|
1
|
Masters GA, Krilov L, Bailey HH, Brose MS,
Burstein H, Diller LR, Dizon DS, Fine HA, Kalemkerian GP, Moasser
M, et al: Clinical cancer advances 2015: Annual report on progress
against cancer from the American society of clinical oncology. J
Clin Oncol. 33:786–809. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ferlay J, Shin HR, Bray F, Forman D,
Mathers C and Parkin DM: Estimates of worldwide burden of cancer in
2008: GLOBOCAN 2008. Int J Cancer. 127:2893–2917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Miller AJ and Mihm MC Jr: Melanoma. N Engl
J Med. 355:51–65. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Boyle GM: Therapy for metastatic melanoma:
An overview and update. Expert Rev Anticancer Ther. 11:725–737.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Sullivan R, LoRusso P, Boerner S and
Dummer R: Achievements and challenges of molecular targeted therapy
in melanoma. Am Soc Clin Oncol Educ Book. 2015:177–186. 2015.
View Article : Google Scholar
|
|
6
|
Kalia M: Biomarkers for personalized
oncology: Recent advances and future challenges. Metabolism. 64(3):
Suppl 1. S16–S21. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Moschos SJ and Pinnamaneni R: Targeted
therapies in melanoma. Surg Oncol Clin N Am. 24:347–358. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang K, Huang C and Nice E: Recent
advances in proteomics: Towards the human proteome. Biomed
Chromatogr. 28:848–857. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bougnoux AC and Solassol J: The
contribution of proteomics to the identification of biomarkers for
cutaneous malignant melanoma. Clin Biochem. 46:518–523. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Byrum SD, Larson SK, Avaritt NL, Moreland
LE, Mackintosh SG, Cheung WL and Tackett AJ: Quantitative
proteomics identifies activation of hallmark pathways of cancer in
patient melanoma. J Proteomics Bioinform. 6:43–50. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Manzini MC, Perez KR, Riske KA, Bozelli JC
Jr, Santos TL, da Silva MA, Saraiva GK, Politi MJ, Valente AP,
Almeida FC, et al: Peptide: lipid ratio and membrane surface charge
determine the mechanism of action of the antimicrobial peptide
BP100. Conformational and functional studies. Biochim Biophys Acta.
1838:1985–1999. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gustafsson OJ, Arentz G and Hoffmann P:
Proteomic developments in the analysis of formalin-fixed tissue.
Biochim Biophys Acta. 1854:559–580. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Quesada-Calvo F, Bertrand V, Longuespée R,
Delga A, Mazzucchelli G, Smargiasso N, Baiwir D, Delvenne P,
Malaise M, De Pauw-Gillet MC, et al: Comparison of two FFPE
preparation methods using label-free shotgun proteomics:
Application to tissues of diverticulitis patients. J Proteomics.
112:250–261. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Tanca A, Abbondio M, Pisanu S, Pagnozzi D,
Uzzau S and Addis MF: Critical comparison of sample preparation
strategies for shotgun proteomic analysis of formalin-fixed,
paraffin-embedded samples: Insights from liver tissue. Clin
Proteomics. 11:282014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Linge A, Kennedy S, O'Flynn D, Beatty S,
Moriarty P, Henry M, Clynes M, Larkin A and Meleady P: Differential
expression of fourteen proteins between uveal melanoma from
patients who subsequently developed distant metastases versus those
who did not. Invest Opthalmol Vis Sci. 53:4634–4643. 2012.
View Article : Google Scholar
|
|
16
|
Ostasiewicz P, Zielinska DF, Mann M and
Wiśniewski JR: Proteome, phosphoproteome, and N-glycoproteome are
quantitatively preserved in formalin-fixed paraffin-embedded tissue
and analyzable by high-resolution mass spectrometry. J Proteome
Res. 9:3688–3700. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Dowling P, Hughes DJ, Larkin AM, Larkin
AM, Meiller J, Henry M, Meleady P, Lynch V, Pardini B, Naccarati A,
et al: Elevated levels of 14-3-3 proteins, serotonin, gamma enolase
and pyruvate kinase identified in clinical samples from patients
diagnosed with colorectal cancer. Clin Chim Acta. 441:133–141.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
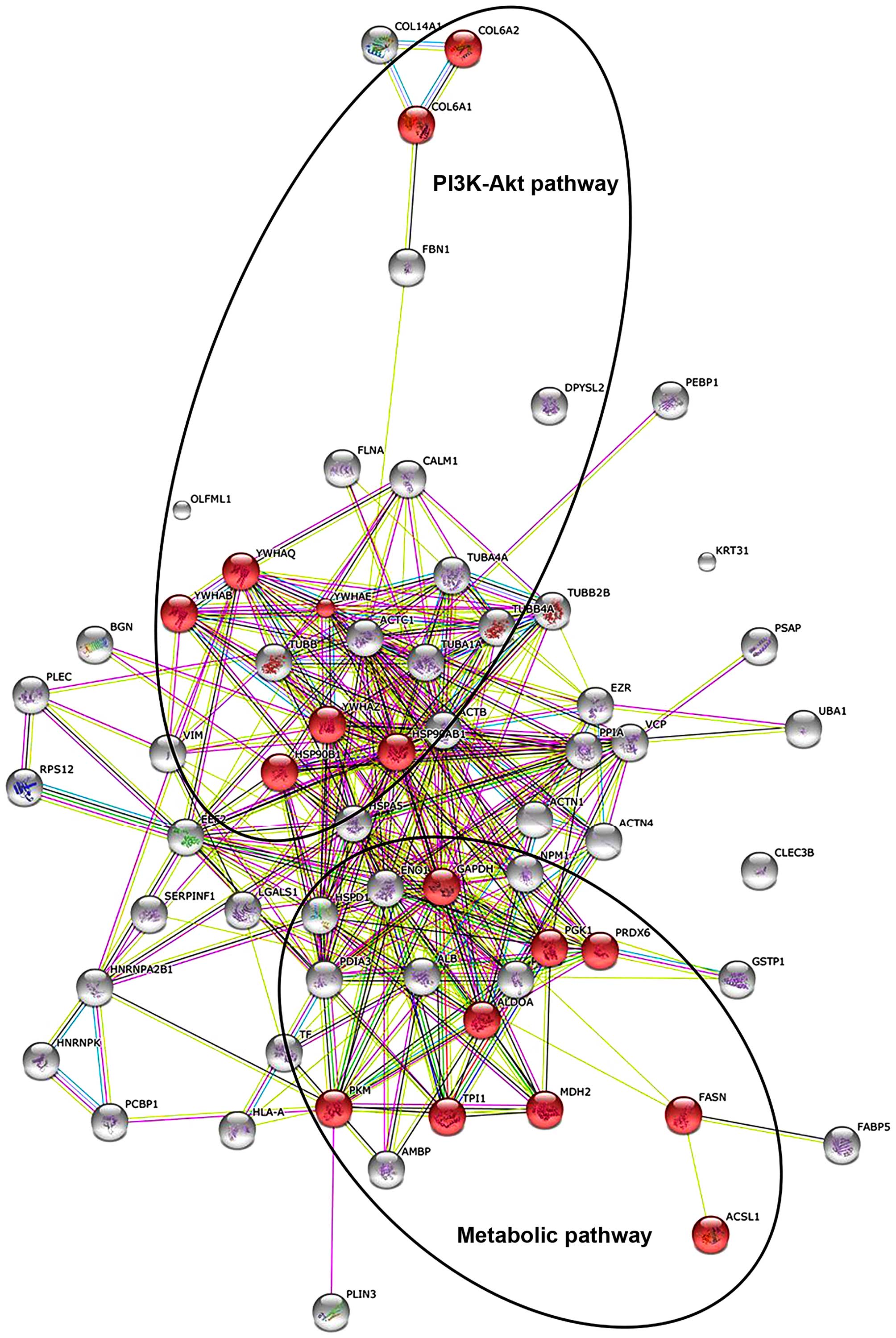
Franceschini A, Szklarczyk D, Frankild S,
Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C
and Jensen LJ: STRING v9.1: Protein-protein interaction networks,
with increased coverage and integration. Nucleic Acids Res.
41:(Database Issue). D808–D815. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Shen K, Sun J, Cao X, Zhou D and Li J:
Comparison of different buffers for protein extraction from
formalin-fixed and paraffin-embedded tissue specimens. PLoS One.
10:e01426502015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Warburg O, Wind F and Negelein E: THE
metabolism of tumors in the body. J Gen Physiol. 8:519–530. 1927.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Böhme I and Bosserhoff AK: Acidic tumor
microenvironment in human melanoma. Pigment Cell Melanoma Res. May
27–2016.(Epub ahead of print). View Article : Google Scholar
|
|
22
|
Han T, Kang D, Ji D, Wang X, Zhan W, Fu M,
Xin HB and Wang JB: How does cancer cell metabolism affect tumor
migration and invasion? Cell Adh Migr. 7:395–403. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Fu QF, Liu Y, Fan Y, Hua SN, Qu HY, Dong
SW, Li RL, Zhao MY, Zhen Y, Yu XL, et al: Alpha-enolase promotes
cell glycolysis, growth, migration, and invasion in non-small cell
lung cancer through FAK-mediated PI3K/AKT pathway. J Hematol Oncol.
8:222015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wong N, Ojo D, Yan J and Tang D: PKM2
contributes to cancer metabolism. Cancer Lett. 356:184–191. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Eigenbrodt E, Basenau D, Holthusen S,
Mazurek S and Fischer G: Quantification of tumor type M2 pyruvate
kinase (Tu M2-PK) in human carcinomas. Anticancer Res.
17:3153–3156. 1997.PubMed/NCBI
|
|
26
|
Lüftner D, Mesterharm J, Akrivakis C,
Geppert R, Petrides PE, Wernecke KD and Possinger K: Tumor type M2
pyruvate kinase expression in advanced breast cancer. Anticancer
Res. 20:5077–5082. 2000.PubMed/NCBI
|
|
27
|
Karachaliou N, Papadaki C, Lagoudaki E,
Trypaki M, Sfakianaki M, Koutsopoulos A, Mavroudis D, Stathopoulos
E, Georgoulias V and Souglakos J: Predictive value of BRCA1, ERCC1,
ATP7B, PKM2, TOPOI, TOPΟ-IIA, TOPOIIB and C-MYC genes in patients
with small cell lung cancer (SCLC) who received first line therapy
with cisplatin and etoposide. PLoS One. 8:e746112013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Desai S, Ding M, Wang B, Lu Z, Zhao Q,
Shaw K, Yung WK, Weinstein JN, Tan M and Yao J: Tissue-specific
isoform switch and DNA hypomethylation of the pyruvate kinase PKM
gene in human cancers. Oncotarget. 5:8202–8210. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Falkenius J, Lundeberg J, Johansson H,
Tuominen R, Frostvik-Stolt M, Hansson J and Brage S Egyhazi: High
expression of glycolytic and pigment proteins is associated with
worse clinical outcome in stage III melanoma. Melanoma Res.
23:452–460. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Rossato FA, Zecchin KG, La Guardia PG,
Ortega RM, Alberici LC, Costa RA, Catharino RR, Graner E, Castilho
RF and Vercesi AE: Fatty acid synthase inhibitors induce apoptosis
in non-tumorigenic melan-a cells associated with inhibition of
mitochondrial respiration. PLoS One. 9:e1010602014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tzivion G, Dobson M and Ramakrishnan G:
FoxO transcription factors; Regulation by AKT and 14-3-3 proteins.
Biochim Biophys Acta. 1813:1938–1945. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hopperton KE, Duncan RE, Bazinet RP and
Archer MC: Fatty acid synthase plays a role in cancer metabolism
beyond providing fatty acids for phospholipid synthesis or
sustaining elevations in glycolytic activity. Exp Cell Res.
320:302–310. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Menendez JA and Lupu R: Fatty acid
synthase and the lipogenic phenotype in cancer pathogenesis. Nat
Rev Cancer. 7:763–777. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Swinnen JV, Roskams T, Joniau S, Van
Poppel H, Oyen R, Baert L, Heyns W and Verhoeven G: Overexpression
of fatty acid synthase is an early and common event in the
development of prostate cancer. Int J Cancer. 98:19–22. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Rossi S, Graner E, Febbo P, Weinstein L,
Bhattacharya N, Onody T, Bubley G, Balk S and Loda M: Fatty acid
synthase expression defines distinct molecular signatures in
prostate cancer. Mol Cancer Res. 1:707–715. 2003.PubMed/NCBI
|
|
36
|
Wang Y, Kuhajda FP, Li JN, Pizer ES, Han
WF, Sokoll LJ and Chan DW: Fatty acid synthase (FAS) expression in
human breast cancer cell culture supernatants and in breast cancer
patients. Cancer Lett. 167:99–104. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Bauerschlag DO, Maass N, Leonhardt P,
Verburg FA, Pecks U, Zeppernick F, Morgenroth A, Mottaghy FM, Tolba
R, Meinhold-Heerlein I and Bräutigam K: Fatty acid synthase
overexpression: Target for therapy and reversal of chemoresistance
in ovarian cancer. J Transl Med. 13:1462015. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Cerne D, Zitnik IP and Sok M: Increased
fatty acid synthase activity in non-small cell lung cancer tissue
is a weaker predictor of shorter patient survival than increased
lipoprotein lipase activity. Arch Med Res. 41:405–409. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Innocenzi D, Alò PL, Balzani A, Sebastiani
V, Silipo V, La Torre G, Ricciardi G, Bosman C and Calvieri S:
Fatty acid synthase expression in melanoma. J Cutan Pathol.
30:23–28. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Kapur P, Rakheja D, Roy LC and Hoang MP:
Fatty acid synthase expression in cutaneous melanocytic neoplasms.
Mod Pathol. 18:1107–1112. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Seguin F, Carvalho MA, Bastos DC, Agostini
M, Zecchin KG, Alvarez-Flores MP, Chudzinski-Tavassi AM, Coletta RD
and Graner E: The fatty acid synthase inhibitor orlistat reduces
experimental metastases and angiogenesis in B16-F10 melanomas. Br J
Cancer. 107:977–987. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Flavin R, Peluso S, Nguyen PL and Loda M:
Fatty acid synthase as a potential therapeutic target in cancer.
Future Oncol. 6:551–562. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Yang YA, Han WF, Morin PJ, Chrest FJ and
Pizer ES: Activation of fatty acid synthesis during neoplastic
transformation: Role of mitogen-activated protein kinase and
phosphatidylinositol 3-kinase. Exp Cell Res. 279:80–90. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Van de Sande T, De Schrijver E, Heyns W,
Verhoeven G and Swinnen JV: Role of the phosphatidylinositol
3′-kinase/PTEN/Akt kinase pathway in the overexpression of fatty
acid synthase in LNCaP prostate cancer cells. Cancer Res.
62:642–646. 2002.PubMed/NCBI
|
|
45
|
Wang D and Sul HS: Insulin stimulation of
the fatty acid synthase promoter is mediated by the
phosphatidylinositol 3-kinase pathway. Involvement of protein
kinase B/Akt. J Biol Chem. 273:25420–25426. 1998.
|
|
46
|
Wang HQ, Altomare DA, Skele KL, Poulikakos
PI, Kuhajda FP, Di Cristofano A and Testa JR: Positive feedback
regulation between AKT activation and fatty acid synthase
expression in ovarian carcinoma cells. Oncogene. 24:3574–3582.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Chen B, Tardell C, Higgins B, Packman K,
Boylan JF and Niu H: BRAFV600E negatively regulates the AKT pathway
in melanoma cell lines. PLoS One. 7:e425982012. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Stahl JM, Sharma A, Cheung M, Zimmerman M,
Cheng JQ, Bosenberg MW, Kester M, Sandirasegarane L and Robertson
GP: Deregulated Akt3 activity promotes development of malignant
melanoma. Cancer Res. 64:7002–7010. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Meier F, Schittek B, Busch S, Garbe C,
Smalley K, Satyamoorthy K, Li G and Herlyn M: The RAS/RAF/MEK/ERK
and PI3K/AKT signaling pathways present molecular targets for the
effective treatment of advanced melanoma. Front Biosci.
10:2986–3001. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
50
|
Daitoku H, Sakamaki J and Fukamizu A:
Regulation of FoxO transcription factors by acetylation and
protein-protein interactions. Biochim Biophys Acta. 1813:1954–1960.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Freeman AK and Morrison DK: 14-3-3
proteins: Diverse functions in cell proliferation and cancer
progression. Semin Cell Dev Biol. 22:681–687. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Tzivion G, Gupta VS, Kaplun L and Balan V:
14-3-3 proteins as potential oncogenes. Semin Cancer Biol.
16:203–213. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Neal CL, Xu J, Li P, Mori S, Yang J, Neal
NN, Zhou X, Wyszomierski SL and Yu D: Overexpression of 14-3-3ζ in
cancer cells activates PI3K via binding the p85 regulatory subunit.
Oncogene. 31:897–906. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Schultz J, Ibrahim SM, Vera J and Kunz M:
14-3-3sigma gene silencing during melanoma progression and its role
in cell cycle control and cellular senescence. Mol Cancer.
8:532009. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Vera J, Schultz J, Ibrahim S, Raatz Y,
Wolkenhauer O and Kunz M: Dynamical effects of epigenetic silencing
of 14-3-3sigma expression. Mol Biosyst. 6:264–273. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Ko BS, Jan YJ, Chang TC, Liang SM, Chen
SC, Liu TA, Wu YM, Wang J and Liou JY: Upregulation of focal
adhesion kinase by 14-3-3ε via NFκB activation in hepatocellular
carcinoma. Anticancer Agents Med Chem. 13:555–562. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Xia H, Nho RS, Kahm J, Kleidon J and Henke
CA: Focal adhesion kinase is upstream of phosphatidylinositol
3-kinase/Akt in regulating fibroblast survival in response to
contraction of type I collagen matrices via a beta 1 integrin
viability signaling pathway. J Biol Chem. 279:33024–33034. 2004.
View Article : Google Scholar : PubMed/NCBI
|