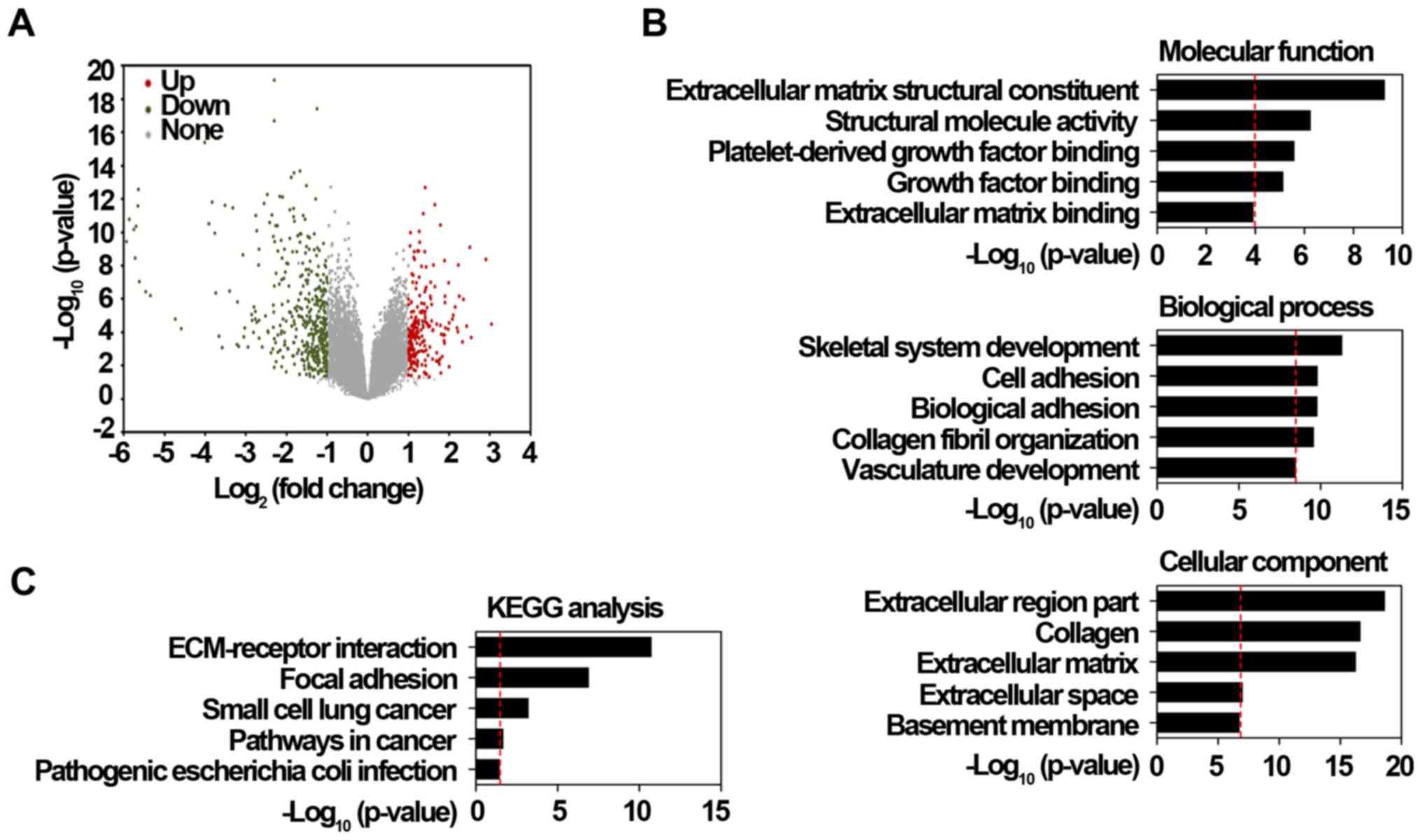
significantly downregulated in cancer tissues (Fig. 1A). To explore the potential roles
of these significantly upregulated DEGs, GO and KEGG pathway
analyses, which can provide valuable insights regarding protein
function, were performed (23). As
shown in Fig. 1B, GO results
showed that these significantly upregulated DEGs were largely
associated with extracellular region part, collagen, extracellular
matrix, and cell or biological adhesion processes. KEGG results
also consistently revealed that these DEGs were highly enriched in
extracellular matrix-receptor interaction and focal adhesion
pathways (Fig. 1C). Based on
further DEGs analysis with the STRING database, the PPI network of
these DEGs, which contained 505 nodes and 1,207 edges, was
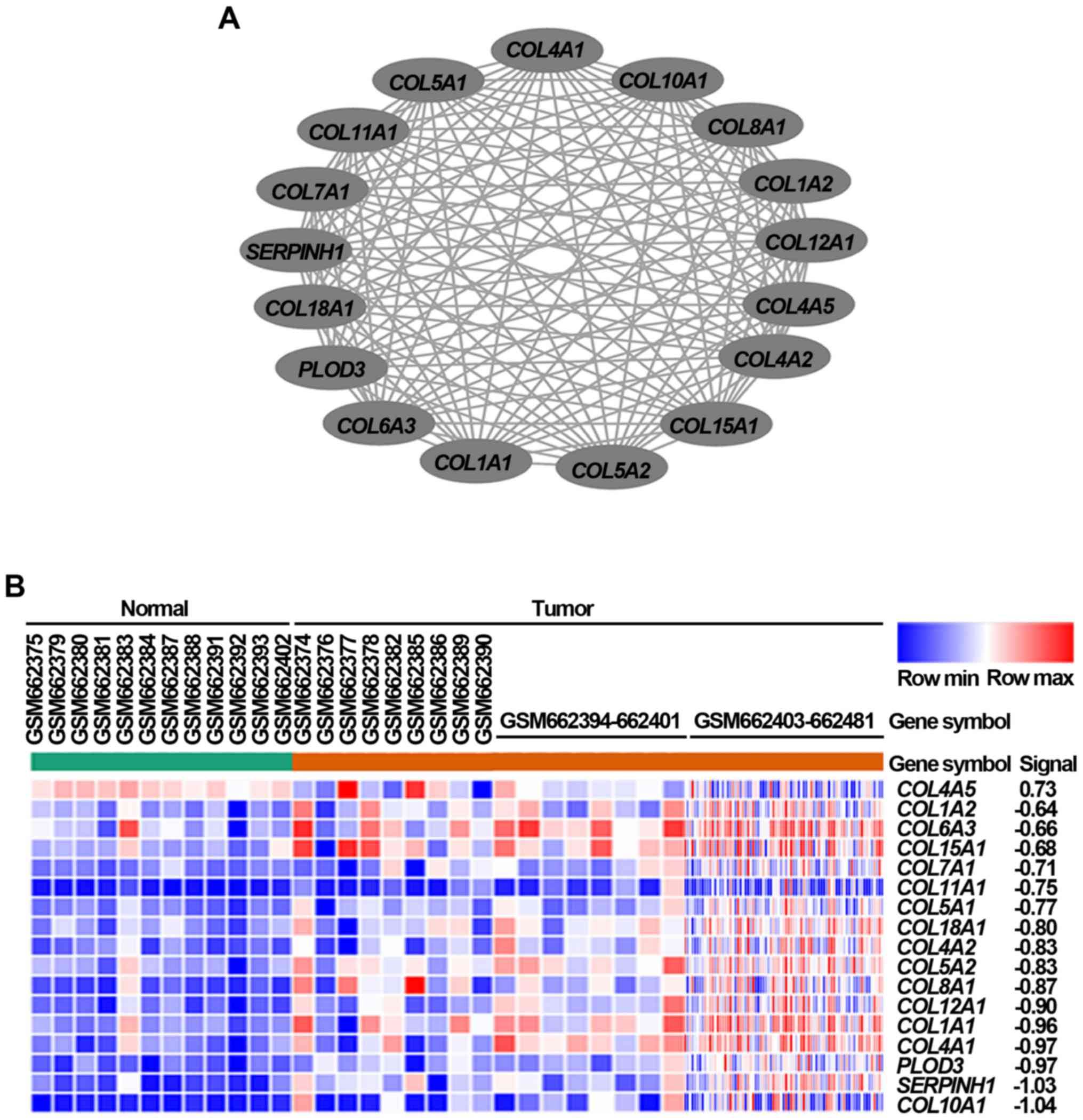
subsequently constructed. Using the MCODE plug-in in Cytoscape, we
obtained the module with the highest score (Fig. 2A), and also performed a cluster
analysis of these genes in the module (Fig. 2B). The module with the highest
score was selected based on these DEGs identified from gastric
cancer tissues via bioinformatics methods.
Expression of COL4A1, overexpressed in
gastric cancer tissues, is also upregulated in
trastuzumab-resistant gastric cancer cells
To further explore the potential genes contributing
to trastuzumab resistance, we retrieved GSE77346 data, a microarray
dataset consisted of one trastuzumab-sensitive gastric cancer cell
line and four trastuzumab-resistant gastric cancer cell lines
(12). After screening the
significantly upregulated DEGs in trastuzumab-resistant cancer
cells, COL4A1, one of hub genes in the selected module
(Fig. 2A), was also found to be
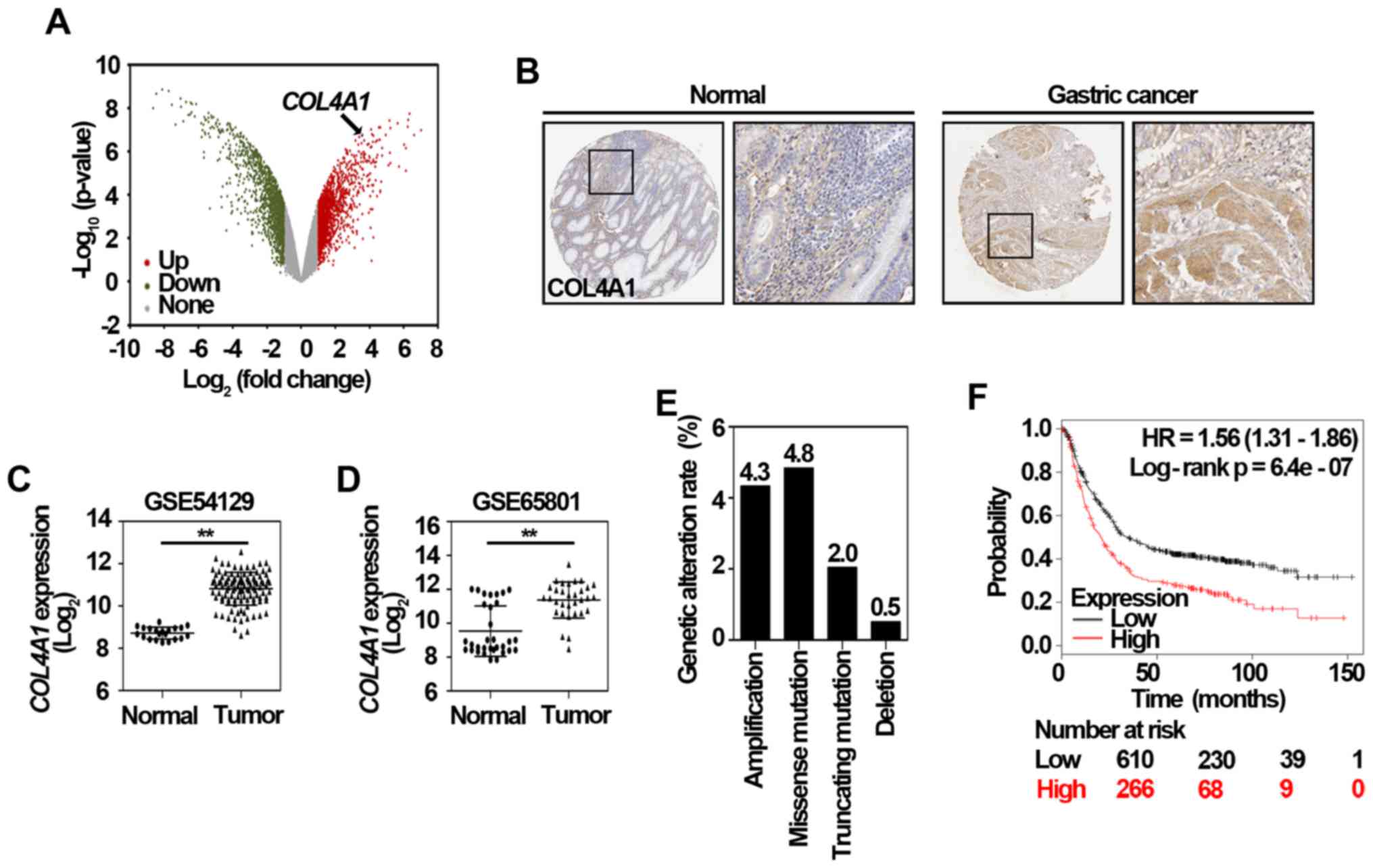
significantly upregulated in trastuzumab-resistant cells (Fig. 3A), suggesting COL4A1 might
be important in both tumorigenesis and trastuzumab resistance.
Since the gene expression is not always consistent with its protein
amount (24), further validation
of COL4A1 protein level in clinical gastric cancer tissues
is quite necessary. By employing the Human Protein Atlas database,
an online tool analyzing protein level from clinical specimens, we
observed that COL4A1 was positively expressed in gastric cancer
tissue, but negatively expressed in normal gastric tissue (Fig. 3B). Two other datasets, including
GSE54129 and GSE65801 were also used to validate the mRNA level of
COL4A1, which was indeed significantly upregulated in
clinical gastric cancer samples (Fig.
3C and D). To investigate potential regulation mechanisms of
COL4A1 in gastric cancer, the genomic alteration of
COL4A1 in The Cancer Genome Atlas (TCGA) cohort was analyzed
using the cBioPortal (www.cbioportal.org/) (25). As shown in Fig. 3E, the amplification, missense
mutation, truncating mutation, and deletion of COL4A1
accounted for 4.3, 4.8, 2.0, and 0.5% of stomach adenocarcinoma
cases, respectively. The prognostic value of COL4A1 in patients
with gastric cancer was analyzed using the KM plotter according to
the low and high expression of COL4A1. As shown in Fig. 3F, the high mRNA level of
COL4A1 [HR 1.56 (1.31–1.86)] was associated with poor
overall survival of gastric cancer patients. Altogether, these data
suggest that COL4A1 is a potential gene candidate that
promotes the development of gastric cancer and subsequent
trastuzumab resistance.
Drug resistance function of COL4A1 is
validated by protein/gene interactions and biological process
annotation analyses
As a user-friendly web interface for functional
prediction of genes, GeneMANIA has been widely used as an effective
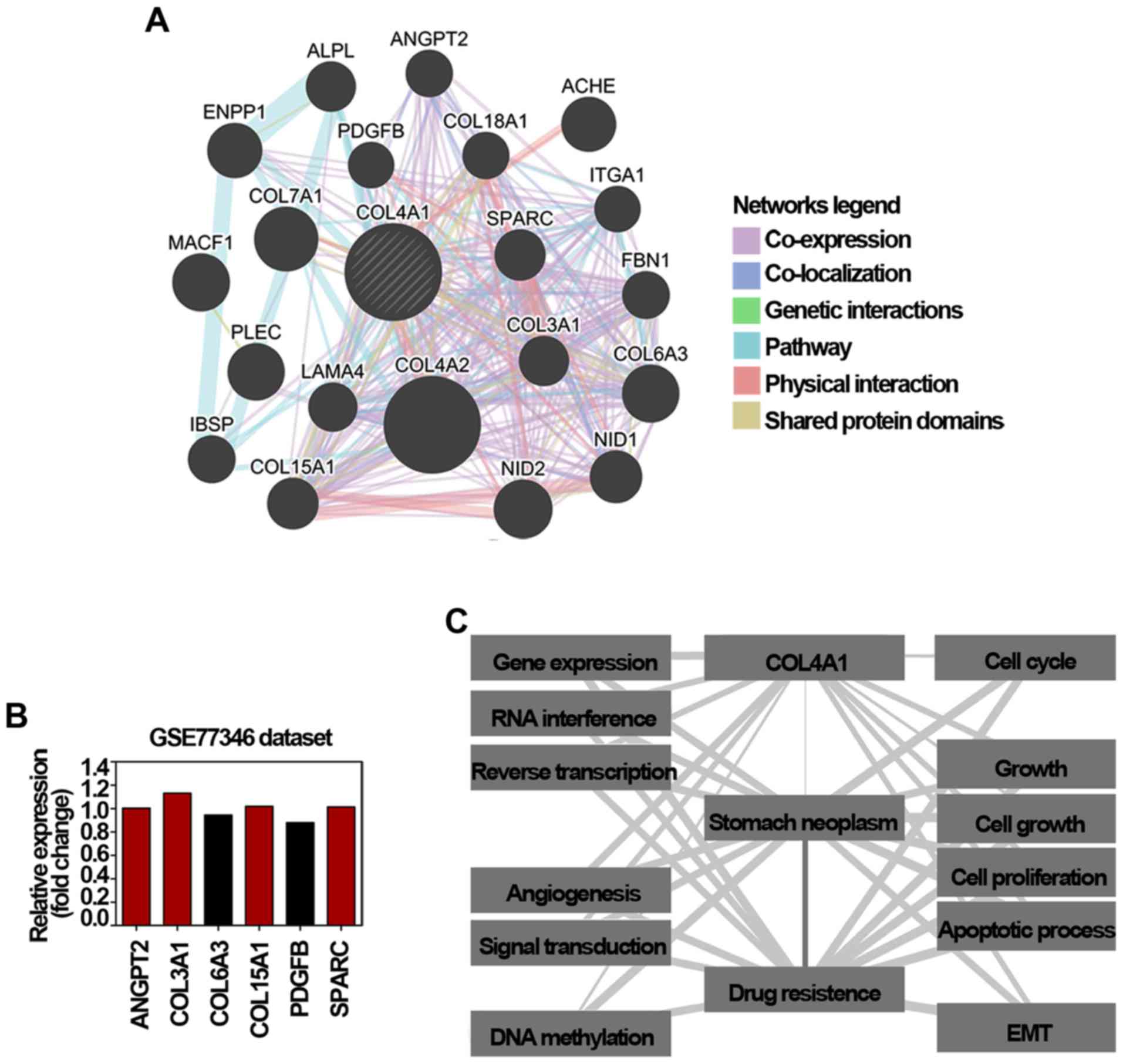
tool to predict and explore drug resistance-related genes (18). Accordingly, COL4A1 showed
interactions with 20 proteins/genes; among these 6 genes were
involved in conferring drug resistance in cancer (Fig. 4A). The mRNA expression of these 6
genes in gastric cancer cells and trastuzumab-resistant gastric
cancer cells were also evaluated using the microarray data of
GSE77346. As shown in Fig. 4B, the
mRNA level of angiopoietin 2 (ANGPT2), COL3A1,
COL15A1, and secreted protein acidic and cysteine rich
(SPARC) were upregulated in trastuzumab-resistant gastric
cancer cells. Among these genes, COL4A1 co-expressed and
co-localized with ANGPT2, collagen type VI α 3 chain
(COL6A3), and SPARC, respectively. It has been
reported bevacizumab resistance in glioblastoma and doxorubin
resistance in liver cancer are largely mediated by an enforcement
of ANGPT2/TIE2 signaling (26,27).
Additionally, overexpression of COL6A3, one of the most
highly upregulated genes in oxaliplatin and cisplatin resistant
ovarian cancer cells, has also been recognized to confer cisplatin
resistance in sensitive cancer cells (28). Moreover, the accumulation of
intracellular SPARC can drive imatinib resistance in chronic
myelogenous leukemia cells (29),
and also modulate cisplatin resistance via modulating the Let-7f-1
miRNA/HMGB1 signaling in medulloblastoma cells (30). COL3A1 and COL15A1,
two types of fibrillar collagen highly expressed in
chemotherapeutic drugs resistant ovarian cancer cells (31), were also exhibited to co-express,
share protein domains, and co-localize with COL4A1 (Fig. 4A). Additionally, COL4A1 also
was co-expressed, shared pathways, and had physical interactions
with platelet derived growth factor subunit B (PDGFB).
Akt/PDGF-B signaling can regulate Akt, and thus confer the
hypoxia-induced cisplatin resistance in liver cancer cells
(32). PDGFB may contribute to the
resistant phenotype and sustain signaling through MAPK and Akt in
breast cancer cells (33). The
Coremine Medical database is a freely available online tool for
obtaining information on health, medicine, and biology (33); thus, we used this database to
annotate the biological process of COL4A1. As shown in
Fig. 4C, 12 biological processes
significantly associated with COL4A1, gastric cancer, and
drug resistance (P<0.01) were annotated. Considering that the
close relationships of COL4A1 with these processes and the
close relationships of the 12 processes with gastric cancer and
drug resistance, COL4A1 might be involved in the development
of drug resistance in gastric cancer via its effect on these
biological processes. In detail, cell growth related biological
processes (including 4 cell growth, cell proliferation, and
apoptotic process), gene expression regulation-related (including
gene expression, RNA interference, and reverse transcription), and
especially, the epithelial-mesenchymal transition (EMT) biological
process were identified to be closely associated with the
development of COL4A1 in the gastric cancer drug resistance.
Therefore, these results suggest that COL4A1 may confer drug
resistance in gastric cancer via regulating cell growth, gene
expression, and in particular, epithelial mesenchymal transition
(EMT) processes.
Drug resistance function of COL4A1 is
further validated by analyzing functionality of miRNAs that target
COL4A1 mRNA
Post-transcriptional gene expression regulated by
miRNAs is important for multiple cellular processes during
development and pathogenesis (34). Amplification and overexpression of
tumor-promoting miRNAs or genetic loss of tumor-suppressing miRNAs
are tightly correlated with the development of cancer and
chemotherapeutic resistance (35).
Therefore, studying target genes of miRNAs is a focus of interest
due to the diagnostic and therapeutic relevance, and the gene
functions can also be predicted according to functionality of these
miRNAs targeting the gene (36).
To obtain the miRNAs targeting COL4A1 mRNA, the miRNA-mRNA
interaction analysis was performed by using the miRWalk. We
identified 553 miRNAs predicted to transcriptionally target
COL4A1 mRNA, indicating the regulation of COL4A1 by
miRNAs. These miRNAs predicted by at least 8 of 9 prediction tools
were then submitted to pathway enrichment using DIANA miRPath
(20). As shown in Table I, the top 5 highly associated
pathway were ECM-receptor interaction, TGF-β signaling pathway,
viral carcinogenesis, proteoglycans in cancer, and Hippo signaling
pathway, which are almost reported to be related with drug
resistance (37–41). It has been established TGFβ
signaling can activate autophagy process, and thereby lead to the
oxaliplatin resistance in colorectal cancer (38). In addition, the activation of Hippo
signaling also contributes to the drug resistance and cancer
relapse (42). YAP, an effector of
hippo signaling, has also been reported to alter clinical response
of EGFR-tyrosine kinase inhibitors in lung cancer patients
(43). Furthermore, it has also
been reported that inactivation of Hippo pathway can restore
gemcitabine sensitivity among a variety of cancers (44). In the present study, as shown in
Table II, 9 of top 10 miRNAs that
target COL4A1 mRNA were tightly associated with drug
resistance in cancers (45–51).
For instance, loss of intracellular miR-29b promotes cisplatin
resistance in gastric cancer. Consistently, ectopic overexpression
of miR-29b in cholangiocarcinoma cells can confer gemcitabine
sensitivity to HuH28 cells (45).
MiR-506 overexpression can confer hydroxycamptothecin resistance in
colon cancer cells by inhibiting PPARα expression (50). Collectively, these data provide
further supports for the drug resistance function of COL4A1
in gastric cancer.
 | Table I.Top 8 enriched pathways regulated by
microRNAs that target collagen type IV α1 mRNA and their
associations with drug resistance in gastric cancer. |
Table I.
Top 8 enriched pathways regulated by
microRNAs that target collagen type IV α1 mRNA and their
associations with drug resistance in gastric cancer.
| Author, year | miRNAs
(hsa-miR-) | KEGG pathway | P-value | Regulation of drug
resistance in cancers | (Refs.) |
|---|
| Wu et al,
2017 | 29b-3p, 124-3p,
148a-3p, 29a-3p, 152-3p, 148b-3p, 506-3p, 628-5p, 29c-3p, 767-5p,
637, 33a-5p, 33b-5p, 203a, 374a-5p, 300, 106a-5p, 98-5p, 381-3p,
let-7b-5p, 7c-5p, 7b-3p, 7a-5p | ECM-receptor
interaction |
1.14×10‒15 | Yes | (37) |
| Sun et al,
2017 |
| TGF-β signaling
pathway |
6.58×10‒11 | Yes | (38) |
| – |
| Viral
carcinogenesis |
9.72×10‒11 | None | – |
| Lanzi et al,
2017 |
| Proteoglycans in
cancer |
3.94×10‒10 | Yes | (39) |
| Gujral et
al, 2017 |
| Hippo signaling
pathway |
6.66×10‒10 | Yes | (44) |
| Lee et al,
2015 |
| Cell cycle |
1.01×10‒9 | Yes | (40) |
| Wallerand et
al, 2010 |
| Adherens
junction |
2.99×10‒8 | Yes | (41) |
| – |
| Pathways in
cancer |
3.46×10‒8 | Cancer pathway | – |
 | Table II.Top 11 microRNAs targeting collagen
type IV α1 mRNA predicted by microRNA-mRNA interactions, and their
drug resistance-associated functions in cancer. |
Table II.
Top 11 microRNAs targeting collagen
type IV α1 mRNA predicted by microRNA-mRNA interactions, and their
drug resistance-associated functions in cancer.
|
|
| miRNA-mRNA
prediction tools |
|
|
|---|
|
|
|
|
|
|
|---|
| Author, year | miRNA (hsa-) | A | B | C | D | E | F | G | H | I | Drug
resistance-related functions of miRNAs in cancers | (Refs.) |
|---|
| Okamoto et
al, 2013 | miR-29b-3p | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Drug
resistance-related | (45) |
| Liu et al,
2016 | miR-124-3p | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Drug
resistance-related | (46) |
| Chen et al,
2017 | miR-148a-3p | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Drug
resistance-related | (47) |
| Zhong et al,
2013 | miR-29a-3p | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Drug
resistance-related | (48) |
| Chen et al,
2017 | miR-152-3p | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Drug
resistance-related | (47) |
| Sui et al,
2015 | miR-148b-3p | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | Drug
resistance-related | (49) |
| Tong et al,
2011 | miR-506-3p | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | Drug
resistance-related | (50) |
| – | miR-628-5p | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | Drug
resistance-related | – |
| Zhang et al,
2013 | miR-29c-3p | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | Drug
resistance-related | (51) |
| – | miR-767-5p | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Drug
resistance-related | – |
| – | miR-637 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | Drug
resistance-related | – |
Discussion
In the present study, based on the bioinformatics
analysis of two microarray datasets including GSE26899 and
GSE77346, COL4A1, which was overexpressed in gastric cancer
tissues and trastuzumab-resistant gastric cancer cells, was
identified as a potential gene that promotes gastric cancer and
trastuzumab resistance. By combining the protein/gene interactions,
biological process annotation, and miRNAs-mRNA interaction
analyses, we showed that COL4A1 may confer trastuzumab
resistance in gastric cancer.
Traditional chemotherapy and newly targeted therapy
are two important methods of cancer treatment; however, the
clinical efficacy of both is largely limited due to the occurrence
of subsequent drug resistance (52). Mechanisms underlying drug
resistance, such as alteration of drug targets or metabolism and
genetic mutation (11) have been
elucidated. Similar to that in breast cancer, the HER2-positivity
rate in gastric cancer can range from 20 to 42% (53). However, although the benefits of
trastuzumab against HER2-positive gastric cancer have been formally
established (3,53), trastuzumab resistance is almost
inevitable and eventually leads to the therapeutic failure
(4). Interfering with the
combination of trastuzumab and HER2 (54) and constitutive dimerization of the
HER2 receptor (55) have been
recognized as causes of trastuzumab resistance. Recently, abnormal
expression of tumorigenesis or drug resistance-related genes, such
as phosphatidylinositol 3-kinase/Akt signaling (56) and insulin like growth factor 1
receptor (57), has been also
identified as the important mechanisms causing trastuzumab
resistance. However, a comprehensive study of specific molecular
mutation underlying trastuzumab resistance is still needed. Drug
resistance-related genes identified from induced drug-resistant
cancer cells may simply be a result of transcriptional changes that
may be irrelevant to resistance mechanisms (58). Thus, targeting these genes usually
fail to translate into clinical practice (21). It has been recently shown that the
ability to acquire drug resistance can arise even before the
malignant transformation stage. For instance, overexpression of
telomerase or inactivation of p53 can contribute to the
drug-resistant phenotype in pre-tumorigenic models (22). Insulin-like growth factor signaling
has also been deemed a factor that promotes both the development of
various tumors and their resistance to chemotherapy (59,60).
This evidence thus reminds us that the genes playing a key role in
both tumor development and subsequent drug resistance likely
produce the essential molecule for drug resistance. Herein, through
retrieving microarray datasets and employing subsequent
bioinformatics analysis, COL4A1 was validated as an
important gene that drives trastuzumab resistance in gastric
cancer.
The heterotrimers formed by COL4A1 and COL4A2
presents in almost all the basement membranes, which are a
specialized form of the extracellular matrix. Besides that the
basement membranes can mediate tissue compartmentalization and
transfer environmental signals to epithelial cells (61), it is also an important structural
and functional component of blood vessels. Accordingly, mutations
in COL4A1 are pleiotropic and contribute to many diseases,
such as myopathy, hemorrhagic stroke, and tumor progression
(62). Upregulated COL4A1 produced
by bladder cancer cells plays pivotal roles in tumor invasion via
induction of tumor budding, while overexpression of COL4A1 also
contributes to breast cancer cells proliferation, which indicates
targeting COL4A1 can be an attractive approach for cancer treatment
(63,64). Besides, COL4A1 has also been
identified as one of biomarkers for prognosis of intrahepatic
cholangiocarcinoma (65). Given
that an interaction between PDGFB and HIF-1α and an interplay
between SPARC, BCL-2, and caspase-8 have been recognized to augment
chemotherapy-induced apoptosis, and thereby inducing resistance
(32,66). Herein, our results suggest
COL4A1 can drive trastuzumab resistance in gastric cancer
via multiple mechanisms, such as cell proliferation and
miRNAs-mediated post-transcriptional modification. Consistent with
our results, a previous study also reported the sustained EMT
phenotype induced by prolonged trastuzumab treatment may lead to
trastuzumab resistance in gastric cancer cells (67). Small molecules that targeting drug
resistance-related genes have shown promising clinical efficacy in
cancer treatment. For instance, erbB3 overexpression drives
paclitaxel resistance in breast cancer, MM-121/SAR256212, an
erbB3-targeted antibody, was thus designed and shows augmented
effect on paclitaxel resistance (68). Activation of the PI3K pathway
frequently occurs in cancers and leads to drug resistance, but
clinical benefits of PI3K inhibitors have been modest to date.
Fortunately, LEE011, a specific CDK 4/6 inhibitor currently under
clinical development, has shown promising effects against PI3K
inhibition resistance (69).
HER2-positive patients receiving trastuzumab may inevitably develop
resistance due to excessive activation of the PI3K/AKT pathway. A
phase 1 clinical trial has been performed to evaluate the efficacy
of the combination of an allosteric AKT inhibitor (MK-2206) and
trastuzumab in patients with HER2-positive solid tumors;
interestingly, MK-2206 is safe, and reversed the trastuzumab
resistance in HER2-overexpressing patients (70). Collectively, this evidence suggests
that identification of drug-resistant genes through bioinformatics
methods and subsequent design of small molecule drugs may have
great potential.
In conclusion, our results show that COL4A1
may confer trastuzumab resistance in gastric cancer via multiple
mechanisms based on bioinformatics analysis. However, further
investigations elucidating the drug resistance function of
COL4A1 in trastuzumab-resistant gastric cancer models are
necessary.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
LG conceived the project and designed the research
plan. RH and WG performed the experiments and analyzed the data
with input from BS who assisted in experimental design and data
analyses. RH and LG wrote the manuscript, and all authors read and
approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Corso S and Giordano S: How can gastric
cancer molecular profiling guide future therapies? Trends Mol Med.
22:534–544. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bang YJ, Van Cutsem E, Feyereislova A,
Chung HC, Shen L, Sawaki A, Lordick F, Ohtsu A, Omuro Y, Satoh T,
et al: Trastuzumab in combination with chemotherapy versus
chemotherapy alone for treatment of HER2-positive advanced gastric
or gastro-oesophageal junction cancer (ToGA): A phase 3,
open-label, randomised controlled trial. Lancet. 376:687–697. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Aprile G, Giampieri R, Bonotto M, Bittoni
A, Ongaro E, Cardellino GG, Graziano F, Giuliani F, Fasola G,
Cascinu S and Scartozzi M: The challenge of targeted therapies for
gastric cancer patients: The beginning of a long journey. Exp Opin
Investig Drugs. 23:925–942. 2014. View Article : Google Scholar
|
|
5
|
Ritter CA, Perez-Torres M, Rinehart C,
Guix M, Dugger T, Engelman JA and Arteaga CL: Human breast cancer
cells selected for resistance to trastuzumab in vivo overexpress
epidermal growth factor receptor and ErbB ligands and remain
dependent on the ErbB receptor network. Clin Cancer Res.
13:4909–4919. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Gottesman MM, Lavi O, Hall MD and Gillet
JP: Toward a better understanding of the complexity of cancer drug
resistance. Annu Rev Pharmacol Toxicol. 56:85–102. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Garofalo M and Croce CM: MicroRNAs as
therapeutic targets in chemoresistance. Drug Resist Updat.
16:47–59. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lu Y, Lemon W, Liu PY, Yi Y, Morrison C,
Yang P, Sun Z, Szoke J, Gerald WL, Watson M, et al: A gene
expression signature predicts survival of patients with stage I
non-small cell lung cancer. PLoS Med. 3:e4672006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Chen F, Xiang CX, Zhou Y, Ao XS, Zhou DQ,
Peng P, Zhang HQ, Liu HD and Huang X: Gene expression profile for
predicting survival of patients with meningioma. Int J Oncol.
46:791–797. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kulasingam V and Diamandis EP: Strategies
for discovering novel cancer biomarkers through utilization of
emerging technologies. Nat Clin Pract Oncol. 5:588–599. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yin F, Liu X, Li D, Wang Q, Zhang W and Li
L: Tumor suppressor genes associated with drug resistance in
ovarian cancer (Review). Oncol Rep. 30:3–10. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Piro G, Carbone C, Cataldo I, Di
Nicolantonio F, Giacopuzzi S, Aprile G, Simionato F, Boschi F,
Zanotto M, Mina MM, et al: An FGFR3 autocrine loop sustains
acquired resistance to trastuzumab in gastric cancer patients. Clin
Cancer Res. 22:6164–6175. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Li H, Yu B, Li J, Su L, Yan M, Zhang J, Li
C, Zhu Z and Liu B: Characterization of differentially expressed
genes involved in pathways associated with gastric cancer. PLoS
One. 10:e01250132015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Barrett T, Wilhite SE, Ledoux P,
Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH,
Sherman PM, Holko M, et al: NCBI GEO: Archive for functional
genomics data sets-update. Nucleic Acids Res. 41:D991–D995. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hulsegge I, Kommadath A and Smits MA:
Globaltest and GOEAST: Two different approaches for Gene Ontology
analysis. BMC Proc. 3 Suppl 4:pp. S102009; View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Kanehisa M and Goto S: KEGG: Kyoto
encyclopedia of genes and genomes. Nucleic Acids Res. 28:27–30.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Gyorffy B, Lánczky A and Szállási Z:
Implementing an online tool for genome-wide validation of
survival-associated biomarkers in ovarian-cancer using microarray
data from 1287 patients. Endocr Relat Cancer. 19:197–208. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Warde-Farley D, Donaldson SL, Comes O,
Zuberi K, Badrawi R, Chao P, Franz M, Grouios C, Kazi F, Lopes CT,
et al: The GeneMANIA prediction server: Biological network
integration for gene prioritization and predicting gene function.
Nucleic Acids Res. 38:W214–W220. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Dweep H and Gretz N: miRWalk2.0: A
comprehensive atlas of microRNA-target interactions. Nat Methods.
12:6972015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Vlachos IS, Zagganas K, Paraskevopoulou
MD, Georgakilas G, Karagkouni D, Vergoulis T, Dalamagas T and
Hatzigeorgiou AG: DIANA-miRPath v3.0: Deciphering microRNA function
with experimental support. Nucleic Acids Res. 43:W460–W466. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Borst P and Wessels L: Do predictive
signatures really predict response to cancer chemotherapy? Cell
cycle (Georgetown, Tex.). 9:4836–4840. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Yague E, Arance A, Kubitza L, O'Hare M,
Jat P, Ogilvie CM, Hart IR, Higgins CF and Raguz S: Ability to
acquire drug resistance arises early during the tumorigenesis
process. Cancer Res. 67:1130–1137. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Xing Z, Chu C, Chen L and Kong X: The use
of Gene Ontology terms and KEGG pathways for analysis and
prediction of oncogenes. Biochim Biophys Acta. 1860:2725–2734.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Maier T, Guell M and Serrano L:
Correlation of mRNA and protein in complex biological samples. FEBS
Lett. 583:3966–3973. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:pl12013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Labussiere M, Cheneau C, Prahst C, Gállego
Pérez-Larraya J, Farina P, Lombardi G, Mokhtari K, Rahimian A,
Delattre JY, Eichmann A and Sanson M: Angiopoietin-2 May be
involved in the resistance to bevacizumab in recurrent
glioblastoma. Cancer Invest. 34:39–44. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Li T, Liu Z, Jiang K and Ruan Q:
Angiopoietin2 enhances doxorubin resistance in HepG2 cells by
upregulating survivin and Ref-1 via MSK1 activation. Cancer Lett.
337:276–284. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sherman-Baust CA, Weeraratna AT, Rangel
LB, Pizer ES, Cho KR, Schwartz DR, Shock T and Morin PJ: Remodeling
of the extracellular matrix through overexpression of collagen VI
contributes to cisplatin resistance in ovarian cancer cells. Cancer
Cell. 3:377–386. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Fenouille N, Puissant A, Dufies M, Robert
G, Jacquel A, Ohanna M, Deckert M, Pasquet JM, Mahon FX, Cassuto
JP, et al: Persistent activation of the Fyn/ERK kinase signaling
axis mediates imatinib resistance in chronic myelogenous leukemia
cells through upregulation of intracellular SPARC. Cancer Res.
70:9659–9670. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Pannuru P, Dontula R, Khan AA, Herbert E,
Ozer H, Chetty C and Lakka SS: miR-let-7f-1 regulates SPARC
mediated cisplatin resistance in medulloblastoma cells. Cell
Signal. 26:2193–2201. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Januchowski R, Zawierucha P, Rucinski M
and Zabel M: Microarray-based detection and expression analysis of
extracellular matrix proteins in drugresistant ovarian cancer cell
lines. Oncol Rep. 32:1981–1990. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Lau CK, Yang ZF, Ho DW, Ng MN, Yeoh GC,
Poon RT and Fan ST: An Akt/hypoxia-inducible
factor-1alpha/platelet-derived growth factor-BB autocrine loop
mediates hypoxia-induced chemoresistance in liver cancer cells and
tumorigenic hepatic progenitor cells. Clin Cancer Res.
15:3462–3471. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
de Leeuw N, Dijkhuizen T, Hehir-Kwa JY,
Carter NP, Feuk L, Firth HV, Kuhn RM, Ledbetter DH, Martin CL, van
Ravenswaaij-Arts CM, et al: Diagnostic interpretation of array data
using public databases and internet sources. Hum Mutat. 33:930–940.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Sayed D and Abdellatif M: MicroRNAs in
development and disease. Physiol Rev. 91:827–887. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Lin S and Gregory RI: MicroRNA biogenesis
pathways in cancer. Nat Rev Cancer. 15:321–333. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Blenkiron C and Miska EA: miRNAs in
cancer: Approaches, aetiology, diagnostics and therapy. Hum Mol
Genet. 16:R106–R113. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wu X, Wang H, Lian Y, Chen L, Gu L, Wang J
and Huang Y, Deng M, Gao Z and Huang Y: GTSE1 promotes cell
migration and invasion by regulating EMT in hepatocellular
carcinoma and is associated with poor prognosis. Sci Rep.
7:51292017. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Sun C, Wang FJ, Zhang HG, Xu XZ, Jia RC,
Yao L and Qiao PF: miR-34a mediates oxaliplatin resistance of
colorectal cancer cells by inhibiting macroautophagy via
transforming growth factor-beta/Smad4 pathway. World J
Gastroenterol. 23:1816–1827. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Lanzi C, Zaffaroni N and Cassinelli G:
Targeting heparan sulfate proteoglycans and their modifying enzymes
to enhance anticancer chemotherapy efficacy and overcome drug
resistance. Curr Med Chem. 24:2860–2886. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lee B, Sandhu S and McArthur G: Cell cycle
control as a promising target in melanoma. Curr Opin Oncol.
27:141–150. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Wallerand H, Robert G, Pasticier G, Ravaud
A, Ballanger P, Reiter RE and Ferrière JM: The
epithelial-mesenchymal transition-inducing factor TWIST is an
attractive target in advanced and/or metastatic bladder and
prostate cancers. Urol Oncol. 28:473–479. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Yu FX, Zhao B and Guan KL: Hippo pathway
in organ size control, tissue homeostasis and cancer. Cell.
163:811–828. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Lee JE, Park HS, Lee D, Yoo G, Kim T, Jeon
H, Yeo MK, Lee CS, Moon JY, Jung SS, et al: Hippo pathway effector
YAP inhibition restores the sensitivity of EGFR-TKI in lung
adenocarcinoma having primary or acquired EGFR-TKI resistance.
Biochem Biophys Res Commun. 474:154–160. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Gujral TS and Kirschner MW: Hippo pathway
mediates resistance to cytotoxic drugs. Proc Natl Acad Sci USA.
114:pp. E3729–E3738. 2017; View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Okamoto K, Miyoshi K and Murawaki Y:
miR-29b, miR-205 and miR-221 enhance chemosensitivity to
gemcitabine in HuH28 human cholangiocarcinoma cells. PLoS One.
8:e776232013. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Liu YX, Wang L, Liu WJ, Zhang HT, Xue JH,
Zhang ZW and Gao CJ: MiR-124-3p/B4GALT1 axis plays an important
role in SOCS3-regulated growth and chemo-sensitivity of CML. J
Hematol Oncol. 9:692016. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Chen MJ, Cheng YM, Chen CC, Chen YC and
Shen CJ: MiR-148a and miR-152 reduce tamoxifen resistance in ER+
breast cancer via downregulating ALCAM. Biochem Biophys Res Commun.
483:840–846. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zhong S, Li W, Chen Z, Xu J and Zhao J:
MiR-222 and miR-29a contribute to the drug-resistance of breast
cancer cells. Gene. 531:8–14. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Sui C, Meng F, Li Y and Jiang Y: miR-148b
reverses cisplatin-resistance in non-small cell cancer cells via
negatively regulating DNA (cytosine-5)-methyltransferase 1 (DNMT1)
expression. J Transl Med. 13:1322015. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Tong JL, Zhang CP, Nie F, Xu XT, Zhu MM,
Xiao SD and Ran ZH: MicroRNA 506 regulates expression of PPAR alpha
in hydroxycamptothecin-resistant human colon cancer cells. FEBS
Lett. 585:3560–3568. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Zhang JX, Qian D, Wang FW, Liao DZ, Wei
JH, Tong ZT, Fu J, Huang XX, Liao YJ, Deng HX, et al: MicroRNA-29c
enhances the sensitivities of human nasopharyngeal carcinoma to
cisplatin-based chemotherapy and radiotherapy. Cancer Lett.
329:91–98. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Holohan C, Van Schaeybroeck S, Longley DB
and Johnston PG: Cancer drug resistance: An evolving paradigm. Nat
Rev Cancer. 13:714–726. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Albarello L, Pecciarini L and Doglioni C:
HER2 testing in gastric cancer. Adv Anat Pathol. 18:53–59. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Price-Schiavi SA, Jepson S, Li P, Arango
M, Rudland PS, Yee L and Carraway KL: Rat Muc4 (sialomucin complex)
reduces binding of anti-ErbB2 antibodies to tumor cell surfaces, a
potential mechanism for herceptin resistance. Int J Cancer.
99:783–791. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Castiglioni F, Tagliabue E, Campiglio M,
Pupa SM, Balsari A and Menard S: Role of exon-16-deleted HER2 in
breast carcinomas. Endocr Relat Cancer. 13:221–232. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Serra V, Markman B, Scaltriti M, Eichhorn
PJ, Valero V, Guzman M, Botero ML, Llonch E, Atzori F, Di Cosimo S,
et al: NVP-BEZ235, a dual PI3K/mTOR inhibitor, prevents PI3K
signaling and inhibits the growth of cancer cells with activating
PI3K mutations. Cancer Res. 68:8022–8030. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Nahta R, Yuan LX, Zhang B, Kobayashi R and
Esteva FJ: Insulin-like growth factor-I receptor/human epidermal
growth factor receptor 2 heterodimerization contributes to
trastuzumab resistance of breast cancer cells. Cancer Res.
65:11118–11128. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Boyer J, Allen WL, McLean EG, Wilson PM,
McCulla A, Moore S, Longley DB, Caldas C and Johnston PG:
Pharmacogenomic identification of novel determinants of response to
chemotherapy in colon cancer. Cancer Res. 66:2765–2777. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Denduluri SK, Idowu O, Wang Z, Liao Z, Yan
Z, Mohammed MK, Ye J, Wei Q, Wang J, Zhao L and Luu HH:
Insulin-like growth factor (IGF) signaling in tumorigenesis and the
development of cancer drug resistance. Genes Dis. 2:13–25. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Casa AJ, Dearth RK, Litzenburger BC, Lee
AV and Cui X: The type I insulin-like growth factor receptor
pathway: A key player in cancer therapeutic resistance. Front
Biosci. 13:3273–3287. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
61
|
Sado Y, Kagawa M, Kishiro Y, Sugihara K,
Naito I, Seyer JM, Sugimoto M, Oohashi T and Ninomiya Y:
Establishment by the rat lymph node method of epitope-defined
monoclonal antibodies recognizing the six different alpha chains of
human type IV collagen. Histochem Cell Biol. 104:267–275. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Kalluri R: Basement membranes: Structure,
assembly and role in tumour angiogenesis. Nat Rev Cancer.
3:422–433. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Miyake M, Hori S, Morizawa Y, Tatsumi Y,
Toritsuka M, Ohnishi S, Shimada K, Furuya H, Khadka VS, Deng Y, et
al: Collagen type IV alpha 1 (COL4A1) and collagen type XIII alpha
1 (COL13A1) produced in cancer cells promote tumor budding at the
invasion front in human urothelial carcinoma of the bladder.
Oncotarget. 8:36099–36114. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Jin R, Shen J, Zhang T, Liu Q, Liao C, Ma
H, Li S and Yu Z: The highly expressed COL4A1 genes contributes to
the proliferation and migration of the invasive ductal carcinomas.
Oncotarget. 8:58172–58183. 2017.PubMed/NCBI
|
|
65
|
Sulpice L, Rayar M, Desille M, Turlin B,
Fautrel A, Boucher E, Llamas-Gutierrez F, Meunier B, Boudjema K,
Clément B and Coulouarn C: Molecular profiling of stroma identifies
osteopontin as an independent predictor of poor prognosis in
intrahepatic cholangiocarcinoma. Hepatology. 58:1992–2000. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Rahman M, Chan AP, Tang MJ and Tai IT:
Correction: A peptide of SPARC interferes with the interaction
between caspase8 and Bcl2 to resensitize chemoresistant tumors and
enhance their regression in vivo. PLoS One. 10:e01272262015.
View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Yang Z, Guo L, Liu D, Sun L, Chen H, Deng
Q, Liu Y, Yu M, Ma Y, Guo N and Shi M: Acquisition of resistance to
trastuzumab in gastric cancer cells is associated with activation
of IL-6/STAT3/Jagged-1/Notch positive feedback loop. Oncotarget.
6:5072–5087. 2015.PubMed/NCBI
|
|
68
|
Wang S, Huang J, Lyu H, Cai B, Yang X, Li
F, Tan J, Edgerton SM, Thor AD, Lee CK and Liu B: Therapeutic
targeting of erbB3 with MM-121/SAR256212 enhances antitumor
activity of paclitaxel against erbB2-overexpressing breast cancer.
Breast Cancer Res. 15:R1012013. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Vora SR, Juric D, Kim N, Mino-Kenudson M,
Huynh T, Costa C, Lockerman EL, Pollack SF, Liu M, Li X, et al: CDK
4/6 inhibitors sensitize PIK3CA mutant breast cancer to PI3K
inhibitors. Cancer Cell. 26:136–149. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Hudis C, Swanton C, Janjigian YY,
Mino-Kenudson M, Huynh T, Costa C, Lockerman EL, Pollack SF, Liu M,
Li X, et al: A phase 1 study evaluating the combination of an
allosteric AKT inhibitor (MK-2206) and trastuzumab in patients with
HER2-positive solid tumors. Breast Cancer Res. 15:R1102013.
View Article : Google Scholar : PubMed/NCBI
|