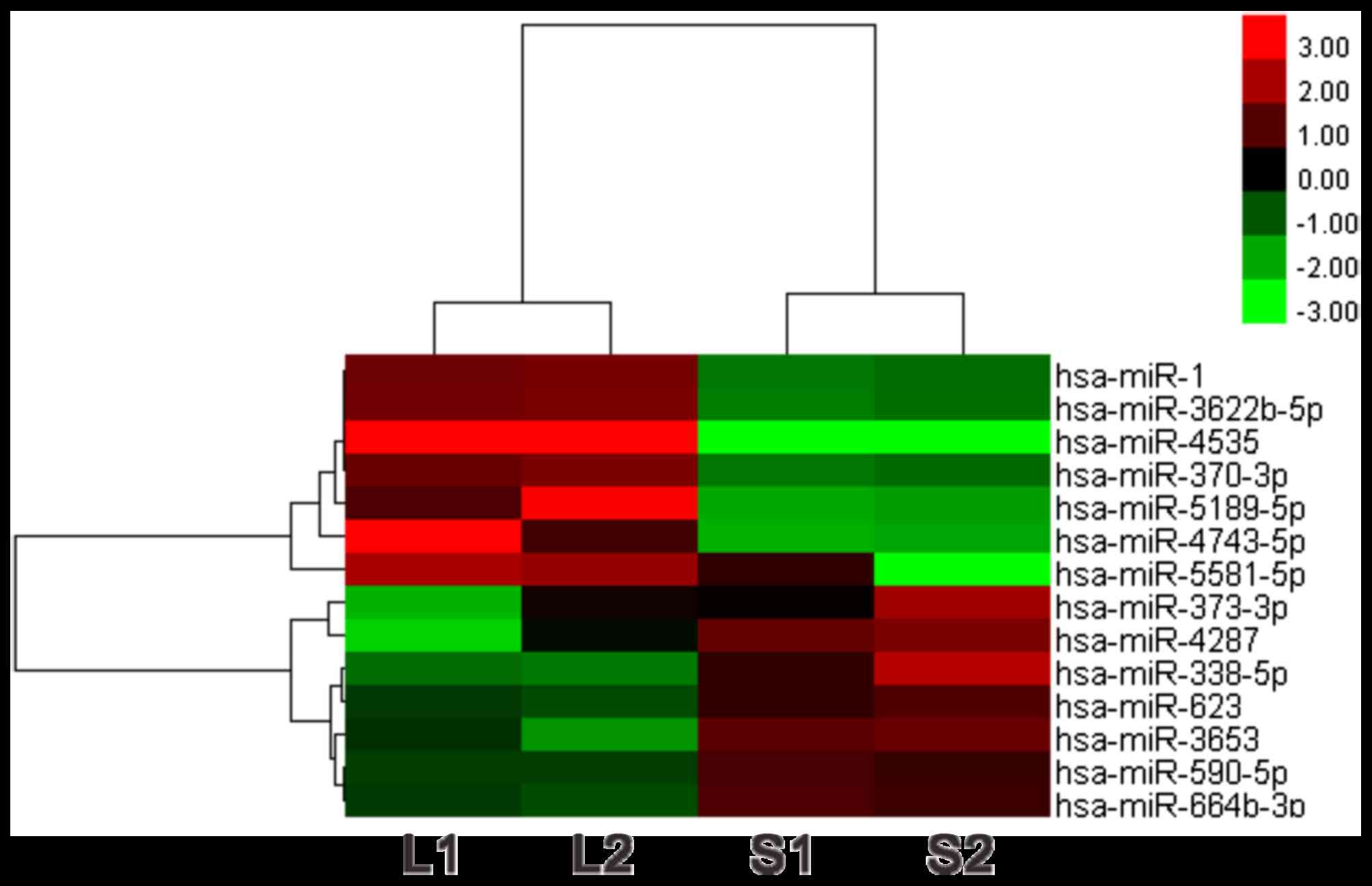
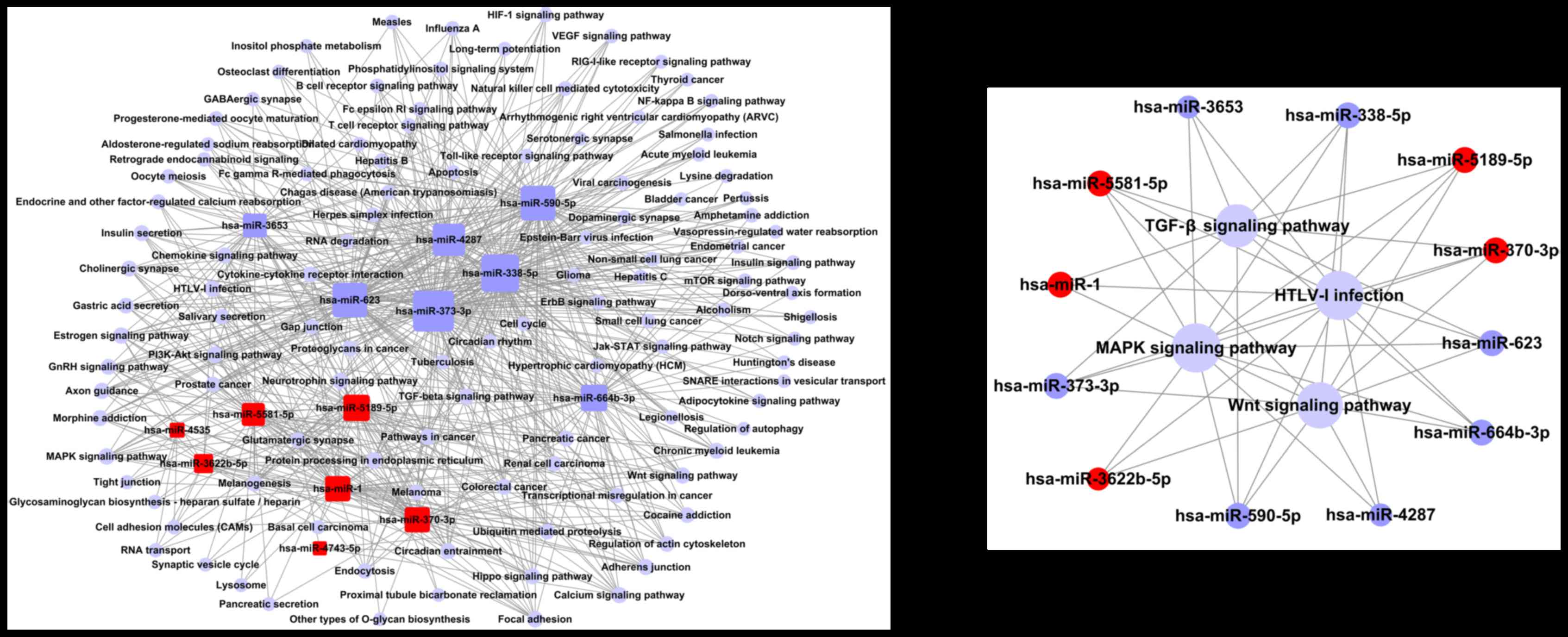
| Up-regulated | hsa-miR-1 | ABCA1, ABHD2, ABI2,
ABL2, ACER2, ADAM12, ADAR, AKAP11, AMOT, AMOTL2, ANKIB1, ANKRD29,
ANKRD34B, ANO1, ANP32B, ANXA4, AP3D1, API5, ARF3, ARHGEF18, ARID2,
ASH2L, ASPH, BCL11A, BDNF, BET1, BLCAP, BMPR1B, BOLL, BSCL2, BSN,
BZRAP1, C1RL, CAGE1, CALN1, CAPRIN1, CASK, CDC42, CEBPZ, CHM,
CLCN3, CLTC, CNN3, COIL, COL4A3, CPEB1, CREB5, CREM, DDX5, DHX15,
DICER1, DLG4, DNAJC5, E2F5, EHMT2, EIF1AX, EIF4E, EML3, EPB41L4B,
ETS1, FAM107B, FAM126A, FAM134A, FAM155A, FAM168B, FAM46C, FAM63B,
FAM91A1, FBXL14, FBXL20, FBXO22, FNDC3A, FOXP1, FRS2, FZD4, G6PD,
GABBR2, GAS2L1, GCH1, GDF6, GJA1, GLCCI1, GLIS2, GMFB, GNPTAB,
HACE1, HIAT1, HIGD1A, HMBOX1, HMGN1, HNRNPK, HNRNPU, HOOK1, HOXB4,
HP1BP3, HS3ST3B1, HSP90B1, HSPD1, JARID2, KCNJ2, KCTD10, KDM5C,
KDSR, KIAA1462, KTN1, LARP4, LASP1, LIN7C, LPPR4, LRCH1, LRRC8A,
MAGI2, MAP3K1, MATR3, MEIS1, MEOX2, MET, MGAT4A, MIER1, MMD2, MMP8,
MON2, MXD1, NAB1, NAMPT, NBEA, NCOA3, NDRG3, NET1, NFAT5, NR3C1,
NR4A3, NRP1, NUP50, NXT2, OSBPL7, OSBPL8, OTX2, PABPC4L, PAX6,
PAX7, PDE7A, PDGFA, PDIK1L, PFKFB2, PHAX, PHIP, PHLDA1, PKD2,
PLEKHO2, POGK, PPIB, PREX1, PRIC285, PRKRIR, PTPLAD1, PTPN2, PTPRK,
RAB43, RARB, RNF138, RNF141, RNF165, RNF213, RSBN1L, RUNX1, SEC22C,
SEC23B, SEC63, SELT, SFRP1, SH3PXD2B, SH3TC2, SLC10A7, SLC16A6,
SLC25A22, SLC25A30, SLC25A36, SLC29A3, SLC35B4, SLC35F1, SLC37A3,
SLC39A9, SLC7A11, SLC8A2, SMAD4, SMAP1, SNED1, SNX13, SNX2, SOX9,
SPRED1, SS18, STC2, STX12, SULF1, TAGLN2, TMED5, TMEM135, TMEM178,
TMSB4X, TNKS2, TNPO1, TNRC6B, TNS3, TPPP, TRAPPC3, TRHDE, TRIM2,
TTC3, TTC7B, UBE2H, UBE4A, UBR5, UTRN, VAMP2, WIPF2, WNK3, WSCD2,
YPEL2, ZBTB4, ZBTB41, ZC3HAV1, ZFP91, ZNF148, ZNF236, ZNF652,
ZZZ3 |
|
|
hsa-miR-3622b-5p | ANKRD52, ATRNL1,
BEND4, CADM4, CBX5, CCDC34, CCDC97, CNKSR2, COL5A3, CPNE5, DCX,
DVL3, EDEM3, EXTL3, FAM126B, FAM20B, FBXL20, FKBP5, FOXP3, GRIK2,
HUWE1, KCTD20, KIAA0317, KIAA1239, KLF12, LARP1, LCORL, LOXL4,
LPPR2, MAP3K3, MBOAT2, MIB1, MUM1L1, MYO1D, NDRG3, NTRK2, NUCKS1,
NUP98, PAX6, PDE7B, PHF20L1, PHF21A, PTP4A1, PVRL1, PXT1, QKI,
RIMKLA, SH3TC2, SLC1A2, SNTB2, SP2, SRGAP3, SSH2, STAG2, TBC1D14,
TCF20, TRIM46, TRIM66, TSGA10, TSPAN11, VPS53, ZBTB7B |
|
| hsa-miR-4535 | APBA1, CHD6,
CLDN19, DNAJB12, EEF1A2, FKBP4, MAT2A, MYH7B, NDST1, PARVA, PTCD1,
RIBC1, SCN2B, SPIN3, SPOPL, TUB |
|
| hsa-miR-370-3p | ABCG4, ABR, ACCN4,
ACOX1, ACTR1A, ACVR2B, ADCY5, AFF1, ANGEL1, ANKH, ANKRD52, ARCN1,
ARF3, ASB10, ATP11A, ATP1A2, ATXN7L3, BAG4, BMF, BSN, C1QTNF6,
CCDC64, CCL21, CDC42EP4, CFL1, CFLAR, CHD2, CHRNA7, CIT, CNGB1,
CRLF1, CYB5B, CYP2U1, DES, DGCR14, DHX35, DMRTB1, DNAJB1, DNAJC11,
DND1, EML1, ENAH, ENOX2, FAM102A, FAM123B, FAM164C, FAM168B, FBLN5,
FBXO46, FGF11, FGF7, FOSL2, FOXO1, GADD45B, GSG1L, HDAC4, HEMK1,
HHIPL1, HIF1AN, HNRNPUL2, HPS5, HSPA12A, HTR4, IKZF4, INO80, IPPK,
IVD, JMY, KCNJ11, KIAA2018, KIF1B, KLC2, KLF12, KLHL18, KRT80,
LPHN2, MAP2K7, MOCS1, MRPS25, MTL5, NAPG, NCDN, NCOA5, NEK9, NF1,
NFASC, NLGN2, NTRK2, ODF2, OPA3, ORAI2, PACS1, PAPL, PCDH10,
PCDH11X, PCDH19, PCLO, PDE7A, PHF19, PLEKHA6, PLEKHM1, POLR2F,
POMT2, PPARGC1B, PRDM10, PRLR, PRRX1, PTCD1, PXMP4, RAB11A,
RAP1GDS1, RAPGEF1, RAPGEFL1, RBBP4, SAP30BP, SEMA6A, SH3BP2, SHE,
SLC10A7, SLC46A1, SLC4A4, SMURF1, SOX12, SPRYD3, ST3GAL3, ST6GAL1,
STK35, SYNGR1, SYNPO2, TGFBR2, TM9SF4, TMCO7, TMEM127, TMEM154,
TMEM184A, TMEM40, TNRC6C, TP53I11, TRIM33, TRIOBP, TRIT1, UBE2R2,
UBTF, USP37, USP47, USP5, VANGL1, VSTM2L, WDTC1, WNT10B, ZBTB39,
ZBTB42, ZC3H7B, ZC3HAV1, ZCCHC17, ZCCHC24, ZDHHC5, ZMYND11, ZNF148,
ZNF185, ZNF37A, ZNF605, ZNF704 |
|
|
hsa-miR-5189-5p | AAK1, ACVR1B,
ACVR2A, ADAT2, ADCY2, ADRBK1, AGBL4, AHI1, AKT2, ANKS6, ARHGAP19,
ARID4A, ARRDC3, ASTN1, ASXL1, ATG4B, ATOH8, ATP2B2, ATP6V0D1,
ATP8A1, BAIAP2L2, BAK1, BMP8B, BMPR2, BRAP, BRD4, BTRC, CA5B,
CACNA1I, CACNB3, CALM3, CBFA2T2, CCBE1, CCDC69, CCDC76, CD300LG,
CDH24, CHL1, CNP, CPLX2, CSMD2, CSRNP1, CYP26B1, DACT3, DCBLD2,
DCHS1, DENND1A, DIS3L2, DNAJC5G, DTX1, DUOX1, EEFSEC, ELFN2,
FAM105B, FAM120C, FAM155B, FAM53C, FBXL19, FBXL20, FBXO33, FBXO41,
FGD3, FGF14, FOXN3, FOXP1, GGT5, GLG1, GLUL, GNA12, GRIN1, GRIP2,
GRK6, HEG1, HM13, HOXA13, IGFN1, KATNB1, KCNK2, KCTD15, KIF21B,
KPNA6, LCNL1, LHFPL4, LHX6, LHX8, LMAN2, LRRTM3, LYPLA2, LZTR1,
M6PR, MAP3K3, MAPK1IP1L, MARCKSL1, MCTS1, MDGA1, MECP2, MLLT6,
MMP19, MPP2, MYO1D, NAGS, NAT8L, NAV1, NDRG4, NFYA, NIPSNAP1, NOL3,
NPTX1, NRP1, NUAK1, NUP43, OBFC1, OFD1, OSBPL7, PBX1, PCDH11X,
PHF15, PHF21A, PIGA, PLEKHM3, PODXL, POLDIP2, POLR2F, PPIL6, PPME1,
PRMT2, PROSC, PTOV1, RAB11FIP3, RAB11FIP4, RAB11FIP5, RAB22A,
RC3H1, REM1, RFX1, RHBDL3, RIMS3, RNF157, RNMTL1, RUNX3, SEC14L1,
SEMA3G, SENP5, SETBP1, SH3PXD2B, SHANK2, SHANK3, SLC17A5, SLC23A2,
SLC26A9, SLC30A6, SLC38A3, SLC6A4, SMARCD1, SNCA, SNX27, SRF,
SRRM1, SS18L1, ST3GAL3, ST7L, STAC3, STK4, STX1B, SUV39H1, SV2A,
SYNPO, SYT9, TBC1D13, TCF7L1, TFAP2A, THRB, TLN2, TMEM79, TRIM10,
TRIM16, TRIM44, TRIM9, TSG101, TSPAN18, TSR2, TTBK1, USH1G, USP54,
VAMP2, VCPIP1, VPS39, WBSCR17, WDR37, WDTC1, WTAP, XYLT1, YEATS2,
ZBTB7B, ZER1, ZHX3, ZNF76, ZNRF1 |
|
|
hsa-miR-4743-5p | AKT1S1, ARL3,
GRIN1, HIC1, NCDN, OLFML2A, SCRT2, ZDHHC8 |
|
|
hsa-miR-5581-5p | APLNR, ATP6V1A,
ATP8A1, BRD4, BTG4, CABP7, CADM1, CALN1, CAPN1, CCDC62, CCL22,
CDON, CHRNB2, CLDN2, CLIC4, CPEB2, CR2, CSNK1D, CTNND2, DCLK2,
DRP2, FAF2, FAM13A, FGA, FNDC5, FOXP1, GGA1, GMEB1, GRIN1, HIF3A,
HNRNPA3, IFFO2, IL4R, IPO7, ITPKB, KCNK3, KPNA6, LAMC1, LHX6, LIPH,
MMP19, MTHFR, MYO5A, NACC1, NCOA3, NEDD4L, NTN1, NWD1, PARP16,
PHF8, PHOSPHO1, PLCB3, PNKD, RALGPS1, RECQL5, RIMKLA, RMND5A,
RNF169, SH3PXD2A, SHROOM4, SLC26A9, SYNGR3, SYT11, TBRG1, TGFBR1,
THSD7A, TP53I11, TPM3, TRIM47, TUB, UBAP2L, UBXN7, UNC119, VEZF1,
ZKSCAN2, ZNF304, ZNF576, ZNF608, ZNF629, ZXDC |
| Down-regulated | hsa-miR-373-3p | A2LD1, AAK1, ABCA1,
ABHD3, ABI2, ABL2, ACBD5, ACVR1C, ADAM9, ADAMTS18, AFF2, AGAP2,
AHNAK, AKAP5, AKTIP, ANKRD13C, ANKRD50, ANKRD52, ANO6, AP1M1,
APBB2, ARHGAP30, ARHGEF10, ARHGEF18, ARHGEF3, ARHGEF7, ARID4A,
ARID4B, ARL4A, ASAP1, ASB1, ASF1B, ASH1L, ATP2B2, ATXN1, BAHD1,
BAMBI, BCAT1, BCL11A, BCL11B, BCL2L11, BCL6B, BMPR2, BNC2, BNIP3L,
BRMS1L, BRWD1, BSCL2, BTBD7, BVES, CAMSAP1, CAMTA1, CASC4, CC2D1A,
CCDC88A, CCND1, CCND2, CD44, CDC25A, CDC40, CDCA7, CFL2, CHD9,
CLIP4, CNN1, CNOT6, CORIN, CREB1, CRK, CROT, CXCL12, CXCL14,
CYB561D1, CYBB, CYBRD1, CYP26B1, DDHD1, DENND5B, DERL2, DGKE,
DIRC2, DLGAP2, DMTF1, DNAJA2, DNAJC27, DPP3, DPP8, DPYSL5, DYNLT3,
EDNRB, EGLN1, EIF4B, ELAVL2, ENDOD1, EPHA2, EPHA5, EPHA7, ERO1LB,
EZH1, FAM102B, FAM117A, FAM18B2, FAM40B, FAM46C, FBXL4, FBXO10,
FBXO41, FGD4, FGD5, FLT1, FMNL3, FOXK2, FOXO3, FRMD4A, FRMD4B,
FYCO1, FZD6, GAB2, GALNT10, GALNT3, GATAD2B, GATC, GDA, GLIS3, GLS,
GNB5, GNG12, GNPDA2, GOLGA1, GPR12, GPR137C, GPR180, GUCY1A3,
HAUS8, HDAC4, HEG1, HIP1, HIPK3, HK1, HLF, HMGXB3, HN1, HNRNPUL2,
HOOK3, HP1BP3, IGDCC3, IKZF2, IL28RA, IL8, INO80D, IPO7, IQSEC1,
IRAK2, IRAK4, IRF2, IRF9, ISM1, ITGB8, JUB, KDM2A, KIAA0226,
KIAA0240, KIAA0513, KIAA1522, KIAA1549, KIAA1737, KIF3B, KLF12,
KLF13, KLF3, KLHL28, KREMEN1, KSR2, LEF1, LEFTY1, LEFTY2, LHX6,
LHX8, LIF, LMO3, LRIT1, LUC7L2, LYPD6, LYRM2, LYSMD3, LYST, MAML1,
MAP1B, MAP3K1, MAP3K14, MAP3K2, MBD2, MBNL2, MBNL3, MCCD1, MCL1,
MDM4, MECP2, MED13L, MFAP3L, MIB1, MICAL3, MKNK2, MKRN1, MLL, MLL3,
MLLT6, MNT, MRPS25, MSL1, MTCH2, MTF1, MTMR3, MTUS1, MYO1D,
NAPEPLD, NCOA3, NCOA7, NECAP1, NEK9, NFATC3, NFIB, NFYA, NHLRC2,
NHLRC3, NNAT, NPAS3, NR2C1, NR2C2, OCRL, ODF2, OPCML, ORMDL3,
OSBPL5, OSTM1, OTUD7B, PAFAH2, PAG1, PAK2, PAM, PAN3, PARP8, PBX3,
PCDH7, PCGF5, PDCD4, PDLIM5, PFN2, PGBD5, PHACTR4, PHC3, PHF6,
PHKA1, PHYHIPL, PIP4K2A, PKD2, PKN2, PLAG1, PLCL1, POFUT1, POLK,
POU6F1, PPARA, PPARGC1B, PPP1R10, PPP1R9A, PPP6C, PRDM16, PRDM8,
PRKACB, PRMT6, PRRT2, PRRX1, PSD3, PSEN1, PTGDR, PTPDC1, RAB11A,
RAB11FIP1, RAB11FIP5, RAB22A, RABEP1, RAD18, RAD23B, RALGDS,
RAPGEF2, RAPGEF5, RAPGEFL1, RASSF2, RBL1, RBMS2, RDX, RELA, RELL1,
RGL1, RGMA, RHOC, RIMKLA, RNF180, RNF216, RNF38, RNF6, RORA,
RPS6KA2, RRAGD, RSBN1, RSBN1L, RSF1, RSRC2, RUNX2, RYR2, SAMD12,
SAR1B, SASH1, SBF1, SCD5, SCN2A, SCN5A, SCRT2, SDC1, SETBP1, SETD7,
SHCBP1, SIK1, SIPA1L3, SLC14A1, SLC16A12, SLC16A9, SLC35E1,
SLC38A1, SLC39A6, SLC46A3, SLC6A9, SMARCC2, SNRK, SNTB2, SNX30,
SNX5, SNX9, SOS1, SPRED1, SS18L1, SSX2IP, ST3GAL5, ST8SIA2, STX16,
SUV420H2, SYAP1, SYDE1, SYNC, SYNPO2, TANC2, TAOK2, TAPT1, TARDBP,
TBCEL, TCEB3, TET2, TET3, TGFBR2, TIAM1, TMCC1, TMEM100, TMTC2,
TMUB2, TNRC18, TNRC6B, TNRC6C, TNS1, TOX, TRAPPC2, TRHDE, TRIM2,
TRIM44, TRIM66, TRPS1, TRPV6, TSEN34, TSHZ3, TTC9, TTPAL, TUSC2,
UBASH3B, UBE2B, UBE2J1, UBE2Q2, UBE2R2, UBE2W, UBN1, UBN2, UHRF1,
UHRF1BP1, ULK1, UNK, UNKL, UPF3A, USP24, USP42, USP46, USP53, VSX1,
WDR26, WDR37, WDR45, WEE1, WIPF2, YTHDF3, ZBTB11, ZBTB41, ZBTB43,
ZBTB44, ZBTB47, ZBTB7A, ZCCHC24, ZDHHC8, ZDHHC9, ZFP91, ZFYVE26,
ZKSCAN1, ZMYND11, ZNF148, ZNF2, ZNF236, ZNF25, ZNF292, ZNF362,
ZNF385A, ZNF436, ZNF473, ZNF512B, ZNF518A, ZNF566, ZNF597, ZNF697,
ZNF862, ZNFX1” |
|
| hsa-miR-4287 | AKT2, AP3M2, APLN,
ASTN1, ATG9A, BAHD1, BHLHE41, BSDC1, BTG2, CALB1, CAMK2A, CAMK2B,
CCDC113, CECR6, COL17A1, CRTC2, DDX3X, DDX3Y, DNAJC21, EHF, EIF2S1,
ENC1, EYA3, FAM117B, FAM76A, GCC1, GRAMD4, HELZ, HUNK, IGSF9B,
KCNA6, KCNK10, KIAA1210, KLF12, KPNA6, KRT80, MDM1, MFAP3L, MID1,
NARG2, NBN, NCAN, NFASC, OPCML, ORAI3, OSBP, PDE1B, PHF23, PI4K2A,
PIK3C2B, PMEPA1, POLD3, RAB1B, RGL1, RIPK1, ROBO2, SGCZ, SGTB,
SH3BP2, SH3RF2, TIGD3, TIMM17B, TOX2, UBN2, VBP1, ZNF48, ZNRF3,
CREB1, CRK, CROT, CXCL12, CXCL14, CYB561D1, CYBB, CYBRD1, CYP26B1,
DDHD1, DENND5B, DERL2, DGKE, DIRC2, DLGAP2, DMTF1, DNAJA2, DNAJC27,
DPP3, DPP8, DPYSL5, DYNLT3, EDNRB, EGLN1, EIF4B, ELAVL2, ENDOD1,
EPHA2, EPHA5, EPHA7, ERO1LB, EZH1, FAM102B, FAM117A, FAM18B2,
FAM40B, FAM46C, FBXL4, FBXO10, FBXO41, FGD4, FGD5, FLT1, FMNL3,
FOXK2, FOXO3, FRMD4A, FRMD4B, FYCO1, FZD6, GAB2, GALNT10, GALNT3,
GATAD2B, GATC, GDA, GLIS3, GLS, GNB5, GNG12, GNPDA2, GOLGA1, GPR12,
GPR137C, GPR180, GUCY1A3, HAUS8, HDAC4, HEG1, HIP1, HIPK3, HK1,
HLF, HMGXB3, HN1, HNRNPUL2, HOOK3, HP1BP3, IGDCC3, IKZF2, IL28RA,
IL8, INO80D, IPO7, IQSEC1, IRAK2, IRAK4, IRF2, IRF9, ISM1, ITGB8,
JUB, KDM2A, KIAA0226, KIAA0240, KIAA0513, KIAA1522, KIAA1549,
KIAA1737, KIF3B, KLF12, KLF13, KLF3, KLHL28, KREMEN1, KSR2, LEF1,
LEFTY1, LEFTY2, LHX6, LHX8, LIF, LMO3, LRIT1, LUC7L2, LYPD6, LYRM2,
LYSMD3, LYST, MAML1, MAP1B, MAP3K1, MAP3K14, MAP3K2, MBD2, MBNL2,
MBNL3, MCCD1, MCL1, MDM4, MECP2, MED13L, MFAP3L, MIB1, MICAL3,
MKNK2, MKRN1, MLL, MLL3, MLLT6, MNT, MRPS25, MSL1, MTCH2, MTF1,
MTMR3, MTUS1, MYO1D, NAPEPLD, NCOA3, NCOA7, NECAP1, NEK9, NFATC3,
NFIB, NFYA, NHLRC2, NHLRC3, NNAT, NPAS3, NR2C1, NR2C2, OCRL, ODF2,
OPCML, ORMDL3, OSBPL5, OSTM1, OTUD7B, PAFAH2, PAG1, PAK2, PAM,
PAN3, PARP8, PBX3, PCDH7, PCGF5, PDCD4, PDLIM5, PFN2, PGBD5,
PHACTR4, PHC3, PHF6, PHKA1, PHYHIPL, PIP4K2A, PKD2, PKN2, PLAG1,
PLCL1, POFUT1, POLK, POU6F1, PPARA, PPARGC1B, PPP1R10, PPP1R9A,
PPP6C, PRDM16, PRDM8, PRKACB, PRMT6, PRRT2, PRRX1, PSD3, PSEN1,
PTGDR, PTPDC1, RAB11A, RAB11FIP1, RAB11FIP5, RAB22A, RABEP1, RAD18,
RAD23B, RALGDS, RAPGEF2, RAPGEF5, RAPGEFL1, RASSF2, RBL1, RBMS2,
RDX, RELA, RELL1, RGL1, RGMA, RHOC, RIMKLA, RNF180, RNF216, RNF38,
RNF6, RORA, RPS6KA2, RRAGD, RSBN1, RSBN1L, RSF1, RSRC2, RUNX2,
RYR2, SAMD12, SAR1B, SASH1, SBF1, SCD5, SCN2A, SCN5A, SCRT2, SDC1,
SETBP1, SETD7, SHCBP1, SIK1, SIPA1L3, SLC14A1, SLC16A12, SLC16A9,
SLC35E1, SLC38A1, SLC39A6, SLC46A3, SLC6A9, SMARCC2, SNRK, SNTB2,
SNX30, SNX5, SNX9, SOS1, SPRED1, SS18L1, SSX2IP, ST3GAL5, ST8SIA2,
STX16, SUV420H2, SYAP1, SYDE1, SYNC, SYNPO2, TANC2, TAOK2, TAPT1,
TARDBP, TBCEL, TCEB3, TET2, TET3, TGFBR2, TIAM1, TMCC1, TMEM100,
TMTC2, TMUB2, TNRC18, TNRC6B, TNRC6C, TNS1, TOX, TRAPPC2, TRHDE,
TRIM2, TRIM44, TRIM66, TRPS1, TRPV6, TSEN34, TSHZ3, TTC9, TTPAL,
TUSC2, UBASH3B, UBE2B, UBE2J1, UBE2Q2, UBE2R2, UBE2W, UBN1, UBN2,
UHRF1, UHRF1BP1, ULK1, UNK, UNKL, UPF3A, USP24, USP42, USP46,
USP53, VSX1, WDR26, WDR37, WDR45, WEE1, WIPF2, YTHDF3, ZBTB11,
ZBTB41, ZBTB43, ZBTB44, ZBTB47, ZBTB7A, ZCCHC24, ZDHHC8, ZDHHC9,
ZFP91, ZFYVE26, ZKSCAN1, ZMYND11, ZNF148, ZNF2, ZNF236, ZNF25,
ZNF292, ZNF362, ZNF385A, ZNF436, ZNF473, ZNF512B, ZNF518A, ZNF566,
ZNF597, ZNF697, ZNF862, ZNFX1 |
|
| hsa-miR-338-5p | AAK1, ADAMTS17,
ADARB2, AEBP2, AMMECR1, APPL1, ARFGAP3, ARID2, ARNT, ATAD1, ATF7,
ATP2C1, ATRX, AUTS2, B4GALT6, BAZ1B, BCL11B, BCL2L11, BTG3, CADM2,
CALM3, CAST, CCDC140, CCNT2, CD28, CD82, CD9, CDK5R1, CDYL2,
CHST12, CLIC4, CLTC, CNR1, CNTN4, CPEB4, CPNE3, CREB3L1, CRIM1,
CSNK1G1, CUL3, DGKG, DICER1, DLAT, DMXL2, DNAJC6, DNM3, DYRK4,
EML1, EP300, EPAS1, EPHA7, ERRFI1, EXOC5, FAM126A, FAM129B,
FAM135B, FAM177A1, FAM18B2, FMNL2, FNDC3B, FOXJ3, FUT9, GATAD2B,
GREM2, GRIA4, GRM7, GTF3C2, GUCY1A3, HCN1, HDAC9, HIF1A, HIPK2,
HSPA12A, IKZF1, IMPACT, INO80D, IREB2, JMJD1C, KAL1, KDM5B,
KIAA1024, KIAA1467, KLF11, KLHL14, KLHL6, KLRAQ1, KRAS, LMO4, LRP1,
MACF1, MBNL1, MBNL2, MCTS1, MEF2C, MIPOL1, MKL2, MLL4, MLLT4, MN1,
MON2, MPPED2, NCK2, NCOA3, NDFIP1, NPAS4, NRP1, NUDT4, NUFIP2,
OCIAD1, ONECUT2, PARD6B, PCDH17, PCDH20, PCGF5, PCNX, PELI1, PHC3,
PHIP, PKN2, PLAGL2, PLEKHA5, PPARGC1A, PPM1B, PPP2R5A, PRDM10,
PRLR, PTCHD1, PTGS1, R3HDM2, RAB14, RAB1A, RAB22A, RAB6B, RAP2C,
RAPGEF5, RAPH1, RCOR1, RICTOR, RND3, RNF138, RORA, SAMD12, SBNO1,
SEC16B, SEMA6A, SERTAD2, SIRT1, SKP1, SLC4A7, SLIT1, SLMAP, SNTB1,
SOX6, SPAST, SPOP, SSX2IP, STAG2, SUB1, SYNCRIP, SYPL1, TAF4,
TANC2, TARDBP, TBX18, TBX2, TCERG1L, TEAD1, TET2, TLK1, TRA2B,
TRAF3, TRPM7, TSHZ3, UBE2N, UBR2, USP25, WASF1, WDFY3, WWC3,
ZBTB44, ZFAND5, ZNF292 |
|
| hsa-miR-623 | AAK1, ACSM2A,
ADARB2, AGPAT4, ALPL, AP3M2, APPL1, ATG9A, BAHD1, CACNA1C, CAMK2B,
CCDC117, CCDC3, CELSR3, CLUAP1, CORO2A, CRTC2, CXCL12, DCLK1,
DCLK3, DSEL, ECE1, EGFLAM, EIF1, ELAVL2, EZR, FAM126A, FAM134C,
FOXN2, GATAD2B, GLIS3, HAS3, HGSNAT, HLCS, HM13, HMGA2, HOXC10,
HOXC9, IGF2R, ILDR2, KIAA1199, KPNA1, MAPK1, MECP2, MEIS1, MFSD11,
MON2, MTMR7, NIPBL, NMT1, NR3C2, NTRK2, NTRK3, OBFC2A, ODZ4, PCMT1,
PDE4A, PI4KB, PLCD4, POLD3, PRIMA1, RBM24, RHOBTB3, RIMKLA, RIN3,
RNF144A, RNF169, RPRD2, SECISBP2L, SH3PXD2A, SH3TC2, SIGLEC1, SKI,
SLC12A2, SLC44A5, SNX13, SUPT16H, TAOK2, TET3, TNFRSF8, TNRC6B,
TPM3, TRIM31, TRPS1, ZMIZ1 |
|
| hsa-miR-3653 | ACVR1C, ADCY2,
AEBP2, AMIGO2, ATP1B4, ATRNL1, ATXN7, BMP3, BMPR2, BNC2, BRD3,
BRPF3, BRWD3, BTG1, CCDC88A, CPEB4, DBT, DGCR2, DIXDC1, DUSP19,
EFNB3, ESRRB, FAM107B, FASLG, GALNT2, GJC1, GPC2, GPC6, HCFC2,
HIPK3, KIAA0947, KIAA2018, KLHL28, LPCAT2, LRRTM2, MED12L, MKLN1,
MYSM1, NCOA1, NIP7, ODZ3, PCDH11X, PDE11A, PHLDA1, PI15, PRPF4B,
R3HDM1, RBBP4, SEC62, SERBP1, SORT1, SPATA5, SV2B, TMEM215,
TMEM50B, TRIM67, TRPM8, VPS33A, YAF2, ZADH2, ZDHHC21, ZFAND5, ZFY,
ZNF280C, ZNF507, ZYG11B |
|
| hsa-miR-590-5p | ARHGAP24, ARHGEF12,
ARMCX1, BAHD1, BMP3, BMPR2, CADM1, CCL22, CEP68, CNOT6, CREB5,
DAG1, DSC2, EIF2C4, EIF4EBP2, ELF2, ENAH, EPHA4, FAM13A, FAM3C,
FASLG, FBXO28, FGD4, FGF1, FRS2, GABRB2, GATAD2B, GLCCI1, GPR64,
ITGB8, JHDM1D, JPH1, KCNT2, KLF12, KLHDC5, LCORL, LRRC57, MATN2,
MBNL1, MICALL1, MTMR12, NELL2, NFAT5, NFIB, OSR1, PAG1, PAIP2B,
PAN3, PBRM1, PCBP2, PDZD2, PER2, PGRMC2, PIK3R1, PLAG1, PLEKHA1,
PPP1R3B, PTPN9, RAB22A, RASGRP1, RAVER2, RBPJ, RECK, RFFL, RP2,
RPRD1A, SATB1, SECISBP2L, SESTD1, SETD1B, SKI, SLC7A6, SNTB2,
SNX29, SPRY2, ST3GAL6, STAG2, TAGAP, TBX2, TET1, TGFB2, TGFBR2,
TNRC6B, UBE2D3, UBN2, UBR3, YOD1, ZCCHC3, ZNF704 |
|
|
hsa-miR-664b-3p | AASS, ABCE1, ABI2,
ACVR2B, ADD3, AKNA, APAF1, APC, ARPM1, ASB13, ATG7, ATP2C1, BACH1,
BACH2, BCAS1, BDNF, BHLHB9, BNC2, CA5B, CACHD1, CCNC, CENPL, COPA,
CREG2, DCP2, DENND4C, DIP2B, DPY19L1, EDAR, ERBB4, ETNK1, FAM114A1,
FAM8A1, FBXW2, FNDC3A, FRMD4A, FZD5, GPRASP2, HIPK2, HMGA2, INTS6,
JPH1, KCTD21, KLF12, KLHDC10, LDB3, LMTK3, MED1, MSR1, MTCH2, MTR,
MYCBP, MYO1D, N4BP2, NDFIP1, NETO1, NFIB, NIPAL3, NOTCH2, NSL1,
NUFIP2, PAPD5, PARVA, PDE4D, PDYN, PELI2, PGM3, PHIP, PI15, PKP1,
PRDM10, PRDM15, PRKAA1, PRKAA2, PRLR, QKI, RANBP9, RAPGEF6, RIBC1,
RS1, RSU1, SALL4, SAMD12, SH2D4B, SMAD3, SNX30, STT3A, TAF4,
TGOLN2, TMEM215, TMEM26, TRAM2, TXLNA, UTP23, VAMP1, VAMP4, WDFY1,
WWC2, YKT6, ZFHX3, ZFP28, ZFX, ZNF24 |