Introduction
Lung cancer is a type of malignant tumor, which
causes serious damage to human health (1). In previous years, the morbidity and
mortality rates of lung cancer have significantly increased, and
the morbidity and mortality rates of men with lung cancer are the
highest among malignant tumor types, whereas among women it is the
second highest (2,3). Metastasis occurs readily in lung
cancer and can transfer to numerous regions of the body, causing
serious and life-threatening complications, leading to normal
tissue destruction (4,5). However, the incidence of lung cancer
metastasis is relatively complex and the pathogenesis remains to be
fully elucidated.
Transmembrane 7 superfamily member 4 (TM7SF4) is a
type of transmembrane protein encoded by the TM7SF4 gene, which is
present predominantly in dendritic cells, and is involved in
biological processes, which include cell fusion, cell
differentiation and immunity homeostasis (5,6). It
has been shown that TM7SF4 is abnormally expressed in thyroid
cancer, breast cancer and several other diseases, and accumulated
evidence suggests that TM7SF4 is key in a variety of prevalent
types of cancer (7). In addition,
TM7SF4 has a marked effect on the occurrence of tumor development,
and the roles of TM7SF4 in these prevalent types of cancer have
attracted significant attention, however, its role in the molecular
pathogenesis underlying lung cancer remains to be elucidated
(7,8).
Phosphatidylinositol 3-kinase (PI3K) and AKT consist
of multiple isoforms, and the PI3K/Akt pathway can regulate
cellular processes as diverse as cell growth, survival,
proliferation and migration (9,10).
PI3K/AKT has frequently been reported in investigations of
signaling pathways in various types of cancer, including breast
cancer, thyroid cancer, ovarian cancer and lung cancer (11–13).
The aim of the present study was to examine the
correlation between the expression of TM7SF4 and cell proliferation
and migration in lung cancer. Initially, the expression levels of
TM7SF4 were examined and compared between the A549 lung cancer cell
line and normal cell lines using reverse transcription-quantitative
polymerase chain reaction (RT-qPCR) analysis. The results confirmed
its high expression of TM7SF4 in lung cancer. Accordingly, the
proliferation and migration of lung cancer cells, and its possible
molecular mechanism were investigated. The results demonstrated
that the inhibition of TM7SF4 decreased cell viability and
migration, whereas the overexpression of TM7SF4 increased cell
proliferation. Therefore, it was concluded that TM7SF4 promoted
cell viability and migration. For further analysis, the signaling
pathway of TM7SF4 in the regulation of lung cancer cells was
investigated, and the results revealed that TM7SF4 promoted cell
viability by modulating activation of the PI3K/Akt pathway in the
A549 cells. Accordingly, the overexpression of TM7SF4 promoted the
expression of phosphorylated (p-)PI3K, p-AKT and p-mammalian target
of rapamycin (mTOR). Taken together, these results provide
opportunities, a theoretical basis and novel insights for further
investigation and the clinical development of novel treatment
strategies for lung cancer.
Materials and methods
Cell lines and cell transfection
The BEAS-2B human normal lung epithelial cell line
and A549 lung cancer cell line were obtained from the American Type
Culture Collection (Manassas, VA, USA). All cell lines were
cultured at 5% CO2 and at 37°C according to the
manufacturer's protocol. The TM7SF4 overexpression and silencing
vectors were constructed by Sangon Biotech Co., Ltd. (Shanghai,
China). A silencing vector containing no silenced TM7SF4 sequence
was transfected into the A549 cells as a control. Cell transfection
was performed using Lipofectamine 2000 reagent according to the
manufacturer's protocol (Invitrogen; Thermo Fisher Scientific,
Inc., Waltham, MA, USA) and then incubated for various durations,
including 25, 50, 75 and 100 h.
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
assay
The cells were cultured on 12-well plates to a
density of 5×104 cells/well. For the MTT assays, the
cells were cultured on 96-well culture plates, and seeded at
5×103 cells/well. The cells were then incubated for
various durations, as stated above. Cell viability was assayed by
adding 20 µl of 10 mg/ml MTT (Sigma-Aldrich; Merck Millipore,
Darmstadt, Germany) to 0.2 ml of culture medium, followed by
incubation for 3 h at 37°C. The medium was then removed, and the
MTT formazan product was dissolved in 150 µl DMSO, followed by
measuring the optical density at 590 nm with a Multiskan EX
microplate reader (Thermo Fisher Scientific, Inc.). Three
independent assays were performed (14).
Cell migration and invasion assay
Cell migration and invasion were evaluated using
Transwell migration chambers (8 µm pore size; Corning Incorporated,
Corning, NY, USA). The membranes for the invasion assay were coated
with a diluted ECM solution (Sigma-Aldrich; Merck Millipore) and
then air-dried at 4°C. Following transfection, the cells
(5×104 cells/well) were seeded with serum-free medium in
the upper portion of a chamber. Medium containing 10% FBS
(Invitrogen; Thermo Fisher Scientific, Inc.) was added to the lower
chamber, served as a chemoattractant. After 24 or 48 h of
incubation at 37°C, the non-invaded cells on the top of the
membrane were scraped and removed using cotton swabs, whereas the
invaded cells were fixed, stained with Diff-Quik and then counted
using light microscopy.
RT-qPCR analysis
Total RNA was extracted from the tissue samples or
cultured cells using TRIzol reagent (Takara Bio Inc., Otsu, Japan).
The concentration and purity of the isolated RNA was then
determined using an SMA 400 UV-VIS spectrophotometer (Merinton,
Shanghai, China). Purified RNA (0.5 µg/µl) was then mixed with
nuclease-free water and used for cDNA synthesis with the
PrimerScript first strand cDNA synthesis kit (Invitrogen; Thermo
Fisher Scientific, Inc.). The expression levels of targets in the
cells were measured in an Eppendorf Mastercycler (Brinkman
Instruments, Westbury, NY, USA) using the SYBR ExScriptqRT-PCR kit
(Takara Biotechnology Co., Ltd., Dalian, China) at a standard final
volume of 20 µl, which contained the following: 1.5 µl cDNA, 10 µl
SYBR Premix EX Taq, 1 µl of forward primer (10 µm), 1 µl reverse
primer (10 µm) and 6.5 µl ddH2O with 30 cycles. The PCR
profile was run under the following cycling conditions: An initial
predenaturation step at 95°C for 5 min, followed by 40 cycles of
denaturation at 95°C for 30 sec, annealing at 53°C for 30 sec,
extension at 72°C for 1 min and a final extension at 72°C for 10
min. Each reaction was performed in triplicate, and the
2−ΔΔCq method (15) was
used to determine the relative gene expression levels. Melting
curve analysis of the amplification products was performed at the
end of each PCR to confirm that only one product was amplified and
detected. Glyceraldehyde 3-phosphate dehydrogenase was selected as
the internal control for mRNA or long non-coding RNAs. The primers
used for target amplification are in Table I.
 | Table I.Primers used for target
amplification. |
Table I.
Primers used for target
amplification.
| Primer | Forward (5′-3′) | Reverse (5′-3′) |
|---|
| TM7SF4 |
GTAAAACGACGGCCAGTTCGTCATCTTGGGACACGTAG |
CTTTCTTTAGGAGTCGGCCAG |
| GADPH |
TGTTGCCATCAATGACCCCTT |
CTCCACGACGTACTCAGCG |
| PI3K | CACCGCATTTGTCGT | CTCCCACTTCTACGC |
| AKT |
GTATGCTGGCAGAGTAGGAGAAC |
CAGGTAACATCAGAGACAGACACA |
| mTOR |
AGGCCGCATTGTCTCTATCAA |
GCAGTAAATGCAGGTAGTCATCCA |
Western blot analysis
The cells were washed once with PBS and lysed in
radioimmunoprecipitation assay buffer (Sangon Biotech Co., Ltd.,
Shanghai, China) containing phenylmethanesufonyl fluoride
(Sigma-Aldrich; Merck Millipore), and centrifuged at 8,000 × g for
10 min at 4°C. Supernatant was collected for the measurement of
protein concentrations using a bicinchoninic acid assay kit
(Pierce; Thermo Fisher Scientific, Inc.). A total of 50 µg protein
in each sample was boiled for 10 min in SDS sample buffer,
separated on a 12% gel and subjected to SDS-PAGE, prior to transfer
onto nitrocellulose membranes (Whatman GmbH, Dassel, Germany).
Membranes were blocked in 5% non-fat dry milk in TBST for 1 h at
room temperature. Subsequently, membranes were incubated with the
following primary antibodies: TM7SF4 (catalog no. ab96809), PI3K
(catalog no. ab182651), AKT (catalog no. ab81283), mTOR (catalog
no. ab87540) and GAPDH (catalog no. ab181603), at 1:1,000 dilution
overnight at 4°C. These antibodies were purchased from Abcam,
Cambridge, UK. The membranes were then washed with TBST and
incubated with horseradish peroxidase-conjugated goat anti-rabbit
secondary antibody (catalog no. ab6721) and goat anti-mouse
secondary antibody (catalog no. ab6789) (1:2,000 dilution) for 2 h
at room temperature. Protein bands were visualized using the
WEST-ZOL-plus (iNtRON Biotechnology, Seoul, Korea) western blot
detection system (16). The
intensity of protein bands was quantified using Image J software
(version 1.46; National Institutes of Health, Bethesda, MD,
USA).
Statistical analysis
All experiments in the present study were performed
three times independently. Data are expressed as the mean ±
standard deviation and were analyzed using GraphPrism Prism 5.0
software (GraphPad Software, Inc., San Diego, CA, USA). An
independent sample t-test was used for paired data significance
calculation. Tukey's post hoc test was used to calculate the
differences among groups. P<0.05 was considered to indicate a
statistically significant difference.
Results
TM7SF4 expressed at high levels in
lung cancer A549 cells
The results of the RT-qPCR analysis and western blot
analysis revealed the expression of TM7SF4 at the mRNA level and
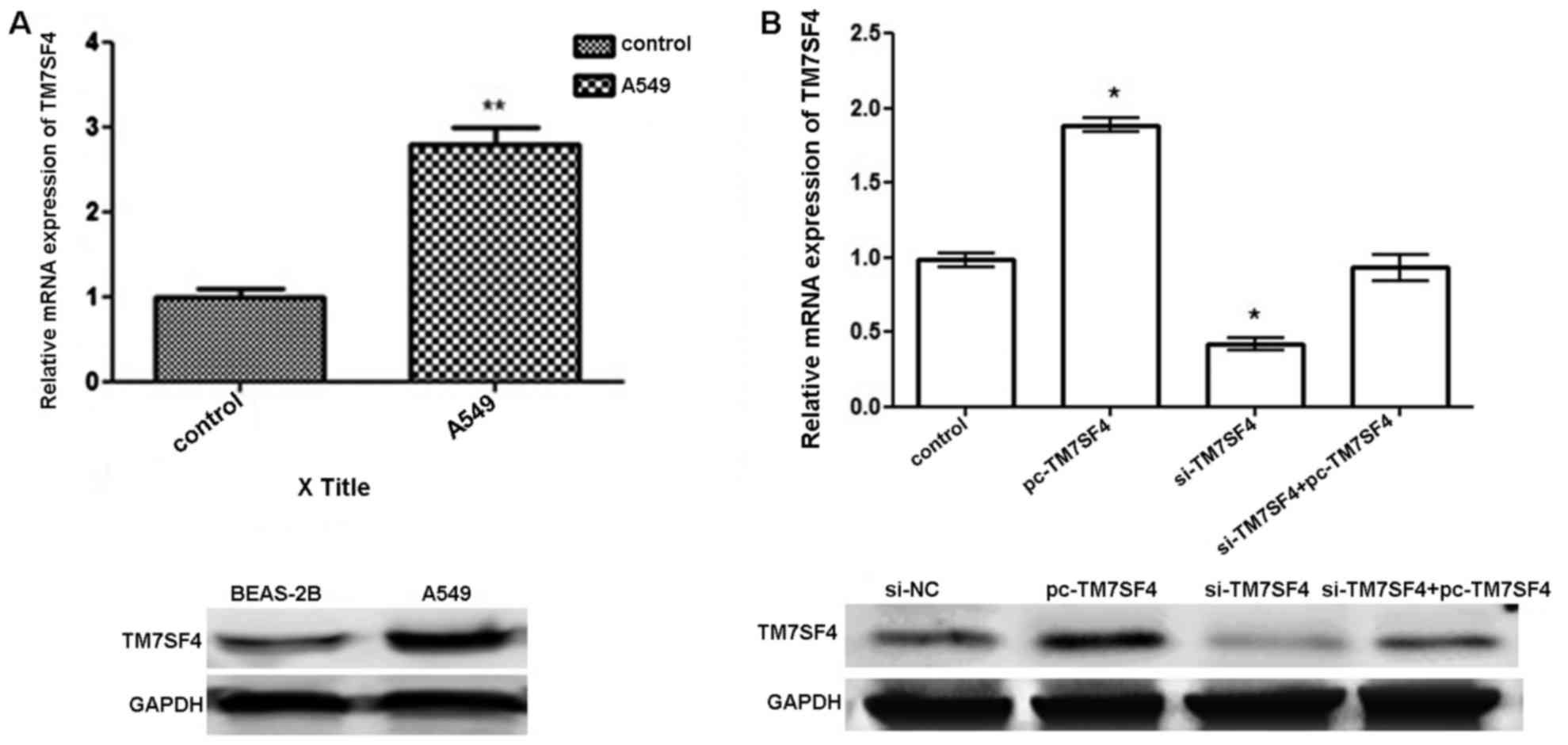
protein level, respectively. As shown in Fig. 1A, the expression of TM7SF4 was
significantly upregulated in the A549 cells, compared with the
normal lung tissues and cell lines (P<0.01). As shown in
Fig. 1B, the transfection with
pc-TM7SF4 effectively upregulated the expression level of TM7SF4 in
the A549 at the mRNA and protein levels. The transfection of cells
with si-TM7SF4 successfully downregulated the expression level of
TM7SF4 in the A549 cells.
Suppression of TM7SF4 inhibits cell
proliferation
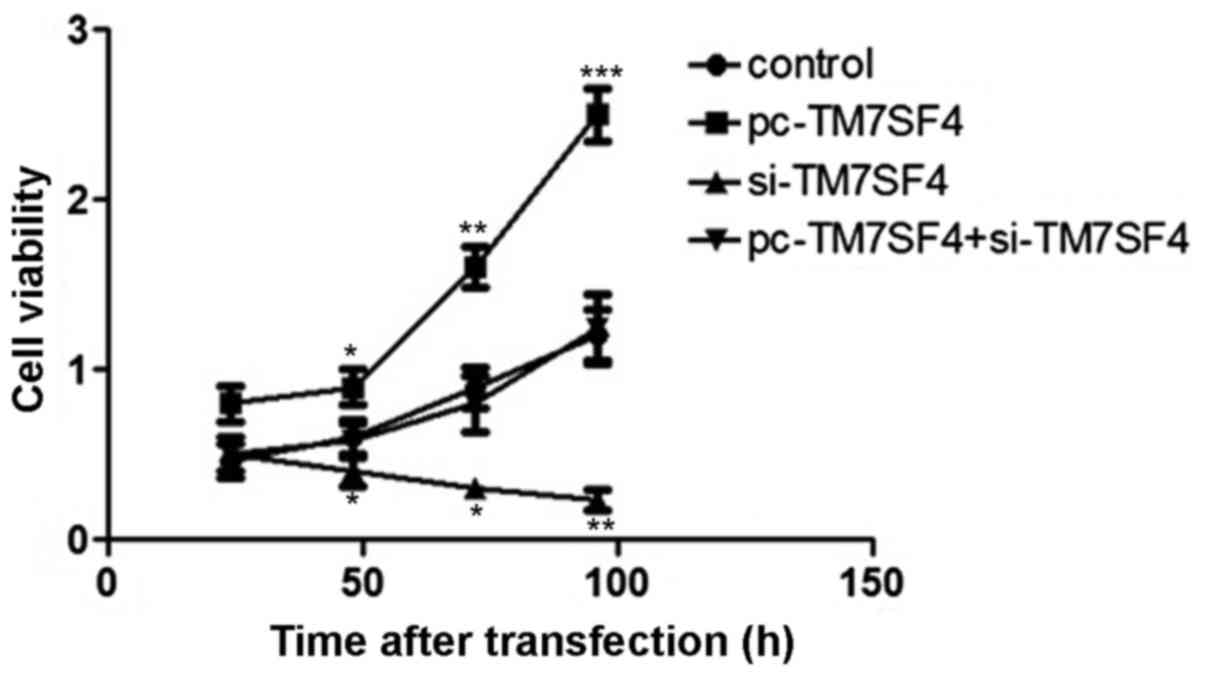
To determine the effect of the expression of TM7SF4
on A549 cell viability, an MTT assay was used to determine the
proliferation rate of A549 cells following 25, 50, 75 and 100 h of
transfection. The results showed that regulation of the expression
of TM7SF4 stimulated the proliferation of A549 cells. The
transfection of cells with si-TM7SF4 decreased A549 viability,
compared with cells in the blank group (P<0.01; Fig. 2).
Suppression of TM7SF4 prevents A549
cell migration
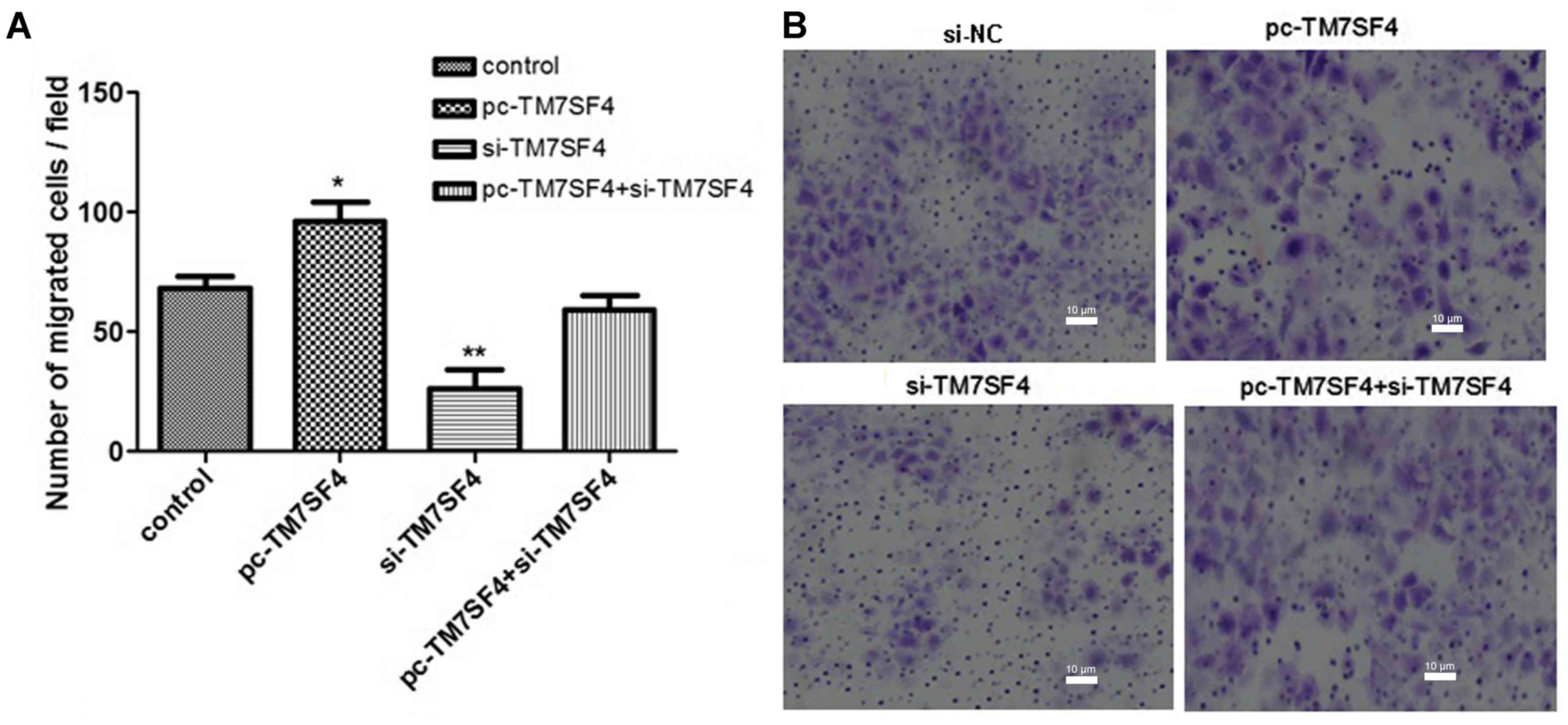
In the subsequent experiments, A Transwell assay was
used to examine the effects of TM7SF4 on A549 cell migration. The
results, as shown in Fig. 3A and
B, confirmed that the silencing of TM7SF4 significantly
inhibited the migration ability of the A549 cells (P<0.01).
TM7SF4 regulates cell proliferation
and migration by targeting the PI3K/AKT/mTOR pathway
To further determine the possible molecular
mechanism underlying the effect of the abnormal expression of
TM7SF4 on A549 cell biological processes, the expression levels of
PI3K/AKT/mTOR signaling pathway-associated proteins were examined
in the cells from each group. As shown in Fig. 4A, the overexpression of TM7SF4
increased the expression levels of p-PI3K, p-AKT and p-mTOR
(P<0.01), therefore, TM7SF4 may be associated with p-PI3K/AKT
pathway activation. Subsequent experiments using western blot
analysis were performed to examine the expression of associated
proteins. The results, as shown in Fig. 4B, confirmed that TM7SF4 regulated
cell migration and invasion through the p-PI3K/AKT pathway.
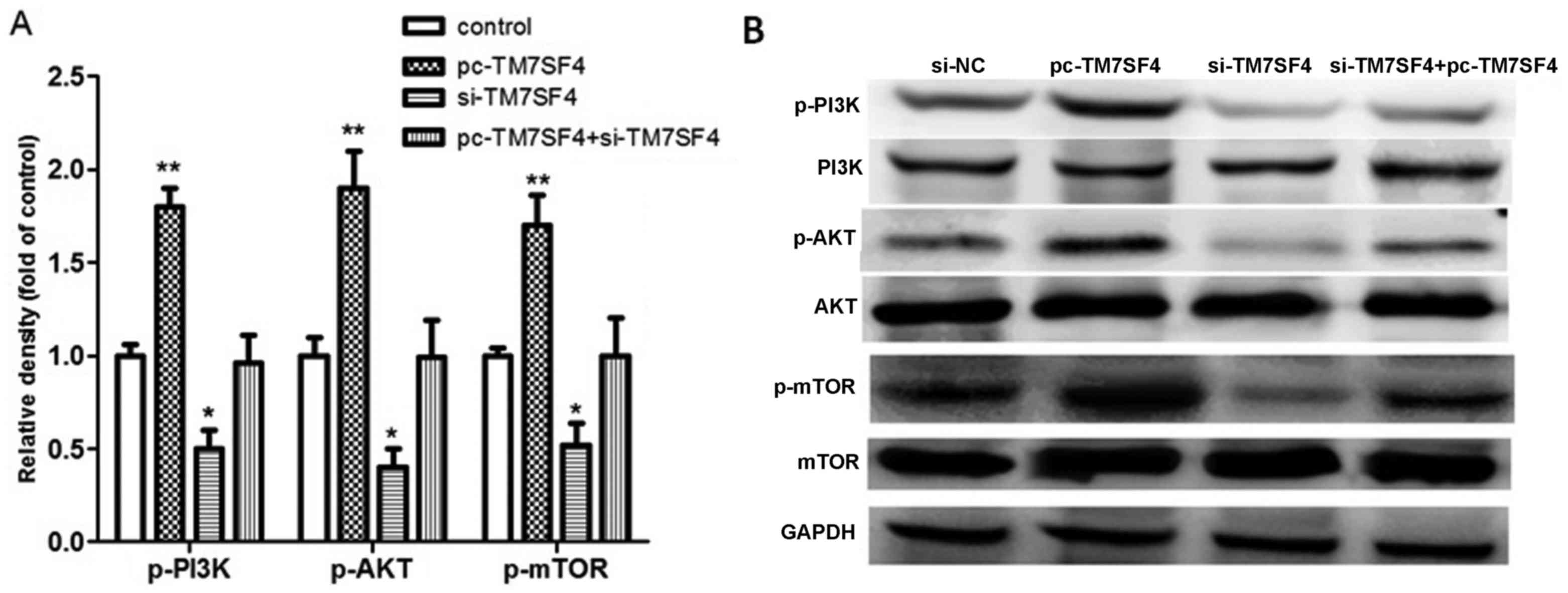 | Figure 4.Signaling pathway evaluation using
reverse transcription-quantitative polymerase chain reaction and
western blot analyses. (A) Detection of relative densities of
p-PI3K, p-AKT and p-mTOR in the si-TM7SF4, pc-TM7SF4 and
si-TM7SF4+pc-TM7SF4 groups. (B) Western blot analysis of p-PI3K,
PI3K, p-AKT, AKT, p-mTOR and mTOR. *P<0.05 and **P<0.01 vs.
control. TM7SF4, transmembrane 7 superfamily member 4; si, small
interfering RNA; GADPH, glyceraldehyde 3-phosphate dehydrogenase;
PI3K, phosphatidylinositol 3-kinase; mTOR, mammalian target of
rapamycin; p-, phosphorylated; NC, negative control. |
Discussion
Lung cancer is one of the leading causes of
mortality without an effective treatment strategy, the prevalence
and mortality rates of which continue to increase rapidly worldwide
(17,18). As a result, it is important to
elucidate the molecular mechanisms underlying the promotion of cell
proliferation, migration and signaling pathways in lung cancer
cells for elucidation of treatments and therapeutic strategies.
TM7SF4 encodes a seven-pass transmembrane protein,
and this protein regulates immunological functions,
osteoclastogenesis and myeloid differentiation (19). TM7SF4 has been reported be
important in Paget's disease of bone, papillary thyroid cancer and
breast cancer, however, the association between TM7SF4 and lung
cancer has not been reported (6,20).
The present study investigated the expression of TM7SF4 in lung
cancer and provided the first confirmation, to the best of our
knowledge, that TM7SF4 was expressed at a high level in lung cancer
cells, determined using RT-qPCR analysis.
The present study examined the association between
TM7SF4 and cell viability, and migration. TM7SF4 was found to
promote the viability and migration of the A549 cells, therefore,
the possibly pathways were subsequently investigated.
Alterations of signaling pathways are important in
the regulation of multiple cellular functions of lung cancer,
including cell growth and proliferation (21–24).
A previous study by Zhu et al confirmed that the AKT
signaling pathway is involved in the intrinsic apoptosis of non
small-cell lung cancer cells (25). In another report, microRNA-223 was
identified as a potential therapeutic target for overcoming
epidermal growth factor receptor-tyrosine kinase inhibitor
resistance, owing to its function in inducing activation of the
PI3K/AKT/mTOR signaling pathway in PC9/ER and PC9/CD133+ cells,
which is responsible for the resistance of PC9/ER and PC9/CD133+
cells to erlotinib (26).
Additionally, a study by Wan et al suggested that
insufficient RFA activates tumor growth in vitro and in
vivo via PI3K/AKT/mTOR signals (27).
In the present study, the AKT signaling pathway in
lung cancer cells was investigated. Using the method of gene
silencing, it was found that silencing TM7SF4 inhibited the
proliferation and metastasis of lung cancer through regulating
activation of the PI3K/AKT/mTOR signaling pathways. In particular,
the results showed that the expression levels of p-PI3K, p-AKT and
p-mTOR were activated when TM7SF4 was overexpressed, whereas the
inhibition of TM7SF4 inhibited this response. Therefore, TM7SF4 was
found to be important in the proliferation and metastasis of lung
cancer. Taken together, the present study confirmed that TM7SF4 was
upregulated in A549 lung cancer cells, and that the downregulation
of TM7SF4 may have certain suppressive roles in the development and
metastasis of lung cancer through suppressing activation of the
PI3K/AKT/mTOR signaling pathway. These findings confirmed that
TM7SF4 may be closely involved in the progression and development
of lung cancer, and may be a novel therapeutic target for this
disease. Insufficient mechanistic understanding has hindered the
prognosis of lung cancer, however, the present study indicated a
novel potential therapeutic approach to improve success in treating
lung cancer via targeting the identified PIKT/AKT/mTOR signaling
pathway. The present study provides a foundation for further
elucidation of the role of TM7SF4 in lung cancer. Specific elements
of the underlying mechanism require further validation
experiments.
Acknowledgements
This study was supported by the Natural Science
Foundation of Shandong Province, China (grant no.
2009ZRB14066).
References
|
1
|
D'Addario G, Früh M, Reck M, Baumann P,
Klepetko W and Felip E; ESMO Guidelines Working Group, . Metastatic
non-small-cell lung cancer: ESMO clinical practice guidelines for
diagnosis, treatment and follow-up. Ann Oncol. 21 Suppl
5:v116–v119. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ferlay J, Steliarova-Foucher E,
Lortet-Tieulent J, Rosso S, Coebergh JW, Comber H, Forman D and
Bray F: Cancer incidence and mortality patterns in Europe:
Estimates for 40 countries in 2012. Eur J Cancer. 49:1374–1403.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Sherwood JL, Corcoran C, Brown H, Sharpe
AD, Musilova M and Kohlmann A: Optimised pre-analytical methods
improve KRAS mutation detection in circulating tumour DNA (ctDNA)
from patients with non-small cell lung cancer (NSCLC). PLoS One.
11:e01501972016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Quéré G, Descourt R, Robinet G, Autret S,
Raguenes O, Fercot B, Alemany P, Uguen A, Férec C, Quintin-Roué I
and Le Gac G: Mutational status of synchronous and metachronous
tumor samples in patients with metastatic non-small-cell lung
cancer. BMC Cancer. 16:2102016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zequn N, Xuemei Z, Wei L, Zongjuan M,
Yujie Z, Yanli H, Yuping Z, Xia M, Wei W, Wenjing D, et al: The
role and potential mechanisms of LncRNA-TATDN1 on metastasis and
invasion of non-small cell lung cancer. Oncotarget. 7:18219–18228.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Donáth J, Speer G, Kósa JP, Árvai K, Balla
B, Juhász P, Lakatos P and Poór G: Polymorphisms of CSF1 and TM7SF4
genes in a case of mild juvenile Paget's disease found using
next-generation sequencing. Croat Med J. 56:145–151. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chung PY, Beyens G, de Freitas F, Boonen
S, Geusens P, Vanhoenacker F, Verbruggen L, Van Offel J, Goemaere
S, Zmierczak HG, et al: Indications for a genetic association of a
VCP polymorphism with the pathogenesis of sporadic Paget's disease
of bone, but not for TNFSF11 (RANKL) and IL-6 polymorphisms. Mol
Genet Metab. 103:287–292. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Valerio MS, Herbert BA, Griffin AC III,
Wan Z, Hill EG and Kirkwood KL: MKP-1 signaling events are required
for early osteoclastogenesis in lineage defined progenitor
populations by disrupting RANKL-induced NFATc1 nuclear
translocation. Bone. 60:16–25. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Safdari Y, Khalili M, Ebrahimzadeh MA,
Yazdani Y and Farajnia S: Natural inhibitors of PI3K/AKT signaling
in breast cancer: Emphasis on newly-discovered molecular mechanisms
of action. Pharmacol Res. 93:1–10. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ye Y, Tang X, Sun Z and Chen S:
Upregulated WDR26 serves as a scaffold to coordinate PI3K/AKT
pathway-driven breast cancer cell growth, migration, and invasion.
Oncotarget. 7:17854–17869. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
McAuliffe PF, Meric-Bernstam F, Mills GB
and Gonzalez-Angulo AM: Deciphering the Role of PI3K/Akt/mTOR
pathway in breast cancer biology and pathogenesis. Clin Breast
Cancer. 10 Suppl 3:S59–S65. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Fu J, Lv H, Guan H, Ma X, Ji M, He N, Shi
B and Hou P: Metallothionein 1G functions as a tumor suppressor in
thyroid cancer through modulating the PI3K/Akt signaling pathway.
BMC Cancer. 13:4622013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Fumarola C, Bonelli MA, Petronini PG and
Alfieri RR: Targeting PI3K/AKT/mTOR pathway in non small cell lung
cancer. Biochemical Pharmacol. 90:197–207. 2014. View Article : Google Scholar
|
|
14
|
Lu L, Li C, Li D, Wang Y, Zhou C, Shao W,
Peng J, You Y, Zhang X and Shen X: Cryptotanshinone inhibits human
glioma cell proliferation by suppressing STAT3 signaling. Mol Cell
Biochem. 381:273–282. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Nam KS, Oh S, Lee KM, Yoo SA and Shin I:
CD44 regulates cell proliferation, migration, and invasion via
modulation of c-Src transcription in human breast cancer cells.
Cell Signal. 27:1882–1894. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Imogen L and Gillham CM: Chemotherapy for
lung cancer. N Engl J Med. 346:14982002. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Shaw AT, Ou SH, Bang YJ, Camidge DR,
Solomon BJ, Salgia R, Riely GJ, Varella-Garcia M, Shapiro GI, Costa
DB, et al: Crizotinib in ROS1-rearranged non-small-cell lung
cancer. N Engl J Med. 372:1963–1971. 2014. View Article : Google Scholar
|
|
19
|
Chung PY, Beyens G, Boonen S, Papapoulos
S, Geusens P, Karperien M, Vanhoenacker F, Verbruggen L, Fransen E,
Van Offel J, et al: The majority of the genetic risk for Paget's
disease of bone is explained by genetic variants close to the CSF1,
OPTN, TM7SF4, and TNFRSF11A genes. Hum Genet. 128:615–626. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Albagha OM, Wani S and Ralston SH:
Identification of a functional variant in the TM7SF4 gene that is
associated with susceptibility to Paget's disease of bone. Bone.
48:S882011. View Article : Google Scholar
|
|
21
|
Mino-Kenudson M and Mark EJ: Reflex
testing for epidermal growth factor receptor mutation and
anaplastic lymphoma kinase fluorescence in situ hybridization in
non-small cell lung cancer. Arch Pathol Lab Med. 135:655–664.
2011.PubMed/NCBI
|
|
22
|
Feng N, Luo J and Guo X: Silybin
suppresses cell proliferation and induces apoptosis of multiple
myeloma cells via the PI3K/Akt/mTOR signaling pathway. Mol Med Rep.
13:3243–3248. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhang G, Wang C, Sun M, Li J, Wang B, Jin
C, Hua P, Song G, Zhang Y, Nguyen LL, et al: Cinobufagin inhibits
tumor growth by inducing intrinsic apoptosis through AKT signaling
pathway in human nonsmall cell lung cancer cells. Oncotarget.
7:28935–28946. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhao XZ, Liu Y, Zhou LJ, Wang ZQ, Wu ZH
and Yang XY: Role of estrogen in lung cancer based on the estrogen
receptor-epithelial mesenchymal transduction signaling pathways.
Onco Targets Ther. 8:2849–2863. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhu Q, Liang X, Dai J and Guan X:
Prostaglandin transporter, SLCO2A1, mediates the invasion and
apoptosis of lung cancer cells via PI3K/AKT/mTOR pathway. Int J
Clin Exp Pathol. 8:9175–9181. 2015.PubMed/NCBI
|
|
26
|
Hu J, Boeri M, Sozzi G, Liu D, Marchianò
A, Roz L, Pelosi G, Gatter K, Pastorino U and Pezzella F: Gene
signatures stratify computed tomography screening detected lung
cancer in high-risk populations. Ebiomedicine. 2:829–840. 2015.
View Article : Google Scholar
|
|
27
|
Wan J, Wu W, Chen Y, Kang N and Zhang R:
Insufficient radiofrequency ablation promotes the growth of
non-small cell lung cancer cells through PI3K/Akt/HIF-1α signals.
Acta Biochim Biophys Sin (Shanghai). 48:371–377. 2016. View Article : Google Scholar : PubMed/NCBI
|