|
1
|
Li C, Miao R, Liu S, Wan Y, Zhang S, Deng
Y, Bi J, Qu K, Zhang J and Liu C: Down-regulation of miR-146b-5p by
long noncoding RNA MALAT1 in hepatocellular carcinoma promotes
cancer growth and metastasis. Oncotarget. 8:28683–28695.
2017.PubMed/NCBI
|
|
2
|
Lu PH, Chen MB, Liu YY, Wu MH, Li WT, Wei
MX, Liu CY and Qin SK: Identification of sphingosine kinase 1
(SphK1) as a primary target of icaritin in hepatocellular carcinoma
cells. Oncotarget. 8:22800–22810. 2017.PubMed/NCBI
|
|
3
|
He R, Gao L, Ma J, Peng Z, Zhou S, Yang L,
Feng Z, Dang Y and Chen G: The essential role of MTDH in the
progression of HCC: A study with immunohistochemistry, TCGA,
meta-analysis and in vitro investigation. Am J Transl Res.
9:1561–1579. 2017.
|
|
4
|
Mo Z, Zheng S, Lv Z, Zhuang Y, Lan X, Wang
F, Lu X, Zhao Y and Zhou S: Senescence marker protein 30 (SMP30)
serves as a potential prognostic indicator in hepatocellular
carcinoma. Sci Rep. 6:393762016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Huang WT, Wang HL, Yang H, Ren FH, Luo YH,
Huang CQ, Liang YY, Liang HW, Chen G and Dang YW: Lower expressed
miR-198 and its potential targets in hepatocellular carcinoma: A
clinicopathological and in silico study. Onco Targets Ther.
9:5163–5180. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Xu Y, Qi Y, Luo J, Yang J, Xie Q, Deng C,
Su N, Wei W, Shi D, Xu F, et al: Hepatitis B Virus X protein
stimulates proliferation, wound closure and inhibits apoptosis of
HuH-7 cells via CDC42. Int J Mol Sci. 18:182017. View Article : Google Scholar
|
|
8
|
Yu C, Cao Q, Chen P, Yang S, Gong X, Deng
M, Ruan B and Li L: Tissue transglutaminase 2 exerts a
tumor-promoting role in hepatitis B virus-related hepatocellular
carcinoma. Tumour Biol. 37:16269–16274. 2016. View Article : Google Scholar
|
|
9
|
Gong X, Wei W, Chen L, Xia Z and Yu C:
Comprehensive analysis of long non-coding RNA expression profiles
in hepatitis B virus-related hepatocellular carcinoma. Oncotarget.
7:42422–42430. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Dengler M, Staufer K, Huber H, Stauber R,
Bantel H, Weiss KH, Starlinger P, Pock H, Klöters-Plachky P,
Gotthardt DN, et al: Soluble Axl is an accurate biomarker of
cirrhosis and hepatocellular carcinoma development: Results from a
large scale multicenter analysis. Oncotarget. 8:46234–46248.
2017.PubMed/NCBI
|
|
11
|
Ye Q, Qian BX, Yin WL, Wang FM and Han T:
Association between the HFE C282Y, H63D polymorphisms and the risks
of non-alcoholic fatty liver disease, liver cirrhosis and
hepatocellular carcinoma: An updated systematic review and
meta-analysis of 5,758 cases and 14,741 controls. PLoS One.
11:e01634232016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Joshi K, Kohli A, Manch R and Gish R:
Alcoholic liver disease: High risk or low risk for developing
hepatocellular carcinoma? Clin Liver Dis. 20:563–580. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Fairman J, Liu KH and Menne S: Prevention
of liver tumor formation in woodchucks with established
hepatocellular carcinoma by treatment with cationic liposome-DNA
complexes. BMC Cancer. 17:1722017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Hectors SJ, Wagner M, Bane O, Besa C,
Lewis S, Remark R, Chen N, Fiel MI, Zhu H, Gnjatic S, et al:
Quantification of hepatocellular carcinoma heterogeneity with
multiparametric magnetic resonance imaging. Sci Rep. 7:24522017.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Xie X, Pan J, Wei L, Wu S, Hou H, Li X and
Chen W: Gene expression profiling of microRNAs associated with UCA1
in bladder cancer cells. Int J Oncol. 48:1617–1627. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sun XJ, Wang Q, Guo B, Liu XY and Wang B:
Identification of skin-related lncRNAs as potential biomarkers that
involved in Wnt pathways in keloids. Oncotarget. 8:34236–34244.
2017.PubMed/NCBI
|
|
17
|
Ma G, Tang M, Wu Y, Xu X, Pan F and Xu R:
LncRNAs and miRNAs: Potential biomarkers and therapeutic targets
for prostate cancer. Am J Transl Res. 8:5141–5150. 2016.
|
|
18
|
Wilusz JE: Long noncoding RNAs: Re-writing
dogmas of RNA processing and stability. Biochim Biophys Acta.
1859:128–138. 2016. View Article : Google Scholar
|
|
19
|
Yuan X, Wang J, Tang X, Li Y, Xia P and
Gao X: Berberine ameliorates nonalcoholic fatty liver disease by a
global modulation of hepatic mRNA and lncRNA expression profiles. J
Transl Med. 13:242015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wei Y and Niu B: Role of MALAT1 as a
prognostic factor for survival in various cancers: A systematic
review of the literature with meta-analysis. Dis Markers.
2015:1646352015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wei Y and Zhang X: Transcriptome analysis
of distinct long non-coding RNA transcriptional fingerprints in
lung adenocarcinoma and squamous cell carcinoma. Tumour Biol.
37:16275–16285. 2016. View Article : Google Scholar
|
|
22
|
Li Y, Li W, Liang B, Li L, Wang L, Huang
H, Guo S, Wang Y, He Y, Chen L, et al: Identification of cancer
risk lncRNAs and cancer risk pathways regulated by cancer risk
lncRNAs based on genome sequencing data in human cancers. Sci Rep.
6:392942016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Li SP, Xu HX, Yu Y, He JD, Wang Z, Xu YJ,
Wang CY, Zhang HM, Zhang RX, Zhang JJ, et al: LncRNA HULC enhances
epithelial-mesenchymal transition to promote tumorigenesis and
metastasis of hepatocellular carcinoma via the miR-200a-3p/ZEB1
signaling pathway. Oncotarget. 7:42431–42446. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Huang ZL, Chen RP, Zhou XT, Zhan HL, Hu
MM, Liu B, Wu GD and Wu LF: Long non-coding RNA MEG3 induces cell
apoptosis in esophageal cancer through endoplasmic reticulum
stress. Oncol Rep. 37:3093–3099. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang Y, Zhang Q, Zhang M, Yuan M, Wang Z,
Zhang J, Zhou X, Zhang Y, Lin F, Na H, et al: DC - SIGNR by
influencing the lncRNA HNRNPKP2 upregulates the expression of CXCR4
in gastric cancer liver metastasis. Mol Cancer. 16:782017.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Li J, Zhang Z, Xiong L, Guo C, Jiang T,
Zeng L, Li G and Wang J: SNHG1 lncRNA negatively regulates
miR-199a-3p to enhance CDK7 expression and promote cell
proliferation in prostate cancer. Biochem Biophys Res Commun.
487:146–152. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Xue F, Liu Y, Chu H, Wen Y, Yan L, Tang Q,
Xiao E, Zhang D and Zhang H: eIF5A2 is an alternative pathway for
cell proliferation in cetuximab-treated epithelial hepatocellular
carcinoma. Am J Transl Res. 8:4670–4681. 2016.PubMed/NCBI
|
|
28
|
Deng LL, Chi YY, Liu L, Huang NS, Wang L
and Wu J: LINC00978 predicts poor prognosis in breast cancer
patients. Sci Rep. 6:379362016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Guo S, Chen W, Luo Y, Ren F, Zhong T, Rong
M, Dang Y, Feng Z and Chen G: Clinical implication of long
non-coding RNA NEAT1 expression in hepatocellular carcinoma
patients. Int J Clin Exp Pathol. 8:5395–5402. 2015.PubMed/NCBI
|
|
30
|
Zhu P, Wang Y, Wu J, Huang G, Liu B, Ye B,
Du Y, Gao G, Tian Y, He L, et al: LncBRM initiates YAP1 signalling
activation to drive self-renewal of liver cancer stem cells. Nat
Commun. 7:136082016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Cao C, Sun J, Zhang D, Guo X, Xie L, Li X,
Wu D and Liu L: The long intergenic noncoding RNA UFC1, a target of
MicroRNA 34a, interacts with the mRNA stabilizing protein HuR to
increase levels of beta-catenin in HCC cells. Gastroenterology.
148:415–426. e4182015. View Article : Google Scholar
|
|
32
|
Zhou M, Zhang XY and Yu X: Overexpression
of the long non-coding RNA SPRY4-IT1 promotes tumor cell
proliferation and invasion by activating EZH2 in hepatocellular
carcinoma. Biomed Pharmacother. 85:348–354. 2017. View Article : Google Scholar
|
|
33
|
Xu LM, Chen L, Li F, Zhang R, Li Z-Y, Chen
F-F and Jiang X-D: Over-expression of the long non-coding RNA
HOTTIP inhibits glioma cell growth by BRE. J Exp Clin Cancer Res.
35:1622016. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhang X, Tang W, Li R, He R, Gan T, Luo Y,
Chen G and Rong M: Downregulation of microRNA-132 indicates
progression in hepatocellular carcinoma. Exp Ther Med.
12:2095–2101. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Jin W, Chen L, Cai X, Zhang Y, Zhang J, Ma
D, Cai X, Fu T, Yu Z, Yu F, et al: Long non-coding RNA TUC338 is
functionally involved in sorafenib-sensitized hepatocarcinoma cells
by targeting RASAL1. Oncol Rep. 37:273–280. 2017. View Article : Google Scholar
|
|
36
|
Yin X, Zheng SS, Zhang L, Xie XY, Wang Y,
Zhang BH, Wu W, Qiu S and Ren ZG: Identification of long noncoding
RNA expression profile in oxaliplatin-resistant hepatocellular
carcinoma cells. Gene. 596:53–88. 2017. View Article : Google Scholar
|
|
37
|
Lin L, Zhang YD, Chen ZY, Chen Y and Ren
CP: The clinicopathological significance of miR-149 and PARP-2 in
hepatocellular carcinoma and their roles in chemo/radiotherapy.
Tumour Biol. 37:12339–12346. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Colombo T, Farina L, Macino G and Paci P:
VT1: A rising star among oncogenic long noncoding RNAs. BioMed Res
Int. 2015:3042082015. View Article : Google Scholar
|
|
39
|
Liu FT, Xue QZ, Zhang Y, Hao TF, Luo HL
and Zhu PQ: Long non-coding RNA HOXA transcript at the distal tip
as a putative biomarker of metastasis and prognosis: A
meta-analysis. Clin Lab. 62:2091–2098. 2016. View Article : Google Scholar
|
|
40
|
Yang Y, Qian J, Xiang Y, Chen Y and Qu J:
The prognostic value of long noncoding RNA HOTTIP on clinical
outcomes in breast cancer. Oncotarget. 8:6833–6844. 2017.
|
|
41
|
Chen Z, He A, Wang D, Liu Y and Huang W:
Long noncoding RNA HOTTIP as a novel predictor of lymph node
metastasis and survival in human cancer: A systematic review and
meta-analysis. Oncotarget. 8:14126–14132. 2017.
|
|
42
|
Chen X, Han H, Li Y, Zhang Q, Mo K and
Chen S: Upregulation of long noncoding RNA HOTTIP promotes
metastasis of esophageal squamous cell carcinoma via induction of
EMT. Oncotarget. 7:84480–84485. 2016.PubMed/NCBI
|
|
43
|
Li Z, Zhao L and Wang Q: Overexpression of
long non-coding RNA HOTTIP increases chemoresistance of
osteosarcoma cell by activating the Wnt/β-catenin pathway. Am J
Transl Res. 8:2385–2393. 2016.
|
|
44
|
Ye H, Liu K and Qian K: Overexpression of
long noncoding RNA HOTTIP promotes tumor invasion and predicts poor
prognosis in gastric cancer. Onco Targets Ther. 9:2081–2088.
2016.PubMed/NCBI
|
|
45
|
Tsang FH, Au SL, Wei L, Fan DN, Lee JM,
Wong CC, Ng IO and Wong CM: Long non-coding RNA HOTTIP is
frequently up-regulated in hepatocellular carcinoma and is targeted
by tumour suppressive miR-125b. Liver Int. 35:1597–1606. 2015.
View Article : Google Scholar
|
|
46
|
Xu X, Wang X, Fu B, Meng L and Lang B:
Differentially expressed genes and microRNAs in bladder carcinoma
cell line 5637 and T24 detected by RNA sequencing. Int J Clin Exp
Pathol. 8:12678–12687. 2015.
|
|
47
|
Subramanian Y, Kaliyappan K and
Ramakrishnan KS: Facile hydrothermal synthesis and characterization
of Co2GeO4/r-GO@C ternary nanocomposite as
negative electrode for Li-ion batteries. J Colloid Interface Sci.
498:76–84. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Fu L, Xu Y, Hou Y, Qi X, Zhou L, Liu H,
Luan Y, Jing L, Miao Y, Zhao S, et al: Proteomic analysis indicates
that mitochondrial energy metabolism in skeletal muscle tissue is
negatively correlated with feed efficiency in pigs. Sci Rep.
7:452912017. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Sang Y, Zhou F, Wang D, Bi X, Liu X, Hao
Z, Li Q and Zhang W: Up-regulation of long non-coding HOTTIP
functions as an oncogene by regulating HOXA13 in non-small cell
lung cancer. Am J Transl Res. 8:2022–2032. 2016.PubMed/NCBI
|
|
50
|
Dai J, Wu H, Zhang Y, Gao K, Hu G, Guo Y,
Lin C and Li X: Negative feedback between TAp63 and Mir-133b
mediates colorectal cancer suppression. Oncotarget. 7:87147–87160.
2016.PubMed/NCBI
|
|
51
|
Wu H, Zhou J, Zeng C, Wu D, Mu Z, Chen B,
Xie Y, Ye Y and Liu J: Curcumin increases exosomal TCF21 thus
suppressing exosome-induced lung cancer. Oncotarget. 7:87081–87090.
2016.PubMed/NCBI
|
|
52
|
Bornstein S, Schmidt M, Choonoo G, Levin
T, Gray J, Thomas CR Jr, Wong M and McWeeney S: IL-10 and integrin
signaling pathways are associated with head and neck cancer
progression. BMC Genomics. 17:382016. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Zeng JH, Xiong DD, Pang YY, Zhang Y, Tang
RX, Luo DZ and Chen G: Identification of molecular targets for
esophageal carcinoma diagnosis using miRNA-seq and RNA-seq data
from The Cancer Genome Atlas: A study of 187 cases. Oncotarget.
8:35681–35699. 2017.PubMed/NCBI
|
|
54
|
Rhodes DR, Kalyana-Sundaram S, Mahavisno
V, Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ,
Kincead-Beal C, Kulkarni P, et al: Oncomine 3.0: Genes, pathways,
and networks in a collection of 18,000 cancer gene expression
profiles. Neoplasia. 9:166–180. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Adler P, Kolde R, Kull M, Tkachenko A,
Peterson H, Reimand J and Vilo J: Mining for coexpression across
hundreds of datasets using novel rank aggregation and visualization
methods. Genome Biol. 10:R1392009. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Tang W, Liao Z and Zou Q: Which
statistical significance test best detects oncomiRNAs in cancer
tissues? An exploratory analysis. Oncotarget. 7:85613–85623.
2016.PubMed/NCBI
|
|
57
|
Li CQ, Huang GW, Wu ZY, Xu YJ, Li XC, Xue
YJ, Zhu Y, Zhao JM, Li M, Zhang J, et al: Integrative analyses of
transcriptome sequencing identify novel functional lncRNAs in
esophageal squamous cell carcinoma. Oncogenesis. 6:e2972017.
View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Dinh TA, Vitucci EC, Wauthier E, Graham
RP, Pitman WA, Oikawa T, Chen M, Silva GO, Greene KG, Torbenson MS,
et al: Comprehensive analysis of The Cancer Genome Atlas reveals a
unique gene and non-coding RNA signature of fibrolamellar
carcinoma. Sci Rep. 7:446532017. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Seiler R, Black PC, Thalmann G, Stenzl A
and Todenhöfer T: Is The Cancer Genome Atlas (TCGA) bladder cancer
cohort representative of invasive bladder cancer? Urol Oncol.
35:458.e1–458.e7. 2017. View Article : Google Scholar
|
|
60
|
Gao H, Wang H and Yang W: Identification
of key genes and construction of microRNA-mRNA regulatory networks
in multiple myeloma by integrated multiple GEO datasets using
bioinformatics analysis. Int J Hematol. 106:99–107. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Tan W, Song Y, Mo C, Jiang S and Wang Z:
Analysis of gene expression profile microarray data in complex
regional pain syndrome. Mol Med Rep. 16:3371–3378. 2017.PubMed/NCBI
|
|
62
|
Mirsafian H, Ripen AM, Leong WM, Chear CT,
Bin Mohamad S and Merican AF: Transcriptome profiling of monocytes
from XLA patients revealed the innate immune function dysregulation
due to the BTK gene expression deficiency. Sci Rep. 7:68362017.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
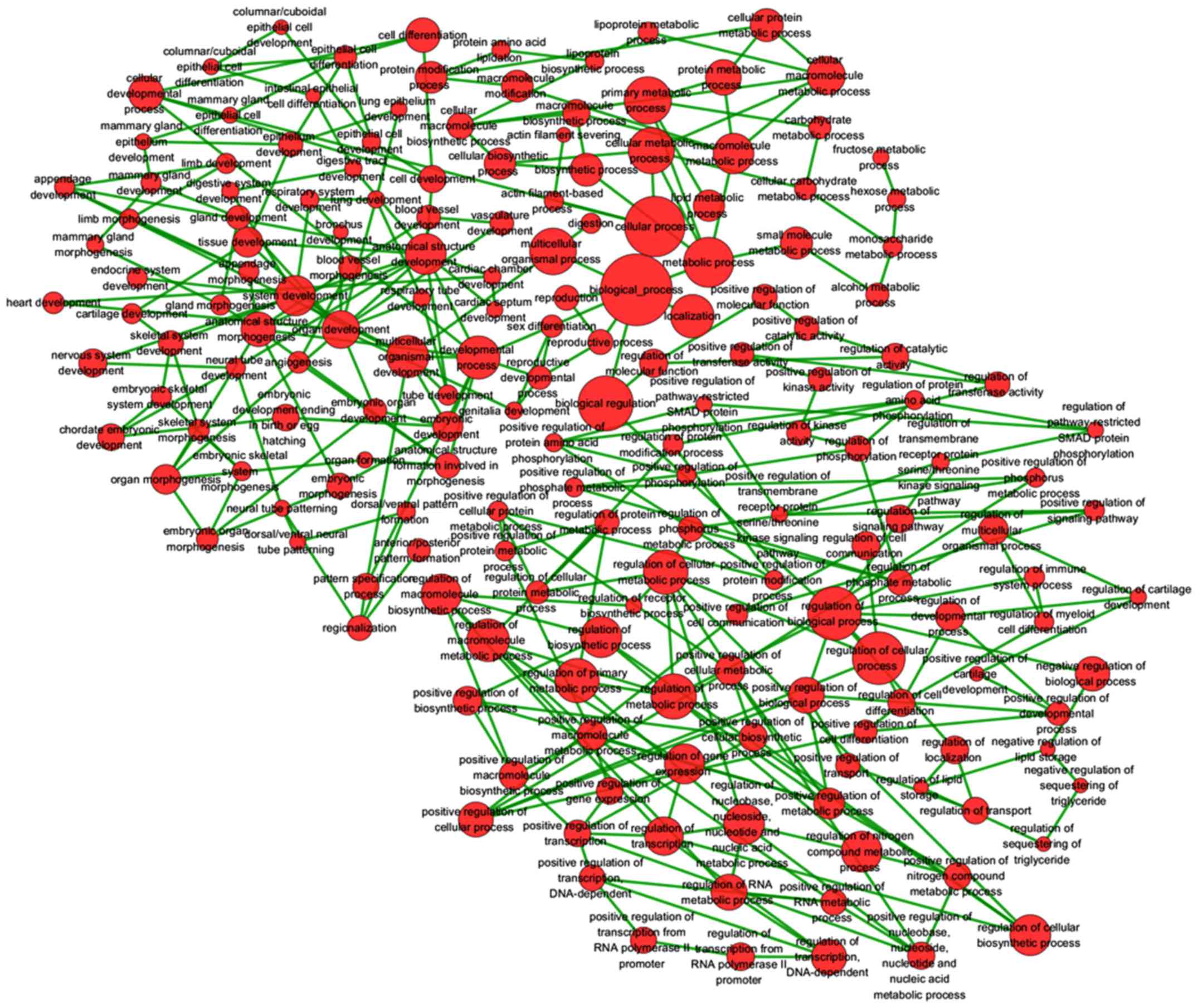
Zhao Z, Bai J, Wu A, Wang Y, Zhang J, Wang
Z, Li Y, Xu J and Li X: Co-LncRNA: Investigating the lncRNA
combinatorial effects in GO annotations and KEGG pathways based on
human RNA-Seq data. Database (Oxford). Sep 10–2015.Epub ahead of
print. View Article : Google Scholar
|
|
64
|
Franceschini A, Szklarczyk D, Frankild S,
Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering
C, et al: STRING v9.1: Protein-protein interaction networks, with
increased coverage and integration. Nucleic Acids Res.
41D:D808–D815. 2013.
|
|
65
|
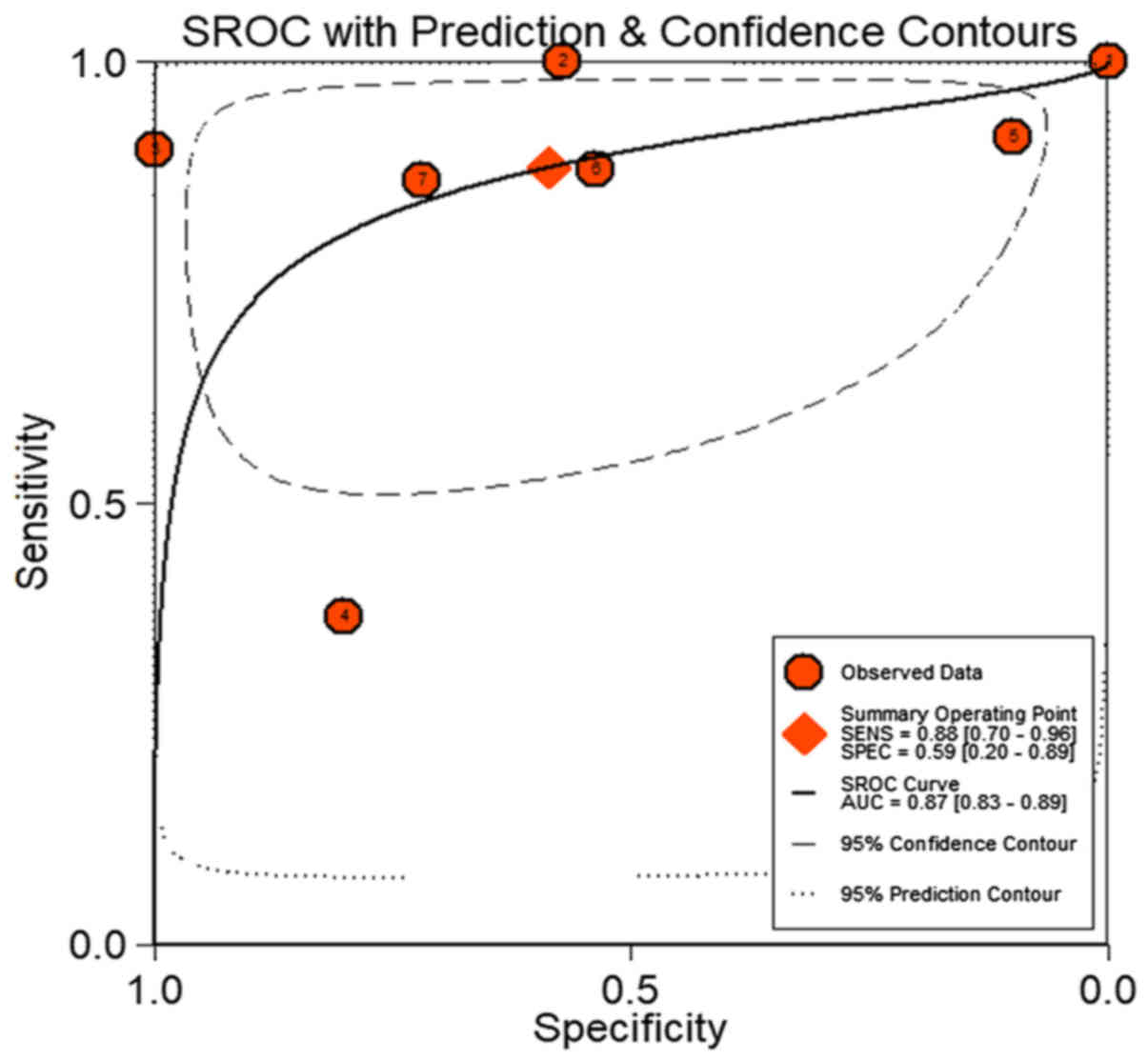
Zamora J, Abraira V, Muriel A, Khan K and
Coomarasamy A: Meta-DiSc: A software for meta-analysis of test
accuracy data. BMC Med Res Methodol. 6:312006. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Shi X, Sun M, Liu H, Yao Y and Song Y:
Long non-coding RNAs: A new frontier in the study of human
diseases. Cancer Lett. 339:159–166. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Xiong DD, Feng ZB, Cen WL, Zeng JJ, Liang
L, Tang RX, Gan XN, Liang HW, Li ZY, Chen G, et al: The clinical
value of lncRNA NEAT1 in digestive system malignancies: A
comprehensive investigation based on 57 microarray and RNA-seq
datasets. Oncotarget. 8:17665–17683. 2017.PubMed/NCBI
|
|
68
|
Lu S, Zhou J, Sun Y, Li N, Miao M, Jiao B
and Chen H: The noncoding RNA HOXD-AS1 is a critical regulator of
the metastasis and apoptosis phenotype in human hepatocellular
carcinoma. Mol Cancer. 16:1252017. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Xu LC, Chen QN, Liu XQ, Wang XM, Chang QM,
Pan Q, Wang L and Wang YL: Up-regulation of LINC00161 correlates
with tumor migration and invasion and poor prognosis of patients
with hepatocellular carcinoma. Oncotarget. 8:56168–56173.
2017.PubMed/NCBI
|
|
70
|
Wang Y, Hu Y, Wu G, Yang Y, Tang Y, Zhang
W, Wang K, Liu Y, Wang X and Li T: Long noncoding RNA PCAT-14
induces proliferation and invasion by hepatocellular carcinoma
cells by inducing methylation of miR-372. Oncotarget.
8:34429–34441. 2017.PubMed/NCBI
|
|
71
|
Quagliata L, Matter MS, Piscuoglio S,
Arabi L, Ruiz C, Procino A, Kovac M, Moretti F, Makowska Z,
Boldanova T, et al: Long noncoding RNA HOTTIP/HOXA13 expression is
associated with disease progression and predicts outcome in
hepatocellular carcinoma patients. Hepatology. 59:911–923. 2014.
View Article : Google Scholar :
|
|
72
|
Ge Y, Yan X, Jin Y, Yang X, Yu X, Zhou L,
Han S, Yuan Q and Yang M: MiRNA-192 [corrected] and miRNA-204
directly suppress lncRNA HOTTIP and interrupt GLS1-mediated
glutaminolysis in hepatocellular carcinoma. PLoS Genet.
11:e10057262015. View Article : Google Scholar
|
|
73
|
Swets JA: Measuring the accuracy of
diagnostic systems. Science. 240:1285–1293. 1988. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Lu G, Zhang G, Zheng X, Zeng Y, Xu Z, Zeng
W and Wang K: c9, t11- conjugated linoleic acid induces HCC cell
apoptosis and correlation with PPAR-γ signaling pathway. Am J
Transl Res. 7:2752–2763. 2015.
|
|
75
|
Li J, Huang Q, Long X, Zhang J, Huang X,
Aa J, Yang H, Chen Z and Xing J: CD147 reprograms fatty acid
metabolism in hepatocellular carcinoma cells through
Akt/mTOR/SREBP1c and P38/PPARα pathways. J Hepatol. 63:1378–1389.
2015. View Article : Google Scholar : PubMed/NCBI
|