Introduction
Escherichia coli, a commensal bacteria of the
gastrointestinal tract in humans and animals, is a common cause of
community and hospital-acquired urinary tract infections and varies
in its susceptibility to antimicrobials (1–3). At
present, misuse of fluoroquinolones and third-generation
cephalosporins has led to an increasing number of drug-resistant
strains of Escherichia coli in China (4).
Previous studies have shown that 65% of clinical
isolates of Escherichia coli produce extended-spectrum
β-lactamases (ESBLs) in China (5,6).
ESBL-producing strains are resistant to β-lactams, fluoroquinolones
and aminoglycosides (7).
Cefotaxime (CTX)-M-type enzymes are the most common types of ESBLs
(8) and are the predominant ESBLs
in Enterobacteriaceae in China, causing hospital- and
community-acquired infections (4).
In addition, a number of studies have indicated that
integrons have developed a highly efficient mechanism for spreading
antibiotic resistance determinants (9,10).
Class 1 integrons play a crucial role in the dissemination of
antibiotic resistance in Gram-negative bacteria and occur
frequently in Escherichia coli by carrying and capturing
genes via site-specific recombination catalyzed by specific
integrase genes (11–14). Class 1 integrons aid in gene
transference in the capture and distribution of gene cassettes
among clinical Gram-negative bacillus (15).
To date, the correlation between class 1 integrons
and ESBLs of Escherichia coli has not been evaluated.
Therefore, the current study focused on analyzing the prevalence of
class 1 integrons and CTX-M-type enzymes in clinical isolates of
Escherichia coli in Chengdu, China between 2009 and 2011 to
determine the correlation between class 1 integrons and ESBLs.
Materials and methods
Samples
A total of 97 non-duplicated clinical Escherichia
coli isolates were collected from the sputum of patients from
the Chengdu No. 7 People’s Hospital (Sichuan, China) between 2009
and 2011 and were identified using the Microscan WalkAWay-40
(Siemens, Erlangen, Germany). Written informed consent was obtained
from the patients.
Isolate susceptibility
Isolate susceptibility was determined by the disc
diffusion technique on Mueller-Hinton agar plates (Oxoid Ltd.,
Basingtoke, Hampshire, UK) in accordance with CLSI guidelines
(16). The following reagents were
used: 10 μg ampicillin (AMP), 100 μg piperacillin (PIP), 30 μg
ceftazidime (CAZ), 30 μg cefepime (FEP), 30 μg ceftriaxone (CRO),
35 μg aztreonam (ATM), 5 μg ciprofloxacin (CIP), 30 μg tetracycline
(TCY), 10/10 μg ampicillin-sulbactam (SAM), 100/10 μg
piperacillin-tazobactam (TZP), 30 μg cefotaxime (CTX), 75 μg
cefoperazone (CFP), 10 μg imipenem (IPM), 10 μg tobramycin (TOB),
10 μg gentamicin (GEN) and 23.75/1.25 μg
trimethoprim-sulfamethoxazole (SXT; all Oxoid Ltd.). Escherichia
coli ATCC25922 and Staphylococcus aureus ATCC25923 were
used as reference strains for susceptibility testing.
A phenotypic confirmatory test was performed with 30
μg CTX, 30/10 μg cefotaxime-clavulanic acid, 30 μg CAZ and 30/10 μg
ceftazidime-clavulanic acid (all Becton-Dickinson, Franklin Lakes,
NJ, USA) disks on Mueller-Hinton agar. The results were analyzed as
previously described (16).
Escherichia coli ATCC 25922, Klebsiella pneumoniae
ATCC 700603 and Pseudomonas aeruginosa ATCC 27853 were used
as controls.
Primers used to amplify CTX-M genes, intI1
and conserved segments are presented in Table I with their corresponding cycling
conditions.
 | Table IPrimer sequences and PCR. |
Table I
Primer sequences and PCR.
| Primers | Primer sequence
(5′-3′) | PCR conditions | Reference | Expected size,
bp |
|---|
CTX-M-F
CTX-M-R |
TGTTGTTAGGAAGTGTGCCGC
TCGTTGGTGGTGCCATAGTC | 1 cycle of 3 min at
94°C, 35 cycles of 1 min at 94°C, 1 min at 55°C and 2 min at 72°C
and 1 cycle of 7 min at 72°C. | 17 | 687 |
IntI1-F
IntI1-R |
GGGTCAAGGATCTGGATTTCG
ACATGGGTGTAAATCATCGTC | 1 cycle of 5 min at
94°C, 30 cycles of 30 sec at 94°C, 1 min at 63°C and 40 sec at 72°C
and 1 cycle of 5 min at 72°C. | 18 | 484 |
5′CS
3′CS |
GGCATCCAAGCAGCAAGC
AAGCAGACTTGACCTGAT | 1 cycle of 3 min at
94°C, 35 cycles of 30 sec at 94°C, 30 sec at 55°C and 1 min at 72°C
and 1 cycle of 5 min at 72°C. | 19 | - |
Polymerase chain reaction (PCR)
Each PCR was carried out in a 25-μl volume using 1.5
units Taq DNA polymerase (Promega Corporation, Madison, WI,
USA) in the reaction buffer provided, which contained 2.5 mM
MgCl2, 50 μM each deoxynucleoside triphosphate, 0.4 μM
selected primer and 2 μl DNA template. Each PCR product (10 μl) was
subjected to electrophoresis on 1.2% agarose gel.
Amplification was performed by a Tpersonal
Thermocycler (Biometra, Göttingen, Germany). PCR products were
sequenced using an ABI3730 Sequencer (Applied Biosystems, Foster
City, CA, USA) and the sequences were compared with the reported
sequences from GenBank.
Results
Antibiotic resistance rates
Antibiotic resistance rates were as follows: TCY,
82.5%; PIP, 79.4%; GEN, 73.2%; AMP and CIP, 64.9% each; SXT, 62.9%;
CTX, 47.4%; CRO, 43.3%; TOB, 40.2%; CFP, 39.2%; CAZ, FEP and ATM
34.0% each; SAM, 9.3%; TZP, 7.2%; and IPM, 2.1%.
All isolates, with the exception of 3, were
sensitive to FEP. Among them, AMP, TCY, aminoglycoside,
fluoroquinolones and folic acid metabolic pathway inhibitor
resistance was >60% and cephalosporin resistance was ~30%.
Notably, 2% of isolates exhibited resistance to IPM.
CTX-M type β-lactamase production
Based on the phenotypic confirmatory test, 31
isolates (32%) were found to be producers of ESBLs. All isolates
that tested positive for ESBLs were also multidrug resistant, with
a statistically significant difference in resistance against 14
antibacterial drugs between positive and negative isolates
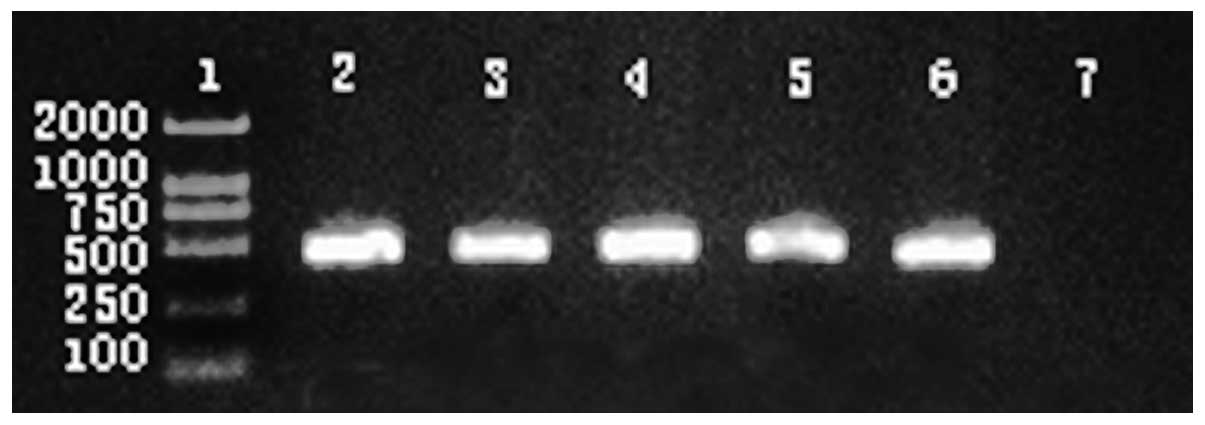
(P<0.05; Table II). CTX-M-type
β-lactamase was tested in Escherichia coli isolates
(Fig. 1) and detected in 19
isolates (61.5%).
 | Table IIComparison of resistance for positive
and negative strains of extended spectrum β-lactamases. |
Table II
Comparison of resistance for positive
and negative strains of extended spectrum β-lactamases.
| ESBLs positive
(n=31) | ESBLs negative
(n=66) | |
|---|
|
|
| |
|---|
| Paper disk of
antimicrobial agent | Resistance, % | Isolates, n | Resistance, % | Isolates, n | P-value |
|---|
| CTX | 74.2 | 23 | 34.8 | 23 | 0.000a |
| GEN | 87.1 | 27 | 66.7 | 44 | 0.027a |
| TOB | 54.8 | 17 | 33.3 | 22 | 0.037a |
| CFP | 58.1 | 18 | 30.3 | 20 | 0.009a |
| SXT | 90.3 | 28 | 50.0 | 33 | 0.000a |
| CAZ | 58.1 | 18 | 22.7 | 15 | 0.001a |
| AMP | 96.8 | 30 | 50.0 | 33 | 0.000a |
| FEP | 61.3 | 19 | 21.2 | 14 | 0.000a |
| TCY | 90.3 | 28 | 78.8 | 52 | 0.133 |
| CIP | 90.3 | 28 | 53.0 | 35 | 0.000a |
| IPM | 6.5 | 2 | 0.0 | 0 | 0.100 |
| PIP | 93.5 | 29 | 72.7 | 48 | 0.014a |
| SAM | 29.0 | 9 | 0.0 | 0 | 0.000a |
| TZP | 19.4 | 6 | 3.2 | 1 | 0.004a |
| CRO | 67.7 | 21 | 31.8 | 21 | 0.001a |
| ATM | 51.6 | 16 | 25.8 | 17 | 0.012a |
Gene cassettes in class 1 integrons
Of the 97 isolates tested, the intI1 gene was
detected in 69 isolates (71.1%; Fig.
2) with a statistically significant resistance to 9
antibacterial drugs identified between positive and negative
isolates (P<0.05; Table III).
Among these class 1 integron gene-positive strains, conserved
segments were amplified in 65 isolates (94.2%). The amplification
products sequenced were 100% identical to the reported sequence
from GenBank. Six conserved segments were detected in the 65
isolates (Fig. 3). Sequence
analysis was identical to the following known sequences:
dfr2d (549 bp; accession no, HQ902143), aadA1 (934
bp; accession no, HQ874618), aacC4-cmlA1 (2,327 bp;
accession no, HM175867), dfrA17-aadA5 (1,593 bp;
accession no, JN108894) and no gene cassette arrays (155 bp;
accession no, FM998811), as presented in Table IV. Four gene cassette arrangements
were found in 65/69 intI1-positive isolates (Table IV). The gene cassette arrangements
were as follows: aacC4-cmlA1 (1.5%), dfr2d
(18.45%), dfrA17-aadA5 (73.8%), aadA1 (10.8%) and
negative control (7.7%). The variable region of the following
integrons is presented in Fig. 3:
dfr2d (549 bp), aadA1 (934 bp),
aacC4+cmlA1 (2,327 bp), dfrA17+aadA5
(1,593 bp) and no gene cassette arrays (155 bp). Among them, 8
(12.3%) of the Escherichia coli isolates carried two
integrons and 57 (87.7%) carried one integron.
 | Table IIIComparison of resistance for positive
and negative strains of class 1 integrons. |
Table III
Comparison of resistance for positive
and negative strains of class 1 integrons.
| Integron-positive
(n=69) | Integron-negative
(n=28) | |
|---|
|
|
| |
|---|
| Antimicrobial
agent | Resistance, % | Isolates, n | Resistance, % | Isolates, n | P-value |
|---|
| CTX | 55.1 | 38 | 28.6 | 8 | 0.015a |
| GEN | 76.8 | 53 | 64.3 | 18 | 1.156 |
| TOB | 50.7 | 35 | 14.3 | 4 | 0.001a |
| CFP | 50.7 | 35 | 10.7 | 3 | 0.000a |
| SXT | 75.4 | 52 | 32.1 | 9 | 0.000a |
| CAZ | 37.7 | 26 | 25.0 | 7 | 0.169 |
| AMP | 73.9 | 51 | 42.9 | 12 | 0.004a |
| FEP | 39.1 | 27 | 21.4 | 6 | 0.074 |
| TCY | 85.5 | 59 | 75.0 | 21 | 0.173 |
| CIP | 73.9 | 51 | 42.9 | 12 | 0.004a |
| IPM | 14.5 | 1 | 14.5 | 1 | 0.496 |
| PIP | 88.4 | 61 | 57.1 | 16 | 0.001a |
| SAM | 13.0 | 9 | 0.0 | 0 | 0.040a |
| TZP | 7.2 | 5 | 7.1 | 2 | 0.676 |
| CRO | 53.6 | 37 | 17.9 | 5 | 0.001a |
| ATM | 36.2 | 25 | 28.6 | 8 | 0.317 |
 | Table IVType and arrangement of gene cassette
contained in class 1 integrons. |
Table IV
Type and arrangement of gene cassette
contained in class 1 integrons.
| Sample |
|---|
|
|
|---|
| Items | 1 | 2 | 3 | 4 | 5 | 6 |
|---|
| Integron
numbers | 1D | 3D | 5D | 7D | 9D | 3D+5D |
| Strains of the
total amount | 1 | 4 | 40 | 7 | 5 | 8 |
| Variable length of
integrons, bp | 2327 | 549 | 1593 | 934 | 155 | 549+1593 |
| Total ratio, % | 1.5 | 6.15 | 61.5 | 10.8 | 7.7 | 12.3 |
| Containing
resistance genes | aacC4-cmlA1 | dfr2d | dfrA17-aadA5 | aadA1 | - | dfr2d
-dfrA17-aadA5 |
Discussion
As shown in Table
II, resistance (%) was detected in all Escherichia coli
isolates of study. Higher resistance to 14 antimicrobial agents was
detected in ESBL-positive isolates compared with ESBL-negative
isolates (P<0.05). Resistance to AMP and PIP decreased depending
on the inhibition of ESBLs by enzyme inhibitors.
In the current study, a total of 31 isolates (32.0%)
producing ESBLs were identified among the 97 Escherichia
coli isolates with the higher prevalence of CTX-M (Fig. 1; 61.3%, 19/31), consistent with a
previous study (20). These
observations indicate that the CTX-M group is dominant in
Chengdu.
The aacC4, aadA1 and aadA5
genes encode resistance to aminoglycosides, cmlA1 encodes
resistance to chloramphenicols and dfr2d and dfrA17
encode resistance to trimethoprim. The gene cassette array
dfrA17+aadA5 is commonly used to detect class 1
integrons (21,22). The prevalence in the present study
was lower than that observed previously by Ozgumus et
al(23), which showed that all
class 1 integron-bearing Escherichia coli contained the
aadA5 gene cassette, conferring resistance to streptomycin
and spectinomycin. The gene cassette with the lowest detection rate
in the present study, aacC4+cmlA1, is infrequent in
other studies.
Antimicrobial resistance phenotypes were studied in
all isolates and the percentages of resistance detected were as
follows (Table III; %
integron-positive/% integron-negative isolates): CTX (55.1/28.6),
GEN (76.8/64.3), TOB (50.7/14.3), CFP (50.7/10.7), SXT (75.4/32.1),
CAZ (37.7/25.0), AMP (73.9/42.9), FEP (39.1/21.4), TCY (39.1/21.4),
CIP (73.9/42.9), IPM (14.5/14.5), PIP (88.4/57.1), SAM (13.0/0),
TZP (7.2/7.1), CRO (53.6/17.9) and ATM (36.2/28.6).
In addition, two isolates were resistant to IPM, and
a higher percentage of resistance to 9 antimicrobial agents
(Table III) was detected among
integron-positive isolates compared with integron-negative isolates
(P<0.05). The percentage of multi-resistant strains detected was
62.3% (43/69) among integron-positive isolates and 25.0% (7/28)
among integron-negative isolates (P<0.05). These observations
are in agreement with the hypothesis that class 1 integrons are
important in the resistance of Escherichia coli to
penicillins, third-generation cephalosporins, ciprofloxacin,
aminoglycosides and monocyclic β-lactam antibiotic.
Elevated percentages of resistance were observed in
a number of β-lactam drugs among ESBL-positive isolates compared
with ESBLs-negative isolates. The percentages of resistance were as
follows (% ESBL-positive/% ESBL-negative isolates): CTX
(74.2/34.8), GEN (87.1/66.7), TOB (54.8/33.3), CFP (58.1/30.3), SXT
(90.3/50.0), CAZ (58.1/22.7), AMP (96.8/50.0), FEP (61.3/21.2), TCY
(90.3/78.8), CIP (90.3/53.0), IPM (6.5/0.0), PIP (93.5/72.7), SAM
(29.0/0.0), TZP (19.4/3.2), CRO (51.6/25.8) and ATM (51.6/25.8).
The percentage of multi-resistant strains detected was 80.6%
(25/31) among ESBL-positive isolates and 37.9% (25/66) among
ESBL-negative isolates (P<0.05). The resistance to
third-generation cephalosporins observed was consistent with the
existence of ESBLs, as reported by Birgy et al(24).
The resistance profiles of isolates with ESBLs and
class 1 integrons are equal. We hypothesize that the presence of
the genes resistant to SXT, GEN and TOB in the variable region and
ESBLs cause resistance in Escherichia coli isolates to the
aforementioned antibacterial drugs. In the current study, 26/31
(83.9%) producers of ESBLs were identified to contain class 1
integrons.
The present study indicates that class 1 integrons
contributed to the multidrug resistance of Escherichia coli.
Class 1 integrons are important for the transfer of resistance
genes (25), as the integrons
carry antimicrobial-resistant gene cassettes and specific
resistance genes correspond to gene cassettes that are detected in
clinical isolates of Gram-negative bacteria (26).
The distribution of ESBLs and class 1 integrons in
Escherichia coli is prevalent with drug resistance in
Chengdu. According to the results of the present study, the
presence of class 1 integrons and ESBLs together mediates the
resistance of Escherichia coli isolates to the majority of
antibacterial agents. Based on our results, we hypothesize that the
combined treatment of ESBLs and class 1 integron may offer a new
perspective for treating resistant Escherichia coli.
Acknowledgements
The authors thank the Department of Pharmacology of
Preclinical and Forensic Medical College of Sichuan University.
References
|
1
|
Sabharwal ER: Antibiotic susceptibility
patterns of uropathogens in obstetric patients. N Am J Med Sci.
4:316–319. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kang HY, Jeong YS, Oh JY, Tae SH, Choi CH,
Moon DC, Lee WK, Lee YC, Seol SY, Cho DT and Lee JC:
Characterization of antimicrobial resistance and class 1 integrons
found in Escherichia coli isolates from humans and animals
in Korea. J Antimicrob Chemother. 55:639–644. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Seputiené V, Povilonis J, Ruzauskas M,
Pavilonis A and Suziedéliené E: Prevalence of trimethoprim
resistance genes in Escherichia coli isolates of human and
animal origin in Lithuania. J Med Microbiol. 59:315–322. 2010.
|
|
4
|
Xiao YH, Giske CG, Wei ZQ, Shen P, Heddini
A and Li LJ: Epidemiology and characteristics of antimicrobial
resistance in China. Drug Resist Updat. 14:236–250. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Jiang HX, Tang D, Liu YH, Zhang XH, Zeng
ZL, Xu L and Hawkey PM: Prevalence and characteristics of
β-lactamase and plasmid-mediated quinolone resistance genes in
Escherichia coli isolated from farmed fish in China. J
Antimicrob Chemother. 67:2350–2253. 2012.
|
|
6
|
Chen YH, Hsueh PR, Badal RE, Hawser SP,
Hoban DJ, Bouchillon SK, Ni Y and Paterson DL: Antimicrobial
susceptibility profiles of aerobic and facultative Gram-negative
bacilli isolated from patients with intra-abdominal infections in
the Asia-Pacific region according to currently established
susceptibility interpretive criteria. J Infect. 62:280–291. 2011.
View Article : Google Scholar
|
|
7
|
Wu H, Liu BG, Liu JH, Pan YS, Yuan L and
Hu GZ: Phenotypic and molecular characterization of CTX-M-14
extended-spectrum β-lactamase and plasmid-mediated ACT-like AmpC
β-lactamase produced by Klebsiella pneumoniae isolates from
chickens in Henan Province, China. Genet Mol Res. 11:3357–3364.
2012.
|
|
8
|
Paterson DL and Bonomo RA:
Extended-spectrum beta-lactamases: a clinical update. Clin
Microbiol Rev. 18:657–686. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Han N, Sheng D and Xu H: Role of
Escherichia coli strain subgroups, integrons, and
integron-associated gene cassettes in dissemination of
antimicrobial resistance in aquatic environments of Jinan, China.
Water Sci Technol. 66:2385–2392. 2012.
|
|
10
|
Hall RM: Integrons and gene cassettes:
hotspots of diversity in bacterial genomes. Ann NY Acad Sci.
1267:71–78. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Woodford N, Carattoli A, Karisik E,
Underwood A, Ellington MJ and Livermore DM: Complete nucleotide
sequences of plasmids pEK204, pEK499 and pEK516, encoding CTX-M
enzymes in three major Escherichia coli lineages from the
United Kingdom, all belonging to the international O25:H4-ST131
clone. Antimicrob Agents Chemother. 53:4472–4482. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Bennett PM: Plasmid encoded antibiotic
resistance: acquisition and transfer of antibiotic resistance genes
in bacteria. Br J Pharmacol. 153(Suppl 1): S347–S357. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Guérin E, Jové T, Tabesse A, Mazel D and
Ploy MC: High-level gene cassette transcription prevents integrase
expression in class 1 integrons. J Bacteriol. 193:5675–5682.
2011.PubMed/NCBI
|
|
14
|
van Essen-Zandbergen A, Smith H, Veldman K
and Mevius D: Occurrence and characteristics of class 1, 2 and 3
integrons in Escherichia coli, Salmonella and
Campylobacter spp. in the Netherlands. J Antimicrob
Chemother. 59:746–750. 2007.PubMed/NCBI
|
|
15
|
Nardelli M, Scalzo PM, Ramírez MS, Quiroga
MP, Cassini MH and Centrón D: Class 1 integrons in environments
with different degrees of urbanization. PLoS One. 7:e392232012.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Clinical and Laboratory Standards
Institute. Performance standards for antimicrobial susceptibility
testing standards: nineteenth informational supplement M100-S19.
2009 Accessed January, 2013
|
|
17
|
Wu LT, Tsou MF, Wu HJ, Chen HE, Chuang YC
and Yu WL: Survey of CTX-M-3 extended-spectrum beta-lactamase
(ESBL) among cefotaxime-resistant Serratia marcescens at a
medical center in middle Taiwan. Diagn Microbiol Infect Dis.
49:125–129. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Costa D, Poeta P, Sáenz Y, Coelho AC,
Matos M, Vinué L, Rodrigues J and Torres C: Prevalence of
antimicrobial resistance and resistance genes in faecal
Escherichia coli isolates recovered from healthy pets. Vet
Microbiol. 127:97–105. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Hsu SC, Chiu TH, Pang JC, Hsuan-Yuan CH,
Chang GN and Tsen HY: Characterisation of antimicrobial resistance
patterns and class 1 integrons among Escherichia coli and
Salmonella enterica serovar Choleraesuis strains
isolated from humans and swine in Taiwan. Int J Antimicrob Agents.
27:383–391. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wu S, Dalsgaard A, Hammerum AM, Porsbo LJ
and Jensen LB: Prevalence and characterization of plasmids carrying
sulfonamide resistance genes among Escherichia coli from
pigs, pig carcasses and human. Acta Vet Scand. 52:472010.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lim KT, Yasin R, Yeo CC, Puthucheary S and
Thong KL: Characterization of multidrug resistant ESBL-producing
Escherichia coli isolates from hospitals in Malaysia. J
Biomed Biotechnol. 2009:1656372009.PubMed/NCBI
|
|
22
|
Yang CM, Lin MF, Lin CH, Huang YT, Hsu CT
and Liou ML: Characterization of antimicrobial resistance patterns
and integrons in human fecal Escherichia coli in Taiwan. Jpn
J Infect Dis. 62:177–181. 2009.PubMed/NCBI
|
|
23
|
Ozgumus OB, Celik-Sevim E, Alpay-Karaoglu
S, Sandalli C and Sevim A: Molecular characterization of antibiotic
resistant Escherichia coli strains isolated from tap and
spring waters in a coastal region in Turkey. J Microbiol.
45:379–387. 2007.PubMed/NCBI
|
|
24
|
Birgy A, Cohen R, Levy C, Bidet P,
Courroux C, Benani M, Thollot F and Bingen E: Community faecal
carriage of extended-spectrum beta-lactamase-producing
Enterobacteriaceae in french children. BMC Infect Dis. 12:3152012.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ramirez MS, Parenteau TR, Centron D and
Tolmasky ME: Functional characterization of Tn1331 gene cassettes.
J Antimicrob Chemother. 62:669–673. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Bennett PM: Integrons and gene cassettes:
a genetic construction kit for bacteria. J Antimicrob Chemother.
43:1–4. 1999. View Article : Google Scholar : PubMed/NCBI
|