Introduction
Tumor tissues are different from normal tissues in
many aspects, mainly due to differences in vasculature (1). In tumor tissues, immature vasculature
forms a microenvironment of hypoxia and malnutrition (2). Tumor cells change their metabolism to
be able to survive and proliferate in these harsh conditions. High
rates of glucose consumption and lactate production, even in the
presence of oxygen, have been observed in various tumor cells; this
phenomenon is known as the Warburg effect (3). Since Warburg's study, the metabolism
of normal and tumor tissues has been studied, and differences are
expected to serve as a basis for new therapeutic targets for
refractory cancers.
Folate plays a pivotal role in the one-carbon
metabolism needed for methylation reactions, nucleotide synthesis,
and DNA repair (4,5). Methylation of DNA and histones is a
well known epigenetic mechanism, and it is the most common
molecular alteration in cancer cells (6). Furthermore, cells with high rates of
proliferation, like tumor cells and embryogenic cells, require the
synthesis of abundant proteins, lipids and nucleotides (7). Due to these features, it was proposed
that folate may be associated with carcinogenesis (8). Jain et al reported that
expression of the mitochondrial glycine biosynthetic pathway was
significantly correlated with cancer cell proliferation, based on
the consumption and release of 219 metabolites from NCI-60 cancer
cell lines. Moreover, high expression levels of three mitochondrial
glycine (also folate) metabolism enzymes, serine hydroxymethyl
transferase (SHMT2), methylenetetrahydrofolate dehydrogenase
(MTHFD2) and tetrahydrofolate synthase (MTHFD1L), were associated
with greater mortality in patients with breast cancer (9). The expression of mitochondrial folate
metabolic enzymes was reported to be a determinant of the response
to methotrexate, and the dihydrofolate reductase inhibitor is used
most often as an anticancer drug in clinical settings (10). Although previous studies have shown
that mitochondrial folate metabolic enzymes correlated with some
types of solid tumors, their importance in colorectal cancer
remains unclear (11).
The aim of the present study was to assess the
importance of mitochondrial folate metabolic enzymes in human
colorectal cancer. Our results indicated that expression levels of
mitochondrial folate metabolic enzymes, including SHMT2, MTHFD2 and
aldehyde dehydrogenase 1 family member L2 (ALDH1L2) were
upregulated in human colorectal tumor tissues compared to normal
tissues. Moreover, immunohistochemical analysis showed that the
combined high expression of SHMT2, MTHFD2 and ALDH1L2 (triple high
expression) was more highly associated with a poor prognosis than
the individual expression levels. Moreover, a multivariate analysis
indicated that triple high expression was independently associated
with recurrence-free survival. We concluded that mitochondrial
folate metabolic enzymes could provide a potential therapeutic
strategy against colorectal cancer.
Materials and methods
The Cancer Genome Atlas tissue
samples
The colorectal adenocarcinoma data were generated by
the Cancer Genome Atlas (TCGA) Research Network: http://cancergenome.nih.gov/. Messenger RNA expression
and DNA methylation were compared between normal colorectal tissue
and cancer with Wanderer software (12). We analyzed exon transcript values
(log2 RPKM) of chr12:57624586-57624783 for SHMT2,
chr2:74425690-74425869 for MTHFD2 and chr12:105413564-105418257 for
ALDH1L2. The mean methylation β values of all the
HumanMethylation450 probes inside the genes were evaluated. The
mutation frequency was assessed in 220 colorectal tumor samples
with sequencing data through the cBioPortal for Cancer Genomics
(http://cbioportal.org) (13,14).
Prognosis of patients with colorectal
cancer
The published GSE17536 database (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE17536),
which included microarray data for 177 patients, was used to screen
for genes related to the prognosis of patients with colorectal
cancer, as described previously (15). The patient data were divided into
two groups (low- and high-expression groups) based on the mean
levels of SHMT2, MTHFD2 and ALDH1L2 expression. Then, we evaluated
the relevance of these gene expression levels to overall
survival.
Tissue samples
Colorectal cancer tissue samples were obtained from
117 consecutive patients that underwent surgery at the Osaka
University Hospital, between 2006 and 2009. Written informed
consent for the use of resected specimens was provided by all
patients. The present study was approved by the Institutional
Review Board (the approved protocol numbers were #08226 and #213).
None of the patients had received chemotherapy or radiotherapy
before surgery. A total of 108 patients underwent curative surgery
and 9 patients underwent palliative surgery. Samples were fixed in
buffered formalin at 4°C overnight, processed through graded
ethanol solutions, and embedded in paraffin. Medical and pathology
studies were reviewed to collect information regarding various
clinicopathological parameters, including age, gender,
carcinoembryonic antigen (CEA), carbohydrate antigen 19-9
(CA-19-9), histological type, depth of tumor invasion, lymph node
metastasis, distant metastasis, pTNM stage (according to the TNM
Classification System of Malignant Tumors, Seventh Edition)
(16), venous invasion and
lymphatic invasion.
Immunohistochemical analysis
Immunohistochemical analysis was performed with the
Vectastain Elite ABC kit (Vector Laboratories, Burlingame, CA, USA)
as previously described (17).
Briefly, tissue sections (3.5 µm thick) were sliced from
paraffin-embedded tissue blocks. After deparaffinization in xylene
and dehydration in graded ethanol solutions, tissue sections were
heated at 110°C for 15 min in 10 mM citrate buffer (pH 6.0),
followed by incubation with 0.3% hydrogen peroxide for 20 min to
block endogenous peroxidase activity. Subsequently, the sections
were incubated with SHMT2 antibody (ab64417; 1:300 dilution; rabbit
polyclonal), MTHFD2 antibody (ab176016; 1:100 dilution; rabbit
polyclonal) and ALDH1L2 antibody (ab170176; 1:200 dilution; rabbit
polyclonal) (all from Abcam, Cambridge, MA, USA) at 4°C overnight.
Then, sections were incubated with a biotinylated secondary
antibody solution for 30 min, followed by incubation with ABC
reagent for 30 min. After development with diaminobenzidine,
sections were counterstained with hematoxylin. As a negative
control, phosphate-buffered saline was used instead of the
antibodies. Representative positive colorectal cancer tissues were
used as a positive control. The analyses were performed
independently, in a blinded manner, by two pathologists.
Evaluation of immunostaining
The staining score was defined as the intensity of
staining that covered the largest region. We first performed
immunostaining for 20 colorectal cancer samples randomly chosen to
decide the concentration of antibodies and samples that could be
applied for a positive control. The samples showing an average
staining were used as a positive control. Unstained tissues were
assigned to ‘negative staining (−)’. The tissues stained weaker
than the positive control but stronger than the unstained tissue
were categorized into ‘weak staining (+)’ and the tissues stained
stronger than the positive control were categorized into ‘strong
staining (++)’. To evaluate associations between expression levels
and clinicopathological features or survival, the samples were
divided into two groups, as follows: low expression (−) and high
expression (+ and ++) for SHMT2 and ALDH1L2, or low expression (−
and +) and high expression (++) for MTHFD2.
Statistical analysis
Data were expressed as the mean ± standard deviation
(SD). We performed statistical analyses with JMP Pro 10 software
(SAS Institute, Cary, NC, USA). Statistically significant
differences were determined with Fisher's exact test.
Recurrence-free survival and overall survival were estimated with
the Kaplan-Meier method, and the statistical significance was
evaluated with the log-rank test. Variables identified as
significant in univariate analyses were entered into the Cox
proportional hazards regression models. A P-value <0.05 was
taken to indicate a significant difference.
Results
Upregulation of enzymes associated
with mitochondrial folate metabolic pathway in colorectal
cancer
Reportedly, enzymes associated with the
mitochondrial folate metabolic pathway show upregulated expression
in cancer (18). To confirm
expression in normal colorectal tissue and colorectal cancer, we
analyzed the data from the TCGA Research Network. We found that the
mRNA expression levels of SHMT2, MTHFD2 and ALDH1L2 associated with
mitochondrial folate metabolism were higher in colorectal cancer
than in normal colorectal tissue (P<0.001, P<0.001 and
P=0.681, respectively) (Fig. 1A).
Several studies have indicated that gene mutations change the
activity and function of metabolic enzymes. For example, mutations
in isocitrate dehydrogenase 1 resulted in a new enzymatic function:
catalysis of the NADPH-dependent reduction of α-ketoglutarate to
R(−)-2-hydroxyglutarate (19). A
mutation in mitochondrial aldehyde dehydrogenase 2, which is
essential for alcohol detoxification, induced structural
alterations that changed its enzymatic activity (20). Therefore, we assessed the importance
of mutations in SHMT2, MTHFD2 and ALDH1L2 in colorectal tumor
samples. The mutation frequency was <2% (Fig. 1B). Methylation of these genes was
also evaluated, based on the data from the TCGA Research Network.
The methylation frequency was similar in normal and tumor
colorectal tissues (Fig. 1C). These
findings showed that expression of SHMT2, MTHFD2 and ALDH1L2 was
associated with the physiological characteristics of colorectal
cancer.
Relevance of SHMT2, MTHFD2 and ALDH1L2
mRNA expression in colorectal cancer with respect to prognosis
Previous studies showed a correlation between the
expression levels of mitochondrial folate metabolism enzymes and
the prognosis of some solid tumors. Therefore, we analyzed
microarray data from patients with colorectal cancer in the
GSE17536 database (11,21). The overall survival rate in patients
with high expression levels of MTHFD2 and ALDH1L2 was significantly
lower than in patients with low expression levels of MTHFD2 and
ALDH1L2 (P<0.001 and P=0.027, respectively) (Fig. 2). We also found a trend for lower
overall survival rate in patients with high expression of SHMT2
compared to patients with low expression of SHMT2 (P=0.353).
Immunohistochemistry for SHMT2, MTHFD2
and ALDH1L2 in colorectal cancer
To study the importance of SHMT2, MTHFD2 and ALDH1L2
further, we collected colorectal cancer samples and analyzed the
expression of SHMT2, MTHFD2 and ALDH1L2 at protein level with
immunohistochemistry. The median follow-up duration of patients
with colorectal cancer was 4.9 years (range; 0.2–6.6 years). SHMT2,
MTHFD2 and ALDH1L2 expression was observed in the glandular cells
of colorectal cancer (Fig. 3).
Fig. 3B, E and H shows the samples
that were used as a positive control. Subsequently, associations
between the expression levels of these genes and prognosis were
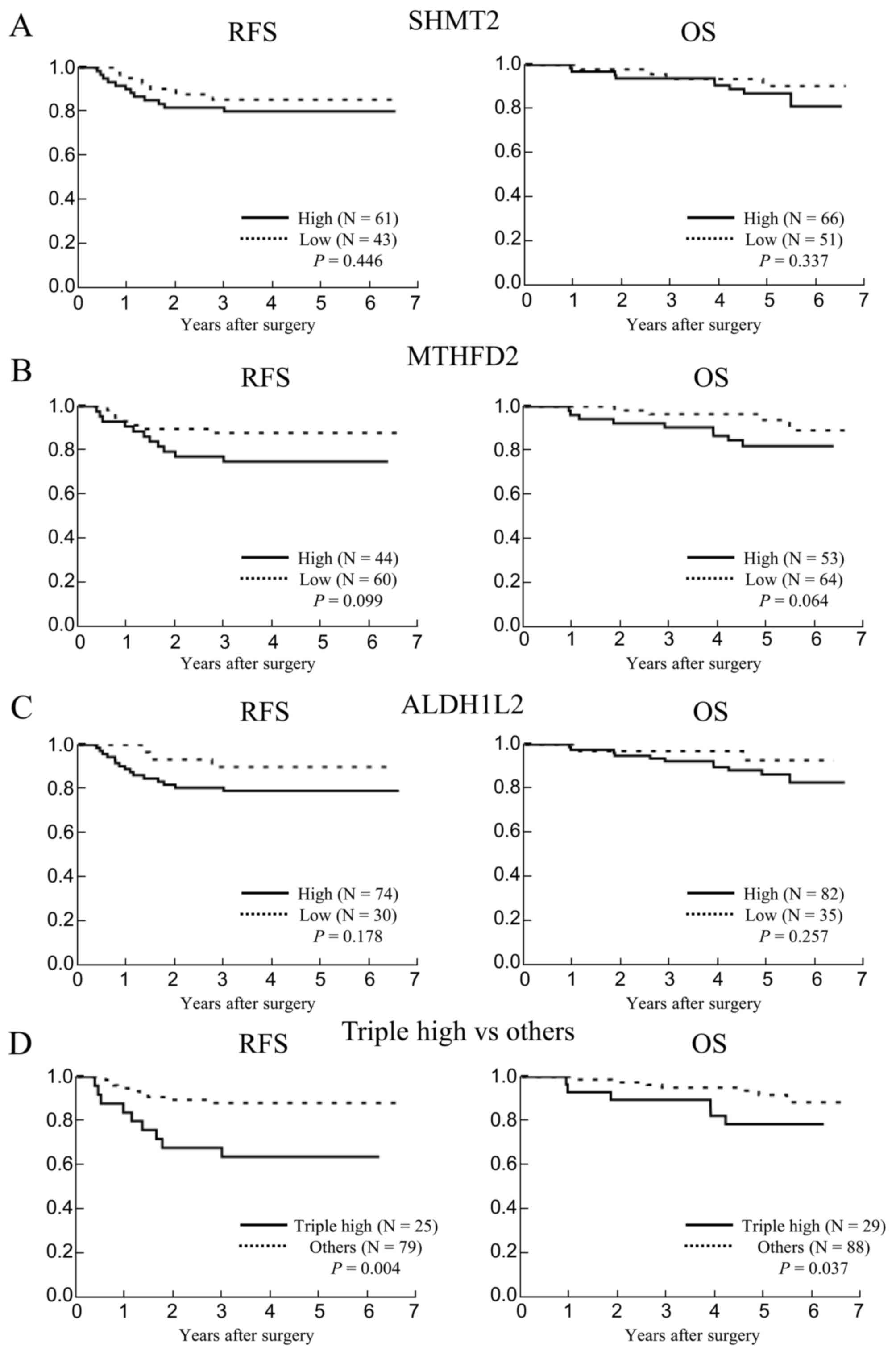
evaluated. Rates of recurrence-free and overall survival in
patients with high expression of SHMT2, MTHFD2 and ALDH1L2 tended
to be lower than in patients with low expression of SHMT2, MTHFD2
and ALDH1L2, but these differences were not significant (P=0.446
and P=0.337, P=0.099 and P=0.064, P=0.178 and P=0.257,
respectively) (Fig. 4A-C). Notably,
high expression of SHMT2, MTHFD2 and ALDH1L2 (triple high) was more
highly associated with poor prognosis than increased expression
levels of each individual gene (Fig.
4D). These results indicated that malignancy in colorectal
cancer was influenced by activation of the mitochondrial folate
metabolic pathway.
 | Figure 3.Immunohistochemical detection of
protein expression. (A-C) SHMT2, (D-F) MTHFD2 and (G-I) ALDH1L2
expression levels varied in colorectal cancer tissues. Examples are
shown for (A, D and G) negative staining (−), (B, E and H) weak
staining (+), and (C, F and I) strong staining (++). Scale bar
indicates, 100 µm. SHMT2, serine hydroxymethyl transferase; MTHFD2,
methylenetetrahydrofolate dehydrogenase; ALDH1L2, aldehyde
dehydrogenase 1 family member L2. |
Impact of triple high expression on
recurrence-free survival
Based on our findings that the expression levels of
enzymes associated with the mitochondrial folate metabolic pathway
could influence colorectal cancer malignancy, we assessed the
relevance of the expression of three mitochondrial folate enzymes
to clinicopathological features. There were no significant
association between the individual expression of MTHFD2 and ALDH1L2
and patient and tumor characteristics (data not shown). In regards
to SHMT2, its expression was negatively associated with lymph node
metastasis, lymphatic invasion and stage (P=0.015, P=0.021 and
P=0.010, respectively, data not shown). Interestingly, triple high
expression did not significantly correlate with patient background
characteristics, including gender, age, CEA and CA19-9; or with
tumor characteristics, including histological type, depth of tumor
invasion, lymph node metastasis, distant metastasis, lymphatic
invasion, venous invasion and stage (Table I). To determine associations between
triple high expression and the most important factors (P<0.100),
we included depth of tumor invasion, lymph node metastasis,
lymphatic invasion, venous invasion, stage, and SHMT2, MTHFD2 and
ALDH1L2 expression in a multivariate analysis for recurrence-free
survival (Table II). This
multivariate analysis showed that triple high expression was only
independently associated with recurrence-free survival
(P=0.017).
 | Table I.Immunohistochemical detection results
for SHMT2, MTHFD2 and ALDH1L2 in colorectal cancer. |
Table I.
Immunohistochemical detection results
for SHMT2, MTHFD2 and ALDH1L2 in colorectal cancer.
| Clinicopathological
factors | Classification | N | Triple high (N) | Others (N) | P-value |
|---|
| Patient
background |
|
|
|
|
|
|
Gender | Male | 71 | 17 | 54 | 0.829 |
|
| Female | 46 | 12 | 34 |
|
| Age | <65 | 57 | 16 | 41 | 0.522 |
|
| ≥65 | 60 | 13 | 47 |
|
| CEA | <3.1 | 66 | 16 | 50 | 1.000 |
|
| ≥3.1 | 51 | 13 | 50 |
|
|
CA19-9 | <40 | 101 | 23 | 78 | 0.221 |
|
| ≥40 | 16 | 6 | 10 |
|
| Tumor
characteristics |
|
|
|
|
|
|
Histological typea | tub1, tub2, pap | 109 | 26 | 83 | 0.407 |
|
| por, muc | 8 | 3 | 5 |
|
| Depth of
tumor invasion | Tis, T1, T2 | 49 | 11 | 38 | 0.669 |
|
| T3, T4 | 68 | 18 | 50 |
|
| Lymph
node metastasis | Positive | 49 | 13 | 36 | 0.829 |
|
| Negative | 68 | 16 | 52 |
|
| Distant
metastasis | Positive | 13 | 4 | 9 | 0.734 |
|
| Negative | 104 | 25 | 79 |
|
|
Lymphatic invasion | Positive | 85 | 22 | 63 | 0.811 |
|
| Negative | 32 | 7 | 25 |
|
| Venous
invasion | Positive | 29 | 8 | 21 | 0.805 |
|
| Negative | 88 | 21 | 67 |
|
|
Stage | 0, I, II | 62 | 14 | 48 | 0.669 |
|
| III, IV | 55 | 15 | 40 |
|
 | Table II.Univariate and multivariate analyses
of recurrence-free survival. |
Table II.
Univariate and multivariate analyses
of recurrence-free survival.
|
|
|
| Univariate
analysis | Multivariate
analysis |
|---|
|
|
|
|
|
|
|---|
| Clinicopathological
factors | Classification | N | HRa | P-value | HRa | P-value |
|---|
| Patient
background |
|
|
|
|
|
|
|
Gender | Male | 65 | 1.69 | 0.305 |
|
|
|
|
Female | 39 | (0.64–5.25) |
|
|
|
|
Age | <65 | 47 | 1.51 | 0.384 |
|
|
|
| ≥65 | 57 | (0.59–3.95) |
|
|
|
|
CEA | <3.1 | 62 | 2.07 | 0.123 |
|
|
|
| ≥3.1 | 42 | (0.82–5.43) |
|
|
|
|
CA19-9 | <40 | 93 | 1.26 | 0.766 |
|
|
|
| ≥40 | 11 | (0.20–4.43) |
|
|
|
| Tumor
characteristics |
|
|
|
|
|
|
|
Histological typeb | tub1, tub2,
pap | 97 | 1.26 | 0.817 |
|
| por, muc | 7 | (0.26–22.66) |
|
|
|
| Depth
of tumor invasion | Tis, T1, T2 | 49 | 3.59 | 0.013 | 1.97 | 0.297 |
|
| T3, T4 | 55 | (1.29–12.68) |
| (0.57–8.25) |
|
| Lymph
node metastasis | Positive | 38 | 3.17 | 0.015 | 1.03 | 0.977 |
|
| Negative | 66 | (1.25–8.63) |
| (0.17–19.66) |
|
|
Lymphatic invasion | Positive | 73 | 3.88 | 0.033 | 1.51 | 0.629 |
|
| Negative | 31 | (1.10–24.55) |
| (0.30–11.17) |
|
| Venous
invasion | Positive | 22 | 3.14 | 0.027 | 1.80 | 0.291 |
|
| Negative | 82 | (1.15–7.99) |
| (0.60–5.32) |
|
|
Stage | 0, I, II | 62 | 3.43 | 0.011 | 1.66 | 0.672 |
|
| III | 42 | (1.33–9.86) |
| (0.08–11.92) |
|
| SHMT2,
MTHFD2 | Triple high | 25 | 3.55 | 0.009 | 3.21 | 0.017 |
| and
ALDH1L2c | Others | 79 | (1.39–9.11) |
| (1.24–8.31) |
|
Discussion
We found that expression levels of the mitochondrial
folate metabolic enzymes, SHMT2, MTHFD2 and ALDH1L2, were
upregulated in colorectal tumor tissues and that triple high
expression was associated with a poor prognosis. The folate
metabolic pathway comprises a cycle mediated by SHMT2, MTHFD2 and
ALDH1L2 (Fig. 5). Our results
indicated that activation of the mitochondrial folate metabolic
pathway without stagnation could offer advantages for survival and
proliferation in colorectal cancer. SHMT2 negatively regulates the
activity of pyruvate kinase, and subsequently, it decreases carbon
flux into the TCA cycle; this activity provides a survival benefit
for glioma cells under hypoxic conditions (22). An analysis of 515 tumor-derived cell
lines showed that patients with significant upregulation of folate
metabolism genes tended to benefit from treatment with methotrexate
(10). In addition to its
importance in methylation reactions, nucleotide synthesis, and DNA
repair, the mitochondrial folate metabolic pathway also influences
the biological characteristics of tumors, which contribute to tumor
malignancy. Interfering with activation of the mitochondrial folate
metabolic pathway could be a potential therapeutic strategy for
treating refractory colorectal cancer.
 | Figure 5.Schematic overview of the folate
metabolic pathway mediated by SHMT2, MTHFD2 and ALDH1L2. (Left)
Cytosolic folate metabolism; (right) mitochondrial folate
metabolism. Arrows that cross the line indicate metabolites that
can travel between compartments. ALDH1L2, aldehyde dehydrogenase 1
family member L2; CH2-THF, methylenetetrahydrofolate; CHO-THF,
formyl-tetrahydrofolate; DHFR, dihydrofolate reductase; Gly,
glycine; MTHFD1, methylenetetrahydrofolate dehydrogenase 1;
MTHFD1L, methylenetetrahydrofolate dehydrogenase 1-like; MTHFD2,
methylenetetrahydrofolate dehydrogenase 2; HCHO, formaldehyde; Ser,
serine; SHMT1, serine hydroxymethyltransferase 1; SHMT2, serine
hydroxymethyltransferase 2; THF, tetrahydrofolate. |
Reportedly, only mitochondrial, not cytosolic,
folate enzymes were significantly associated with rapid cancer cell
proliferation and prognosis in breast cancer (9). To elucidate the importance of folate
enzymes, including dihydrofolate reductase, MTHFD1 and SHMT1, the
prognosis of patients with colorectal cancer was assessed based on
data from the published GSE17536 database. Rates of disease-free
and overall survival were not significantly different between
groups with high and low dihydrofolate reductase expression
(P=0.290 and P=0.906, respectively) or between groups with high and
low MTHFD1 expression (P=0.252 and P=0.279, respectively) (data not
shown). Conversely, low SHMT1 expression was significantly
associated with a short overall survival rate (P=0.033) (data not
shown). These data supported the hypothesis that mitochondrial
folate metabolism may be more important than cytosolic folate
metabolism in human colorectal cancer. The detailed function and
mechanism of cytosolic folate metabolism in tumor cells remain
unclear. Additional studies are needed to address whether the
function of cytosolic folate enzymes differs from that of
mitochondrial folate enzymes.
Few studies have described the regulation of
mitochondrial folate enzymes. Reportedly, mammalian target of
rapamycin complex 1 (mTORC1) positively regulates the expression of
MTHFD2 via activating transcription factor 4 (23). It is well known that activation of
mTORC1 is correlated with nutrient, energy and redox sensing
processes (24). Therefore, the
expression of mitochondrial folate enzymes may also be regulated by
environmental factors, including oxidative stress, growth factors
and amino acids. mTORC1 controls several anabolic metabolism
pathways required for cell proliferation and growth. This function
is thought to depend partially on mitochondrial folate metabolism,
which is associated with nucleotide synthesis, because nucleotides
are necessary for cell proliferation and growth.
The formyl-tetrahydrofolate synthase activity of
MTHFD1L was reported to play an important role in the proliferation
of breast cancer cells (9). MTHFD1L
also participates in the cycle of the mitochondrial folate pathway
(Fig. 5). We assessed the
relationship between MTHFD1L expression and prognosis of patients
with colorectal cancer, based on data from the GSE17536 database.
However, we found that MTHFD1L expression was not significantly
related to the prognosis of patients (data not shown). Although
ALDH1L2 and MTHFD1L catalyze the same reaction, which converts
formyl-tetrahydrofolate to tetrahydrofolate, they may have
different physiological effects in different types of cancer.
Moreover, in different cancer types, the functions of metabolic
enzymes may be diverse. Studying the mechanism of metabolic enzymes
in different cancer types will promote the development of specific
cancer therapies.
Acknowledgements
This study was supported in part by a Grant-in-Aid
for Scientific Research from the Ministry of Education, Culture,
Sports, Science and Technology; a Grant-in-Aid from the Third
Comprehensive 10-year Strategy for Cancer Control, Ministry of
Health, Labor and Welfare; a grant from the Kobayashi Cancer
Research Foundation; a grant from the Princess Takamatsu Cancer
Research Fund, Japan; a grant from the National Institute of
Biomedical Innovation; and a grant from the Osaka University Drug
Discovery Funds. A.H. is a research fellow of the Japan Society for
the Promotion of Science. Partial support was received from Taiho
Pharmaceutical Co., Ltd. (H.I., J.K. and M.M.), Chugai Co., Ltd.,
Yakult Honsha Co., Ltd., Merck Co., Ltd., Takeda Science Foundation
and Takeda Medical Research Foundation (M.K., M.M., N.N. and H.I.)
through institutional endowments.
References
|
1
|
Brown JM and Giaccia AJ: The unique
physiology of solid tumors: Opportunities (and problems) for cancer
therapy. Cancer Res. 58:1408–1416. 1998.PubMed/NCBI
|
|
2
|
Daşu A, Toma-Daşu I and Karlsson M:
Theoretical simulation of tumour oxygenation and results from acute
and chronic hypoxia. Phys Med Biol. 48:2829–2842. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Warburg O: On the origin of cancer cells.
Science. 123:309–314. 1956. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Choi SW and Mason JB: Folate status:
Effects on pathways of colorectal carcinogenesis. J Nutr.
132:(Suppl). 2413S–2418S. 2002.PubMed/NCBI
|
|
5
|
Duthie SJ and Hawdon A: DNA instability
(strand breakage, uracil misincorporation, and defective repair) is
increased by folic acid depletion in human lymphocytes in vitro.
FASEB J. 12:1491–1497. 1998.PubMed/NCBI
|
|
6
|
Widschwendter M and Jones PA: DNA
methylation and breast carcinogenesis. Oncogene. 21:5462–5482.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Pike ST, Rajendra R, Artzt K and Appling
DR: Mitochondrial C1-tetrahydrofolate synthase (MTHFD1L) supports
the flow of mitochondrial one-carbon units into the methyl cycle in
embryos. J Biol Chem. 285:4612–4620. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ventrella-Lucente LF, Unnikrishnan A,
Pilling AB, Patel HV, Kushwaha D, Dombkowski AA, Schmelz EM,
Cabelof DC and Heydari AR: Folate deficiency provides protection
against colon carcinogenesis in DNA polymerase beta
haploinsufficient mice. J Biol Chem. 285:19246–19258. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Jain M, Nilsson R, Sharma S, Madhusudhan
N, Kitami T, Souza AL, Kafri R, Kirschner MW, Clish CB and Mootha
VK: Metabolite profiling identifies a key role for glycine in rapid
cancer cell proliferation. Science. 336:1040–1044. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Vazquez A, Tedeschi PM and Bertino JR:
Overexpression of the mitochondrial folate and glycine-serine
pathway: A new determinant of methotrexate selectivity in tumors.
Cancer Res. 73:478–482. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Liu F, Liu Y, He C, Tao L, He X, Song H
and Zhang G: Increased MTHFD2 expression is associated with poor
prognosis in breast cancer. Tumour Biol. 35:8685–8690. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Díez-Villanueva A, Mallona I and Peinado
MA: Wanderer, an interactive viewer to explore DNA methylation and
gene expression data in human cancer. Epigenetics Chromatin.
8:222015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:pl12013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Koseki J, Colvin H, Fukusumi T, Nishida N,
Konno M, Kawamoto K, Tsunekuni K, Matsui H, Doki Y, Mori M, et al:
Mathematical analysis predicts imbalanced IDH1/2 expression
associates with 2-HG-inactivating β-oxygenation pathway in
colorectal cancer. Int J Oncol. 46:1181–1191. 2015.PubMed/NCBI
|
|
16
|
Edge S, Byrd DR, Compton CC, Fritz AG,
Greene FL and Trotti A: AJCC Cancer Staging Handbook: From the AJCC
Cancer Staging Manual. Springer; New York: 2011
|
|
17
|
Miyo M, Yamamoto H, Konno M, Colvin H,
Nishida N, Koseki J, Kawamoto K, Ogawa H, Hamabe A, Uemura M, et
al: Tumour-suppressive function of SIRT4 in human colorectal
cancer. Br J Cancer. 113:492–499. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Nilsson R, Jain M, Madhusudhan N, Sheppard
NG, Strittmatter L, Kampf C, Huang J, Asplund A and Mootha VK:
Metabolic enzyme expression highlights a key role for MTHFD2 and
the mitochondrial folate pathway in cancer. Nat Commun. 5:31282014.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Dang L, White DW, Gross S, Bennett BD,
Bittinger MA, Driggers EM, Fantin VR, Jang HG, Jin S, Keenan MC, et
al: Cancer-associated IDH1 mutations produce 2-hydroxyglutarate.
Nature. 462:739–744. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Jin S, Chen J, Chen L, Histen G, Lin Z,
Gross S, Hixon J, Chen Y, Kung C, Chen Y, et al: ALDH2(E487K)
mutation increases protein turnover and promotes murine
hepatocarcinogenesis. Proc Natl Acad Sci USA. 112:9088–9093. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lehtinen L, Ketola K, Mäkelä R, Mpindi JP,
Viitala M, Kallioniemi O and Iljin K: High-throughput RNAi
screening for novel modulators of vimentin expression identifies
MTHFD2 as a regulator of breast cancer cell migration and invasion.
Oncotarget. 4:48–63. 2013.PubMed/NCBI
|
|
22
|
Kim D, Fiske BP, Birsoy K, Freinkman E,
Kami K, Possemato RL, Chudnovsky Y, Pacold ME, Chen WW, Cantor JR,
et al: SHMT2 drives glioma cell survival in ischaemia but imposes a
dependence on glycine clearance. Nature. 520:363–367. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Ben-Sahra I, Hoxhaj G, Ricoult SJ, Asara
JM and Manning BD: mTORC1 induces purine synthesis through control
of the mitochondrial tetrahydrofolate cycle. Science. 351:728–733.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Csibi A, Fendt SM, Li C, Poulogiannis G,
Choo AY, Chapski DJ, Jeong SM, Dempsey JM, Parkhitko A, Morrison T,
et al: The mTORC1 pathway stimulates glutamine metabolism and cell
proliferation by repressing SIRT4. Cell. 153:840–854. 2013.
View Article : Google Scholar : PubMed/NCBI
|