Introduction
MicroRNAs (miRNAs) are a very important regulatory
RNA molecules that influence gene function by inhibiting
translation or inducing the degradation of target mRNA (1). Importantly, expression imbalances
caused by miRNA regulation can contribute to the pathogenesis of
various tumors (2). For example,
miR-21 has been implicated in breast, colon, prostate and thyroid
cancers, among others (2). The
potent effect of miRNAs on disease processes indicates a need for
studying their expression and function, with the aim of
establishing a solid foundation for the clinical treatment of
tumors (3–5).
Another miRNA, miR-125b, has been shown to exhibit
abnormal expression in a variety of tumor tissues; however, its
effects are not consistent across tissue types (6–8). In
prostate cancer, the high expression of miR-125b can promote tumor
proliferation (9), but, in breast
cancer, miR-125b can significantly inhibit tumor proliferation
(10). A certain study found that
this miRNA exhibited altered expression in gastric cancer according
to disease progression (11).
However, no other reports on miR-125b expression and its effect on
gastric cancer have been identified.
Gastric cancer occurs with high incidence in Asian
populations and thus has been an important focus of cancer research
(12). In this study, real-time
PCR was used to detect miR-125b expression in 50 cases of gastric
cancer tissues and corresponding adjacent normal tissues. To
determine the effect of miR-125b on the proliferation and apoptosis
of gastric cancer cells, the miR-125b mimic was transfected into
the gastric cancer cell line, HGC-27. Our results reveal the
potential functions of miR-125b in gastric cancer and may pave the
way for its application in clinical diagnosis and treatment.
Materials and methods
Specimens
Specimens of both gastric carcinoma tissues and
corresponding adjacent normal tissues (>5 cm away from cancer
tissues) were collected from 50 patients who received surgical
resection of gastric cancer from January 2010 to December 2011 in
Xijing Hospital, the Fourth Military Medical University, Xi’an,
China. Gastric cancer diagnoses were confirmed by post-operative
pathological biopsy. No patients had received pre-operative
radiotherapy, chemotherapy or endocrine therapy. The age range of
the patients was 32–70 years and the mean age was 53.9±9.6 years.
During surgery, the cancer and adjacent normal tissues were cut,
100 mg of which were washed with 1X PBS, placed in cryovials and
kept in a refrigerator at −80°C.
Cell culture and transfection
The HGC-27 human gastric cancer cells were provided
by the Cell Bank of the Shanghai Institute of Cell Biology (Chinese
Academy of Sciences, Shanghai, China) and placed in DMEM medium
(Gibco-BRL, Carlsbad, CA, USA) containing 10% FBS (PAA
Laboratories, Pasching, Austria), 1×105 IU/l penicillin
and 1×105 IU/l streptomycin and conventionally cultured
in an incubator at 37°C, 5% CO2. One day prior to
transfection, the cell culture medium was transferred at a density
of 1×104 cells/ml to prepare for transfection during the
logarithmic growth phase. Cells were washed once with serum-free
medium, then resuspended in complete medium without antibiotics at
a cell concentration of 4×105 in a volume of 1,500 μl.
Suspensions were transferred onto a 6-well plate and placed in DMEM
overnight without serum and antibiotics. Transfections were
performed with DharmaconFECT1 (Dharmacon, Inc., Lafayette, CO,
USA), using 100 pmol mimic-miR-125b (Dharmacon Inc.) per well in
the transfection group and 100 pmol negative control (Dharmacon
Inc.) per well in the control group. After 6 h of culture at 37°C,
5% CO2, 500 μl of 10% FBS-supplemented medium were added
and the cells were continuously cultured for 72 h.
Detecting cell proliferation with
CCK-8
Suspensions containing 5×104 cells/ml
were prepared and 100 μl of suspension were added to a 96-well
plate. Cells were continuously cultured at 37°C. At 24, 48 and 72 h
after culture, 10 μl CCK-8 solution (Dojindo, Kunamoto, Japan) were
added to each well before an additional incubation at 37°C for 4 h.
The optical density of the suspensions was then determined by
absorbance at 450 nm.
Detecting cell apoptosis with
FITC-Annexin V
The cells were collected into 10-ml centrifuge tubes
and washed with incubation buffer. Following centrifugation at 800
rpm for 5 min, the cells were resuspended in 100 μl FITC-Annexin
V-PI (BD Biosciences, Franklin Lakes, NJ, USA) labeling solution
and incubated at room temperature in the dark for 15 min. Following
another centrifugation at 800 rpm for 5 min, the cells were washed
with incubation buffer. Fluorescent dye solution was added and the
cells were incubated at 4°C for 20 min in the dark with occasional
mixing. The excitation wavelength for flow cytometry was 488 nm;
FITC was detected at 515 nm and PI was detected at >560 nm. Data
were plotted in bivariate flow cytometry scatter plots: the lower
left quadrant (FITC−/PI−) shows living cells;
the upper right quadrant (FITC+/PI+) shows
non-living cells, namely, necrotic cells; the lower right quadrant
(FITC+/PI−) indicates apoptotic cells.
Quantitative real-time PCR
TRIzol reagent (Invitrogen Life Technologies,
Carlsbad, CA, USA) was used according to the manufacturer’s
instructions to extract the total cellular RNA from the tissue
specimens; RNA purity and concentration were determined by UV
spectrophotometry. MiRNA 1st-strand cDNA was synthesized by
stem-loop primer reverse transcription reaction (Promega Corp.,
Madison, WI, USA). The following primer sequences were used:
Hsa-miR-125b RT primer,
5′-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACTCACAAG-3′;
Hsa-miR-125b sense primer, 5′-GGATTCCCTGAGACCCTAAC-3′; Hsa-miR-125b
antisense primer, 5′-GTGCAGGGTCCGAGGT-3′; U6 RT primer,
5′-AAAATATGGAACGCTTCACGAATTTG-3′; U6 sense primer,
5′-CTCGCTTCGGCAGCACATATACT- 3′; U6 antisense primer,
5′-ACGCTTCACGAATTTGCGTGTC-3′. The real-time quantitative PCR kit
(Takara Bio., Inc., Shiga, Japan) was used to facilitate the
reactions. Reactions were performed under the following thermal
cycling conditions: 94°C for 20 sec; and 40 cycles of 94°C for 30
sec, 60°C for 45 sec. Samples were amplified in triplicate and
reactions were performed on a Bio-Rad IQ5 PCR instrument (Bio-Rad,
Hercules, CA, USA) which was also used to analyze the results.
Statistical analysis
SPSS17.0 statistical software was applied to
statistical analysis. Measurement data are expressed as the means ±
standard deviation (SD). A two-independent-sample t-test was used
to compare the miR-125b expression between the groups. The above
analyses were performed with two-sided tests, with a test level (α)
of 0.05 and P<0.05 was considered to indicate a statistically
significant difference.
Results
MiR-125b expression in gastric cancer and
adjacent non-cancerous tissues
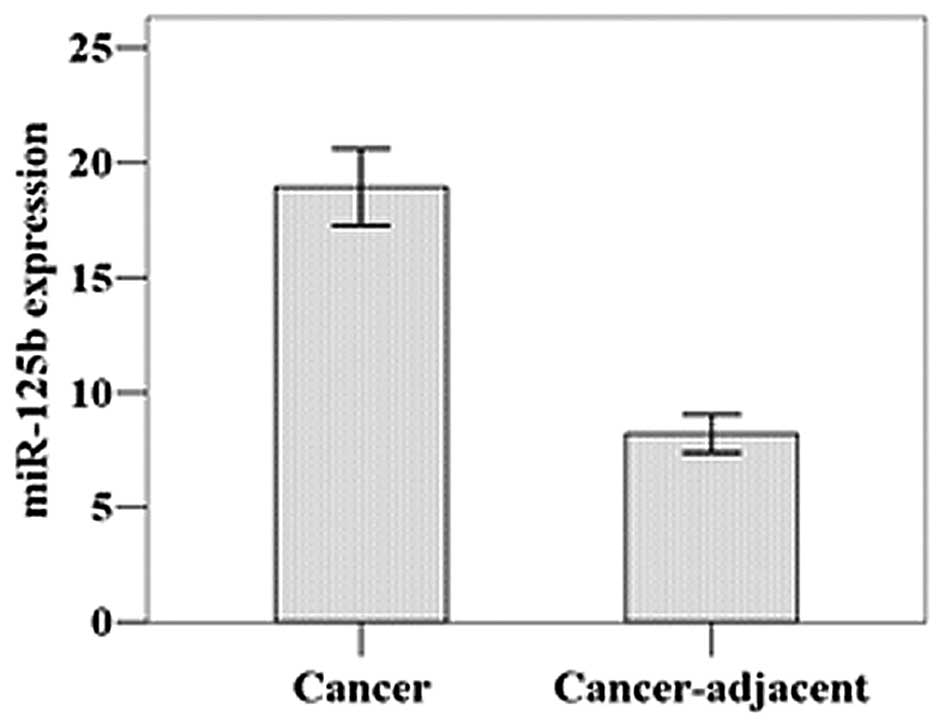
As shown by real-time PCR, the relative mean
miR-125b expression level was 18.94±5.96 in the gastric cancer
tissues and 8.20±2.91 in the adjacent normal tissues (Fig. 1). This difference was statistically
significant (t=11.452, P=0.001).
Effect of miR-125b overexpression on
HGC-27 cell proliferation
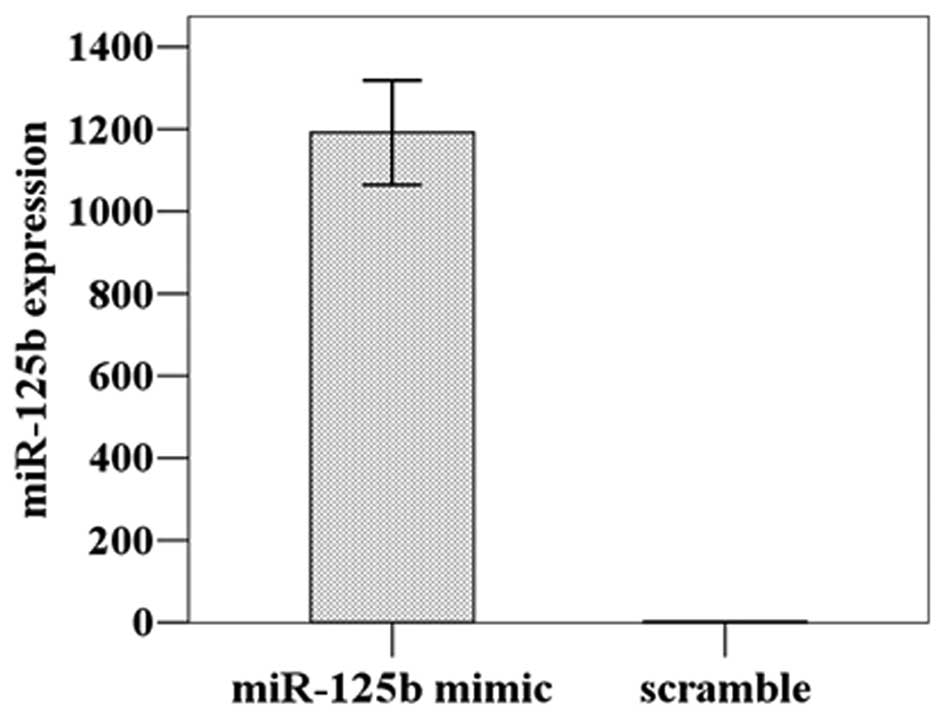
The miR-125b expression level was detected at
1192.00±109.77 in the HGC-27 cells transfected with miR-125b mimic,
a significant increase of approximately 1000-fold compared to the
cells treated with the scrambled miRNA (1.29±0.18) (t=18.788,
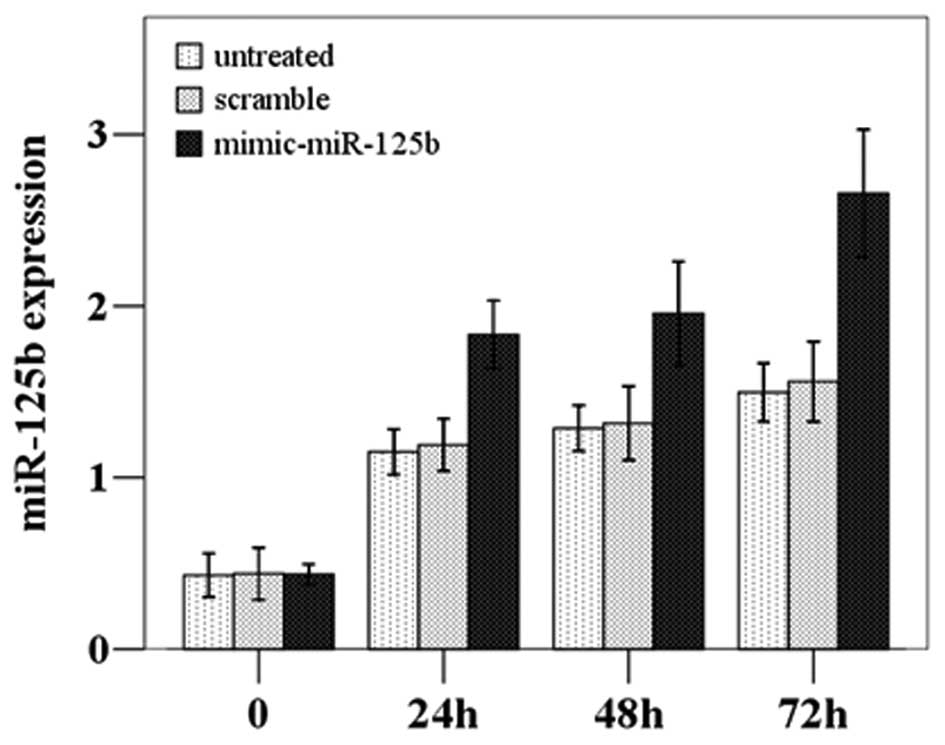
P=0.001); thus, the transfection was successful (Fig. 2). The subsequent CCK8 proliferation
assay demonstrated that the proliferation was significantly
increased in the miR-125b-overexpressing cells compared to the
untreated and scramble-treated cells (F=23.095, P=0.002) (Fig. 3).
Effect of miR-125b overexpression on
HGC-27 cell apoptosis
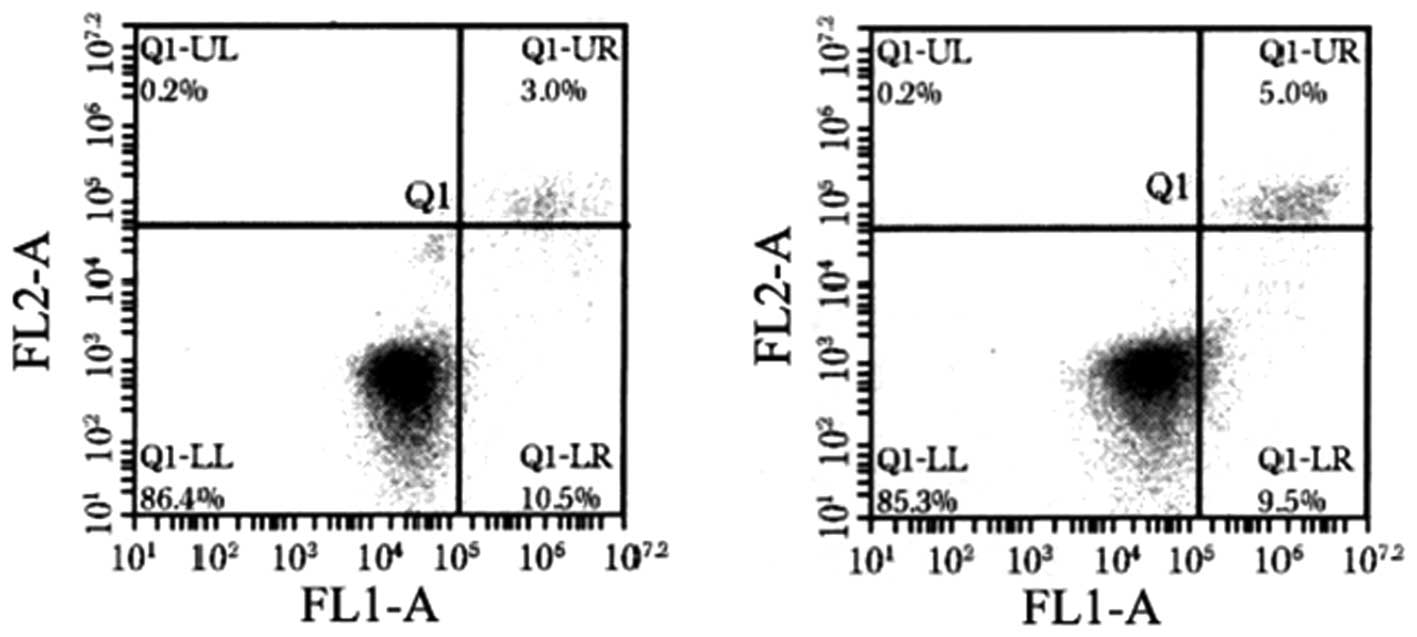
To determine whether the apoptosis of HGC-27 cells
is also dysregulated when miR-125b is overexpressed, the cells were
analyzed by flow cytometry for Annexin V. The proportion of Annexin
V-positive cells was decreased when miR-125b was overexpressed,
which can be observed mainly as the percentage of cells in early
apoptosis (lower right quadrant, Fig.
4). Furthermore, in HGC-27 cells expressing the miR-125b mimic,
the early and late apoptotic rates were different compared to the
scramble-treated and untreated cells (P<0.05, Table I).
 | Table IMean percentage of early and late
apoptotic HGC-27 cells from three independent experiments. |
Table I
Mean percentage of early and late
apoptotic HGC-27 cells from three independent experiments.
| Treatment | Early apoptotic
cells | Late apoptotic
cells |
|---|
| Mimic-miR-125b | 6.47±0.39 | 3.94±0.20 |
| Scramble | 9.04±0.70a | 3.09±0.09a |
| Untreated | 10.64±1.21a,b | 4.34±0.07a,b |
| F-value | 18.909 | 72.337 |
| P-value | 0.003 | 0.001 |
Discussion
The potent ability of miRNAs to regulate a variety
of physiological and pathological processes is well-accepted as a
key component of gene expression regulatory networks in eukaryotes.
Indeed, these molecules exert effects on cell proliferation and
apoptosis, organismal growth and development, hematopoiesis and
organ formation (13). Recent
studies on tumor-associated miRNAs have uncovered roles for a
number of miRNA molecules in tumorigenesis, as well as in cancer
diagnosis and prognosis (2,14).
Our current study on the expression of miR-125b
(previously implicated in other tumors types), in gastric cancer
confirms the results from a previous study, demonstrating its
dysregulation in this type of cancer (11). Furthermore, the overexpression of
this miRNA in gastric cancer cells in vitro revealed the
affects on cellular proliferation and apoptosis; proliferation was
increased and apoptosis decreased when miR-125b was overexpressed.
The dysregulation of these cellular processes by miR-125b may
promote tumor initiation and progression. These results indicate
that miR-125b plays an important role in the pathogenesis of
gastric cancer. Further studies are required to determine whether
miR-125b represents a potential oncogenic target for the diagnosis
and treatment of gastric cancer.
Acknowledgements
This study was supported by the National Science
Foundation of China (grant no. 81000164).
References
|
1
|
Iorio MV and Croce CM: MicroRNAs in
cancer: small molecules with a huge impact. J Clin Oncol.
27:5848–5856. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Nikitina EG, Urazova LN and Stegny VN:
MicroRNAs and human cancer. Exp Oncol. 34:2–8. 2012.
|
|
3
|
Esquela-Kerscher A and Slack FJ:
Oncomirs-microRNAs with a role in cancer. Nat Rev Cancer.
6:259–269. 2006. View
Article : Google Scholar
|
|
4
|
Hobert O: Gene regulation by transcription
factors and microRNAs. Science. 319:1785–1786. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kota J, Chivukula RR, O’Donnell KA, et al:
Therapeutic microRNA delivery suppresses tumorigenesis in a murine
liver cancer model. Cell. 137:1005–1017. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Xu N, Brodin P, Wei T, et al: MiR-125b, a
microRNA downregulated in psoriasis, modulates keratinocyte
proliferation by targeting FGFR2. J Invest Dermatol. 131:1521–1529.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Shi XB, Xue L, Ma AH, Tepper CG, Kung HJ
and White RW: miR-125b promotes growth of prostate cancer xenograft
tumor through targeting pro-apoptotic genes. Prostate. 71:538–549.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Glud M, Rossing M, Hother C, et al:
Downregulation of miR-125b in metastatic cutaneous malignant
melanoma. Melanoma Res. 20:479–484. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shi XB, Xue L, Yang J, et al: An
androgen-regulated miRNA suppresses Bak1 expression and induces
androgen-independent growth of prostate cancer cells. Proc Natl
Acad Sci USA. 104:19983–19988. 2007. View Article : Google Scholar
|
|
10
|
Scott GK, Goga A, Bhaumik D, Berger CE,
Sullivan CS and Benz CC: Coordinate suppression of ERBB2 and ERBB3
by enforced expression of micro-RNA miR-125a or miR-125b. J Biol
Chem. 282:1479–1486. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ueda T, Volinia S, Okumura H, et al:
Relation between microRNA expression and progression and prognosis
of gastric cancer: a microRNA expression analysis. Lancet Oncol.
11:136–146. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lochhead P and El-Omar EM: Helicobacter
pylori infection and gastric cancer. Best Pract Res Clin
Gastroenterol. 21:281–297. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Bartel DP: MicroRNAs: genomics biogenesis,
mechanism and function. Cell. 116:281–297. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Croce C: Introduction to the role of
microRNAs in cancer diagnosis, prognosis and treatment. Cancer J.
18:213–214. 2012. View Article : Google Scholar : PubMed/NCBI
|