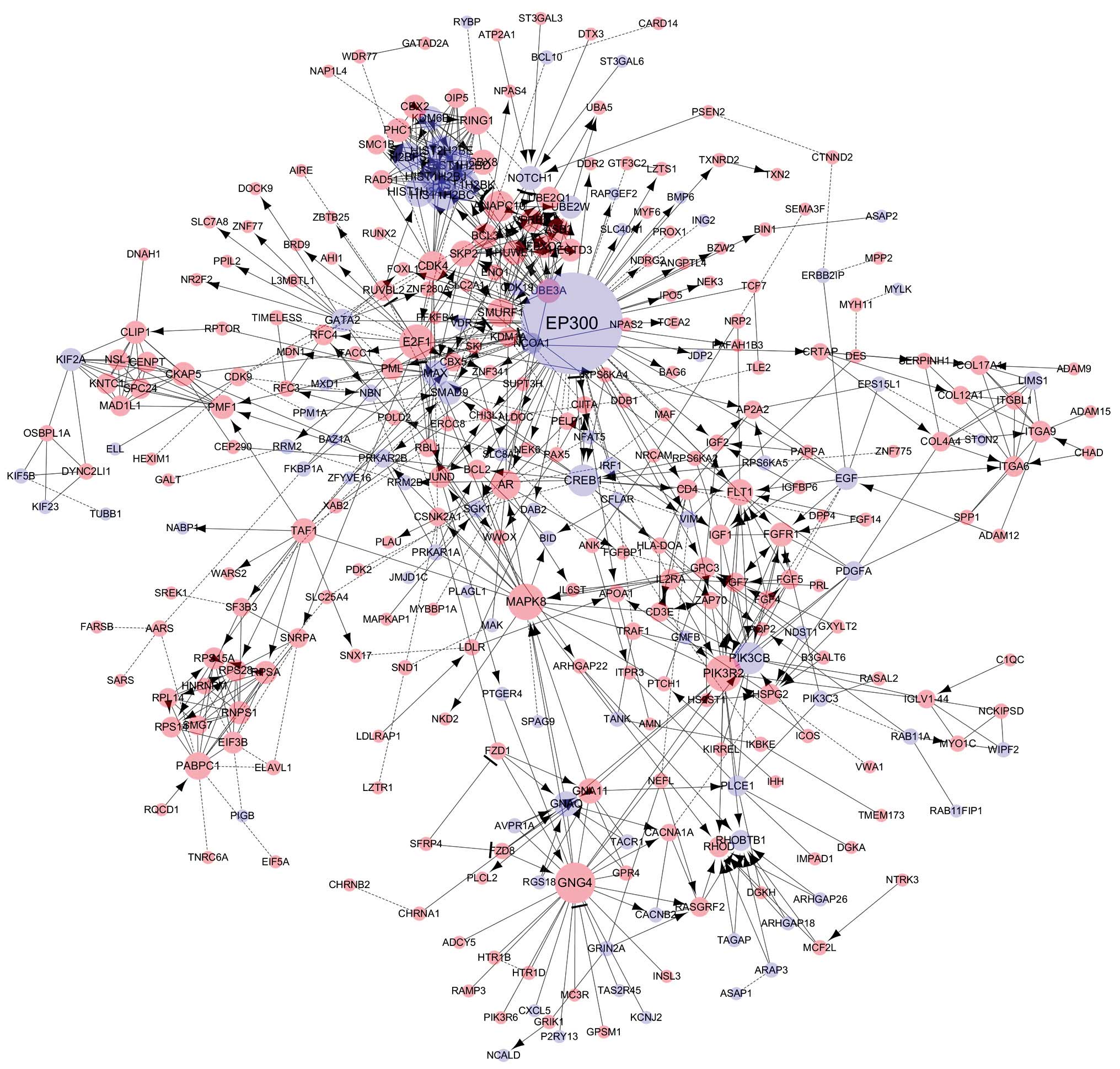
| EP300 | E1A binding protein
p300 | 0.03886829 | 81 | 25 | 56 | Down |
| GNG4 | Guanine nucleotide
binding protein (G protein), γ 4 | 0.00246082 | 26 | 9 | 17 | Up |
| MAPK8 | Mitogen-activated
protein kinase 8 | 0.00373044 | 22 | 12 | 10 | Up |
| PIK3R2 |
Phosphoinositide-3-kinase, regulatory
subunit 2 (p85 β) | 0.00275468 | 22 | 17 | 5 | Up |
| E2F1 | E2F transcription
factor 1 | 0.00881333 | 21 | 5 | 16 | Up |
| CREB1 | CAMP responsive
element binding protein 1 | 0.0020355 | 18 | 4 | 14 | Down |
| PIK3CB |
Phosphoinositide-3-kinase, catalytic, β
polypeptide | 0.00123254 | 18 | 13 | 5 | Down |
| HIST1H2BC | Histone cluster 1,
H2bc | 0.00045679 | 18 | 8 | 10 | Down |
| HIST1H2BK | Histone cluster 1,
H2bk | 0.00044752 | 18 | 11 | 7 | Down |
| HIST1H2BJ | Histone cluster 1,
H2bj | 0.00044752 | 18 | 10 | 8 | Down |
| HIST2H2BE | Histone cluster 2,
H2be | 0.00084908 | 18 | 11 | 7 | Down |
| HIST1H2BD | Histone cluster 1,
H2bd | 0.00039501 | 17 | 8 | 9 | Down |
| ANAPC10 | Anaphase promoting
complex subunit 10 | 0 | 17 | 0 | 17 | Up |
| AR | Androgen receptor
(dihydrotestosterone receptor; testicular feminization; spinal and
bulbar muscular atrophy; kennedy disease) | 0 | 16 | 0 | 16 | Up |
| CDK4 | Cyclin-dependent
kinase 4 | 0.00042396 | 16 | 1 | 15 | Up |
| HIST1H2AC | Histone cluster 1,
H2ac | 0.000341 | 16 | 5 | 11 | Down |
| H2BFS | H2B histone family,
member S | 0.00003574 | 16 | 4 | 12 | Down |
| SMURF1 | SMAD specific E3
ubiquitin protein ligase 1 | 0.00112437 | 15 | 11 | 4 | Up |
| PABPC1 | Poly (A) binding
protein, cytoplasmic 1 | 0.00046983 | 14 | 4 | 10 | Up |
| RING1 | Ring finger protein
1 | 0.00051894 | 14 | 13 | 1 | Up |
| MAX | MYC associated
factor X | 0.00190278 | 13 | 5 | 8 | Down |
| FLT1 | Fms-related
tyrosine kinase 1 (vascular endothelial growth factor/vascular
permeability factor receptor) | 0.00358728 | 13 | 7 | 6 | Up |
| SKP2 | S-phase
kinase-associated protein 2 (p45) | 0.0016332 | 13 | 8 | 5 | Up |
| CBX8 | Chromobox homolog 8
(Pc class homolog, Drosophila) | 0.00101008 | 12 | 1 | 11 | Up |
| TAF1 | TAF1 RNA polymerase
II, TATA box binding protein (TBP)-associated factor, 250 kDa | 0.00086181 | 12 | 11 | 1 | Up |
| BCL3 | B-cell CLL/lymphoma
3 | 0.00123249 | 12 | 1 | 11 | Up |
| GNA11 | Guanine nucleotide
binding protein (G protein), α 11 (Gq class) | 0.00031838 | 12 | 3 | 9 | Up |
| GNAQ | Guanine nucleotide
binding protein (G protein), q polypeptide | 0.00064179 | 12 | 6 | 6 | Down |
| PMF1 | Polyamine-modulated
factor 1 | 0.00076915 | 11 | 9 | 2 | Up |
| CBX2 | Chromobox homolog 2
(Pc class homolog, drosophila) | 0 | 11 | 0 | 11 | Up |
| HUWE1 | HECT, UBA and WWE
domain containing 1 | 0.00050967 | 11 | 4 | 7 | Up |
| KDM6B | Lysine (K)-specific
demethylase 6B | 0 | 11 | 9 | 2 | Down |
| UBE3A | Ubiquitin protein
ligase E3A (human papilloma virus E6-associated protein, Angelman
syndrome) | 0.00000927 | 11 | 10 | 1 | Up |
| FGFR1 | Fibroblast growth
factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer
syndrome) | 0.00350998 | 11 | 6 | 5 | Up |
| PHC1 | Polyhomeotic
homolog 1 (Drosophila) | 0 | 11 | 10 | 1 | Up |
| SMAD9 | SMAD family member
9 | 0.0020102 | 11 | 8 | 3 | Down |
| CKAP5 | Cytoskeleton
associated protein 5 | 0.000139 | 11 | 2 | 9 | Up |
| NOTCH1 | Notch homolog 1,
translocation-associated (Drosophila) | 0.00167253 | 11 | 6 | 5 | Down |
| FBXO3 | F-box protein
3 | 0 | 10 | 2 | 8 | Up |
| UBE2Q1 |
Ubiquitin-conjugating enzyme E2Q
(putative) 1 | 0.00000927 | 10 | 8 | 2 | Up |
| KIF2A | Kinesin heavy chain
member 2A | 0.00010193 | 10 | 4 | 6 | Down |
| UBE2W |
Ubiquitin-conjugating enzyme E2W
(putative) | 0.00000927 | 10 | 8 | 2 | Down |
| HECTD3 | HECT domain
containing 3 | 0 | 10 | 3 | 7 | Up |
| ASB3 | Ankyrin repeat and
SOCS box-containing 3 | 0 | 10 | 1 | 9 | Up |
| EGF | Epidermal growth
factor (β-urogastrone) | 0.00189305 | 10 | 2 | 8 | Down |
| GATA2 | GATA binding
protein 2 | 0.00200163 | 10 | 4 | 6 | Down |
| RPSA | Ribosomal protein
SA | 0.00015167 | 10 | 7 | 3 | Up |
| VPRBP | Vpr (HIV-1) binding
protein | 0 | 10 | 10 | 0 | Up |
| CLIP1 | CAP-GLY domain
containing linker protein 1 | 0.0000556 | 10 | 2 | 8 | Up |
| RNPS1 | RNA binding protein
S1, serine-rich domain | 0.00017607 | 10 | 2 | 8 | Up |
| GPC3 | Glypican 3 | 0.00296725 | 9 | 3 | 6 | Up |
| EIF3B | Eukaryotic
translation initiation factor 3, subunit B | 0 | 9 | 0 | 9 | Up |
| RPS28 | Ribosomal protein
S28 | 0.00004973 | 9 | 6 | 3 | Up |
| RPS15A | Ribosomal protein
S15a | 0.00004973 | 9 | 5 | 4 | Up |
| PML | Promyelocytic
leukemia | 0.00137496 | 9 | 6 | 3 | Up |
| ITGA9 | Integrin, α 9 | 0.00008649 | 9 | 6 | 3 | Up |
| JUND | Jun D
proto-oncogene | 0.00140147 | 9 | 4 | 5 | Up |
| NSL1 | NSL1, MIND
kinetochore complex component, homolog (S. cerevisiae) | 0 | 8 | 6 | 2 | Up |
| RHOD | Ras homolog gene
family, member D | 0.00038457 | 8 | 7 | 1 | Up |
| HSPG2 | Heparan sulfate
proteoglycan 2 | 0.00096904 | 8 | 5 | 3 | Up |
| SPC24 | SPC24, NDC80
kinetochore complex component, homolog (S. cerevisiae) | 0 | 8 | 8 | 0 | Up |
| CD4 | CD4 molecule | 0.00166231 | 8 | 2 | 6 | Up |
| NCOA1 | Nuclear receptor
coactivator 1 | 0.0000556 | 8 | 6 | 2 | Down |
| IGF1 | Insulin-like growth
factor 1 (somatomedin C) | 0.00081239 | 8 | 2 | 6 | Up |
| ITGA6 | Integrin, α 6 | 0.00008649 | 8 | 5 | 3 | Up |
| RPS14 | Ribosomal protein
S14 | 0.00000185 | 8 | 4 | 4 | Up |
| RHOBTB1 | Rho-related BTB
domain containing 1 | 0.00038457 | 8 | 7 | 1 | Down |
| RFC4 | Replication factor
C (activator 1) 4, 37 kDa | 0.00023013 | 8 | 6 | 2 | Up |
| KNTC1 | Kinetochore
associated 1 | 0 | 8 | 4 | 4 | Up |
| CENPT | Centromere protein
T | 0 | 8 | 0 | 8 | Up |
| RUVBL2 | RuvB-like 2 (E.
coli) | 0 | 8 | 8 | 0 | Up |
| IL2RA | Interleukin 2
receptor, α | 0.0002709 | 8 | 2 | 6 | Up |
| MAD1L1 | MAD1 mitotic arrest
deficient-like 1 (yeast) | 0 | 8 | 5 | 3 | Up |
| PLCE1 | Phospholipase C,
epsilon 1 | 0 | 8 | 8 | 0 | Down |
| RASGRF2 | Ras
protein-specific guanine nucleotide-releasing factor 2 | 0.0002085 | 7 | 5 | 2 | Up |
| FGF7 | Fibroblast growth
factor 7 (keratinocyte growth factor) | 0.00004633 | 7 | 2 | 5 | Up |
| SIX5 | SIX homeobox 5 | 1 | 7 | | | Up |
| SMC1B | Structural
maintenance of chromosomes 1B | 0 | 7 | 7 | 0 | Up |
| KDM1A | Lysine (K)-specific
demethylase 1 | 0.0001058 | 7 | 5 | 2 | Up |
| CD3E | CD3e molecule,
epsilon (CD3-TCR complex) | 0 | 7 | 0 | 7 | Up |
| PDGFA | Platelet-derived
growth factor α polypeptide | 0.0009675 | 7 | 4 | 3 | Down |
| COL4A4 | Collagen, type IV,
α 4 | 0 | 7 | 0 | 7 | Up |
| RAD51 | RAD51 homolog (RecA
homolog, E. coli) (S. cerevisiae) | 0 | 7 | 7 | 0 | Up |
| BCL2 | B-cell CLL/lymphoma
2 | 0 | 7 | 0 | 7 | Up |
| SMG7 | Smg-7 homolog,
nonsense mediated mRNA decay factor (C. elegans) | 0 | 7 | 7 | 0 | Up |
| FGF5 | Fibroblast growth
factor 5 | 0.00007058 | 7 | 2 | 5 | Up |
| OIP5 | Opa interacting
protein 5 | 0 | 7 | 7 | 0 | Up |
| RPL14 | Ribosomal protein
L14 | 0 | 7 | 2 | 5 | Up |
| SNRPA | Small nuclear
ribonucleoprotein polypeptide A | 0 | 7 | 7 | 0 | Up |
| COL17A1 | Collagen, type
XVII, α 1 | 0.00126029 | 7 | 1 | 6 | Up |
| FGF4 | Fibroblast growth
factor 4 (heparin secretory transforming protein 1, Kaposi sarcoma
oncogene) | 0 | 6 | 0 | 6 | Up |
| LIMS1 | LIM and senescent
cell antigen-like domains 1 | 0 | 6 | 6 | 0 | Down |
| ITGBL1 | Integrin, β-like 1
(with EGF-like repeat domains) | 0 | 6 | 5 | 1 | Up |
| COL12A1 | Collagen, type XII,
α 1 | 0 | 6 | 0 | 6 | Up |
| RBL1 | Retinoblastoma-like
1 (p107) | 0.00010657 | 6 | 5 | 1 | Up |
| CBX5 | Chromobox homolog 5
(HP1 α homolog, Drosophila) | 0.00021578 | 6 | 1 | 5 | Up |
| PRKAR2B | Protein kinase,
cAMP-dependent, regulatory, type II, β | 0 | 6 | 6 | 0 | Down |
| SF3B3 | Splicing factor 3b,
subunit 3, 130 kDa | 0.0001529 | 6 | 2 | 4 | Up |
| AP2A2 | Adaptor-related
protein complex 2, α 2 subunit | 0 | 6 | 0 | 6 | Up |
| IGLV1-44 | Immunoglobulin
lambda variable 1–44 | 0.00012974 | 6 | 1 | 5 | Up |
| CDK9 | Cyclin-dependent
kinase 9 | 0 | 6 | 0 | 6 | Up |
| CSNK2A1 | Casein kinase 2, α
1 polypeptide | 0.00045515 | 6 | 1 | 5 | Up |
| PELP1 | Proline, glutamic
acid and leucine rich protein 1 | 0.00034009 | 5 | 4 | 1 | Up |
| TRAF1 | TNF
receptor-associated factor 1 | 0 | 5 | 5 | 0 | Up |
| VIM | Vimentin | 0 | 5 | 5 | 0 | Down |
| CRTAP | Cartilage
associated protein | 0.00468901 | 5 | 3 | 2 | Up |
| NBN | Nibrin | 0.00107495 | 5 | 2 | 3 | Down |
| CACNA1A | Calcium channel,
voltage-dependent, P/Q type, α 1A subunit | 0 | 5 | 0 | 5 | Up |
| CIITA | Class II, major
histocompatibility complex, transactivator | 0 | 5 | 0 | 5 | Up |
| IRF1 | Interferon
regulatory factor 1 | 0.00002008 | 5 | 4 | 1 | Down |
| FZD1 | Frizzled homolog 1
(Drosophila) | 0.00061315 | 5 | 1 | 4 | Up |
| SGK1 |
Serum/glucocorticoid regulated kinase
1 | 0 | 5 | 5 | 0 | Down |
| APOA1 | Apolipoprotein
A-I | 0.00117688 | 5 | 1 | 4 | Up |
| ZAP70 | Zeta-chain (TCR)
associated protein kinase 70kDa | 0 | 5 | 5 | 0 | Up |
| DYNC2LI1 | Dynein, cytoplasmic
2, light intermediate chain 1 | 0 | 4 | 0 | 4 | Up |
| OSBPL1A | Oxysterol binding
protein-like 1A | 0 | 4 | 4 | 0 | Up |
| RFC3 | Replication factor
C (activator 1) 3, 38kDa | 0.00014364 | 4 | 2 | 2 | Up |
| SERPINH1 | Serpin peptidase
inhibitor, clade H (heat shock protein 47), member 1, (collagen
binding protein 1) | 0 | 4 | 4 | 0 | Up |
| HNRNPM | Recombinant
Heterogeneous nuclear ribonucleoprotein M | 0.00003243 | 4 | 1 | 3 | Up |
| PIK3C3 |
Phosphoinositide-3-kinase, class 3 | 0.00103422 | 4 | 1 | 3 | Down |
| PRKAR1A | Protein kinase,
cAMP-dependent, regulatory, type I, α (tissue specific extinguisher
1) | 0.00000463 | 4 | 3 | 1 | Down |
| AARS | Alanyl-tRNA
synthetase | 0 | 4 | 0 | 4 | Up |
| FZD8 | Frizzled homolog 8
(Drosophila) | 0 | 4 | 0 | 4 | Up |
| MYO1C | Myosin IC | 0.00003707 | 4 | 1 | 3 | Up |
| RAB11A | RAB11A, member RAS
oncogene family | 0.00047261 | 4 | 3 | 1 | Down |
| HLA-DOA | Major
histocompatibility complex, class II, DO α | 0.00003707 | 4 | 3 | 1 | Up |
| IGF2 | Insulin-like growth
factor 2 (somatomedin A) | 0.00076297 | 4 | 2 | 2 | Up |
| LDLR | Low density
lipoprotein receptor (familial hypercholesterolemia) | 0.00007105 | 4 | 2 | 2 | Up |
| ENO1 | Enolase 1, (α) | 0.00003398 | 4 | 2 | 2 | Up |
| ZNF280A | Zinc finger protein
280A | 0 | 3 | 3 | 0 | Up |
| ARHGAP22 | Rho GTPase
activating protein 22 | 0 | 3 | 0 | 3 | Up |
| DDB1 | Damage-specific DNA
binding protein 1, 127kDa | 0 | 3 | 0 | 3 | Up |
| ERCC8 | Excision repair
cross-complementing rodent repair deficiency, complementation groUp
8 | 0.0000139 | 3 | 1 | 2 | Up |
| VDR | Vitamin D
(1,25-dihydroxyvitamin D3) receptor | 0 | 3 | 3 | 0 | Down |
| POLD2 | Polymerase (DNA
directed), delta 2, regulatory subunit 50kDa | 0.00012047 | 3 | 1 | 2 | Up |
| CACNB2 | Calcium channel,
voltage-dependent, β 2 subunit | 0 | 3 | 1 | 2 | Down |
| PTCH1 | Patched homolog 1
(Drosophila) | 0 | 3 | 3 | 0 | Up |
| SPP1 | Secreted
phosphoprotein 1 (osteopontin, bone sialoprotein I, early
T-lymphocyte activation 1) | 0 | 3 | 3 | 0 | Up |
| KIF5B | Kinesin family
member 5B | 0.00000927 | 3 | 1 | 2 | Down |
| ARAP3 | ArfGAP with RhoGAP
domain, ankyrin repeat and PH domain 3 | 0 | 3 | 0 | 3 | Down |
| SKI | V-ski sarcoma viral
oncogene homolog (avian) | 0.00040774 | 3 | 2 | 1 | Up |
| EPS15L1 | Epidermal growth
factor receptor pathway substrate 15-like 1 | 0.0000278 | 3 | 2 | 1 | Down |
| FKBP1A | FK506 binding
protein 1A, 12kDa | 0.00009267 | 3 | 1 | 2 | Down |
| RPS6KA4 | Ribosomal protein
S6 kinase, 90kDa, polypeptide 4 | 0 | 3 | 3 | 0 | Up |
| BID | BH3 interacting
domain death agonist | 0.00000927 | 3 | 1 | 2 | Down |
| GRIN2A | Glutamate receptor,
ionotropic, N-methyl D-aspartate 2A | 0.0000556 | 3 | 1 | 2 | Down |
| NEFL | Neurofilament,
light polypeptide 68kDa | 0.00001853 | 3 | 1 | 2 | Up |
| ANK2 | Ankyrin 2,
neuronal | 0 | 3 | 0 | 3 | Up |
| TIMELESS | Timeless homolog
(Drosophila) | 0 | 3 | 3 | 0 | Up |
| SND1 | Staphylococcal
nuclease and tudor domain containing 1 | 0 | 3 | 3 | 0 | Up |
| RPS6KA5 | Ribosomal protein
S6 kinase, 90kDa, polypeptide 5 | 0.00006487 | 3 | 2 | 1 | Down |
| RPS6KA2 | Ribosomal protein
S6 kinase, 90kDa, polypeptide 2 | 0 | 3 | 3 | 0 | Up |
| RRM2 | Ribonucleotide
reductase M2 polypeptide | 0.00009267 | 3 | 2 | 1 | Down |
| RRM2B | Ribonucleotide
reductase M2 B (TP53 inducible) | 0 | 3 | 3 | 0 | Down |
| GPR4 | G protein-coUpled
receptor 4 | 0 | 3 | 3 | 0 | Up |
| WIPF2 | WAS/WASL
interacting protein family, member 2 | 0 | 3 | 3 | 0 | Down |
| CFLAR | CASP8 and FADD-like
apoptosis regulator | 0.00104252 | 3 | 1 | 2 | Down |
| ELAVL1 | ELAV (embryonic
lethal, abnormal vision, Drosophila)-like 1 (Hu antigen R) | 0 | 3 | 1 | 2 | Up |
| IKBKE | Inhibitor of κ
light polypeptide gene enhancer in B-cells, kinase epsilon | 0.00011429 | 3 | 1 | 2 | Up |
| ERBB2IP | Erbb2 interacting
protein | 0.00003707 | 3 | 2 | 1 | Down |
| PAX5 | Paired box 5 | 0.00012371 | 3 | 2 | 1 | Up |
| TACR1 | Tachykinin receptor
1 | 0 | 3 | 3 | 0 | Down |
| MCF2L | MCF.2 cell line
derived transforming sequence-like | 0 | 3 | 0 | 3 | Up |
| NRP2 | Neuropilin 2 | 0.00045407 | 3 | 2 | 1 | Up |
| XAB2 | XPA binding protein
2 | 0 | 3 | 3 | 0 | Up |
| WWOX | WW domain
containing oxidoreductase | 0 | 3 | 3 | 0 | Up |
| TANK | TRAF family
member-associated NFKB activator | 0.00001853 | 3 | 2 | 1 | Down |
| AVPR1A | Arginine
vasopressin receptor 1A | 0 | 3 | 0 | 3 | Down |
| NCKIPSD | NCK interacting
protein with SH3 domain | 0 | 3 | 2 | 1 | Up |
| NEK6 | NIMA (never in
mitosis gene a)-related kinase 6 | 0.00010116 | 3 | 1 | 2 | Up |
| CDK19 | cyclin-dependent
kinase 19 | 0 | 3 | 0 | 3 | Down |
| B3GALT6 | UDP-Gal:βGal β
1,3-galactosyltransferase polypeptide 6 | 0 | 2 | 0 | 2 | Up |
| NFAT5 | Nuclear factor of
activated T-cells 5, tonicity-responsive | 0 | 2 | 2 | 0 | Down |
| HS2ST1 | Heparan sulfate
2-O-sulfotransferase 1 | 0.00042627 | 2 | 1 | 1 | Up |
| ARHGAP18 | Rho GTPase
activating protein 18 | 0 | 2 | 0 | 2 | Down |
| TACC1 | Transforming,
acidic coiled-coil containing protein 1 | 0 | 2 | 2 | 0 | Up |
| SNX17 | Sorting nexin
17 | 0.00003398 | 2 | 1 | 1 | Up |
| TPD52L1 | Tumor protein
D52-like 1 | 1 | 2 | | | Up |
| ALDOC | Aldolase C,
fructose-bisphosphate | 0 | 2 | 0 | 2 | Up |
| FGFBP1 | Fibroblast growth
factor binding protein 1 | 0 | 2 | 2 | 0 | Up |
| GRIK1 | Glutamate receptor,
ionotropic, kainate 1 | 0 | 2 | 0 | 2 | Up |
| PTGER4 | Prostaglandin E
receptor 4 (subtype EP4) | 0 | 2 | 2 | 0 | Down |
| TXNRD2 | Thioredoxin
reductase 2 | 0 | 2 | 2 | 0 | Up |
| FOXL1 | Forkhead box
L1 | 0 | 2 | 1 | 1 | Up |
| MXD1 | MAX dimerization
protein 1 | 0 | 2 | 1 | 1 | Down |
| TLE2 | Transducin-like
enhancer of split 2 (E(sp1) homolog, Drosophila) | 0 | 2 | 2 | 0 | Up |
| NRCAM | Neuronal cell
adhesion molecule | 0.00001853 | 2 | 1 | 1 | Up |
| KCNAB1 | Potassium
voltage-gated channel, shaker-related subfamily, β member 1 | 1 | 2 | | | Up |
| UBA5 |
Ubiquitin-activating enzyme E1-domain
containing 1 | 0 | 2 | 0 | 2 | Up |
| TCF7 | Transcription
factor 7 (T-cell specific, HMG-box) | 0.00037994 | 2 | 1 | 1 | Up |
| WDR77 | WD repeat domain
77 | 0 | 2 | 2 | 0 | Up |
| KIF23 | Kinesin family
member 23 | 0 | 2 | 1 | 1 | Down |
| CHRNA1 | Cholinergic
receptor, nicotinic, α 1 (muscle) | 0 | 2 | 0 | 2 | Up |
| AQP2 | Aquaporin 2
(collecting duct) | 0 | 2 | 0 | 2 | Up |
| MAK | Male germ
cell-associated kinase | 0.00000927 | 2 | 1 | 1 | Down |
| SFRP4 | Secreted
frizzled-related protein 4 | 0 | 2 | 2 | 0 | Up |
| TAGAP | T-cell activation
GTPase activating protein | 0 | 2 | 2 | 0 | Down |
| DPYSL5 |
Dihydropyrimidinase-like 5 | 1 | 2 | | | Up |
| CHI3L1 | Chitinase 3-like 1
(cartilage glycoprotein-39) | 0 | 2 | 0 | 2 | Up |
| CEP290 | Centrosomal protein
290kDa | 0 | 2 | 0 | 2 | Up |
| MYH11 | Myosin, heavy chain
11, smooth muscle | 0.00000927 | 2 | 1 | 1 | Up |
| ZNF341 | Zinc finger protein
341 | 0 | 2 | 2 | 0 | Up |
| DES | Desmin | 0 | 2 | 0 | 2 | Up |
| NDST1 |
N-deacetylase/N-sulfotransferase (heparan
glucosaminyl) 1 | 0 | 2 | 2 | 0 | Down |
| SLC8A1 | Solute carrier
family 8 (sodium/calcium exchanger), member 1 | 0.00000927 | 2 | 1 | 1 | Down |
| PAPPA |
Pregnancy-associated plasma protein A,
pappalysin 1 | 0 | 2 | 2 | 0 | Up |
| ITPR3 | Inositol
1,4,5-triphosphate receptor, type 3 | 0.00000927 | 2 | 1 | 1 | Up |
| GMFB | Glia maturation
factor, β | 0.00015136 | 2 | 1 | 1 | Down |
| SLC25A4 | Solute carrier
family 25 (mitochondrial carrier; adenine nucleotide translocator),
member 4 | 0 | 2 | 2 | 0 | Up |
| CTNND2 | Catenin
(cadherin-associated protein), delta 2 (neural | 0 | 2 | 0 | 2 | Up |
| plakophilin-related
arm-repeat protein) | | | | | |
| ADAM12 | ADAM
metallopeptidase domain 12 (meltrin α) | 0 | 2 | 0 | 2 | Up |
| IL6ST | Interleukin 6
signal transducer (gp130, oncostatin M receptor) | 0 | 2 | 2 | 0 | Up |
| IGFBP6 | Insulin-like growth
factor binding protein 6 | 0 | 2 | 2 | 0 | Up |
| GTF3C2 | General
transcription factor IIIC, polypeptide 2, β 110kDa | 0 | 2 | 1 | 1 | Up |
| LZTS1 | Leucine zipper,
putative tumor sUppressor 1 | 0 | 2 | 2 | 0 | Up |
| GSTO2 | Glutathione
S-transferase omega 2 | 0.66666667 | 2 | | | Up |
| NPAS2 | Neuronal PAS domain
protein 2 | 0 | 2 | 2 | 0 | Up |
| ARHGAP26 | Rho GTPase
activating protein 26 | 0 | 2 | 0 | 2 | Down |
| DAB2 | Disabled homolog 2,
mitogen-responsive phosphoprotein (Drosophila) | 0 | 2 | 0 | 2 | Down |
| EIF4E2 | Eukaryotic
translation initiation factor 4E family member 2 | 1 | 2 | | | Up |
| RPTOR | Regulatory
associated protein of MTOR, complex 1 | 0 | 2 | 2 | 0 | Up |
| RASAL2 | RAS protein
activator like 2 | 0 | 2 | 2 | 0 | Up |
| FGF14 | Fibroblast growth
factor 14 | 0 | 2 | 0 | 2 | Up |
| GXYLT2 | Glucoside
xylosyltransferase 2 | 0.00042627 | 2 | 1 | 1 | Up |
| BIN1 | Bridging integrator
1 | 0.00114908 | 2 | 1 | 1 | Up |
| PSEN2 | Presenilin 2
(Alzheimer disease 4) | 0 | 2 | 2 | 0 | Up |
| RGS18 | Regulator of
G-protein signaling 18 | 0 | 2 | 2 | 0 | Down |
| PPM1A | Protein phosphatase
1A (formerly 2C), magnesium-dependent α | 0.00007413 | 2 | 1 | 1 | Down |
| KIRREL | Kin of IRRE like
(Drosophila) | 0 | 2 | 0 | 2 | Up |
| CYP2E1 | Cytochrome P450,
family 2, subfamily E, polypeptide 1 | 0.66666667 | 2 | | | Up |
| BCL10 | B-cell CLL/lymphoma
10 | 0 | 2 | 0 | 2 | Down |
| ICOS | Inducible T-cell
co-stimulator | 0 | 2 | 0 | 2 | Up |
| HTR1B | 5-hydroxytryptamine
(serotonin) receptor 1B | 0 | 2 | 1 | 1 | Up |
| HTR1D | 5-hydroxytryptamine
(serotonin) receptor 1D | 0 | 2 | 2 | 0 | Up |
| MAF | V-maf
musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 0 | 2 | 2 | 0 | Up |
| SUPT3H | SUppressor of Ty 3
homolog (S. cerevisiae) | 0 | 2 | 2 | 0 | Up |
| CHAD | Chondroadherin | 0 | 2 | 0 | 2 | Up |
| CAD | Carbamoyl-phosphate
synthetase 2, aspartate transcarbamylase, and dihydroorotase | 0 | 1 | | | Up |
| CTPS2 | CTP synthase
II | 0 | 1 | | | Up |
| PDCD11 | Programmed cell
death 11 | 0 | 1 | | | Up |
| RBM28 | RNA binding motif
protein 28 | 0 | 1 | | | Up |
| EIF5A | Eukaryotic
translation initiation factor 5A | 0 | 1 | 0 | 1 | Up |
| CPD | Carboxypeptidase
D | 0 | 1 | | | Up |
| DPP4 |
Dipeptidyl-peptidase 4 (CD26, adenosine
deaminase complexing protein 2) | 0 | 1 | 1 | 0 | Up |
| NCALD | Neurocalcin
delta | 0 | 1 | 1 | 0 | Down |
| ZNF169 | Zinc finger protein
169 | 0 | 1 | | | Up |
| GABARAPL1 | GABA(A)
receptor-associated protein like 1 | 0 | 1 | | | Down |
| TECPR2 | Tectonin
β-propeller repeat containing 2 | 0 | 1 | | | Down |
| PPRC1 | Peroxisome
proliferator-activated receptor γ, coactivator-related 1 | 0 | 1 | | | Up |
| GPSM1 | G-protein signaling
modulator 1 (AGS3-like, C. elegans) | 0 | 1 | 1 | 0 | Up |
| MRPL14 | Mitochondrial
ribosomal protein L14 | 0 | 1 | | | Up |
| MRPL4 | Mitochondrial
ribosomal protein L4 | 0 | 1 | | | Up |
| TNRC6A | Trinucleotide
repeat containing 6A | 0 | 1 | 1 | 0 | Up |
| RAPGEF2 | Rap guanine
nucleotide exchange factor (GEF) 2 | 0 | 1 | 1 | 0 | Down |
| KCNA1 | Potassium
voltage-gated channel, shaker-related subfamily, member 1 (episodic
ataxia with myokymia) | 0 | 1 | | | Up |
| PAFAH1B3 | Platelet-activating
factor acetylhydrolase, isoform Ib, γ subunit 29kDa | 0 | 1 | 1 | 0 | Up |
| AIRE | Autoimmune
regulator | 0 | 1 | 0 | 1 | Up |
| P2RY13 | Purinergic receptor
P2Y, G-protein coUpled, 13 | 0 | 1 | 1 | 0 | Down |
| GATAD2A | GATA zinc finger
domain containing 2A | 0 | 1 | 0 | 1 | Up |
| BAG6 | BCL2-associated
athanogene 6 | 0 | 1 | 0 | 1 | Up |
| PFKFB4 |
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase
4 | 0 | 1 | 1 | 0 | Up |
| BZW2 | Basic leucine
zipper and W2 domains 2 | 0 | 1 | 0 | 1 | Up |
| CHRNB2 | Cholinergic
receptor, nicotinic, β 2 (neuronal) | 0 | 1 | 1 | 0 | Up |
| SARS | Seryl-tRNA
synthetase | 0 | 1 | 1 | 0 | Up |
| RYBP | RING1 and YY1
binding protein | 0 | 1 | 1 | 0 | Down |
| ADCY5 | Adenylate cyclase
5 | 0 | 1 | 0 | 1 | Up |
| GALT |
Galactose-1-phosphate
uridylyltransferase | 0 | 1 | 0 | 1 | Up |
| STON2 | Stonin 2 | 0 | 1 | 1 | 0 | Down |
| DGKA | Diacylglycerol
kinase, α 80kDa | 0 | 1 | 0 | 1 | Up |
| PLXNA4 | Plexin A4 | 0 | 1 | | | Up |
| MYLK | Myosin, light chain
kinase | 0 | 1 | 1 | 0 | Down |
| MYF6 | Myogenic factor 6
(herculin) | 0 | 1 | 1 | 0 | Up |
| MDN1 | MDN1, midasin
homolog (yeast) | 0 | 1 | 1 | 0 | Up |
| AMN | Amnionless homolog
(mouse) | 0 | 1 | 0 | 1 | Up |
| DOCK9 | Dedicator of
cytokinesis 9 | 0 | 1 | 0 | 1 | Up |
| PDK2 | Pyruvate
dehydrogenase kinase, isozyme 2 | 0 | 1 | 0 | 1 | Up |
| MOB1B | MOB kinase
activator 1B | 0 | 1 | | | Down |
| SAV1 | Salvador homolog 1
(Drosophila) | 0 | 1 | | | Up |
| PIK3R6 |
phosphoinositide-3-kinase, regulatory
subunit 6 | 0 | 1 | 1 | 0 | Up |
| IMPAD1 | Inositol
monophosphatase domain containing 1 | 0 | 1 | 0 | 1 | Up |
| NR2F2 | Nuclear receptor
subfamily 2, groUp F, member 2 | 0 | 1 | 1 | 0 | Up |
| JMJD1C | Jumonji domain
containing 1C | 0 | 1 | 1 | 0 | Down |
| IPO5 | Importin 5 | 0 | 1 | 1 | 0 | Up |
| PPIL2 | Peptidylprolyl
isomerase (cyclophilin)-like 2 | 0 | 1 | 1 | 0 | Up |
| CYP1A1 | Cytochrome P450,
family 1, subfamily A, polypeptide 1 | 0 | 1 | | | Up |
| ELP2 | Elongation protein
2 homolog (S. cerevisiae) | 0 | 1 | | | Up |
| IKBKAP | Inhibitor of κ
light polypeptide gene enhancer in B-cells, kinase
complex-associated protein | 0 | 1 | | | Down |
| BRD9 | Bromodomain
containing 9 | 0 | 1 | 0 | 1 | Up |
| RAMP3 | Receptor (G
protein-coUpled) activity modifying protein 3 | 0 | 1 | 1 | 0 | Up |
| NAP1L4 | Nucleosome assembly
protein 1-like 4 | 0 | 1 | 1 | 0 | Up |
| NTRK3 | Neurotrophic
tyrosine kinase, receptor, type 3 | 0 | 1 | 1 | 0 | Up |
| CXCL5 | Chemokine (C-X-C
motif) ligand 5 | 0 | 1 | 0 | 1 | Down |
| NDRG2 | NDRG family member
2 | 0 | 1 | 1 | 0 | Up |
| SEMA3F | Sema domain,
immunoglobulin domain (Ig), short basic domain, secreted,
(semaphorin) 3F | 0 | 1 | 1 | 0 | Up |
| RUNX2 | Runt-related
transcription factor 2 | 0 | 1 | 0 | 1 | Up |
| MPP2 | Membrane protein,
palmitoylated 2 (MAGUK p55 subfamily member 2) | 0 | 1 | 1 | 0 | Up |
| SLC40A1 | Solute carrier
family 40 (iron-regulated transporter), member 1 | 0 | 1 | 1 | 0 | Down |
| SREK1 | Splicing regulatory
glutamine/lysine-rich protein 1 | 0 | 1 | 1 | 0 | Up |
| ING2 | Inhibitor of growth
family, member 2 | 0 | 1 | 1 | 0 | Down |
| AKIRIN2 | Akirin 2 | 0 | 1 | | | Down |
| CDH6 | Cadherin 6, type 2,
K-cadherin (fetal kidney) | 0 | 1 | | | Up |
| CDH7 | Cadherin 7, type
2 | 0 | 1 | | | Up |
| KCNC1 | Potassium
voltage-gated channel, Shaw-related subfamily, member 1 | 0 | 1 | | | Up |
| TMEM173 | Transmembrane
protein 173 | 0 | 1 | 1 | 0 | Up |
| ZNF775 | Zinc finger protein
775 | 0 | 1 | 1 | 0 | Up |
| DTX3 | Deltex 3 homolog
(Drosophila) | 0 | 1 | 0 | 1 | Up |
| EIF4G2 | Eukaryotic
translation initiation factor 4 γ, 2 | 0 | 1 | | | Up |
| ASAP1 | ArfGAP with SH3
domain, ankyrin repeat and PH domain 1 | 0 | 1 | 1 | 0 | Down |
| ZNF133 | Zinc finger protein
133 | 0 | 1 | | | Up |
| SPAG9 | Sperm associated
antigen 9 | 0 | 1 | 1 | 0 | Down |
| SLC4A4 | Solute carrier
family 4, sodium bicarbonate cotransporter, member 4 | 0 | 1 | | | Up |
| SLC4A7 | Solute carrier
family 4, sodium bicarbonate cotransporter, member 7 | 0 | 1 | | | Up |
| MYBBP1A | MYB binding protein
(P160) 1a | 0 | 1 | 1 | 0 | Up |
| WARS2 | Tryptophanyl tRNA
synthetase 2, mitochondrial | 0 | 1 | 1 | 0 | Up |
| KCNJ2 | Potassium
inwardly-rectifying channel, subfamily J, member 2 | 0 | 1 | 1 | 0 | Down |
| DGKH | Diacylglycerol
kinase, eta | 0 | 1 | 0 | 1 | Up |
| ANGPTL4 | Angiopoietin-like
4 | 0 | 1 | 0 | 1 | Up |
| MAPKAP1 | Mitogen-activated
protein kinase associated protein 1 | 0 | 1 | 0 | 1 | Up |
| RAB11FIP1 | RAB11 family
interacting protein 1 (class I) | 0 | 1 | 1 | 0 | Down |
| IHH | Indian hedgehog
homolog (Drosophila) | 0 | 1 | 0 | 1 | Up |
| SLC2A1 | Solute carrier
family 2 (facilitated glucose transporter), member 1 | 0 | 1 | 1 | 0 | Up |
| ADAM9 | ADAM
metallopeptidase domain 9 (meltrin γ) | 0 | 1 | 0 | 1 | Up |
| ST3GAL6 | ST3 β-galactoside
α-2,3-sialyltransferase 6 | 0 | 1 | 1 | 0 | Down |
| ST3GAL3 | ST3 β-galactoside
α-2,3-sialyltransferase 3 | 0 | 1 | 1 | 0 | Up |
| PIGB |
Phosphatidylinositol glycan anchor
biosynthesis, class B | 0 | 1 | 1 | 0 | Down |
| SLC7A8 | Solute carrier
family 7 (cationic amino acid transporter, y+ system), member
8 | 0 | 1 | 1 | 0 | Up |
| AHI1 | Abelson helper
integration site 1 | 0 | 1 | 0 | 1 | Up |
| DDR2 | Discoidin domain
receptor family, member 2 | 0 | 1 | 0 | 1 | Up |
| MC3R | Melanocortin 3
receptor | 0 | 1 | 1 | 0 | Up |
| TXN2 | Thioredoxin 2 | 0 | 1 | 0 | 1 | Up |
| PROX1 | Prospero homeobox
1 | 0 | 1 | 1 | 0 | Up |
| NABP1 | Nucleic acid
binding protein 1 | 0 | 1 | 0 | 1 | Down |
| TUBB1 | Tubulin, β 1 | 0 | 1 | 1 | 0 | Down |
| PLCL2 | Phospholipase
C-like 2 | 0 | 1 | 1 | 0 | Up |
| CARD14 | Caspase recruitment
domain family, member 14 | 0 | 1 | 1 | 0 | Up |
| JDP2 | Jun dimerization
protein 2 | 0 | 1 | 1 | 0 | Down |
| ZNF77 | Zinc finger protein
77 | 0 | 1 | 1 | 0 | Up |
| ZBTB25 | Zinc finger and BTB
domain containing 25 | 0 | 1 | 1 | 0 | Up |
| AP4E1 | Adaptor-related
protein complex 4, epsilon 1 subunit | 0 | 1 | | | Up |
| ADAM15 | ADAM
metallopeptidase domain 15 | 0 | 1 | 0 | 1 | Up |
| DNAH1 | Dynein, axonemal,
heavy chain 1 | 0 | 1 | 1 | 0 | Up |
| FARSB | Phenylalanyl-tRNA
synthetase, β subunit | 0 | 1 | 1 | 0 | Up |
| INSL3 | Insulin-like 3
(Leydig cell) | 0 | 1 | 1 | 0 | Up |
| TAS2R45 | Taste receptor,
type 2, member 45 | 0 | 1 | 1 | 0 | Down |
| PRL | Prolactin | 0 | 1 | 1 | 0 | Up |
| ATP2A1 | ATPase, Ca++
transporting, cardiac muscle, fast twitch 1 | 0 | 1 | 0 | 1 | Up |
| EYA4 | Eyes absent homolog
4 (Drosophila) | 0 | 1 | | | Up |
| BMP6 | Bone morphogenetic
protein 6 | 0 | 1 | 0 | 1 | Down |
| MKNK1 | MAP kinase
interacting serine/threonine kinase 1 | 0 | 1 | | | Down |
| TCEA2 | Transcription
elongation factor A (SII), 2 | 0 | 1 | 1 | 0 | Up |
| L3MBTL1 | l (3) mbt-like 1 | 0 | 1 | 1 | 0 | Up |
| ASAP2 | ArfGAP with SH3
domain, ankyrin repeat and PH domain 2 | 0 | 1 | 0 | 1 | Down |
| ZFYVE16 | Zinc finger, FYVE
domain containing 16 | 0 | 1 | 1 | 0 | Down |
| HEXIM1 | Hexamethylene
bis-acetamide inducible 1 | 0 | 1 | 1 | 0 | Up |
| NEK3 | NIMA (never in
mitosis gene a)-related kinase 3 | 0 | 1 | 1 | 0 | Up |
| DDX56 | DEAD
(Asp-Glu-Ala-Asp) box polypeptide 56 | 0 | 1 | | | Up |
| RPF1 | Ribosome production
factor 1 homolog | 0 | 1 | | | Up |
| STX3 | Syntaxin 3 | 0 | 1 | | | Down |
| STX7 | Syntaxin 7 | 0 | 1 | | | Down |
| GATM | Glycine
amidinotransferase (L-arginine:glycine amidinotransferase) | 0 | 1 | | | Up |
| ELL | Elongation factor
RNA polymerase II | 0 | 1 | 1 | 0 | Down |
| EZH1 | Enhancer of zeste
homolog 1 (Drosophila) | 0 | 1 | | | Up |
| JARID2 | Jumonji, AT rich
interactive domain 2 | 0 | 1 | | | Down |
| NKD2 | Naked cuticle
homolog 2 (Drosophila) | 0 | 1 | 1 | 0 | Up |
| ZNF569 | Zinc finger protein
569 | 0 | 1 | | | Up |
| C1QC | Complement
component 1, q subcomponent, C chain | 0 | 1 | 0 | 1 | Up |
| DPYS |
Dihydropyrimidinase | 0 | 1 | | | Up |
| LZTR1 | Leucine-zipper-like
transcription regulator 1 | 0 | 1 | 0 | 1 | Up |
| VWA1 | Von Willebrand
factor A domain containing 1 | 0 | 1 | 1 | 0 | Up |
| PEX14 | Peroxisomal
biogenesis factor 14 | 0 | 1 | | | Up |
| PEX3 | Peroxisomal
biogenesis factor 3 | 0 | 1 | | | Up |
| PLAU | Plasminogen
activator, urokinase | 0 | 1 | 1 | 0 | Up |
| PLAGL1 | Pleiomorphic
adenoma gene-like 1 | 0 | 1 | 1 | 0 | Down |
| LDLRAP1 | Low density
lipoprotein receptor adaptor protein 1 | 0 | 1 | 1 | 0 | Up |
| NPAS4 | Neuronal PAS domain
protein 4 | 0 | 1 | 1 | 0 | Up |
| BAZ1A | Bromodomain
adjacent to zinc finger domain, 1A | 0 | 1 | 0 | 1 | Down |
| RQCD1 | RCD1 required for
cell differentiation 1 homolog (S. pombe) | 0 | 1 | 1 | 0 | Up |
| ZNF337 | Zinc finger protein
337 | 0 | 1 | | | up |