Introduction
Lung cancer remains the leading cause of cancer
mortalities worldwide, particularly, in economically developed
countries, although marginal progress has been made in the
treatment of lung cancer (1). Based
on its histologic features and responses to conventional
therapeutic strategies, lung cancer has been divided into two major
categories: Non-small cell lung cancer (NSCLC; one of the most
lethal types of cancer) and small cell lung cancer. Due to the
incurable nature of lung cancer, NSCLC is considered to be a
terminal illness with a five-year survival rate of ~16% (2). The poor prognosis is predominantly
attributed to metastasis in the early stages of NSCLC. Therefore,
investigating the potential biological markers of cancer invasion
and metastasis is urgently required for guidance on postoperative
surveillance and therapeutic decisions.
Cancer invasion and metastasis is a multistep
process, which involves tumor cells escaping from a primary site,
migrating into the blood or lymphatic system and re-establishing
novel tumors at a distant site. Various signaling molecules that
regulate multiple cellular processes are involved in cell migration
(3). For example, the actin
cytoskeleton is dynamically remodeled during cell migration, and
this reorganization produces the force that is necessary for cell
migration (4). Previous studies
identified that Wiskott-Aldrich syndrome protein family 3 (WASF3;
also termed, WAVE3), is critical for the regulation of actin
cytoskeleton dynamics via the activation of the Arp2/3 complex, and
is involved in cancer cell motility, invasion and metastasis
(5–8). Downregulation of WASF3 has been found
to inhibit the invasion and metastasis of breast cancer cells, and
has been proposed as a metastasis promoter gene (8). In addition, it has been reported that,
compared with lower stage tumors and normal tissue, WASF3
expression is increased in advanced breast and prostate cancer
(9,10). However, currently, little is known
about the expression status of WASF3 and its association with the
clinicopathological features of NSCLC. In the present study, the
mRNA expression levels of WASF3 were analyzed in 38 NSCLC patients
and in matched normal tissues, and the protein expression status in
96 specimens was analyzed using the quantitative polymerase chain
reaction (qPCR) and immunohistochemistry IHC. The correlations
between WASF3 expression patterns and the clinicopathological
features of NSCLC were analyzed. In addition, the association
between WASF3 expression and the five-year overall survival was
evaluated.
Patients and methods
Study population and samples
138 NSCLC patients who were diagnosed with, and
underwent surgical removal of, a primary lesion at the First
Affiliated Hospital of Liaoning Medical University (Liaoning,
China) were included in the present study. None of the patients
underwent radiotherapy or chemotherapy prior to surgery.
Histopathological evaluation was conducted independently by two
pathologists. All of the clinical and follow-up data were based on
studies from the tumor registry office of the First Affiliated
Hospital of Liaoning Medical University. The present study was
approved by the Ethics Committee of Liaoning Medical University and
written informed consent was acquired from each patient.
One-hundred lung cancer tissue samples from these patients were
formalin-fixed, paraffin-embedded and prepared for IHC.
Postoperative follow-up endured for at least five years for 96
patients, while four patients failed to be followed up and were
excluded from the study. Fresh tumor tissues and matched normal
tissues from an additional 38 NSCLC patients were immediately
transferred to liquid nitrogen and stored at −80°C for subsequent
qPCR. Clinicopathological data are summarized in Table I.
 | Table ICharacteristics of the non-small cell
lung cancer patients (n=134). |
Table I
Characteristics of the non-small cell
lung cancer patients (n=134).
| Variable | qPCR analysis
(n=38) | IHC analysis
(n=96) |
|---|
| Gender |
| Female | 23 | 56 |
| Male | 15 | 40 |
| Age, years |
| <60 | 22 | 44 |
| ≥60 | 16 | 52 |
| Histological
subtype |
| Adenocarcinoma | 20 | 52 |
| Squamous cell
carcinoma | 18 | 44 |
| Differentiation
status |
| Well | 14 | 30 |
| Moderate | 12 | 42 |
| Poor | 12 | 24 |
| Lymph node
metastasis |
| Negative | 16 | 42 |
| Positive | 22 | 54 |
| Tumor staging |
| IA–IB | 16 | 32 |
| IIA–IIB | 16 | 44 |
| IIIA | 6 | 20 |
Total RNA extraction and cDNA
synthesis
Total RNA was extracted from the tissues using the
TRIzol RNA kit (Invitrogen Life Technologies, Carlsbad, CA, USA) in
accordance with the manufacturer’s instructions. Ultraviolet
spectroscopy was performed using an Eppendorf BioPhotometer
(Eppendorf, Hamburg, Germany) to determine the RNA concentration
and purity. Total RNA (2 μg) was reverse-transcribed into
first-strand cDNA using the first-strand PrimeScript™ RT Reagent
kit with gDNA Eraser (Takara Bio, Inc., Japan), according to the
manufacturer’s instructions. The final reaction volume was 20
μl.
qPCR
qPCR using the SYBR® Green I (Takara Bio,
Inc.) technique was adopted to examine the WASF3 expression levels
of the tissues from the 38 NSCLC patients. Primers spanning at
least one intron were chosen to minimize inaccuracies due to
genomic DNA contamination. The housekeeping gene,
glyceraldehydes-3-phosphate dehydrogenase (GAPDH) served as an
internal control. The primer sequences used were as follows: Sense,
5′-TGA TAA CTG AGC CAA AGT GGT GAT G-3′ and antisense, 5′-TGG CGT
ATG ATA GCG GCA AG-3′ (PCR product length, 198 bp) for WASF3; and
sense, 5′-GCA CCG TCA AGG CTG AGA AC-3′ and antisense, 5′-TGG TGA
AGA CGC CAG TGG A-3′ (PCR product length, 138 bp) for GAPDH. The
qPCR was run on a Mastercycler® ep realplex (Eppendorf)
using a SYBR® Premix Ex Taq™ kit (Takara Bio, Inc.).
Briefly, PCR (total volume, 20 μl) was performed with 2 μl cDNA,
0.2 μM each primer pair and 10 μl SYBR® Premix Ex Taq
(2× concentration). Following the initial denaturation at 95°C for
30 sec, 40 three-segment cycles consisting of the following
procedure were performed: 5 sec at 95°C; 30 sec at 55°C; and 30 sec
at 72°C. The fluorescence was automatically measured using the
Mastercycler® ep realplex (Eppendorf) during PCR and
during one three-segment cycle of product melting (95°C for 15 sec,
60°C for 15 sec, 95°C for 15 sec). In order to further verify the
amplification of the desired fragments, the PCR products were
assessed via electrophoresis analysis on 3% agarose gel. The
2−ΔΔCT method was used to present the data of the genes
of interest relative to an internal control gene (11,12).
IHC
Paraffin-embedded 4-μm thick tissue sections were
deparaffinized in a series of xylene baths and rehydrated during
graded alcohol washes. All sections were retrieved by microwave
treatment and treated with 3% hydrogen peroxidase for 15 min to
block endogenous peroxidase activity. The sections were
subsequently incubated with the primary anti-WASF3 rabbit antibody
(1:100; ab-110739; Abcam, Cambridge, UK) at 4°C overnight.
Thereafter, the sections were stained with a ready-to-use secondary
anti-rabbit antibody conjugated with horseradish peroxidase
(Zhongshan Biotechnology Inc., Beijing, China) for 30 min at room
temperature. The stained specimens were exposed to a
3,3′-diaminobenzidine kit (Zhongshan Biotechnology Inc., Beijing,
China) at room temperature for 1 min and counterstained with
hematoxylin. The primary antibodies were replaced with
phosphate-buffered saline to serve as the negative controls. All
slides were independently examined and scored by two pathologists,
who were blind to the clinical and pathological data of the
subjects. Cancer cells in at least five fields were counted at a
magnification of ×200 using a Zeiss Imager A1 (Carl Zeriss AG,
Oberkochen, Germany). For the WASF3 IHC assessment, the ratio of
positive cells per specimen and staining intensity were analyzed.
The WASF3 immunoreactivity level was classified by the proportion
of positive cells as follows: 0, <5% positive cells; 1+, 5–30%
positive cells; 2+, >30–50% positive cells; and 3+, >50%
positive cells. In addition, the intensity of WASF3 expression was
scored: 0, Negative to weak; 1, moderate; and 2, strong. The score
was the sum of the intensity and the percentage of positive cells.
A score of ≤1 was applied as a cut-off point for loss of WASF3
expression.
Statistical analysis
A two-sample t-test for independent samples was used
for the continuous variables. Statistical analysis for comparing
between groups regarding categorical data was performed using the
χ2-test. Comparison of more than two groups with
continuous variables was performed with the Kruskal-Wallis one-way
analysis of variance by ranks. The Kaplan-Meier method with
log-rank test was used for comparing survival curves between the
groups. Cox regression (or proportional hazards regression) was
adopted to analyze the effect of various risk factors on survival.
SPSS software version 16.0 (SPSS Inc., Chicago, IL, USA) was used
for statistical analysis, tests were two-sided and P<0.05 was
considered to indicate a statistically significant difference.
Results
mRNA expression of WASF3 in NSCLC and
correlation with clinicopathological features of NSCLC
The mRNA expression levels of WASF3 in 38 NSCLC
patients and in matched normal lung tissue samples were
quantitatively assessed using qPCR and the SYBR® Green I
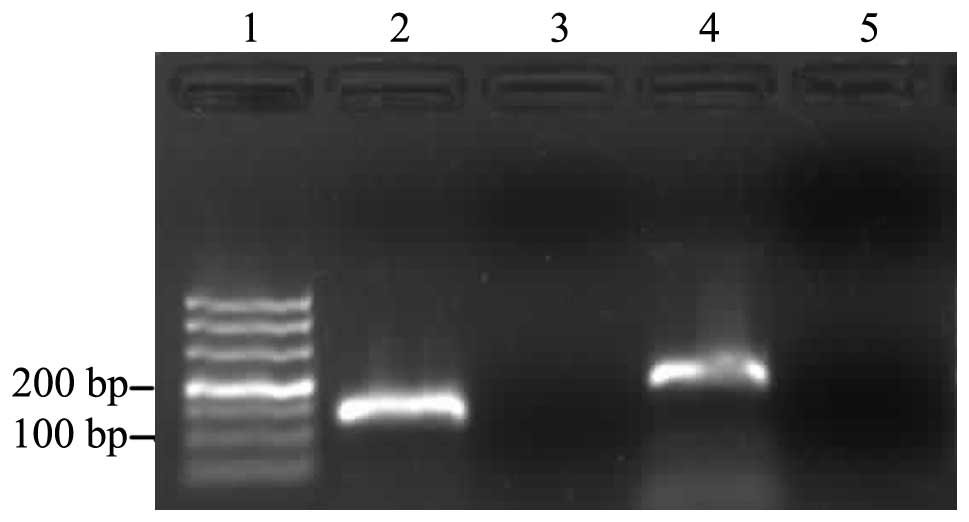
technique. Gel electrophoresis analysis of the amplification
products revealed a single band with the anticipated size for WASF3
(198 bp) and GAPDH (138 bp; Fig.
1). Furthermore, melting curve analysis identified the specific
amplification of the target and reference genes. The mRNA
expression level of WASF3 was markedly (approximately five times)
higher in the NSCLC tissues (4.8373±0.3142) compared with that in
the normal tissues (1.000) (Fig.
2).
The association between WASF3 mRNA expression in the
NSCLC tissues and various clinicopathological features was analyzed
(Table II). The expression of
WASF3 was significantly higher in adenocarcinoma compared with that
in squamous cell carcinoma (P=0.010). There was a significant
correlation observed between WASF3 expression and the tumor stage
(P=0.013). However, significant correlation between WASF3 mRNA
expression, and lymph node metastasis status and differentiation
status (P=0.815 and P=0.214, respectively) was not detected.
 | Table IICorrelation between WASF3 mRNA
expression and the clinicopathological features. |
Table II
Correlation between WASF3 mRNA
expression and the clinicopathological features.
| | WASF3 expression |
|---|
| |
|
|---|
| Variable | Patients, n | Cancer tissuea | P-value |
|---|
| Histological
subtype | | | 0.010 |
| Adenocarcinoma | 20 | 5.5852±0.4402 | |
| Squamous cell
carcinoma | 18 | 4.0062±0.3686 | |
| Lymph node
metastasis | | | 0.815 |
| Negative | 16 | 4.9254±0.5049 | |
| Positive | 22 | 4.7732±0.4094 | |
| Differentiation
status | | | 0.241 |
| Well | 14 | 4.1408±0.5059 | |
| Moderate | 12 | 5.1874±0.6482 | |
| Poor | 12 | 5.2997±0.4434 | |
| Tumor staging | | | 0.013 |
| IA–IB | 16 | 4.1336±0.3875 | |
| IIA–IIB | 16 | 4.8171±0.4960 | |
| IIIA | 6 | 6.7677±0.6696 | |
Expression of the WASF3 protein in NSCLC
and correlation with clinicopathological features
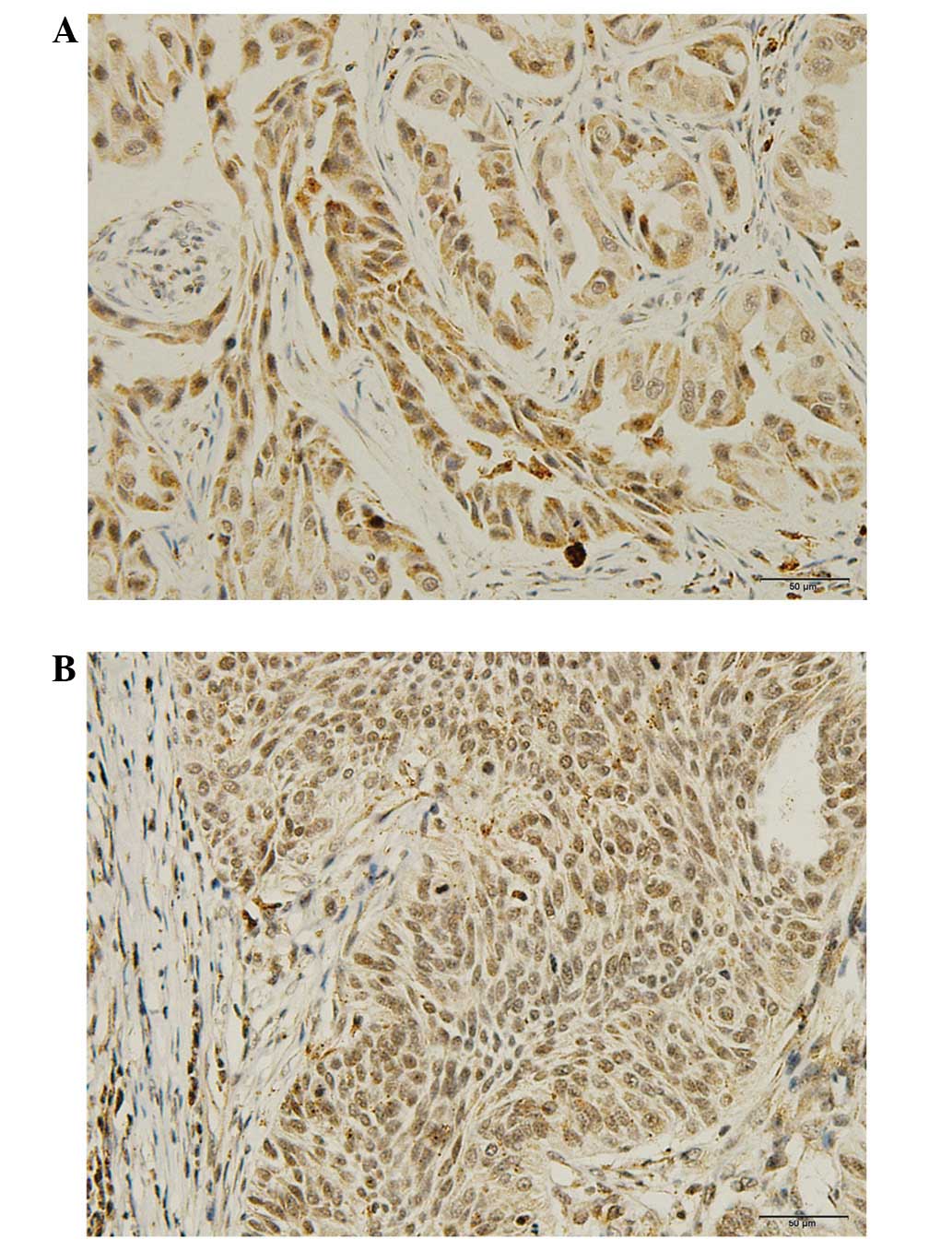
IHC demonstrated that the WASF3 protein was
predominantly localized to the cytoplasm (Fig. 3). Positive WASF3 expression was
observed in 53 (55.2%) cases and negative expression was noted in
43 (44.8%) cases out of the 96 patients. As anticipated, it was
found that WASF3 expression in the NSCLC cases was significantly
correlated with the histological subtype and tumor staging (P=0.001
and P=0.024, respectively), however, was not correlated with
gender, age, differentiation status or lymph node metastasis.
Detailed results are presented in Table III.
 | Table IIIAssociation between WASF3 protein
expression and clinicopathological parameters in non-small cell
lung cancer. |
Table III
Association between WASF3 protein
expression and clinicopathological parameters in non-small cell
lung cancer.
| | WASF3 expression | |
|---|
| |
| |
|---|
| Variable | Patients (n) | Pos. (n=53) | Neg. (n=43) | P-value |
|---|
| Gender | | | | 0.274 |
| Male | 56 | 31 | 25 | |
| Female | 40 | 22 | 18 | |
| Age, years | | | | 0.543 |
| <60 | 44 | 24 | 20 | |
| ≥60 | 52 | 29 | 23 | |
| Histological
subtype | | | | 0.001 |
| Adenocarcinoma | 52 | 37 | 15 | |
| Squamous cell
carcinoma | 44 | 16 | 28 | |
| Differentiation
status | | | | 0.481 |
| Well | 30 | 14 | 16 | |
| Moderate | 42 | 24 | 18 | |
| Poor | 24 | 15 | 9 | |
| Lymph node
metastasis | | | | 0.623 |
| Negative | 42 | 22 | 20 | |
| Positive | 54 | 31 | 23 | |
| Tumor staging | | | | 0.024 |
| IA–IB | 32 | 12 | 20 | |
| IIA–IIB | 44 | 26 | 18 | |
| IIIA | 20 | 15 | 5 | |
Correlation between expression of WASF3
and overall survival
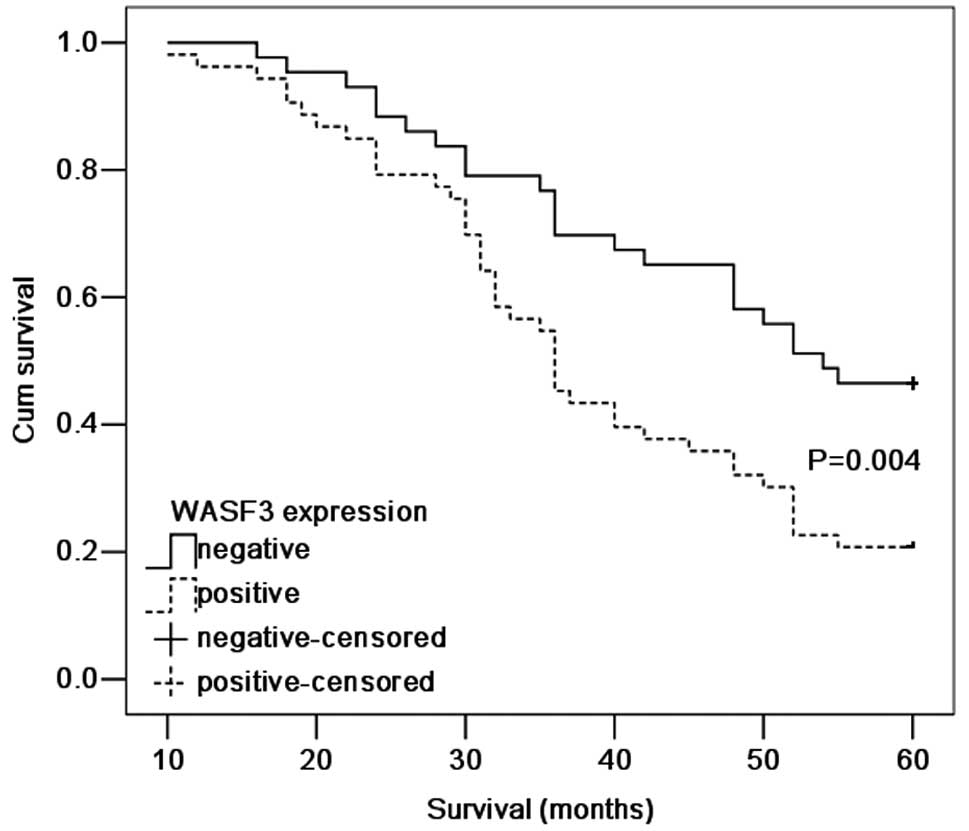
According to the Kaplan-Meier survival analysis,
patients exhibiting WASF3-positive expression had a poorer
prognosis (P=0.004; Fig. 4)
compared with those patients with WASF3-negative expression. The
five-year survival rate for patients with high expression of WASF3
was 20.8% compared with 46.5% in patients exhibiting a low
expression. The Cox proportional hazards model was adopted to
perform univariate and multivariate analysis of survival. As a
result of the univariate analysis, the tumor staging (P=0.005) and
WASF3 expression (P=0.006) were identified to be associated with
overall survival. In the multivariate analysis, the expression of
WASF3 emerged as an independent and significant factor, which was
associated with a poor five-year survival rate (relative risk,
0.463; 95% CI, 0.271–0.792). Detailed results are presented in
Table IV.
 | Table IVCox proportional hazard regression
model analysis. |
Table IV
Cox proportional hazard regression
model analysis.
| Univariate
analysis | Multivariate
analysis |
|---|
|
|
|
|---|
| Variable | Relative risk | 95% CI | P-value | Relative risk | 95% CI | P-value |
|---|
| Gender |
| Male vs.
Female | 1.218 | 0.739–2.007 | 0.440 | 1.400 | 0.842–2.327 | 0.194 |
| Age, years |
| <60 vs.
≥60 | 0.724 | 0.443–1.185 | 0.199 | 0.642 | 0.383–1.075 | 0.092 |
| Histological
subtype |
| AD vs. SCC | 1.070 | 0.657–1.744 | 0.785 | 0.823 | 0.483–1.405 | 0.476 |
| Differentiation
status |
| Well + Moderate
vs. Poor | 0.949 | 0.546–1.650 | 0.852 | 1.090 | 0.606–1.963 | 0.773 |
| Tumor stage |
| I+II vs. III | 0.446 | 0.255–0.780 | 0.005 | 0.420 | 0.232–0.762 | 0.004 |
| Lymph node
metastasis |
| Negative vs.
Positive | 0.740 | 0.452–1.213 | 0.233 | 0.891 | 0.527–1.507 | 0.667 |
| WASF3
expression |
| Negative vs.
Positive | 0.491 | 0.294–0.819 | 0.006 | 0.463 | 0.271–0.792 | 0.005 |
Discussion
Metastasis is accountable for ~90% of mortalities in
patients with solid tumors (13–18)
and is the most problematic issue during cancer treatment.
Mechanistic and clinical studies have clearly demonstrated WASF3 as
a critical component in cancer progression and metastasis (19). Furthermore, recent studies have
reported on the critical role of WASF3 in numerous malignancies,
including prostate (9,20), breast (21) and colon cancer (22). However, there is limited data
available regarding the expression status of WASF3 in NSCLC. In the
present study, the results of IHC and qPCR clearly demonstrate that
WASF3 expression was increased in the tumor tissue samples, which
indicates a potential role for the WASF3 protein in the
pathogenesis of NSCLC. In addition, the present data demonstrated
that WASF3 expression was significantly correlated with the
histological subtype and tumor staging, which is consistent with
previous studies regarding breast and prostate cancer (6,20).
However, a significant association was not observed between WASF3
expression and lymph node metastasis in the present study. This may
be due to the smaller sample size, which did not provide adequate
power to detect such a difference. Furthermore, the present
findings indicated a significant association between WASF3
expression and a reduced overall survival in the univariate and
multivariate analyses. These results indicate that WASF3 may have
prognostic value and may present as a possible therapeutic target
for the treatment of lung cancer.
WASF3, as a member of the Wiskott–Aldrich syndrome
protein/WAVE family of structurally and functionally associated
proteins, is significant in the regulation of actin polymerization
in the cytoskeleton, cell motility and cancer cell invasion
(5,6,8,23–25).
Additionally, WASF3 has been hypothesized to be involved in cancer
invasion in numerous cancer cell lines (5,8–10,20,21,26,27),
which implies that WASF3 may be critical in cancer progression and
metastasis. Fernando et al (20) reported that the expression of WASF3
is stronger in prostate cancer tissues when compared with normal
tissues and the expression of WASF3 was found to be significantly
correlated with advanced human prostate cancer. The findings of the
present study concerning NSCLC have been corroborated by similar
findings in breast cancer patients; Kulkarni et al (21) observed that WASF3 expression was
increased in the tumors of patients who developed distant
metastases and markedly upregulated in the more aggressive
triple-negative breast cancer patients. The data from the present
study indicates that WASF3 may present as a useful biomarker for
cancer progression and metastasis.
A significant correlation between the expression
levels of WASF3 and the stage of lung cancer was observed in the
present study. Furthermore, the patients exhibiting WASF3-positive
expression were associated with a poorer prognosis when compared
with those exhibiting WASF3-negative expression. Notably, the
protein expression of WASF3 was not significantly increased in
patients with lymph node metastases when compared with those
without lymph node metastases. This was not consistent with
previous results regarding breast cancer (21); however, a larger sample size may be
required in future study. Zhang et al (22) found that WASF3 was overexpressed in
colorectal cancer tissues, however, the colorectal cancer patients
with WASF3 expression were associated with a good prognosis. The
findings of the present study conflict with the abovementioned
results regarding colorectal cancer. An explanation for this may be
that there are numerous complex factors, which affect the
interactions that occur in vivo. For example, the p38
mitogen-activated protein kinase (MAPK) signaling pathways, which
are activated by WASF3, may possess a selective advantage in
several types of cancer (28). The
activated p38 MAPK signaling pathway may act as either a tumor
suppressor (29,30) or as a tumor promoter depending on
the type of cancer (31–33). Therefore, further investigation into
the underlying mechanisms are required.
In conclusion, the present study identified that
WASF3 was upregulated in NSCLC tissues, which indicates a potential
role for WASF3 in the pathogenesis of NSCLC. Notably, this result
provides clear support with regard to the function of WASF3 as a
metastasis promoter protein. Furthermore, it was found that WASF3
may serve as a predictive marker of overall survival in NSCLC
patients and may provide a potential target for anti-tumor therapy.
However, in order to fully elucidate the exact function of WASF3 in
NSCLC, further, larger studies are required.
Acknowledgements
The present study was financially supported by the
Youth Science and Technology Foundation of the First Affiliated
Hospital of Liaoning Medical University (Liaoning, China; grant no.
FY2011–02).
References
|
1
|
Collins LG, Haines C, Perkel R and Enck
RE: Lung cancer: diagnosis and management. Am Fam Physician.
75:56–63. 2007.
|
|
2
|
American Cancer Society. Cancer facts
& figures 2012. American Cancer Society, Inc.; Atlanta:
2012
|
|
3
|
Ridley AJ, Schwartz MA, Burridge K, et al:
Cell migration: integrating signals from front to back. Science.
302:1704–1709. 2003.
|
|
4
|
Pollard TD and Borisy GG: Cellular
motility driven by assembly and disassembly of actin filaments.
Cell. 112:453–465. 2003.
|
|
5
|
Sossey-Alaoui K, Ranalli TA, Li X, Bakin
AV and Cowell JK: WAVE3 promotes cell motility and invasion through
the regulation of MMP-1, MMP-3, and MMP-9 expression. Exp Cell Res.
308:135–145. 2005.
|
|
6
|
Sossey-Alaoui K, Li X and Cowell JK:
c-Abl-mediated phosphorylation of WAVE3 is required for
lamellipodia formation and cell migration. J Biol Chem.
282:26257–26265. 2007.
|
|
7
|
Sossey-Alaoui K, Downs-Kelly E, Das M,
Izem L, Tubbs R and Plow EF: WAVE3, an actin remodeling protein, is
regulated by the metastasis suppressor microRNA, miR-31, during the
invasion-metastasis cascade. Int J Cancer. 129:1331–1343. 2011.
|
|
8
|
Sossey-Alaoui K, Safina A, Li X, Vaughan
MM, Hicks DG, Bakin AV and Cowell JK: Down-regulation of WAVE3, a
metastasis promoter gene, inhibits invasion and metastasis of
breast cancer cells. Am J Pathol. 170:2112–2121. 2007.
|
|
9
|
Teng Y, Ren MQ, Cheney R, Sharma S and
Cowell JK: Inactivation of the WASF3 gene in prostate cancer cells
leads to suppression of tumorigenicity and metastases. Br J Cancer.
103:1066–1075. 2010.
|
|
10
|
Teng Y, Liu M and Cowell JK: Functional
interrelationship between the WASF3 and KISS1 metastasis-associated
genes in breast cancer cells. Int J Cancer. 129:2825–2835.
2011.
|
|
11
|
Chen S, Zha X, Yang L, Li B, Liye Z and Li
Y: Deficiency of CD3gamma, delta, epsilon and zeta expression in
T-cells from AML patients. Hematology. 16:31–36. 2011.
|
|
12
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
|
|
13
|
Berx G, Raspé E, Christofori G, Thiery JP
and Sleeman JP: Pre-EMTing metastasis? Recapitulation of
morphogenetic processes in cancer. Clin Exp Metastasis. 24:587–597.
2007.
|
|
14
|
Chiang AC and Massagué J: Molecular basis
of metastasis. N Engl J Med. 359:2814–2823. 2008.
|
|
15
|
Savagner P: Leaving the neighborhood:
molecular mechanisms involved during epithelial-mesenchymal
transition. Bioessays. 23:912–923. 2001.
|
|
16
|
Spaderna S, Schmalhofer O, Hlubek F, Jung
A, Kirchner T and Brabletz T: Epithelial-mesenchymal and
mesenchymal-epithelial transitions during cancer progression. Verh
Dtsch Ges Pathol. 91:21–28. 2007.
|
|
17
|
Nguyen DX and Massagué J: Genetic
determinants of cancer metastasis. Nat Rev Genet. 8:341–352.
2007.
|
|
18
|
Nguyen DX, Bos PD and Massagué J:
Metastasis: from dissemination to organ-specific colonization. Nat
Rev Cancer. 9:274–284. 2009.
|
|
19
|
Sossey-Alaoui K: Surfing the big WAVE:
Insights into the role of WAVE3 as a driving force in cancer
progression and metastasis. Semin Cell Dev Biol. 24:287–297.
2013.
|
|
20
|
Fernando HS, Sanders AJ, Kynaston HG and
Jiang WG: WAVE3 is associated with invasiveness in prostate cancer
cells. Urol Oncol. 28:320–327. 2010.
|
|
21
|
Kulkarni S, Augoff K, Rivera L, McCue B,
Khoury T, Groman A, et al: Increased expression levels of WAVE3 are
associated with the progression and metastasis of triple negative
breast cancer. PLoS One. 7:e428952012.
|
|
22
|
Zhang Y, Guan XY, Dong B, Zhao M, Wu JH,
Tian XY and Hao CY: Expression of MMP-9 and WAVE3 in colorectal
cancer and its relationship to clinicopathological features. J
Cancer Res Clin Oncol. 138:2035–2044. 2012.
|
|
23
|
Suetsugu S and Takenawa T: Regulation of
cortical actin networks in cell migration. Int Rev Cytol.
229:245–286. 2003.
|
|
24
|
Millard TH, Sharp SJ and Machesky LM:
Signalling to actin assembly via the WASP (Wiskott-Aldrich syndrome
protein)-family proteins and the Arp2/3 complex. Biochem J.
380:1–17. 2004.
|
|
25
|
Sossey-Alaoui K, Li X, Ranalli TA and
Cowell JK: WAVE3-mediated cell migration and lamellipodia formation
are regulated downstream of phosphatidylinositol 3-kinase. J Biol
Chem. 280:21748–21755. 2005.
|
|
26
|
Teng Y, Ngoka L, Mei Y, Lesoon L and
Cowell JK: HSP90 and HSP70 proteins are essential for stabilization
and activation of WASF3 metastasis-promoting protein. J Biol Chem.
287:10051–10059. 2012.
|
|
27
|
Condeelis J, Singer RH and Segall JE: The
great escape: when cancer cells hijack the genes for chemotaxis and
motility. Annu Rev Cell Dev Biol. 21:695–718. 2005.
|
|
28
|
Wagner EF and Nebreda AR: Signal
integration by JNK and p38 MAPK pathways in cancer development. Nat
Rev Cancer. 9:537–549. 2009.
|
|
29
|
Engel FB, Schebesta M, Duong MT, Lu G, Ren
S, Madwed JB, et al: p38 MAP kinase inhibition enables
proliferation of adult mammalian cardiomyocytes. Genes Dev.
19:1175–1187. 2005.
|
|
30
|
Hui L, Bakiri L, Stepniak E and Wagner EF:
p38alpha: a suppressor of cell proliferation and tumorigenesis.
Cell Cycle. 6:2429–2433. 2007.
|
|
31
|
Lee RJ, Albanese C, Stenger RJ, Watanabe
G, Inghirami G, Haines GK III, et al: pp60(v-src) induction of
cyclin D1 requires collaborative interactions between the
extracellular signal-regulated kinase, p38, and Jun kinase
pathways. A role for cAMP response element-binding protein and
activating transcription factor-2 in pp60(v-src) signaling in
breast cancer cells. J Biol Chem. 274:7341–7350. 1999.
|
|
32
|
Ricote M, García-Tuñón I, Bethencourt F,
Fraile B, Onsurbe P, Paniagua R and Royuela M: The p38 transduction
pathway in prostatic neoplasia. J Pathol. 208:401–407. 2006.
|
|
33
|
Recio JA and Merlino G: Hepatocyte growth
factor/scatter factor activates proliferation in melanoma cells
through p38 MAPK, ATF-2 and cyclin D1. Oncogene. 21:1000–1008.
2002.
|