Introduction
Under conditions of severe liver injury, or
inhibited hepatocyte proliferation, undifferentiated hepatic
progenitor/stem cells are activated to mediate liver regeneration
(1,2). Progenitor cells are activated via
various signaling and growth factors (2). In humans, proliferating progenitor
cells are observed during chronic liver disease, and the degree of
hepatic progenitor cell activation corresponds with the severity of
liver fibrosis and inflammation (3–6).
Therefore, it is necessary to understand the mechanisms underlying
progenitor activation and differentiation during chronic liver
disease.
Connective tissue growth factor (CTGF) is a
cysteine-rich protein secreted via a 37-amino acid signal sequence.
CTGF is an important mediator of liver fibrosis and its
overexpression correlates with the progression of hepatic fibrosis
(7–9). CTGF also plays an important role in
progenitor activation during liver regeneration triggered by
2-N-acetylaminofluorene/partial hepatectomy (2-AAF/PHx) (10–12). Previous studies conducted in our
laboratory showed that transforming growth factor-β1 (TGF-β1)
inhibits the viability of rat progenitors in a dose-dependent
manner and alters their phenotype (13). Being a downstream mediator of
TGF-β1, CTGF may prove to be an important factor, governing cell
differentiation and regeneration during tissue repair.
To investigate this possibility, we examined the
influence of CTGF on the differentiation of hepatic progenitor
cells in vitro, using WB-F344 cells as a model for rat
hepatic progenitor cells (14,15). WB-F344 cells are characterized by
a high nuclear to cytoplasmic ratio, and they express markers
characteristic of both hepatocytes [α-fetoprotein (AFP) and albumin
(ALB)] and cholangiocytes [cytokeratin-19 (CK-19)]. These cells
retain the ability to differentiate into either hepatocytes or
cholangiocytes under the appropriate conditions (14,16,17).
Materials and methods
Cell culture
WB-F344 cells were plated in Dulbecco’s modified
Eagle’s medium (DMEM) supplemented with 10% (v/v) fetal bovine
serum (FBS) (both from Gibco, Grand Island, NY, USA) (complete
medium). When the cells were 80–90% confluent, they were detached
using trypsin and reseeded at a density of 1×105
cells/ml in 100-mm dishes. WB-F344 cells were cultured overnight in
serum-free medium prior to the experiments. Cells were either
stimulated, or not stimulated, with different concentrations of
CTGF (1 or 5 ng/ml; PeproTech, Rocky Hill, NJ, USA), a CTGF
inhibitor (WP631, 0.01 nM; Alexis Biochemical, San Diego, CA, USA)
and cultured for 24 h. Total mRNA and proteins were then
extracted.
Cell viability
WB-F344 cells (6,000 cells/well) were seeded into
96-well plates and treated with CTGF or WP631 as described above. A
cell viability assay was performed using
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT).
Three duplicate wells were used for each sample. Twenty microliters
of MTT (5 mg/ml) was added to each well and incubated for 4 h. The
supernatant was then removed and 150 μl DMSO was added to
dissolve the formazan crystals. Plates were then transferred to a
plate reader and the absorbance was measured at 450 nm.
Quantitative real-time PCR analysis of
differentiation marker expression
Quantitative real-time PCR was performed using an
ABI PRISM 7300 Sequence Detection (Applied Biosystems, Foster City,
CA, USA) and the ABI Power SYBR-Green PCR Master Mix kit (Applied
Biosystems, Warrington, UK). The primers used for amplification
(Table I) were designed to be
between 60 and 150 bp in length according to the PE Applied
Biosystems guidelines for the comparative CT method. A
final primer concentration of 300 nM was assessed as the optimum
for use under all conditions to ensure that no nonspecific
amplification occurred in the sample wells. RT was carried out for
2 min at 5°C before incubation for 10 min at 95°C to inactivate the
reverse transcriptase, which otherwise interferes with the DNA
polymerase. Forty cycles at 95°C for 15 sec followed by 60°C for 60
sec were performed, and the fluorescence was measured at the end of
each extension cycle. A SYBR-Green dissociation curve was
constructed at the end of the reaction to ensure specificity; the
temperature was increased from 65 to 95°C in 0.1°C/sec increments,
and the fluorescence signals were measured and plotted against the
temperature. The amount of target, normalized to an endogenous
reference [glyceraldehyde 3-phosphate dehydrogenase (GAPDH)] and
relative to a calibrator (WB-F344 cells cultured untreated), was
given by the formula 2−ΔΔCt, as determined by the ABI
PRISM 7300 System Software’s (Applied Biosystems) built-in
algorithm using an adaptive baseline to determine the Ct. The
relative amounts were expressed as the means ± SD from three
independent experiments.
 | Table I.Primers used for reverse transcription
polymerase chain reaction. |
Table I.
Primers used for reverse transcription
polymerase chain reaction.
| Gene | Primer sequences | Product size
(bp) | Accession no. | Annealing
temperature |
|---|
| CTGF | F:
5′-ACCATGCTCGCCTCCGTC-3′ | 1050 | NM_022266 | 58°C |
| R:
5′-GCTTTACGCCATGTCTCCATAC-3′ |
| GAPDH | F:
5′-ATGGCCTTCCGTGTTCCTAC-3′ | 318 | BC059110 | 58°C |
| R:
5′-TTACTCCTTGGAGGCCATGT-3′ |
Differentiation marker expression
assessed by western blotting
Total proteins were extracted using a cellular
protein extraction solution, and protein concentrations were
measured using the BCA protein assay (Pierce, Rockford, IL, USA).
After boiling for 10 min, the lysates were separated on 12%
SDS-PAGE gels. Proteins were transferred to nitrocellulose
membranes (Amersham Biosciences, Piscataway, NJ, USA), which were
blocked in TTBS containing 5% non-fat dried milk for 2 h at room
temperature. The membranes were then incubated with anti-ALB,
anti-AFP, anti-CK-19 (all from R&D System, USA) and anti-CTGF
(Abcam, Hong Kong, China) primary antibodies (Table II) at 4°C overnight. After three
washes, the membranes were incubated with the corresponding
horseradish peroxidase-conjugated rabbit anti-mouse or goat
anti-rabbit secondary antibodies (diluted 1:8,000 and 1:9,000,
respectively) for 1 h. The membranes were then subjected to
chemiluminescence (Pierce Chemical Co.) detection and fluorography
using X-ray film. After signal detection, the membranes were
re-incubated with an anti-β-actin antibody (Sigma-Aldrich, St.
Louis, MO, USA) as a loading control. The experiment was performed
three times.
 | Table II.Antibodies and conditions for western
blot analysis. |
Table II.
Antibodies and conditions for western
blot analysis.
| Primary antibody | Type | Host | Total proteins
(μg) | Working dilution | Source |
|---|
| AFP | Monoclonal | Mouse | 20 | 1:1,000 | R&D MAB1368 |
| CK-19 | Monoclonal | Mouse | 20 | 1:1,000 | R&D MAB3506 |
| ALB | Monoclonal | Mouse | 20 | 1:1,000 | R&D MAB1455 |
| CTGF | Polyclonal | Rabbit | 20 | 1:500 | Abcam ab5097 |
| β-actin | Monoclonal | Mouse | 20 | 1:4,000 | Sigma-Aldrich
A1978 |
Periodic acid-schiff (PAS) staining
Cells were fixed by 4% paraformaldehyde for 30 min
and then washed with dH2O. They were then oxidized in
0.5% periodic acid for 15 min at room temperature, rinsed in
dH2O for 1 min and treated with Schiff’s reagent
(Sigma-Aldrich) for 30 min at room temperature. After rinsing in
dH2O for 2 min, counterstaining was performed with
haematoxylin for 2 min, rinsed in running tap water and observed
under an inverted microscope.
Determination of medium levels of albumin
and cytochrome P450
Medium levels of albumin and cytochrome P450 were
determined using an albumin enzyme-linked immunosorbent assay
system and cytochrome P450 enzyme-linked immunosorbent assay system
(both from Blue Gene, China) according to the instructions supplied
by the manufacturer.
Construction of full-length CTGF gene
plasmids
Full-length CTGF was amplified from the cDNA of
WB-F344 cells using the forward,
5′-atagatctgaccatgctcgcctccgtcg-3′ (BglII site
italicized) and the reverse primer,
5′-atctcgagctttacgccatgtctcc ata-3′ (XhoI site
italicized). This fragment was inserted into the BglII and
XhoI sites of the vector, dl6-95/neo to yield the
recombinant plasmid, dl6-95/CTGF/neo. The recombinant plasmids were
amplified in competitive cells, purified using a Wizard®
Plus SV Miniprep DNA Purification System (Promega) and verified by
DNA sequencing.
Establishment of the stably transfected
WB-F344 cell line and identification
Prior to the transfection experiments, monolayers
were trypsinized and the cells were plated 1×105
cells/well in a 6-well format. After overnight culture, cells were
60–70% confluent. The cells were divided into three groups: a
WB-F344 cell control group; a dl6-95/neo group and a
dl6-95/CTGF/neo group, and transfected with plasmid DNA using
FuGene® HD transfection reagent (9 μl
transfection reagent to 3 g DNA; Roche, Mannheim, Germany)
according to the manufacturer’s protocol. The medium was replaced
with medium containing 10% FBS after 8 h of transfection.
Forty-eight hours later, cells were trypsinized, plated in 2 x
100-mm dishes, and cultured overnight. The medium was then replaced
with fresh complete medium containing G418 (100 μg/ml) for
the stable transfection experiments. The medium was replaced every
3–5 days. Cells were incubated for ~20 days, during which the G418
killed all the control cells, and the dl6-95/neo and
dl6-95/CTGF/neo groups developed stable clones. The medium was then
replaced with medium containing 50 μg/ml G418 for continued
culture until total mRNA or proteins were extracted. The expression
of CTGF mRNA was detected by reverse transcription-polymerase chain
reaction. The sequences of the primers used for PCR are shown in
Table III. The PCR conditions
were: 30 cycles at 94°C for 40 sec, annealing for 40 sec
(temperature shown in Table I)
and 72°C for 60 sec. Hepatic stellate cells were used as a control.
PCR products were analyzed by electrophoresis on 1% agarose
gels.
 | Table III.Primers used for real-time polymerase
chain reaction. |
Table III.
Primers used for real-time polymerase
chain reaction.
| Gene | Primer
sequences | Product size
(bp) | Accession no. |
|---|
| AFP | F:
5′-GGAGAAGTGCTGCAAAGACC-3′ | 120 | NM_012493 |
| R:
5′-TTGTCCTTTCTTCCTCCTGG-3′ |
| ALB | F:
5′-AGAACCAGGCCACTATCTC-3′ | 110 | NM_134326 |
| R:
5′-CAGATCGGCAGGAATGTTGT-3′ |
| CK-19 | F:
5′-CAGCAGTATTGAAGTCCAGC-3′ | 139 | NM_199498 |
| R:
5′-TCAAGCAGGCTTCGGTAGGT-3′ |
| GAPDH | F:
5′-CCTGCCAAGTATGATGACATCAAGA-3′ | 75 | BC059110 |
| R:
5′-GTAGCCCAGGATGCCCTTTAGT-3′ |
Statistical analysis
Data are expressed as means ± SD from three
independent experiments. Differences between mean values of
multiple groups were analyzed using the nonparametric ANOVA test
(SPSS, Inc., Chicago, IL, USA). Comparison between 2 groups was
made using the Student’s t-test. P<0.05 was indicative of a
statistically significant result.
Results
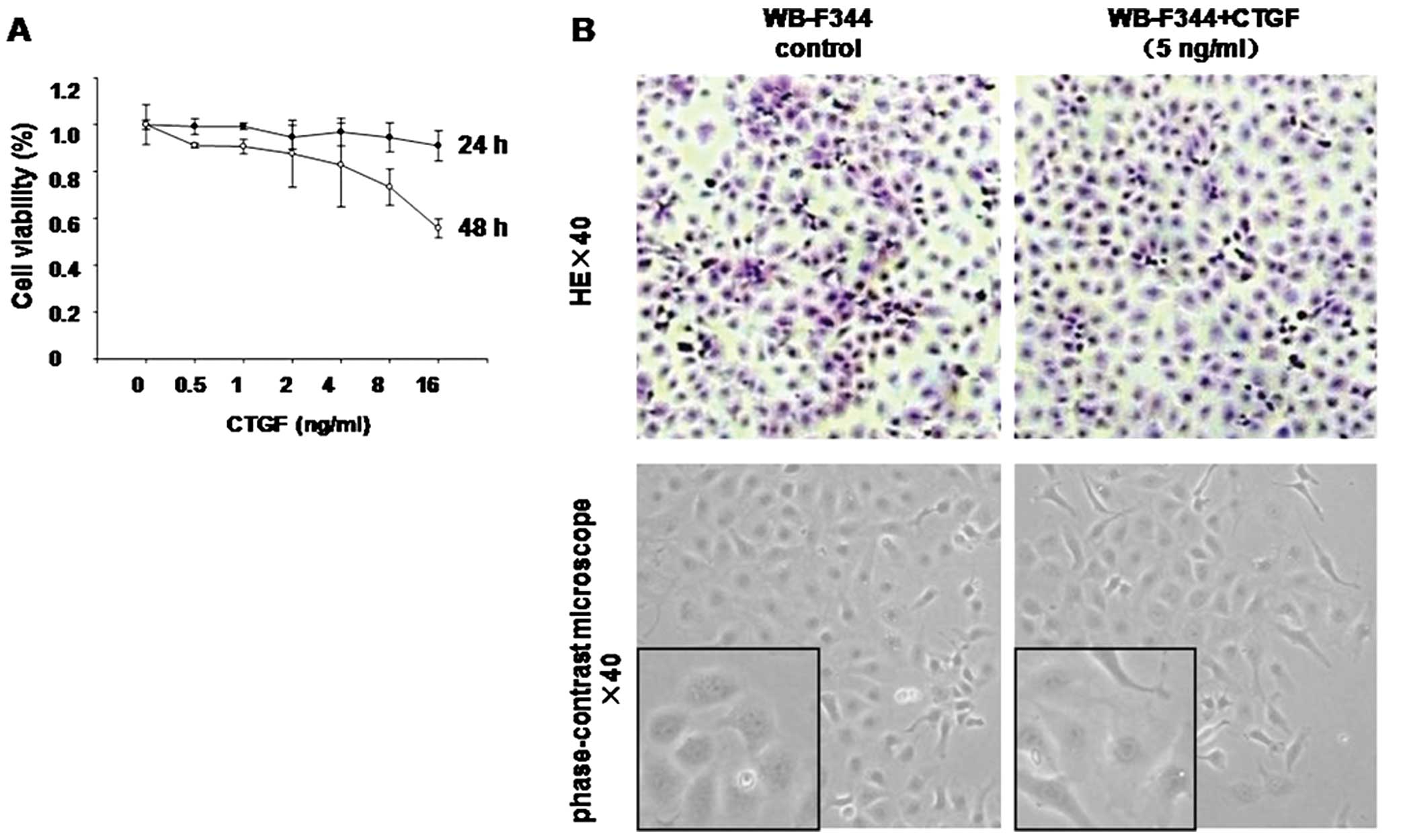
Effect of recombinant CTGF on viability
and morphology of WB-F344 cells
WB-F344 cells were treated with different doses of
rCTGF (0.5, 1, 2, 4, 8 or 16 ng/ml), resulting in cell viabilities
of 99, 99, 95, 97, 95 and 91%, respectively, after 24 h of
incubation. The percentage of viable cells gradually decreased to
91, 91, 88, 83, 73 and 56%, respectively, after 48 h of incubation
(Fig. 1A). To determine the dose
which had the lowest effect on progenitor proliferation, we
selected doses of 1 and 5 ng/ml CTGF and a treatment time of 24
h.
Cultured WB-F344 cells were small and polygonal with
a high nuclear/cytoplasmic ratio. After treatment with CTGF (5
ng/ml) for 24 h, the cells were enlarged and the
nuclear/cytoplasmic ratio was decreased (Fig. 1B).
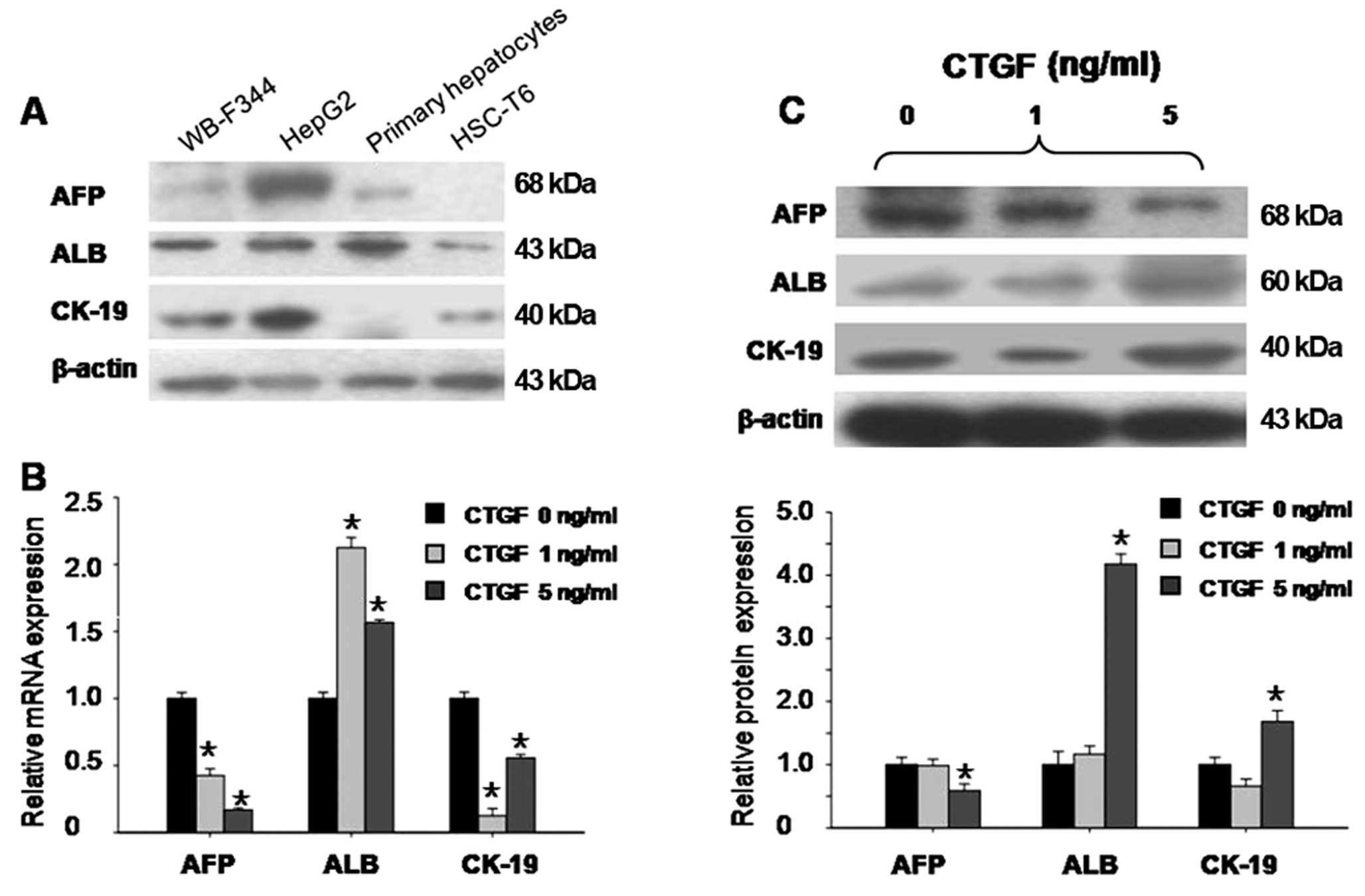
Recombinant CTGF induces the
differentiation of WB-F344 cells into hepatocytes
The WB-F344 cell line is an in vitro model of
hepatic progenitor cells. These progenitors express both AFP and
ALB, which are hepatocyte markers, highly expressed either in the
liver carcinoma cell line HepG2 or in primary isolated rat
hepatocytes. WB-F344 cells also express CK-19, which is a
cholangiocyte marker, also highly expressed by HepG2 cells
(Fig. 2A).
To study whether CTGF alters the expression of the
hepatic progenitor cell markers to induce differentiation, we
observed changes in their expression levels after CTGF treatment.
Treatment with CTGF at 5 ng/ml in serum-free medium for 24 h
resulted in a 0.2-fold decrease in the AFP mRNA level, a 1.6-fold
increase in the ALB mRNA level and a 0.6-fold decrease in the CK-19
mRNA level (in all P<0.05) (Fig.
2B). This was accompanied by a 0.6-fold decrease in AFP protein
levels, a 3.5-fold increase in the ALB protein level, and a
1.6-fold increase in the CK-19 protein level, respectively (in all
P<0.05) (Fig. 2C). CTGF at 1
ng/ml caused only minor changes in the expression levels of the
hepatic progenitor markers at the protein level. These results
indicate that recombinant CTGF induces WB-F344 cell differentiation
into hepatocytes rather than cholangiocytes.
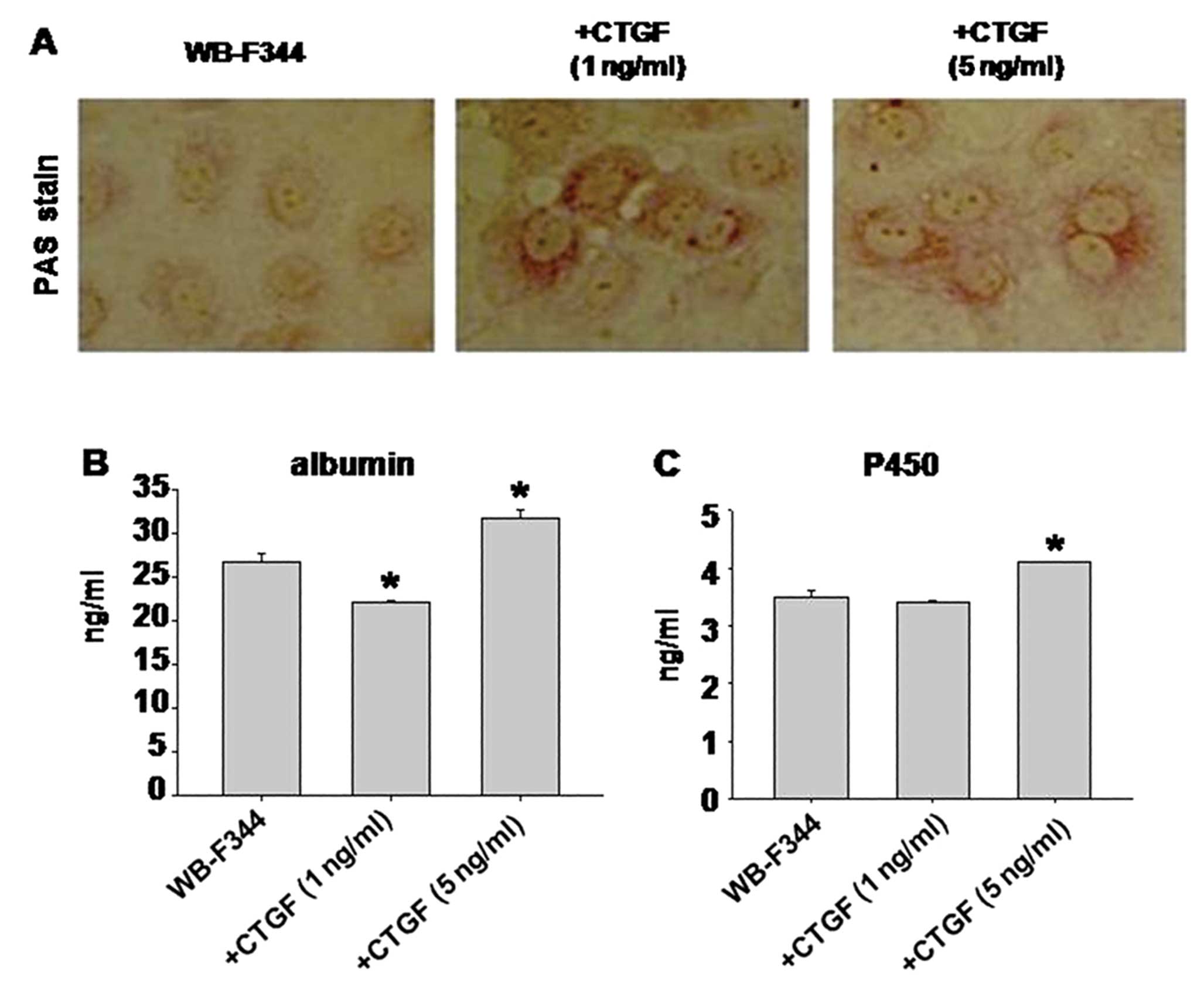
Finally, to confirm whether the differentiated cells
exhibit the functional properties of hepatocytes, we examined the
specific function of hepatocytes, including glycogen storage,
albumin production and cytochrome P450 production. WB-F344 cells
stimulated with CTGF gained the capability of glycogen storage
(Fig. 3A), which was absent in
cells without CTGF. These results were in correlation with the
albumin (Fig. 3B) and cytochrome
P450 (Fig. 3C) production.
Inhibition of CTGF suppresses WB-F344
cell differentiation
To further investigate the role of CTGF in the
differentiation of hepatic progenitor cells, we focused on whether
a chemical inhibitor that blocks CTGF expression suppresses the
differentiation of WB-F344 cells. A chemical inhibitor of CTGF
(WP631) was used.
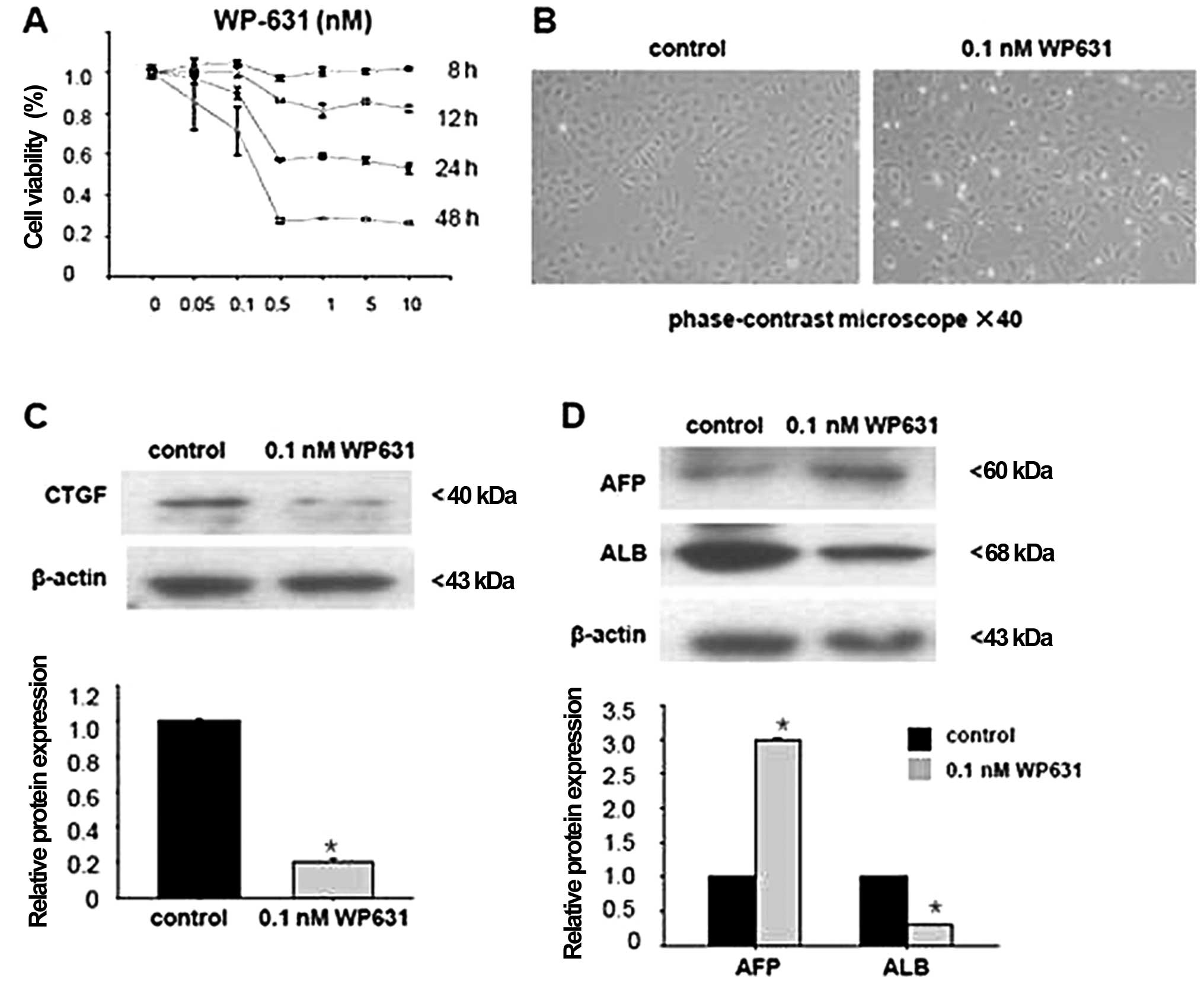
We first determined the optimal concentration of
WP631 by an MTT assay. Treatment with WP631 at ≥0.5 nM for 24 or 48
h reduced cell viability. Thus, we selected 0.1 nM WP631 for the
subsequent experiments, which resulted in cell viability of 90%
after 24 h (Fig. 4A), without
significantly affecting WB-F344 cell morphology (Fig. 4B).
WB-F344 cells were incubated with the inhibitor for
24 h (preliminary time-response experiments indicated that this
afforded maximal inhibition of CTGF expression; data not shown).
Incubation with 0.1 nM WP631 in serum-free medium decreased CTGF
expression 0.4-fold (P<0.05) (Fig.
4C).
Finally, we examined whether inhibition of CTGF
affects WB-F344 cell differentiation towards hepatocytes. Treatment
with 0.1 nM WP631 decreased ABL expression 0.6-fold (P<0.05),
but increased AFP expression 2.1-fold (P<0.05) (Fig. 4D).
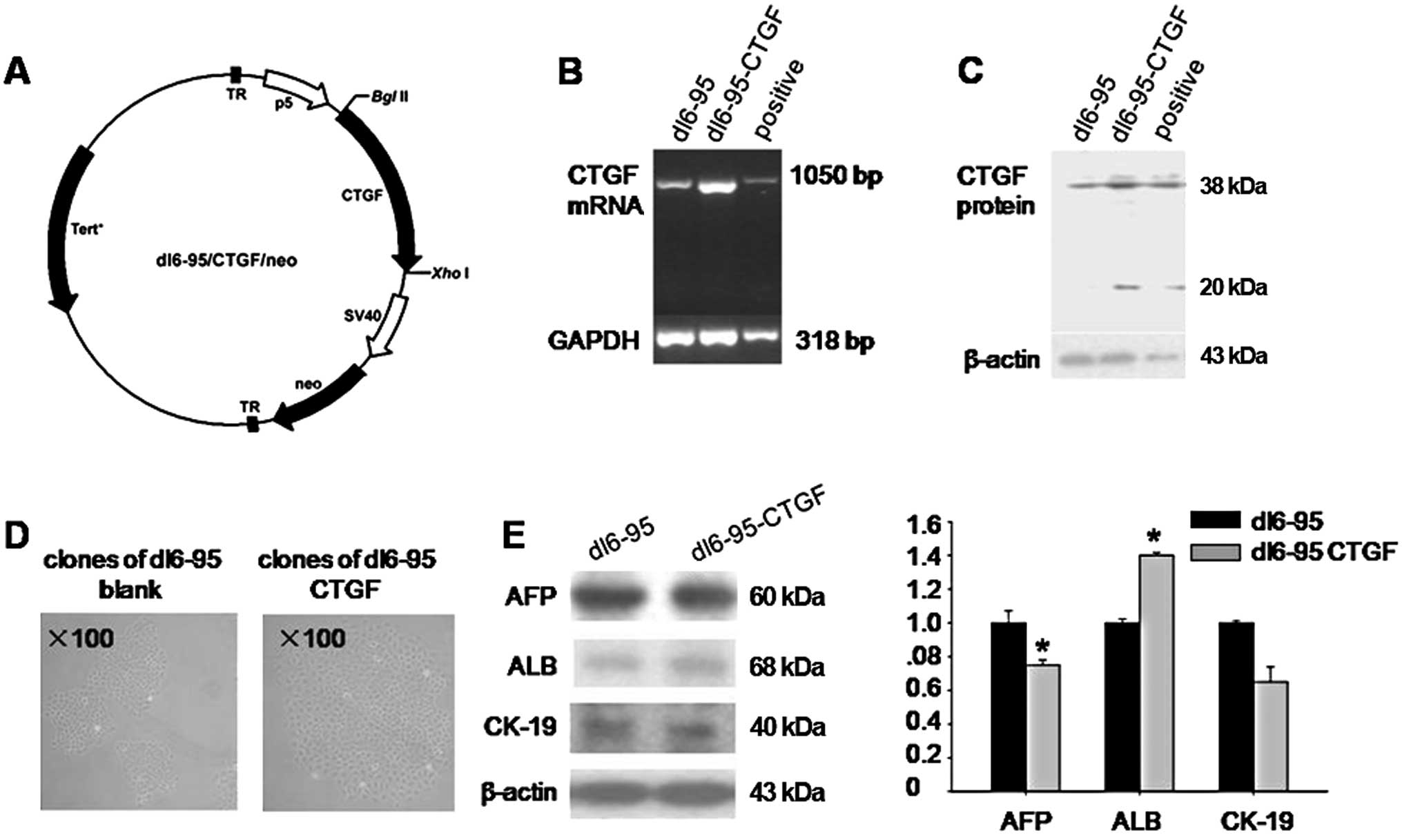
Endogenous CTGF induces WB-F344 cell
differentiation
We next examined whether endogenous CTGF also
induces the differentiation of WB-F344 cells. We isolated CTGF cDNA
from WB-F344 cells and constructed a plasmid carrying the
full-length CTGF gene (details in ‘Materials and methods’)
(Fig. 5A). WB-F344 cells were
transfected with the CTGF plasmid (dl6-95/CTGF/neo) or a blank
plasmid (dl6-95/neo) using the FuGene HD reagent (Fig. 5D). Following the selection of
clones in medium containing G418, the expression of CTGF by stably
transfected WB-F344 cells was analyzed by PCR and western blotting.
The expression of CTGF mRNA and protein was increased in WB-F344
cells transfected with the CTGF plasmid when compared with that in
the control (Fig. 5B and C).
After stable transfection, endogenous CTGF also
altered the phenotype of hepatic progenitor cells. The expression
of AFP decreased in the dl6-95/CTGF/neo group (0.7-fold; P<0.05)
when compared with the dl6-95/neo group. In contrast, expression of
the mature hepatocyte maker, ALB, increased (1.4-fold compared with
the blank plasmid; P<0.05). CK-19 expression did not change
significantly (P>0.05) (Fig.
5E). Endogenous CTGF stimulated WB-F344 cell differentiation to
a lesser degree than recombinant CTGF.
Discussion
Progenitor cell-mediated liver regeneration is a
complex process, involving sequential waves of cytokine secretion
and remodeling of the extracellular matrix (ECM) (2,18).
CTGF, a TGF-β downstream mediator, is involved in many biological
processes, such as cell adhesion, proliferation, survival,
migration and angiogenesis (8,19,20). CTGF may also be involved in liver
regeneration, since a TGF-β1 responsive element is present in the
promoter region of the CTGF gene (21,22). Studies have shown that CTGF
expression increases during liver regeneration occurring after
2AFF/PH and GalN administration in rats (10–12). Taken together, these data indicate
that CTGF may play an important role in progenitor activity during
the liver regeneration phase. However, there is no direct evidence
showing that CTGF affects progenitor differentiation.
The results of the present study indicate that CTGF
induces the differentiation of hepatic progenitor cells into
hepatocytes in vitro. Both stimulation by recombinant CTGF
and overexpression of CTGF induced WB-F344 cells to differentiate
into hepatocytes. CTGF increased the expression of a mature
hepatocyte marker (albumin) and decreased that of a fetal
hepatocyte marker (AFP). Importantly, WB-F344 cells acquired
hepatocyte-specific functions upon treatment with CTGF resulting in
albumin production, cytochrome P450 and glycogen storage
ability
However, induction of WB-F344 cell differentiation
using recombinant CTGF was greater than that noted with endogenous
CTGF. CTGF, a secreted protein, acts via both autocrine and
paracrine cellular circuits to regulate functions such as cell
proliferation, growth and cell differentiation (7,23);
therefore, under certain conditions, CTGF induced by TGF-β,
vascular endothelial growth factor (VEGF), platelet derived growth
factor (PDGF), endothelin-1 or hypoxia may play a role in
differentiation (24).
To further investigate the role of CTGF during the
differentiation of WB-F344 cells, we used an inhibitor of CTGF.
According to previous studies, WP631 is able to suppress CTGF
expression (7,24).
WP631 (dimethanesulfonate) was found to inhibit
Sp1-initiated transcription and was effective in reducing CTGF
expression (25). Sp-1, one of
several general transcription factor-binding sites, is thought to
be present in the promoter region of the CTGF gene (20,22). In our study, WP631 was very potent
at inhibiting CTGF expression in WB-F344 cells and impaired WB-F344
cell differentiation into hepatocytes.
Numerous studies have shown that CTGF plays a
crucial role in the fibrotic remodeling of various organs, and it
has frequently been proposed as a therapeutic target for the
management of fibrotic disorders (9,21).
However, little is known concerning its physiological function. In
this study, we found that CTGF induced the differentiation of
WB-F344 cells into hepatocytes in vitro. It has been
reported that the degree of hepatic progenitor cell activation
corresponds with the severity of liver fibrosis and inflammation
(5,19). CTGF is overexpressed in fibrotic
lesions and the degree of overexpression correlates with the
severity of fibrosis (9,19). Also, activation of hepatic
stellate cells is required for an appropriate oval cell response to
differentiation (4,6,17).
As mentioned above, the ability of CTGF to induce the
differentiation of hepatic progenitor cells and promote liver
regeneration during fibrosis requires further investigation.
Acknowledgements
This study was supported by the
National Natural Science Foundation of China (81270519 and
81100294), the Program for New Century Excellent Talents in the
University (NCET-09-0008) and the Beijing Health System Talents
Plan (2009-3-06).
References
|
1.
|
Michalopoulos GK: Liver regeneration after
partial hepatectomy: critical analysis of mechanistic dilemmas. Am
J Pathol. 176:2–13. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
2.
|
Fausto N, Campbell JS and Riehle KJ: Liver
regeneration. Hepatology. 43:S45–S53. 2006. View Article : Google Scholar
|
|
3.
|
Bird TG, Lorenzini S and Forbes SJ:
Activation of stem cells in hepatic diseases. Cell Tissue Res.
1:283–300. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4.
|
Pintilie DG, Shupe TD, Oh SH, et al:
Hepatic stellate cells’ involvement in progenitor-mediated liver
regeneration. Lab Invest. 90:1199–1208. 2010.
|
|
5.
|
Zheng ZY, Weng SY and Yu Y: Signal
molecule-mediated hepatic cell communication during liver
regeneration. World J Gastroenterol. 15:5776–5783. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6.
|
Roskams T: Relationships among stellate
cell activation, progenitor cells, and hepatic regeneration. Clin
Liver Dis. 12:853–860. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7.
|
Gressner OA and Gressner AM: Connective
tissue growth factor: a fibrogenic master switch in fibrotic liver
diseases. Liver Int. 28:1065–1079. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
8.
|
Tong Z, Chen R, Alt DS, et al:
Susceptibility to liver fibrosis in mice expressing a connective
tissue growth factor transgene in hepatocytes. Hepatology.
50:939–947. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9.
|
Chen L, Charrier AL, Leask A, et al:
Ethanol-stimulated differentiated functions of human or mouse
hepatic stellate cells are mediated by connective tissue growth
factor. J Hepatol. 55:399–406. 2011. View Article : Google Scholar
|
|
10.
|
Pi L, Oh SH, Shupe T and Petersen BE: Role
of connective tissue growth factor in oval cell response during
liver regeneration after 2-AAF/PHx in rats. Gastroenterology.
128:2077–2088. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
11.
|
Ujike K, Shinji T, Hirasaki S, et al:
Kinetics of expression of connective tissue growth factor gene
during liver regeneration after partial hepatectomy and
D-galactosamine-induced liver injury in rats. Biochem Biophys Res
Commun. 277:448–454. 2000. View Article : Google Scholar
|
|
12.
|
Pi L, Ding X, Jorgensen M, et al:
Connective tissue growth factor with a novel fibronectin binding
site promotes cell adhesion and migration during rat oval cell
activation. Hepatology. 47:996–1004. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13.
|
Wang P, Liu T, Cong M, et al: Expression
of extracellular matrix genes in cultured hepatic oval cells: an
origin of hepatic stellate cells through transforming growth factor
beta? Liver Int. 29:575–584. 2009. View Article : Google Scholar
|
|
14.
|
Lázaro CA, Rhim JA, Yamada Y and Fausto N:
Generation of hepatocytes from oval cell precursors in culture.
Cancer Res. 58:5514–5522. 1998.PubMed/NCBI
|
|
15.
|
Snykers S, De Kock J, Rogiers V and
Vanhaecke T: In vitro differentiation of embryonic and adult stem
cells into hepatocytes: state of the art. Stem Cells. 27:577–605.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
16.
|
Shiojiri N, Lemire JM and Fausto N: Cell
lineages and oval cell progenitors in rat liver development. Cancer
Res. 51:2611–2620. 1991.PubMed/NCBI
|
|
17.
|
Nagai H, Terada K, Watanabe G, et al:
Differentiation of liver epithelial (stem-like) cells into
hepatocytes induced by coculture with hepatic stellate cells.
Biochem Biophys Res Commun. 293:1420–1425. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
18.
|
Duncan AW, Dorrell C and Grompe M: Stem
cells and liver regeneration. Gastroenterology. 137:466–481. 2009.
View Article : Google Scholar
|
|
19.
|
Blom IE, Goldschmeding R and Leask A: Gene
regulation of connective tissue growth factor: new targets for
antifibrotic therapy? Matrix Biol. 21:473–482. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20.
|
Gressner OA, Weiskirchen R and Gressner
AM: Evolving concepts of liver fibrogenesis provide new diagnostic
and therapeutic options. Comp Hepatol. 6:72007. View Article : Google Scholar : PubMed/NCBI
|
|
21.
|
Holmes A, Abraham DJ, Chen Y, et al:
Constitutive connective tissue growth factor expression in
scleroderma fibroblasts is dependent on Sp1. J Biol Chem.
278:41728–41733. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
22.
|
Grotendorst GR, Okochi H and Hayashi N: A
novel transforming growth factor beta response element controls the
expression of the connective tissue growth factor gene. Cell Growth
Differ. 7:469–480. 1996.PubMed/NCBI
|
|
23.
|
Tan JT, McLennan SV, Song WW, et al:
Connective tissue growth factor inhibits adipocyte differentiation.
Am J Physiol Cell Physiol. 295:C740–C751. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24.
|
Leask A, Parapuram SK, Shi-Wen X and
Abraham DJ: Connective tissue growth factor (CTGF, CCN2) gene
regulation: a potent clinical bio-marker of fibroproliferative
disease? J Cell Commun Signal. 3:89–94. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25.
|
Gressner OA, Lahme B, Demirci I, et al:
Differential effects of TGF-beta on connective tissue growth factor
(CTGF/CCN2) expression in hepatic stellate cells and hepatocytes. J
Hepatol. 47:699–710. 2007. View Article : Google Scholar : PubMed/NCBI
|