Introduction
Pancreatic cancer remains a leading cause of cancer
deaths in the advanced nation (1,2). The
overall 5-year survival rate is reported to be less than 5%
(3). A reliable and clinically
relevant prognostic biomarker which can stratify the disease is
needed for developing new strategies.
It is a known fact that chromatin, highly condensed
and dynamically structured, can be temporally rearranged so that
specific genes can be expressed or repressed (4). Studies have shown that modification
of chromatin structure is an essential step in gene regulation
primarily mediated by chromatin remodeling proteins. Among these
proteins, histone is known to play a dynamic role in the regulation
of transcription (5–7). Often, transcription is also regulated
by other cofactors, and the balance of chromatin remodeling
activities may be crucial to ensure accurate responses to
developmental or environmental cues and to prevent the transition
of normal cells into cancer cells (8).
The SWItch/sucrose non-fermentable (SWI/SNF) complex
is a major complex of adenosine triphosphate (ATP)-dependent
chromatin remodeling factors and controls the transcriptional
activity of a variety of genes involved in cellular growth and
transformation by altering chromatin structure (9–13).
SWI/SNF complex, originally identified in yeast, is composed of
more than 10 characterized subunits (14,15)
and human SWI/SNF complexes contain one of the two core ATPase
subunits, BRM or BRG1 (13,16–18).
Growing genetic and molecular evidence indicates that specific
subunits of the SWI/SNF complex can act as tumor suppressors
(6,19). However, there is no report on the
relationship between SWI/SNF components expression and the clinical
significance of pancreatic cancer. In this study, we investigated
the expression levels of SWI/SNF components to clarify the clinical
impact of SWI/SNF complex on pancreatic cancer.
Materials and methods
Patients and samples
The surgical specimens of pancreatic cancer tissue
obtained from 68 patients were evaluated. All of the patients had
undergone macroscopically curative resection (R0, 1) at Kanagawa
Cancer Center between July 2006 and April 2010. The
clinicopathological characteristics of these patients are shown in
Table I. In all cases, archival
hematoxylin and eosin-stained (H&E) slides of the primary tumor
were retrieved and reviewed to confirm the pathological features as
well as to select suitable tissue blocks for immunohistochemical
analysis. Informed consent was obtained from each patient. The
Ethics Committees of the Kanagawa Cancer Center approved the
protocol before initiation of the study. We declare no conflicts of
interest.
 | Table I.The clinicopathological
characteristics of all patients. |
Table I.
The clinicopathological
characteristics of all patients.
| Clinicopathological
characteristics | No. of patients
(n=68) |
|---|
| Age | |
| <65 | 30 |
| ≥65 | 38 |
| Sex | |
| Male | 36 |
| Female | 32 |
| Tumor location in
pancreas | |
| Head | 46 |
| Body/tail | 22 |
| Tumor size
(cm) | |
| <4 | 29 |
| ≥4 | 39 |
| Histological
type | |
| Well/mod | 32 |
| Poor | 36 |
| T | |
| T1–3 | 38 |
| T4 | 30 |
| N | |
| N0 | 17 |
| N1 | 51 |
| M | |
| M0 | 53 |
| M1 | 15 |
| Curability of
surgery | |
| R0 | 43 |
| R1 | 25 |
| Stage | |
| 0–III | 53 |
| IV | 15 |
| Adjuvant
gemcitabine | |
| Yes | 42 |
| No | 26 |
Tissue microarrays and
immunohistochemistry
Microarrays consisting of cores, each measuring 2 mm
in diameter, were prepared from formalin-fixed paraffin-embedded
tissue blocks of surgically removed primary tumors. Each tissue
core of the primary tumor was sampled.
Immunohistochemical staining was performed using
commercially available polyclonal rabbit, or mouse antibodies
raised against BRM (Abcam Inc., Cambridge, MA), BRG1 (Santa Cruz
Biotechnology Inc., Santa Cruz, CA), BAF250a (Santa Cruz
Biotechnology Inc.), BAF180 (Sigma-Aldrich Inc., St. Louis, MO),
BAF47 (Santa Cruz Biotechnology Inc.). Tissue microarray blocks
were sectioned at a thickness of 4 μm and mounted on
pre-coated glass slides. The sections were de-paraffinized through
a graded series of xylene and rehydrated through a graded series of
alcohol to distilled water. Endogenous peroxidase was quenched with
3% hydrogen peroxide in methanol at room temperature. The sections
were placed in a 95°C solution of 0.01 M sodium citrate buffer (pH
6.0) for 40 min for antigen retrieval. Normal goat serum (5%) was
then applied for 15 min to block any non-specific protein binding
sites. Primary polyclonal antibodies were applied for 1 h at room
temperature at the following dilutions: anti-BRM at 1:250,
anti-BRG1 at 1:200, anti-BAF250a at 1:100, anti-BAF180 at 1:90 and
BAF47 at 1:300. Immunoreactive proteins were detected using the
Simple Stain MAX-PO (Multi).
All sections were counterstained with Mayer’s
hematoxylin, and negative controls were included in each staining
sequence. The intensity and global level of staining were scored
semi-quantitatively for each tissue microarray by an investigator
blinded to all of the clinicopathological variables. The global
level of staining refers to the percentage of tumor cells that
stained positively for an antibody within each tissue microarray at
×200 magnification using a light microscope.
Scoring of immunohistochemical
reactivity
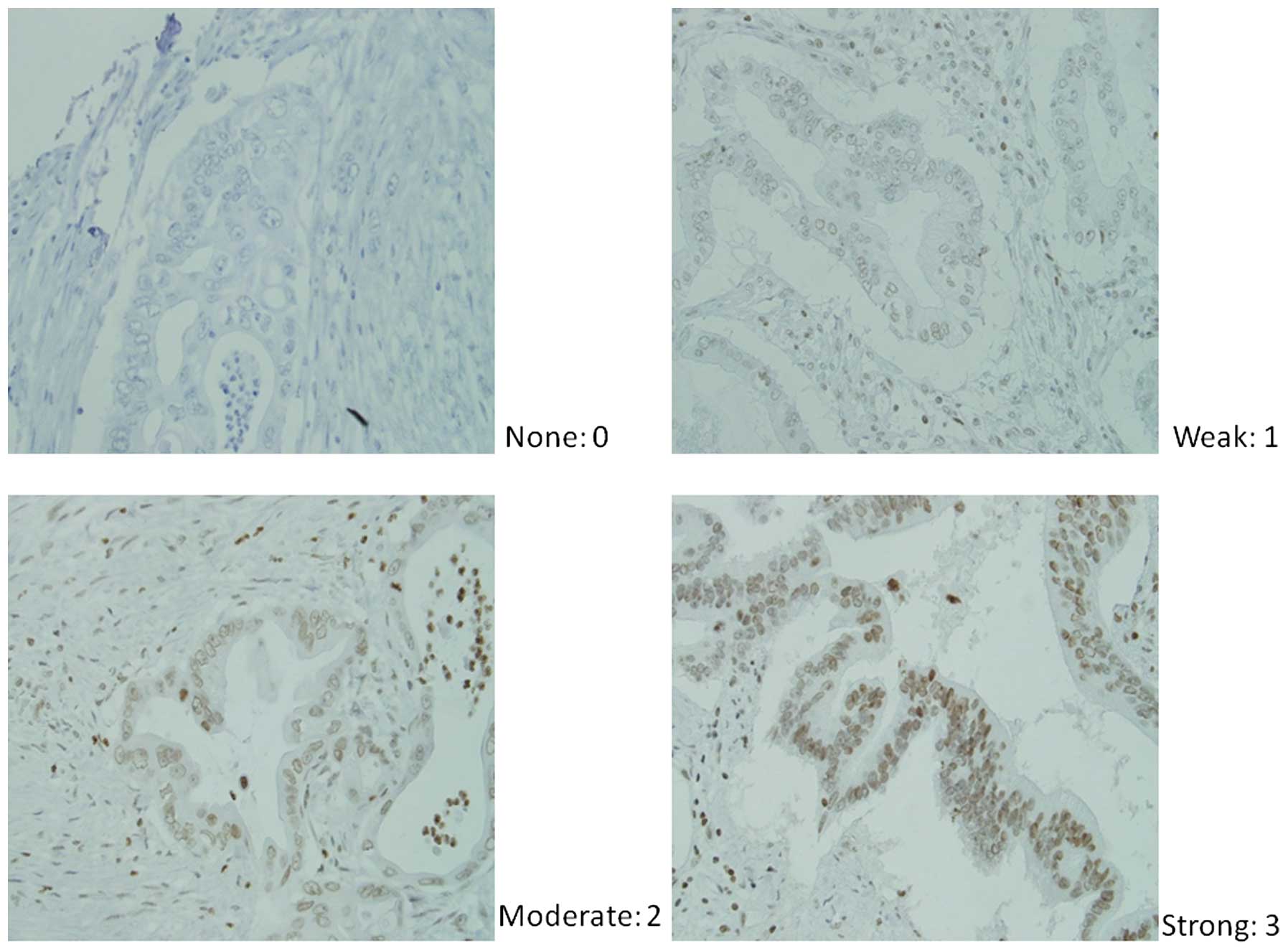
Immunohistochemical scoring was completed using the
modified Histoscore (H-score) (20), which involves a semiquantitative
assessment of both the intensity of staining (graded as: 0,
non-staining; 1, weak; 2, median; or 3, strong using adjacent
normal mucosa as the median) and the percentage of positive cells
(Fig. 1). The range of possible
scores was from 0 to 300. Expression level of each component was
categorized as low or high according to the median value of
H-score.
Statistical analysis
The relationships between the expression level and
the clinicopathological factors were evaluated with the
χ2 test. The postoperative survival rate from the day of
primary tumor resection was analyzed using the Kaplan-Meier method
and any differences in the survival rates were assessed with the
log-rank test. A Cox proportional-hazard model was used for the
multivariate analyses. Differences were considered significant when
P<0.05. The statistical analysis was performed using the PASW
Statistics 18 (SPSS, Inc., Chicago, IL).
Results
Relation of SWI/SNF component expression
to clinicopathological features
The distribution of H-score is showed in Fig. 2. Expression level of the SWI/SNF
components was categorized as low or high according to the median
value of the H-score. Relations between the expression levels of
each component and clinicopathological features were then examined.
Factors implicating significant relations were tumor size, T
factor, M factor, lymphatic invasion, and stage in BRM, histology
and stage in BRG1, tumor size in BAF180, lymphatic invasion in
BAF47, respectively (Table
II).
 | Table II.Relation of SWI/SNF component
expression to clinicopathological factors. |
Table II.
Relation of SWI/SNF component
expression to clinicopathological factors.
| BRM
| BRG1
| BAF250a
| BAF180
| BAF47
|
|---|
| Factors | Low | High | p-value | Low | High | p-value | Low | High | p-value | Low | High | p-value | Low | High | p-value |
|---|
| Age (years) | | | | | | | | | | | | | | | |
| <65/≥65 | 15/19 | 15/19 | 1.000 | 18/19 | 12/22 | 0.143 | 13/21 | 17/17 | 0.329 | 19/15 | 11/23 | 0.051 | 13/21 | 17/17 | 0.329 |
| Gender | | | | | | | | | | | | | | | |
| Male/female | 16/18 | 16/18 | 1.000 | 16/18 | 16/18 | 1.000 | 13/21 | 19/15 | 0.145 | 15/19 | 17/17 | 0.627 | 17/17 | 15/19 | 0.627 |
| Tumor size | | | | | | | | | | | | | | | |
| <4/≥4 cm | 19/15 | 10/24 | 0.027 | 12/22 | 17/17 | 0.220 | 14/20 | 15/19 | 0.806 | 10/24 | 19/15 | 0.027 | 15/19 | 14/20 | 0.806 |
| Histology | | | | | | | | | | | | | | | |
| Well,
mod/poor | 18/16 | 14/20 | 0.331 | 11/23 | 21/13 | 0.015 | 14/20 | 18/16 | 0.331 | 13/21 | 19/15 | 0.145 | 15/19 | 17/17 | 0.627 |
| T | | | | | | | | | | | | | | | |
| T1-3/4 | 25/9 | 13/21 | 0.003 | 23/11 | 15/19 | 0.051 | 17/17 | 21/13 | 0.329 | 19/15 | 19/15 | 1.000 | 20/14 | 18/16 | 0.625 |
| N | | | | | | | | | | | | | | | |
| N0/N1 | 9/25 | 8/26 | 0.779 | 10/24 | 7/27 | 0.401 | 10/24 | 7/27 | 0.401 | 8/26 | 9/25 | 0.779 | 9/25 | 8/26 | 0.779 |
| M | | | | | | | | | | | | | | | |
| M0/M1 | 30/4 | 23/11 | 0.041 | 27/7 | 26/8 | 0.770 | 24/10 | 29/5 | 0.114 | 25/9 | 28/6 | 0.380 | 28/6 | 25/9 | 0.380 |
| Vessel
invasion | | | | | | | | | | | | | | | |
| No/yes | 12/22 | 8/26 | 0.287 | 11/23 | 9/25 | 0.595 | 7/27 | 13/21 | 0.110 | 10/24 | 10/24 | 1.000 | 8/26 | 12/22 | 0.287 |
| Lymphatic
invasion | | | | | | | | | | | | | | | |
| No/yes | 15/19 | 6/28 | 0.018 | 13/21 | 8/26 | 0.189 | 9/25 | 12/22 | 0.431 | 9/25 | 12/22 | 0.431 | 15/19 | 6/28 | 0.018 |
| Stage | | | | | | | | | | | | | | | |
| 0–III/IV | 18/16 | 5/29 | 0.001 | 17/17 | 6/28 | 0.005 | 10/24 | 13/21 | 0.442 | 11/23 | 12/22 | 0.798 | 14/20 | 9/25 | 0.200 |
| Curability | | | | | | | | | | | | | | | |
| R0/R1 | 25/9 | 18/16 | 0.078 | 23/11 | 20/14 | 0.451 | 20/14 | 23/11 | 0.451 | 21/13 | 22/12 | 0.801 | 20/14 | 23/11 | 0.451 |
Analysis of prognostic factors in all
patients
Univariate Cox regression analysis for overall
survival in all patients showed that age, tumor size, histological
type, M factor, curability of the surgery, and expression level of
BRM as well as BAF180 were significant predictors (Table III). On multivariate Cox
proportional hazard analysis, histology, expression level of BRM
and BAF180 were significant independent predictors of overall
survival in patients with pancreatic cancer (Table IV).
 | Table III.Univariate analysis for overall
survival in pancreatic cancer. |
Table III.
Univariate analysis for overall
survival in pancreatic cancer.
| Factors | HR (95% CI) | p-value |
|---|
| Age (years) | | 0.035 |
| <65 | 1.0 | |
| ≥65 | 0.533
(0.293–0.967) | |
| Sex | | 0.632 |
| Male | 1.0 | |
| Female | 0.865
(0.478–1.565) | |
| Tumor size
(cm) | | 0.035 |
| <4 | 1.0 | |
| ≥4 | 1.979
(1.048–3.739) | |
| Histology | | 0.002 |
| Well/mod | 1.0 | |
| Poor | 2.744
(1.429–5.271) | |
| T | | 0.071 |
| T1–3 | 1.0 | |
| T4 | 1.733
(0.955–3.146) | |
| N | | 0.602 |
| N0 | 1.0 | |
| N1 | 1.208
(0.594–2.458) | |
| M | | 0.010 |
| M0 | 1.0 | |
| M1 | 2.329
(1.222–4.439) | |
| Curability of
surgery | | 0.020 |
| R0 | 1.0 | |
| R1 | 2.068
(1.121–3.815) | |
| BRM | | 0.011 |
| Low | 1.0 | |
| High | 2.225
(1.199–4.129) | |
| BRG1 | | 0.601 |
| Low | 1.0 | |
| High | 0.853
(0.471–1.546) | |
| BAF250a | | 0.479 |
| Low | 1.0 | |
| High | 0.807
(0.446–1.461) | |
| BAF180 | | 0.007 |
| Low | 1.0 | |
| High | 0.428
(0.231–0.793) | |
| BAF47 | | 0.226 |
| Low | 1.0 | |
| High | 0.690
(0.378–1.258) | |
 | Table IV.Multivariate analysis for overall
survival in pancreatic cancer. |
Table IV.
Multivariate analysis for overall
survival in pancreatic cancer.
| Factors | HR (95% CI) | p-value |
|---|
| Age | | 0.169 |
| <65 | 1.0 | |
| ≥65 | 0.633
(0.330–1.214) | |
| Tumor size
(cm) | | 0.755 |
| <4 | 1.0 | |
| ≥4 | 1.122
(0.543–2.318) | |
| Histology | | 0.011 |
| Well/Mod | 1.0 | |
| Poor | 2.702
(1.253–5.830) | |
| M | | 0.486 |
| M0 | 1.0 | |
| M1 | 1.381
(0.557–3.424) | |
| Curability of
surgery | | 0.076 |
| R0 | 1.0 | |
| R1 | 1.981
(0.932–4.214) | |
| BRM | | 0.032 |
| Low | 1.0 | |
| High | 2.144
(1.066–4.311) | |
| BAF180 | | 0.041 |
| Low | 1.0 | |
| High | 0.501
(0.258–0.971) | |
Comparison of survival by the status of
BRM and BAF180
The 5-year survival rate of high BRM patients was
9.8%, which was significantly worse than that of low BRM patients
(43.8%) (Fig. 3). Also, the 5-year
survival rate of low BAF180 (8.1%) was significantly worse than
that of high BAF180 patients (40.8%) (Fig. 3).
Hazard analysis of SWI/SNF components in
the patients treated with adjuvant gemcitabine
Multivariate analysis (Table V) and survival analysis (Fig. 4) showed that BRM-high and
BAF180-low were independent prognostic factors for overall survival
in the patients treated with adjuvant gemcitabine.
 | Table V.Multivariate analysis for overall
survival in patients with adjuvant gemcitabine. |
Table V.
Multivariate analysis for overall
survival in patients with adjuvant gemcitabine.
| Factors | HR (95% CI) | p-value |
|---|
| Age | | 0.002 |
| <65 | 1.0 | |
| ≥65 | 0.227
(0.089–0.580) | |
| Tumor size
(cm) | | 0.280 |
| <4 | 1.0 | |
| ≥4 | 0.593
(0.230–1.531) | |
| Histology | | 0.267 |
| Well/Mod | 1.0 | |
| Poor | 1.907
(0.610–5.964) | |
| M | | 0.923 |
| M0 | 1.0 | |
| M1 | 0.947
(0.315–2.847) | |
| Curability of
surgery | | 0.784 |
| R0 | 1.0 | |
| R1 | 1.145
(0.433–3.029) | |
| BRM | | 0.017 |
| Low | 1.0 | |
| High | 3.411
(1.251–9.305) | |
| BAF180 | | 0.016 |
| Low | 1.0 | |
| High | 0.336
(0.138–0.819) | |
Discussion
Chromatin remodeling factors have been the subject
of great interest in oncology. However, little is known about their
role in pancreatic cancer.
The SWI/SNF complexes are large, multi-subunit
complexes containing 10 or more subunits, serving as a master
switch that directs and limits the execution of specific cellular
programs, such as differentiation and growth control (21). Each complex has one of the two
different ATPase as core motor; BRM or BRG1, and subunits which are
referred to as BAFs (BRM- or BRG1-associated factors). The
BRM-containing complex is termed BRM/BAF. The BRG1-containing
complexes are further divided into those that contain the BAF250a
(termed BRG1/BAF) or the BAF180 (termed PBAF). These three types of
complexes are believed to have different molecular functions
(22).
There are several studies reporting that the subunit
of SWI/SNF complex was decreased in cancer tissues. They revealed
the mutation of ARID1A, which codes BAF250a protein, in
about half of ovarian clear cell carcinomas (23,24),
and PBRM1, which codes BAF180, in approximately 40% of renal
cell carcinomas (25). Another
study identified the SWI/SNF chromatin remodeling complex as tumor
suppressor, by mediating retinoblastoma protein (RB)-derived
regulation of the cell cycle (22,26,27).
However, the roles of these subunits in pancreatic cancers are
poorly understood.
In this study, we investigated the expression levels
of 5 key subunits; BRM, BRG1, BAF250a, BAF180, which are the key
subunits when subdividing complex types, and BAF47. There is
established evidence that BAF47 is a tumor suppressor in rhabdoid
tumors (28).
In the analysis of expression level and
clinicopahological features, high BRM was related to worse
clinicopathological features in general, including larger tumor
size, T4 disease, other organ metastasis, lymphatic invasion, and
stage IV disease. Stage IV disease was also correlated to high
BRG1, which is reported to have similar biological function as BRM.
On the other hand, better clinicopathological features were related
to high BAF expression. High BAF180 was related to smaller tumor
size, and high BAF47 was associated with negative lymphatic
invasion.
In addition, our multivariate analysis revealed both
high BRM and low BAF180 were independent prognostic indicators for
poor survival, whereas the expression level of BRG1, BAF250a, and
BAF47 were not related to overall survival.
As a next step, we investigated the prognostic
significance of these factors in the patients with adjuvant
gemcitabine. Gemcitabine remains standard therapy in the adjuvant
and palliative settings for pancreatic cancer (29,30).
However, the response rate of gemcitabine is very low, with only
18% of 1-year survival rate (31).
Developing a novel biomarker, which predicts the response for
gemcitabine, is urgently needed. In the analysis of the patients
with gemcitabine, we reached the same result; both high BRM and low
BAF180 were independent prognostic indicators for poor
survival.
A previous study showed that BRM or BRG1 is lost in
10–20% of the bladder, colon, breast, esophageal, pancreatic and
ovarian cancers by immunohistochemical staining of tissue
microarrays (32). Another study
reported BRM was lost in approximately 15–20% of primary non-small
lung cancers, and silencing of BRM was a prognostic factor for poor
outcome (33,34). Although BRM is supposed to be
involved in many biological functions, these data showed
BRM-containing complexes (BRM/BAF) as tumor suppressor in cancer
tissue.
It is also reported that BRM has a role in trans
cription of CD44 (35), which is
important in the process of tumor-endothelium interactions, cell
migration, cell adhesion, tumor progression and metastasis
(36).
Our result showed that the patient with high BRM had
a significantly worse survival than those without (5-year OS: 9.8
vs. 43.8%, p=0.009), suggesting BRM/BAF in pancreatic cancer may
contribute to tumor progression.
We also revealed the significant relationship
between high BAF180 expression and smaller-sized tumor, and
identified BAF180 as an independent prognostic factor for better
survival in pancreatic cancer.
BAF180 maps to the 3p12 region (37) where allele loss is frequent and
homozygous deletion have been detected in lung and breast cancer
cell lines (38,39). Thus, genes located on this region
have been thought as candidates for tumor suppressors. Actually, it
is reported that BAF180 mutation is associated with carcinogenesis
of breast cancer, and BAF180 suppresses tumorigenesis through its
ability to regulate p21 (40),
which controls the cell cycle (41). Recent research also clarified
BAF180 mutation in clear cell renal cell carcinoma (42). These results suggest the idea that
BAF180-containing complexes (PBAF) suppress tumor progression,
which does not contradict our present results.
BAF250a-containing SWI/SNF complexes (BRG1/BAF) are
reported to have different structure and biological properties from
PBAF (43,44). A previous study showed that BAF250a
was deleted in as many as 30% of renal cell carcinoma and 10% of
breast carcinoma (19,45). These results lead to the concept
that BRG1/BAF appear to have antagonistic effect on cell cycle
progression (46). However, our
data did not show the relationship of BAF250a expression to
clinicopathological features or overall survival in pancreatic
cancer.
Based on this study, we reached the conclusion that
high BRM, and low BAF180 are useful biomarker not only for the
patients with curative resection, but also for those with adjuvant
gemcitabine. Future investigation into biological functions of
SWI/SNF components could lead to better management in pancreatic
cancer.
References
|
1.
|
Parkin DM, Bray FI and Devesa SS: Cancer
burden in the year 2000. The global picture. Eur J Cancer. 37(Suppl
8): S4–S66. 2001.PubMed/NCBI
|
|
2.
|
Jemal A, Siegel R, Ward E, Hao Y, Xu J and
Thun MJ: Cancer statistics, 2009. CA Cancer J Clin. 59:225–249.
2009. View Article : Google Scholar
|
|
3.
|
Hidalgo M: Pancreatic cancer. N Engl J
Med. 362:1605–1617. 2010. View Article : Google Scholar
|
|
4.
|
Reisman DN, Strobeck MW, Betz BL, et al:
Concomitant down-regulation of BRM and BRG1 in human tumor cell
lines: differential effects on RB-mediated growth arrest vs CD44
expression. Oncogene. 21:1196–1207. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
5.
|
Wade PA: Transcriptional control at
regulatory checkpoints by histone deacetylases: molecular
connections between cancer and chromatin. Hum Mol Genet.
10:693–698. 2001. View Article : Google Scholar
|
|
6.
|
Neely KE and Workman JL: The complexity of
chromatin remodeling and its links to cancer. Biochim Biophys Acta.
1603:19–29. 2002.PubMed/NCBI
|
|
7.
|
Klochendler-Yeivin A, Muchardt C and Yaniv
M: SWI/SNF chromatin remodeling and cancer. Curr Opin Genet Dev.
12:73–79. 2002. View Article : Google Scholar
|
|
8.
|
Davis PK and Brackmann RK: Chromatin
remodeling and cancer. Cancer Biol Ther. 2:22–29. 2003. View Article : Google Scholar
|
|
9.
|
Narlikar GJ, Fan HY and Kingston RE:
Cooperation between complexes that regulate chromatin structure and
transcription. Cell. 108:475–487. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
10.
|
Langst G and Becker PB: Nucleosome
mobilization and positioning by ISWI-containing
chromatin-remodeling factors. J Cell Sci. 114:2561–2568.
2001.PubMed/NCBI
|
|
11.
|
Kingston RE and Narlikar GJ: ATP-dependent
remodeling and acetylation as regulators of chromatin fluidity.
Genes Dev. 13:2339–2352. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
12.
|
Burns LG and Peterson CL: The yeast
SWI-SNF complex facilitates binding of a transcriptional activator
to nucleosomal sites in vivo. Mol Cell Biol. 17:4811–4819.
1997.PubMed/NCBI
|
|
13.
|
Wu C: Chromatin remodeling and the control
of gene expression. J Biol Chem. 272:28171–28174. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
14.
|
Breeden L and Nasmyth K: Cell cycle
control of the yeast HO gene: cis- and trans-acting
regulators. Cell. 48:389–397. 1987. View Article : Google Scholar : PubMed/NCBI
|
|
15.
|
Recht J and Osley MA: Mutations in both
the structured domain and N-terminus of histone H2B bypass the
requirement for Swi-Snf in yeast. EMBO J. 18:229–240. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
16.
|
Phelan ML, Sif S, Narlikar GJ and Kingston
RE: Reconstitution of a core chromatin remodeling complex from
SWI/SNF subunits. Mol Cell. 3:247–253. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
17.
|
Tyler JK and Kadonaga JT: The ‘dark side’
of chromatin remodeling: repressive effects on transcription. Cell.
99:443–446. 1999.
|
|
18.
|
Wang W, Cote J, Xue Y, et al: Purification
and biochemical heterogeneity of the mammalian SWI-SNF complex.
EMBO J. 15:5370–5382. 1996.PubMed/NCBI
|
|
19.
|
Decristofaro MF, Betz BL, Rorie CJ,
Reisman DN, Wang W and Weissman BE: Characterization of SWI/SNF
protein expression in human breast cancer cell lines and other
malignancies. J Cell Physiol. 186:136–145. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
20.
|
McCarty KS Jr, Miller LS, Cox EB, Konrath
J and McCarty KS Sr: Estrogen receptor analyses. Correlation of
biochemical and immunohistochemical methods using monoclonal
antireceptor antibodies. Arch Pathol Lab Med. 109:716–721.
1985.
|
|
21.
|
Bourgo RJ, Siddiqui H, Fox S, et al:
SWI/SNF deficiency results in aberrant chromatin organization,
mitotic failure, and diminished proliferative capacity. Mol Biol
Cell. 20:3192–3199. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22.
|
Reisman D, Glaros S and Thompson EA: The
SWI/SNF complex and cancer. Oncogene. 28:1653–1668. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
23.
|
Wiegand KC, Shah SP, Al-Agha OM, et al:
ARID1A mutations in endometriosis-associated ovarian carcinomas. N
Engl J Med. 363:1532–1543. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24.
|
Jones S, Wang TL, Shih Ie M, et al:
Frequent mutations of chromatin remodeling gene ARID1A in ovarian
clear cell carcinoma. Science. 330:228–231. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
25.
|
Varela I, Tarpey P, Raine K, et al: Exome
sequencing identifies frequent mutation of the SWI/SNF complex gene
PBRM1 in renal carcinoma. Nature. 469:539–542. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26.
|
Weissman B and Knudsen KE: Hijacking the
chromatin remodeling machinery: impact of SWI/SNF perturbations in
cancer. Cancer Res. 69:8223–8230. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27.
|
Roberts CW and Orkin SH: The SWI/SNF
complex - chromatin and cancer. Nat Rev Cancer. 4:133–142. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
28.
|
Biegel JA, Kalpana G, Knudsen ES, et al:
The role of INI1 and the SWI/SNF complex in the development of
rhabdoid tumors: meeting summary from the workshop on childhood
atypical teratoid/rhabdoid tumors. Cancer Res. 62:323–328.
2002.PubMed/NCBI
|
|
29.
|
Heinemann V, Boeck S, Hinke A, Labianca R
and Louvet C: Meta-analysis of randomized trials: evaluation of
benefit from gemcitabine-based combination chemotherapy applied in
advanced pancreatic cancer. BMC Cancer. 8:822008. View Article : Google Scholar
|
|
30.
|
El-Rayes BF and Philip PA: A review of
systemic therapy for advanced pancreatic cancer. Clin Adv Hematol
Oncol. 1:430–434. 2003.PubMed/NCBI
|
|
31.
|
Moore MJ, Goldstein D, Hamm J, et al:
Erlotinib plus gemcitabine compared with gemcitabine alone in
patients with advanced pancreatic cancer: a phase III trial of the
National Cancer Institute of Canada Clinical Trials Group. J Clin
Oncol. 25:1960–1966. 2007. View Article : Google Scholar
|
|
32.
|
Glaros S, Cirrincione GM, Muchardt C,
Kleer CG, Michael CW and Reisman D: The reversible epigenetic
silencing of BRM: implications for clinical targeted therapy.
Oncogene. 26:7058–7066. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
33.
|
Reisman DN, Sciarrotta J, Wang W,
Funkhouser WK and Weissman BE: Loss of BRG1/BRM in human lung
cancer cell lines and primary lung cancers: correlation with poor
prognosis. Cancer Res. 63:560–566. 2003.PubMed/NCBI
|
|
34.
|
Fukuoka J, Fujii T, Shih JH, et al:
Chromatin remodeling factors and BRM/BRG1 expression as prognostic
indicators in non-small cell lung cancer. Clin Cancer Res.
10:4314–4324. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
35.
|
Batsche E, Yaniv M and Muchardt C: The
human SWI/SNF subunit Brm is a regulator of alternative splicing.
Nat Struct Mol Biol. 13:22–29. 2006. View
Article : Google Scholar : PubMed/NCBI
|
|
36.
|
Martin TA, Harrison G, Mansel RE and Jiang
WG: The role of the CD44/ezrin complex in cancer metastasis. Crit
Rev Oncol Hematol. 46:165–186. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
37.
|
Xue Y, Canman JC, Lee CS, et al: The human
SWI/SNF-B chromatin-remodeling complex is related to yeast rsc and
localizes at kinetochores of mitotic chromosomes. Proc Natl Acad
Sci USA. 97:13015–13020. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
38.
|
Maitra A, Wistuba II, Washington C, et al:
High-resolution chromosome 3p allelotyping of breast carcinomas and
precursor lesions demonstrates frequent loss of heterozygosity and
a discontinuous pattern of allele loss. Am J Pathol. 159:119–130.
2001. View Article : Google Scholar
|
|
39.
|
Zabarovsky ER, Lerman MI and Minna JD:
Tumor suppressor genes on chromosome 3p involved in the
pathogenesis of lung and other cancers. Oncogene. 21:6915–6935.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
40.
|
Xia W, Nagase S, Montia AG, et al: BAF180
is a critical regulator of p21 induction and a tumor suppressor
mutated in breast cancer. Cancer Res. 68:1667–1674. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
41.
|
Harper JW, Adami GR, Wei N, Keyomarsi K
and Elledge SJ: The p21 Cdk-interacting protein Cip1 is a potent
inhibitor of G1 cyclin-dependent kinases. Cell. 75:805–816. 1993.
View Article : Google Scholar : PubMed/NCBI
|
|
42.
|
Duns G, Hofstra RM, Sietzema JG, et al:
Targeted exome sequencing in clear cell renal cell carcinoma tumors
suggests aberrant chromatin regulation as a crucial step in ccRCC
development. Hum Mutat. 33:1059–1062. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
43.
|
Nie Z, Xue Y, Yang D, et al: A specificity
and targeting subunit of a human SWI/SNF family-related
chromatin-remodeling complex. Mol Cell Biol. 20:8879–8888. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
44.
|
Sekine I, Sato M, Sunaga N, et al: The
3p21 candidate tumor suppressor gene BAF180 is normally expressed
in human lung cancer. Oncogene. 24:2735–2738. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
45.
|
Wang X, Nagl NG Jr, Flowers S, Zweitzig D,
Dallas PB and Moran E: Expression of p270 (ARID1A), a component of
human SWI/SNF complexes, in human tumors. Int J Cancer.
112:6362004. View Article : Google Scholar : PubMed/NCBI
|
|
46.
|
Nagl NG Jr, Zweitzig DR, Thimmapaya B,
Beck GR Jr and Moran E: The c-myc gene is a direct target of
mammalian SWI/SNF-related complexes during
differentiation-associated cell cycle arrest. Cancer Res.
66:1289–1293. 2006. View Article : Google Scholar : PubMed/NCBI
|