Introduction
Gastrointestinal (GI) cancer, including colorectal
cancer (CRC) and gastric cancer (GC), are common malignant tumors
with poor prognosis and high mortality rates among both males and
females in China (1). Following lung
and liver cancer, CRC and GC result in the greatest numbers of
mortalities (2). Despite the use of
colonoscopy and gastroscopy in cancer screening, patients with GI
cancer tend to be diagnosed at an advanced stage (3). The mortality of gastrointestinal cancer
has declined in several Western countries since the introduction of
cancer screening programs. Additionally, the removal of adenomas,
early detection of cancerous lesions and the availability of more
effective therapies for early stage disease have contributed to a
decrease in the mortality rate (4).
Cancer patients diagnosed at an early stage generally receive
prompt treatment and have an improved prognosis compared with those
diagnosed at later stages (5,6). Several
National Comprehensive Cancer Network member institutions advise
the use of molecular screening methods, including
immunohistochemical analysis, for mismatch repair (MMR) protein
expression or microsatellite instability (MSI) analysis on all
newly diagnosed patients with CRCs (7–10). These
recommendations reflect the importance of MMR genes and the MSI
index in cancer management.
DNA repair can repair damaged DNA and maintain
genomic stability. Multiple DNA repair genes, including the MMR and
homologous recombination (HR) genes, are involved in the
development of GI cancer (11,12).
Defective MMR (dMMR) has been revealed to lead to MSI, resulting in
high numbers of somatic mutations in intestinal carcinogenesis
(13,14).
Clinically, MSI evaluation is recommended in
patients with stage II CRC in order to inform treatment
decision-making regarding chemotherapy administration (15). Germline mutations in MMR genes have
been implicated in Lynch syndrome, which is a highly penetrant,
autosomal-dominant inherited cancer predisposition syndrome
characterized by the early onset of cancers in the colon as well as
at extra-colonic sites such as the endometrium, ovaries, stomach,
small intestine, pancreas, urinary tract and brain (16–18).
Furthermore, somatic mutations in MMR genes have been reported to
result in dMMR (19–22). A number of studies have reported
patients with CRC patients harbored germline mutations in HR genes,
including ATM serine/threonine kinase (ATM), BRCA1 DNA
repair associated (BRCA1), BRCA2 DNA repair associated
(BRCA2) and partner and localizer of BRCA2 (PALB2)
(23–26). A previous study (27) reported somatic pathogenic mutations
among all tumor lineages and revealed that the HR deficiency
frequency was ~13% in all solid tumor types (n=53,619) and ~6.3% in
CRC. Another report indicated that 41–50% of ovarian carcinomas are
estimated to exhibit HR deficiency. However, the frequency of HR
deficiency varies according to the method utilized for its
evaluation (germline mutations, somatic mutations or HRD score) and
histological subtype (28).
Previous studies demonstrated the importance of the
MMR and HR genes on GI cancer (29–31).
However, the characteristics of mutations in these genes remain
unclear, and so is the range of effects they may exert on genomic
instability. In the present study, targeted capture and massively
parallel next generation sequencing (NGS) technologies were used to
study the mutations of 183 cancer-promoting genes in 137 patients
with GI cancer, with a particular focus on the mutational status of
MMR and HR genes.
Materials and methods
Patients and collection of clinical
samples
A total of 137 patients with stage III/IV CRC and GC
were included in the current study, which was approved by Ethics
Committee of Second Hospital of Anhui Medical University (Hefei,
China), between May 2017 and May 2018. Written informed consent was
obtained from each patient prior to sample collection. The samples
were obtained from 92 males and 45 females (median age, 60 years;
age range, 27–84 years). Tissues were fixed with 10% formalin at
room temperature for 8 h. A total of 92 formalin-fixed,
paraffin-embedded (FFPE) tumor sections collected following surgery
were retrieved. In addition, 45 blood samples for circulating
cell-free DNA (cfDNA) extraction were collected prior to surgery
from patients receiving no prior chemotherapy or radiotherapy. A
total of 13 genomic DNA fragments were extracted from
patient-matched peripheral blood (when available) and were used as
matched normal controls. All FFPE tumor samples were confirmed to
have >20% tumor cells upon H&E staining (data not shown).
The clinical and pathological characteristics of the patients were
obtained from hospital records. Basic patient information is
summarized in Table I.
 | Table I.Distribution of SNV and indel
mutations in patients with gastrointestinal cancer. |
Table I.
Distribution of SNV and indel
mutations in patients with gastrointestinal cancer.
| Characteristic | Number of
specimens | Number of SNV and
indel mutationsa (mean
± SEM) | P-value |
|---|
| Sex |
|
| 0.8010 |
|
Female | 45 | 210.7±24.04 |
|
|
Male | 92 | 210.0±15.62 |
|
| Age |
|
| 0.2044 |
|
≥55 | 86 | 224.5±17.88 |
|
|
<55 | 51 | 186.2±17.77 |
|
| Tumor type |
|
| 0.9105 |
|
Colorectal | 75 | 213.8±17.54 |
|
|
Gastric | 62 | 205.8±19.77 |
|
| MMR gene mutation
status |
|
|
4.476×10−08 |
|
Inactivation (+)b | 56 | 280.2±20.11 |
|
|
Inactivation (−)c | 81 | 161.8±15.10 |
|
| HR gene mutation
status |
|
|
2.581×10−05 |
|
Inactivation (+)d | 112 | 232.6±15.06 |
|
|
Inactivation (−)e | 25 | 110.0±10.23 |
|
DNA extraction
FFPE tumor sections (40 µm) were treated with 100%
xylene (Sigma-Aldrich; Merck KGaA, Darmstadt, Germany) at room
temperature, followed by 100% ethanol (32). Deparaffinized samples were then
suspended in proteinase K-containing buffer (Thermo Fisher
Scientific, Inc., Waltham, MA, USA). Following extraction using
phenol-chloroform (V:V=1:1; Sigma-Aldrich; Merck KGaA), DNA samples
were treated with ethanol for precipitation and resuspended in
deionized water. For cfDNA extraction from blood samples, Cell-Free
DNA BCT tubes (Streck, Omaha, NE, USA) were used for the collection
of 10 ml blood samples. Validation of the adopted protocols for
this study has been performed previously (33). In brief, 1.2 ml of plasma was
collected from each patient using two stages of centrifugation at
4°C at 1,600 × g for 10 min, prior to cfDNA extraction. cfDNA
extraction was performed with the QIAamp Circulating Nucleic Acid
Kit (Qiagen, Inc., Valencia, CA, USA), according to the methodology
described by the manufacturer. The processing of white blood cell
(WBC) DNA as a control was performed as follows: Using 2 ml of
total peripheral blood, DNA was extracted using the Flexigene DNA
kit (Qiagen, Inc.). Quantification of isolated DNA samples was
performed using a NanoDrop spectrophotometer (Thermo Fisher
Scientific, Inc.), and by fluorimetry, using Qubit dsDNA
high-sensitivity and/or broad-range assay kits (Thermo Fisher
Scientific, Inc.).
Library construction and
sequencing
The FD-180 panel targeted for exon regions of 183
tumor driver genes (not shown) was used for the generation of
sequencing libraries with the Illumina platforms (Kapa Biosystems;
Roche Diagnostics, Basel, Switzerland) and the SeqCap EZ Choice
Library (Roche NimbleGen, Inc., Madison, WI, USA), with DNA
fragments and cfDNA employed in the library construction following
the manufacturer's protocol. DNA sequencing was completed with the
Illumina NextSeq 500 system at a depth of 10,000X (cfDNA) and
3,000X (FFPE). All the operations are carried out in accordance
with the product manual.
Variant calling and analysis
Raw data were processed into clean FASTQ output with
Flexbar (https://sourceforge.net/projects/flexbar/) through
trimming of adapter sequences and the removal of low-quality reads
(average quality score <15). Raw reads were checked for data
quality using FASTQ (www.bioinformatics.babraham.ac.uk). Plots of quality
(Q) scores across all bases in reads indicated the majority of
positions had Q≥20. Q scores are logarithmically related to the
base calling error probabilitie s(P), Q=−10 log10P. A lower base
call accuracy of 99% (Q20) will have an incorrect base call
probability of 1 in 100, meaning that every 100 bp sequencing read
will likely contain an error. Raw reads were then trimmed for
adapter contamination with Trimmomatic (version 0.32; http://usadellab.org/cms/). Leading and trailing
low-quality bases (Q<3) were removed. Reads were also scanned
with a 4-base-wide sliding window and the following bases were cut
when the average Q per base dropped to <15. Finally, only reads
>50 bases were kept for subsequent analysis. Either
patient-matched peripheral blood (when available) or in-batch
pooled FFPE normal controls were used for mutation calling. Paired
clean reads, following Trimmomatic treatment, were aligned against
the reference genome hg19 (hgdownload.soe.ucsc.edu/goldenPath/hg19/bigZips) using
Burrows-Wheelers Aligner (version 0.7.13) (34). Calibration was then performed on the
remaining reads, which were realigned via the Genome Analysis
Toolkit (version 4.0.3.0) (35).
Genome Analysis Toolkit was used to perform base quality score
recalibration (BQSR). BQSR is a process by which machine learning
is applied to a model to score errors empirically and adjust the
quality scores accordingly.
Analysis of the realigned BAM files and detection of
somatic single-nucleotide variants (SNVs) and insertion/deletion
(indel) mutations was performed using MuTect (version 1.7.0)
(36). Normal germline variants were
filtered out using the Single Nucleotide Polymorphism Database
(37) or the Exome Aggregation
Consortium database (http://exac.broadinstitute.org/). Default parameter
settings were used for all programs. The elimination of erroneous
base calls and generation of final mutations was performed by
variation frequency (>0.5%).
Variation analysis
The SNV and indel mutations (including stopgain,
frameshift, splicing, synonymous, non-synonymous and non-frameshift
types) of the 183-gene panel were analyzed in the 137 patients with
GI cancer. MMR and HR genes were then investigated for SNV and
indel mutations, respectively.
MMR and HR genes inactivation
analysis
A total of four MMR (MLH1, MSH2, MSH6 and
PMS2), and 15 HR [(BRCA1, BRCA2, ATM, phosphatase and
tensin homolog (PTEN), BLM RecQ like helicase (BLM),
BRCA1 interacting protein C-terminal helicase 1 (BRIP1), FA
complementation group A (FANCA), FA complementation group C
(FANCC), FA complementation group D2 (FANCD2), FA
complementation group E (FANCE), FA complementation group F
(FANCF), FA complementation group G (FANCG), nibrin
(NBN), PALB2 and Werner syndrome RecQ like helicase
(WRN)] genes were selected and analyzed according to
previous studies (18–24). In the present study, inactivation
mutations included stopgain, frameshift and splicing somatic
mutations. Any MMR or HR gene with an inactivation mutation was
defined as MMR inactivation-positive (MMR+) or HR
inactivation-positive (HR+).
Statistical analysis
The results are presented as mean ± standard
deviation. Statistical significance was tested using the
Kruskal-Wallis test. P<0.05 indicated a statistically
significant difference. A number of SNV and indel mutation sites in
all MMR or HR genes to represent the MMR or HR mutational status
were used. The number of SNV and indel mutation sites of the
183-gene panel in each patient was used to predict the degree of
genomic instability. To investigate the effects of the mutational
status of MMR and HR genes, the Pearson's correlation test between
MMR and HR mutational status and genomic instability was used. All
statistical analyses were performed using GraphPad Prism (version
5.0; GraphPad, San Diego, CA, USA) and SPSS software (version 17.0;
SPSS Inc., Chicago, IL, USA).
Results
Molecular mutation characteristics in
137 patients with GI cancer
To investigate the molecular characteristics of GI
cancer, we analyzed the SNV and indel mutations of 183 genes in 137
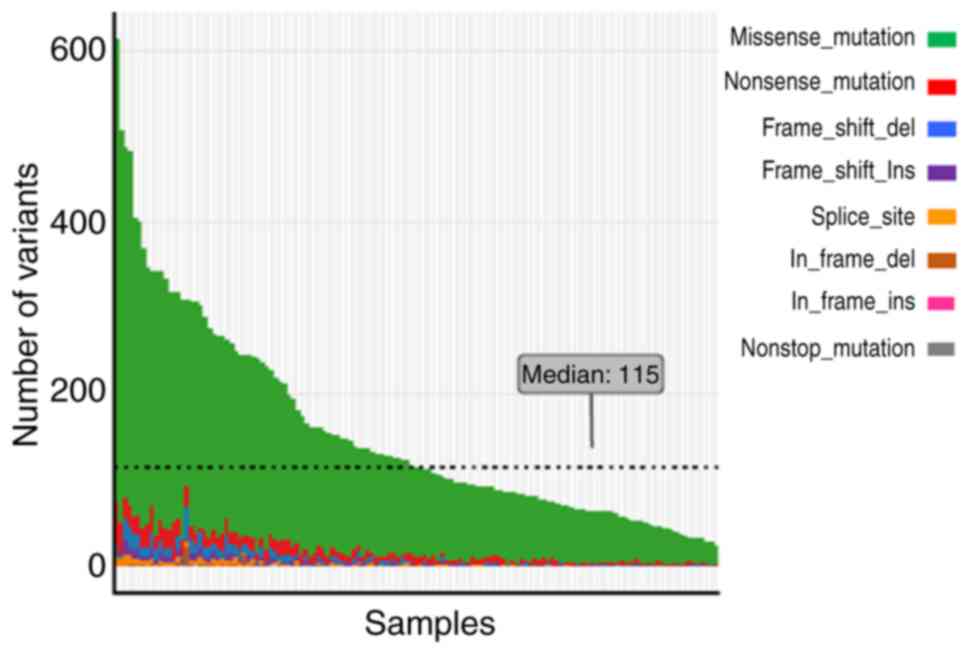
patients using NGS technology. As shown in Fig. 1, the number of mutations among the
137 patients was 24–613 (median, 115). Several MMR gene
inactivation mutations were identified. Certain HR genes had a high
frequency of inactivation mutations including, BRCA2 (32.9%;
n=45), ATM (29.2%; n=40), BRCA1 (17.5%; n=24) and
FANCD2 (15.3%; n=21). These results indicated the different
effects of MMR and HR genes on genomic instability.
The 20 most frequently mutated genes are shown in
Fig. 2A. The mechanistic target of
rapamycin kinase was the most commonly mutated gene, detected in
85% of the patients. Other genes, including PMS2 and
regulator of WNT signaling pathway (APC), also exhibited a
high mutation frequency. Certain HR genes were among the 20 most
commonly mutated genes, including ATM, FANCD2, ATR
serine/threonine kinase (ATR), BRCA2, BRCA1 and
FANCA (Fig. 2A). The
mutational status of MMR and HR genes is illustrated in Fig. 2B. As presented, PMS2 mutation
was detected in 83% of the patients. ATM, FANCD2 and
BRCA2 were also found to have a mutation frequency of ~80%
(Fig. 2B), whereas FANCF had
a relatively lower (29%) mutation frequency.
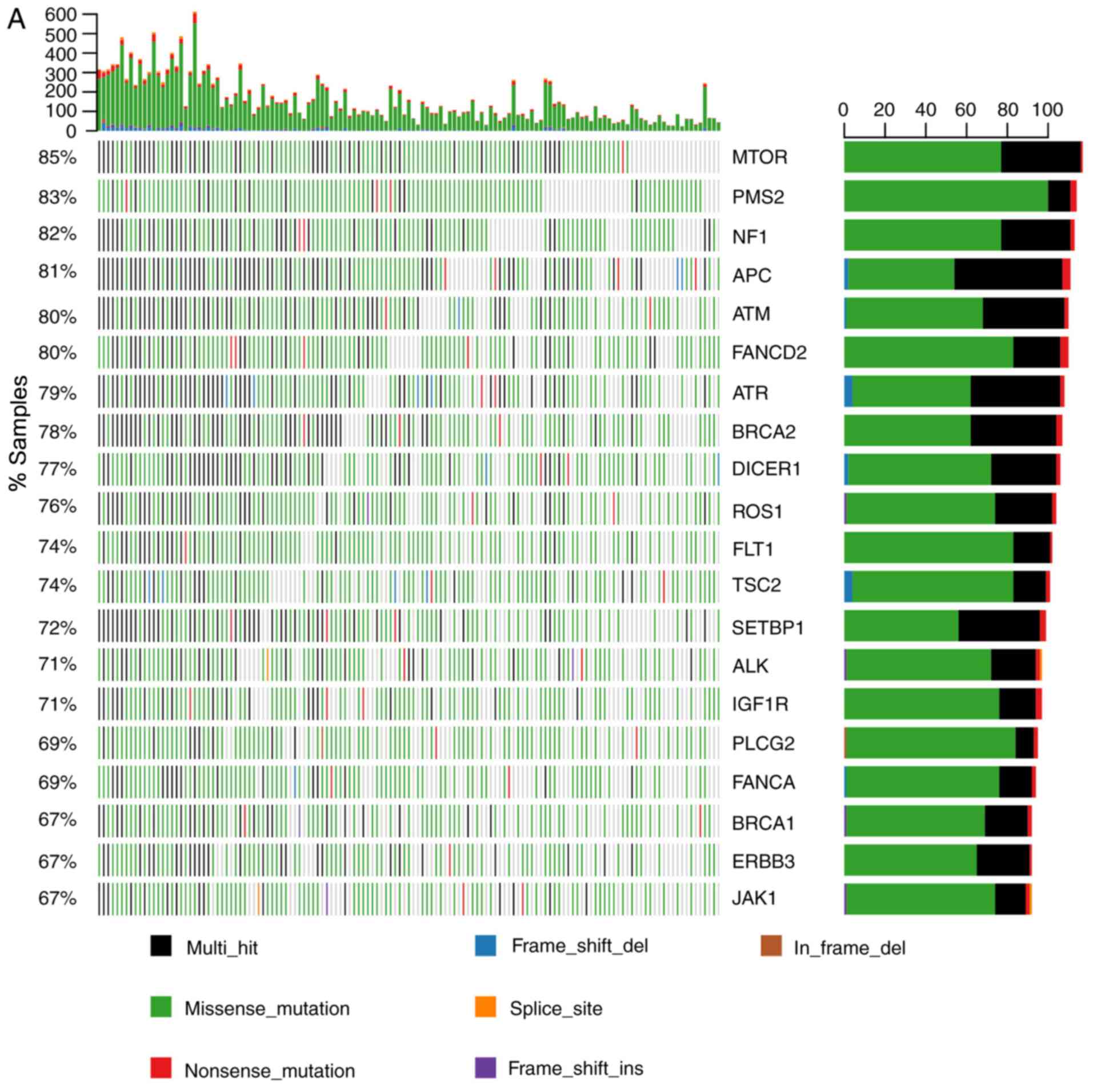 | Figure 2.Molecular characteristics of somatic
mutations in the 137 patients with gastrointestinal cancer. (A) All
SNV and indel mutations
(stopgain/frameshift/splicing/nonsynonymous/synonymous/non-frameshift)
of the 20 most commonly mutated genes in patients and gene mutation
frequency were analyzed. The x-axis represents each of the 137
samples and the y-axis represents the proportion of the gene
mutation samples. The highest numbers of SNV and indel mutations in
each patient; left, gene mutation frequency in the 137 patients;
right, number of patients with gene mutations. (B) The mutational
status of mismatch repair and homologous recombination genes. The
x-axis represents each of the 137 samples; the y-axis represents
the proportion of the gene mutation samples. SNV, single-nucleotide
variant; indel, insertion-deletion; del, deletion; ins, insertion;
MTOR, mechanistic target of rapamycin kinase; PMS2, PMS1 homolog 2
mismatch repair system component; NF1, neurofibromin 1; APC, APC
regulator of WNT signaling pathway; ATM, ATM serine/threonine
kinase; FANCD2, FA complementation group D2; ATR, ATR
serine/threonine kinase; BRCA2, BRCA2 DNA repair associated;
DICER1, dicer 1 ribonuclease III; ROS1, ROS proto-oncogene 1
receptor tyrosine kinase; FLT1, fms related tyrosine kinase 1;
TSC2, TSC complex subunit 2; SETBP1, SET binding protein 1; ALK,
ALK receptor tyrosine kinase; IGF1R, insulin like growth factor 1
receptor; PLCG2, phospholipase C γ 2; FANCA, FA complementation
group A; BRCA1, BRCA1 DNA repair associated; ERBB3, erb-b2 receptor
tyrosine kinase 3; JAK1, Janus kinase 1; BRIP1, BRCA1 interacting
protein C-terminal helicase 1; MSH6, mutS homolog 6; PALB2, partner
and localizer of BRCA2; BLM, BLM RecQ like helicase; WRN, Werner
syndrome RecQ like helicase; MSH2, mutS homolog 2; MLH1, mutL
homolog 1; FANCE, FA complementation group E; FANCG, FA
complementation group G; NBN, nibrin; FANCF, FA complementation
group F. |
Mutational status of MMR and HR genes
positively indicates genomic instability in GI cancer
A Pearson's correlation test between MMR and HR
mutational status and genomic instability was performed. The
results of correlation analysis indicated a significant association
between MMR gene mutational status and genomic instability
(R2=0.703; P=2.2×10−16). Patients harboring
high numbers of SNV and indel mutations in MMR genes had a higher
frequency of mutations in all 183-gene panels (Fig. 3A). A significant correlation was also
observed in the HR group (R2=0.901;
P=2.2×10−16; Fig. 3A).
These results demonstrated that the mutational status of MMR and HR
genes is positively correlated with genomic instability.
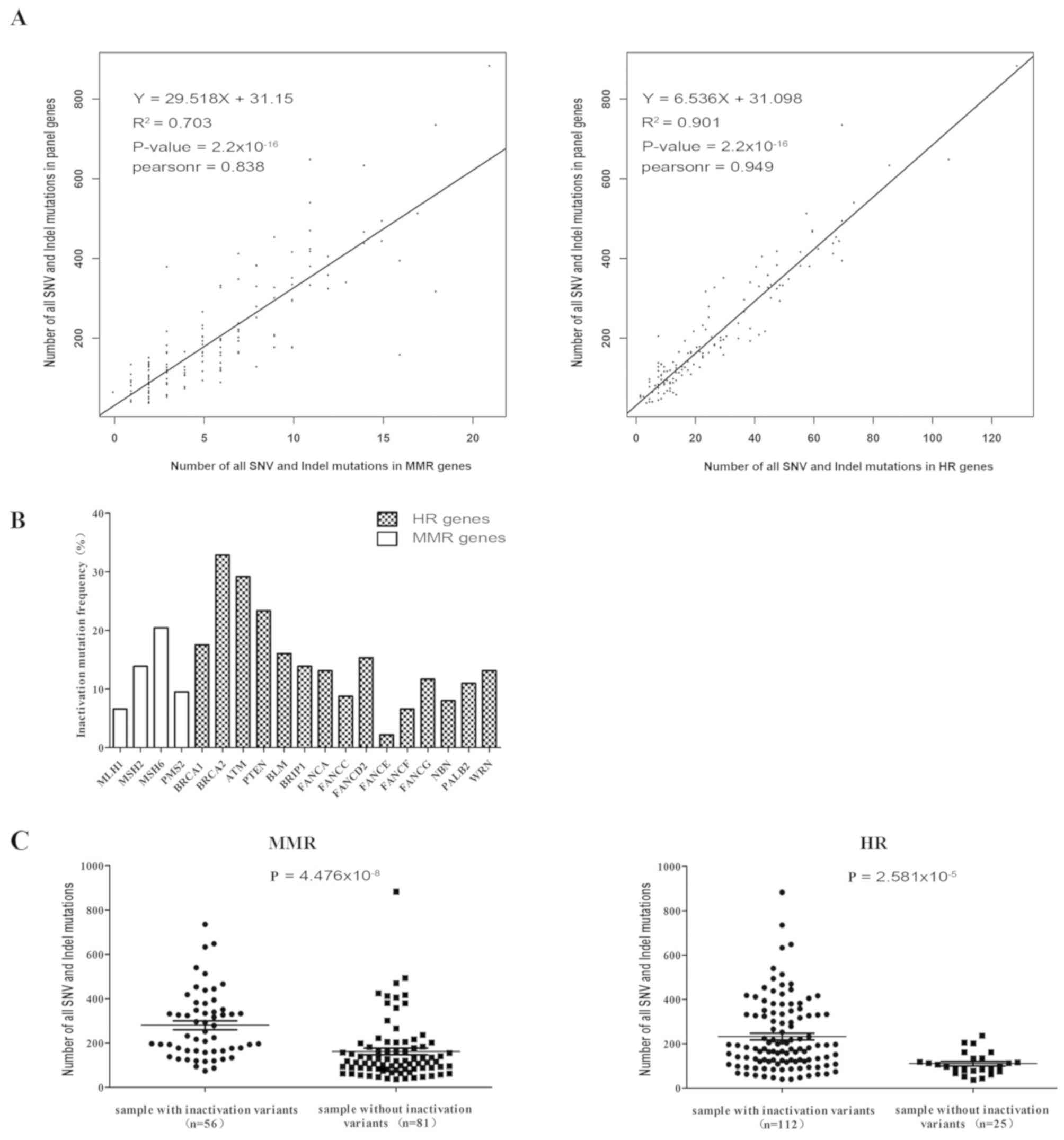 | Figure 3.Effects of MMR and HR gene deficiency
on genomic instability in gastrointestinal cancer. (A) Number of
SNV and indel mutation sites in MMR or HR genes positively
correlated with mutation sites in the 183-gene panel. (B)
Inactivation mutation frequency in MMR and HR genes in the 137
patients with gastrointestinal cancer. (C) Differences in the
number of SNV and indel mutations in a panel of 183 genes between
MMR or HR gene inactivation mutation positive and negative groups.
The numbers in brackets on the x-axis is number of samples with
inactivation mutations. MMR, mismatch repair; HR, homologous
recombination; SNV, single-nucleotide variant; indel,
insertion-deletion; MLH1, mutL homolog 1; MSH2, mutS homolog 2;
MSH6, mutS homolog 6; PMS2, PMS1 homolog 2 mismatch repair system
component; BRCA1, BRCA1 DNA repair associated; BRCA2, BRCA2 DNA
repair associated; ATM, ATM serine/threonine kinase; PTEN,
phosphatase and tensin homolog; BLM, BLM RecQ like helicase; BRIP1,
BRCA1 interacting protein C-terminal helicase 1; FANCA, FA
complementation group A; FANCC, FA complementation group C; FANCD2,
FA complementation group D2; FANCE, FA complementation group E;
FANCF, FA complementation group F; FANCG, FA complementation group
G; NBN, nibrin; PALB2, partner and localizer of BRCA2; WRN, Werner
syndrome RecQ like helicase. |
Prevalence and influence of MMR and HR
deficiency in GI cancer
The current study aimed to summarize the
inactivation mutation ratio of MMR and HR genes in patients with GI
cancer. Different inactivation mutation frequencies of MMR and HR
genes were detected, as shown in Fig.
3B. BRCA2 exhibited a high deficiency frequency (45/137,
32.85%) in GI cancer (Table II).
Other genes, including ATM (29.2%), PTEN (23.36%),
BRCA1 (17.52%), MSH6 (20.44%), FANCD2 (15.33%)
and MSH2 (13.87%) also exhibited high proportions of
inactivation mutations (Fig. 3B,
Table II). The current study
revealed that 40.9% (56/137) of the patients with GI cancer
harbored at least one inactivation mutation site in one or more MMR
genes (Table I). A total of 81.8%
(112/137) of the patients also exhibited HR gene deficiencies
(Table I). Moreover, as shown in
Fig. 3C, patients positive for MMR
gene inactivation mutations had significantly increased genomic
instability compared with negative cases (SNV and indel count,
280.2±20.11 vs. 161.8±15.10, respectively; P<0.0001), as well as
HR gene inactivation mutation-positive patients (SNV and indel
count, 232.6±15.06 vs. 110.0±10.23, respectively; P<0.0001)
(Table I). By contrast, no
significant association between genomic instability and other
clinical characteristics, including sex, age and tumor type, were
identified in GI cancer (Table I).
The results obtained suggested the contribution of MMR and HR gene
deficiency to genomic instability in GI cancer.
 | Table II.Molecular mutation characteristics of
MMR and HR genes in 137 patients with gastrointestinal cancer. |
Table II.
Molecular mutation characteristics of
MMR and HR genes in 137 patients with gastrointestinal cancer.
| Proportion of
samples with MMR and HR genes inactivation mutation | Number of SNV and
indel mutations in 183 panel genesa (mean ± SEM) |
|---|
|
|
|---|
| Gene | Stopgain | Frameshift | Splicing | Proportion of
samples with inactivation mutation % (n) | Inactivation
mutation (+)b
(n)c | Inactivation
mutation (−)d (n) | P-value |
|---|
| MLH1 | 7 | 4 | 0 | 6.57 (9) | 283.56±53.32
(9) | 205.0±13.43
(128) |
0.0576 |
| MSH2 | 8 | 11 | 0 | 13.87 (19) | 276.6±30.99
(19) | 199.5±14.15
(118) |
0.0053 |
| MSH6 | 12 | 17 | 0 | 20.44 (28) | 315.6±32.77
(28) | 183.1±12.99
(109) | <0.0001 |
| PMS2 | 9 | 5 | 0 | 9.49 (13) | 266.0±31.59
(13) | 204.3±14.00
(124) |
0.0263 |
| BRCA1 | 19 | 5 | 0 | 17.52 (24) | 365.6±42.67
(24) | 177.2±10.82
(113) | <0.0001 |
| BRCA2 | 30 | 22 | 0 | 32.85 (45) | 305.3±25.74
(45) | 163.7±12.32
(92) | <0.0001 |
| ATM | 31 | 16 | 0 | 29.2 (40) | 315.1±31.22
(40) | 166.9±10.60
(97) | <0.0001 |
| PTEN | 3 | 30 | 0 | 23.36 (32) | 208.3±24.39
(32) | 210.8±15.42
(105) |
0.9089 |
| BLM | 17 | 6 | 0 | 16.06 (22) | 281.5±27.90
(22) | 196.6±14.34
(115) |
0.0019 |
| BRIP1 | 13 | 7 | 0 | 13.87 (19) | 321.3±33.13
(19) | 192.3±13.57
(118) |
0.0002 |
| FANCA | 7 | 12 | 0 | 13.14 (18) | 378.4±43.08
(18) | 184.7±12.03
(119) | <0.0001 |
| FANCC | 5 | 8 | 0 | 8.76 (12) | 282.1±61.22
(12) | 203.3±13.02
(125) |
0.3094 |
| FANCD2 | 16 | 7 | 0 | 15.33 (21) | 318.0±46.56
(21) | 190.7±12.21
(116) |
0.0076 |
| FANCE | 0 | 3 | 0 | 2.19 (3) | 529.7±62.86
(3) | 203.0±12.65
(134) |
0.0071 |
| FANCF | 5 | 4 | 0 | 6.57 (9) | 268.4±38.93
(9) | 206.1±13.69
(128) |
0.0443 |
| FANCG | 2 | 15 | 0 | 11.68 (16) | 347.6±49.84
(16) | 192.0±12.45
(121) |
0.0023 |
| NBN | 6 | 6 | 0 | 8.03 (11) | 329.4±44.71
(11) | 199.8±13.33
(126) |
0.0045 |
| PALB2 | 9 | 7 | 0 | 10.95 (15) | 248.8±35.28
(15) | 205.5±14.02
(122) |
0.1789 |
| WRN | 11 | 9 | 0 | 13.14 (18) | 367.1±43.27
(18) | 186.5±12.25
(119) | <0.0001 |
Different MMR or HR genes may exert
different effects on genomic instability
Among the 137 patients with GI cancer, 56 were found
to be positive for MMR gene inactivation mutations and 117 were
found to be positive with HR gene inactivation mutations (36 of the
patients were double-positive). Although the results obtaine in the
current study demonstrated that patients positive for either MMR or
HR gene inactivation mutations had higher genomic instability
compared with negative patients, as shown in Fig. 4A, there was no significant difference
between patients with MMR gene inactivation mutations and those
with HR gene deficiencies in the current study (SNV and indel
count, 280.2±20.11 vs. 232.6±15.06, respectively; P=0.019).
Additionally, the positive and negative groups according to the
inactivation mutation status of 4 MMR and 15 HR genes were
compared, and the number of SNV and indel mutations in the 183-gene
panel of patients with any MMR or HR inactivation mutations was
revealed to be higher compared with those without such mutations,
including those positive for inactivation mutations in MSH6,
BRCA1, BRCA2, ATM or FANCD2 (Table II). Additionally, patients with
MLH1, PTEN, FANCC or PALB2 inactivation mutations did
not exhibit significant differences in the number of SNV and indel
mutations compared with patients without inactivation mutations in
these genes (Table II). No
significant effects on genomic instability among the four groups
patients with MLH1, MSH2, MSH6 and PMS2 inactivation
mutations (Fig. 4B). However, the
effect on genomic instability of certain genes, including BRCA1,
BRCA2, ATM, FANCA and BRIP1, was markedly more
pronounced compared with that of PTEN in patients with HR
deficiency (Fig. 4C). These results
reflected the different effect on genomic instability among MMR and
HR genes.
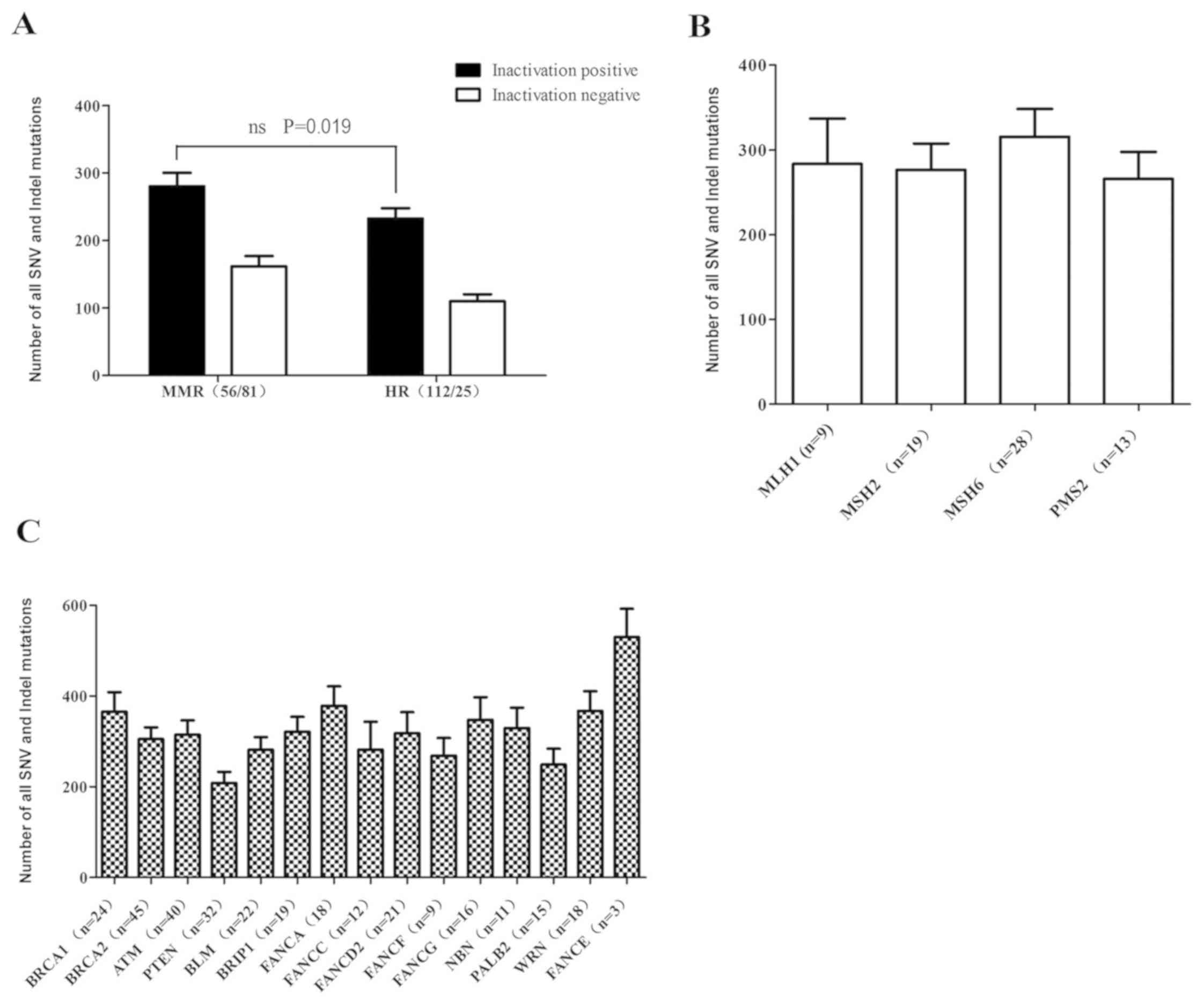 | Figure 4.Different inactivated MMR and HR
genes had different influence on genomic instability. (A) Number of
SNV and indel mutations in 183-gene panel in inactivation positive
groups for MMR and HR genes. Inactivation positive mutations
included individuals with at least one type of
stopgain/frameshift/splicing mutation. Inactivation negative
mutations included individuals with no stopgain/frameshift/splicing
mutations. The numbers in brackets on the x-axis represents the
number of samples with inactivation mutation. There was no
significant difference between patients with MMR gene inactivation
mutations and those with HR gene deficiencies in the current study.
(B) Parallel comparison of the effects of MMR genes on genomic
instability. MMR, mismatch repair; HR, homologous recombination;
SNV, single-nucleotide variant; indel, insertion-deletion; MLH1,
mutL homolog 1; MSH2, mutS homolog 2; MSH6, mutS homolog 6; PMS2,
PMS1 homolog 2 mismatch repair system component. The numbers in
brackets on the x-axis is number of samples with inactivation
mutation. (C) The effects of HR genes on genomic instability.
BRCA1, BRCA1 DNA repair associated; BRCA2, BRCA2 DNA repair
associated; ATM, ATM serine/threonine kinase; PTEN, phosphatase and
tensin homolog; BLM, BLM RecQ like helicase; BRIP1, BRCA1
interacting protein C-terminal helicase 1; FANCA, FA
complementation group A; FANCC, FA complementation group C; FANCD2,
FA complementation group D2; FANCF, FA complementation group F;
FANCG, FA complementation group G; NBN, nibrin; PALB2, partner and
localizer of BRCA2; WRN, Werner syndrome RecQ like helicase; FANCE,
FA complementation group E. The numbers in brackets on the x-axis
represents the number of samples with inactivation mutations. |
Discussion
GI cancer is a leading cause of morbidity and
mortality, and accounts for 25% of new cancer cases and
cancer-associated mortalities worldwide (1,38).
Despite modern improvements in treatment, the majority of patients
with GI cancer have a poor prognosis due to late-stage diagnosis
(3). To date, early screening and
diagnosis based on the understanding of the molecular mechanisms
underlying GI cancer is becoming the best way to benefit patients
with GI cancer (39). MMR deficiency
and MSI in particular, which are suspected to be associated with
the efficacy of immune checkpoint inhibitors (40). Approximately 15% of patients with CRC
exhibit high MSI, mostly as a result of dMMR (41). In the present study, a systematic
analysis of 183 gene mutations in 137 patients with GI cancer was
performed to evaluate novel molecular characteristics in GI cancer.
Furthermore, the effects of MMR and HR mutational status on genomic
instability were investigated.
Several studies have proposed APC as one of
the most prominent tumor promoting genes, regulating the
Wnt/β-catenin signaling pathway and participating in the
tumorigenesis of CRC (42–44). The results obtained in the current
study revealed that the APC gene was mutated in 81% of
patients with GI cancer. Other genes, including PMS2,
neurofibromin 1 and ATM, also exhibited a high mutation
frequency. Notably, seven of the 20 most frequently mutated genes
in the 183-gene panel were DNA repair genes (MMR gene, MSH2;
HR genes, ATM, FANCD2, ATR, BRCA2, FANCA and BRCA1).
This indicated that homologous recombination repair is an important
event during the progression of GI cancer, in addition to MMR. With
the high mutation frequency of the MMR and HR genes, correlation
analysis was performed and a significant association between the
mutational status of MMR or HR genes and genomic instability in GI
cancers was identified. The higher the number of mutations in MMR
or HR genes, the higher the number of mutations in the 183-gene
panel.
The DNA MMR system regulates genetic fidelity, the
accumulation of genetic errors, MSI and intestinal carcinogenesis
(13,14). In the present study, 40.9% (n=56,137)
of patients with GI cancer had one or more MMR gene inactivating
mutations, whereas HR deficiency occurred in 81.8% (112/137) of the
patients. A previous study indicated that 35% of patients with
pancreatic cancer (n=109) harbored pathogenic or likely pathogenic
mutations in HR genes (45). In
addition, no significant association was observed between genomic
instability and other clinical characteristics, including sex, age
and tumor type in GI cancer. One large-scale analysis suggested
that 13% of all tumor types had HR deficiency (n=53,619), including
PTEN (5.8%), BRCA2 (2.8%), BRCA1 (2.6%) and
ATM (1.2%) (27). However,
the data obtained in the current study revealed that MSH6
was the most frequently mutated MMR gene, with an inactivation
mutation in 20.44% of the cases. A high deficiency frequency of
certain HR genes, including BRCA2, ATM, BRCA1, PTEN and
FANCD2, was detected in the current study. These results
demonstrated the molecular characteristics of HR gene inactivation
mutations, in addition to dMMR. Furthermore, almost no splicing
mutations in MMR and HR genes were detected in GI cancer. Further
investigation revealed that patients with MLH1, PTEN, FANCC
or PALB2 inactivation mutations did not exhibit significant
differences in the numbers of SNV and indel mutations compared with
patients without inactivation mutations in these genes.
No notably different effects on genomic instability
among the four groups with MLH1, MSH2, MSH6 and PMS2
inactivation mutations were observed in the current study. However,
the effect on genomic instability of certain genes, including
BRCA1, BRCA2, ATM, FANCA and BRIP1, was more
pronounced compared with that of PTEN in patients with HR
deficiency.
The present study demonstrated the association
between genomic instability and the mutational status of MMR and HR
genes in GI cancer. The results obtained reveal a novel model based
on the mutational status of DNA repair genes for predicting
response to therapy. The higher the number of mutations detected in
DNA repair genes, the more tumor neoantigens may be induced;
however, further research and evaluation of this model is
required.
Acknowledgements
The authors would like to thank Professor Jian Xu
[First Dimension Biosciences (Suzhou) Co., Ltd., Suzhou, China] for
contributing to revising the manuscript.
Funding
No funding was received.
Availability of data and material
The datasets generated and/or analyzed during the
present study are not publicly available due to confidential
personal information that could compromise research participant
privacy.
Authors' contributions
XL, TT and DL designed the study. KH, PJ and WZ
clinically diagnosed the patients, obtained the informed consent,
harvested tissue samples, examined the archives and identified the
cases included in the study, examined the slides and collected
pathological information. HY, XW and PM performed the data
analysis. PM and HY wrote the manuscript. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
The First Affiliated Hospital of Anhui Medical
University Ethics Committee approved the study and all patients
provided written informed consent prior to enrollment into the
project.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Chen W, Zheng R, Baade PD, Zhang S, Zeng
H, Bray F, Jemal A, Yu XQ and He J: Cancer statistics in China,
2015. CA Cancer J Clin. 66:115–132. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Brody H: Colorectal cancer. Nature. 521
(Suppl):S12015. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ng SC and Wong SH: Colorectal cancer
screening in Asia. Br Med Bull. 105:29–42. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Labianca R, Nordlinger B, Beretta GD,
Mosconi S, Mandalà M, Cervantes A and Arnold D; ESMO Guidelines
Working Group, : Early colon cancer: ESMO Clinical Practice
Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 6
(Suppl):vi64–vi72. 2013.
|
|
5
|
Lutgens MW, Oldenburg B, Siersema PD, van
Bodegraven AA, Dijkstra G, Hommes DW, de Jong DJ, Stokkers PC, van
der Woude CJ and Vleggaar FP: Colonoscopic surveillance improves
survival after colorectal cancer diagnosis in inflammatory bowel
disease. Br J Cancer. 101:1671–1675. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lansdorp-Vogelaar I, van Ballegooijen M,
Zauber AG, Habbema JD and Kuipers EJ: Effect of rising chemotherapy
costs on the cost savings of colorectal cancer screening. J Natl
Cancer Inst. 101:1412–1422. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Beamer LC, Grant ML, Espenschied CR,
Blazer KR, Hampel HL, Weitzel JN and MacDonald DJ: Reflex
immunohistochemistry and microsatellite instability testing of
colorectal tumors for Lynch syndrome among US cancer programs and
follow-up of abnormal results. J Clin Oncol. 30:1058–1063. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Burt RW: Who should have genetic testing
for the Lynch syndrome? Ann Intern Med. 155:127–128. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Matloff J, Lucas A, Polydorides AD and
Itzkowitz SH: Molecular tumor testing for Lynch syndrome in
patients with colorectal cancer. J Natl Compr Canc Netw.
11:1380–1385. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ward RL, Hicks S and Hawkins NJ:
Population-based molecular screening for Lynch syndrome:
Implications for personalized medicine. J Clin Oncol. 31:2554–2562.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Broustas CG and Lieberman HB: DNA damage
response genes and the development of cancer metastasis. Radiat
Res. 181:111–130. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Spies M and Fishel R: Mismatch repair
during homologous and homeologous recombination. Cold Spring Harb
Perspect Biol. 7:a0226572015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Vilar E and Gruber SB: Microsatellite
instability in colorectal cancer-the stable evidence. Nat Rev Clin
Oncol. 7:153–162. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Vilar E and Tabernero J: Molecular
dissection of microsatellite instable colorectal cancer. Cancer
Discov. 3:502–511. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ribic CM, Sargent DJ, Moore MJ, Thibodeau
SN, French AJ, Goldberg RM, Hamilton SR, Laurent-Puig P, Gryfe R,
Shepherd LE, et al: Tumor microsatellite-instability status as a
predictor of benefit from fluorouracil-based adjuvant chemotherapy
for colon cancer. N Engl J Med. 349:247–257. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Dudley JC, Lin MT, Le DT and Eshleman JR:
Microsatellite Instability as a Biomarker for PD-1 Blockade. Clin
Cancer Res. 22:813–820. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kastrinos F and Stoffel EM: The history,
genetics, and strategies for cancer prevention in Lynch syndrome.
Clin Gastroenterol Hepatol. 12:715–727.e41-e43. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lynch HT and de la Chapelle A: Genetic
susceptibility to non-polyposis colorectal cancer. J Med Genet.
36:801–818. 1999.PubMed/NCBI
|
|
19
|
Herman JG, Umar A, Polyak K, Graff JR,
Ahuja N, Issa JP, Markowitz S, Willson JK, Hamilton SR, Kinzler KW,
et al: Incidence and functional consequences of hMLH1 promoter
hypermethylation in colorectal carcinoma. Proc Natl Acad Sci USA.
95:6870–6875. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Veigl ML, Kasturi L, Olechnowicz J, Ma AH,
Lutterbaugh JD, Periyasamy S, Li GM, Drummond J, Modrich PL,
Sedwick WD and Markowitz SD: Biallelic inactivation of hMLH1 by
epigenetic gene silencing, a novel mechanism causing human MSI
cancers. Proc Natl Acad Sci USA. 95:8698–8702. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang YC, Lu YP, Tseng RC, Lin RK, Chang
JW, Chen JT, Shih CM and Chen CY: Inactivation of hMLH1 and hMSH2
by promoter methylation in primary non-small cell lung tumors and
matched sputum samples. J Clin Invest. 111:887–895. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hsu HS, Wen CK, Tang YA, Lin RK, Li WY,
Hsu WH and Wang YC: Promoter hypermethylation is the predominant
mechanism in hMLH1 and hMSH2 deregulation and is a poor prognostic
factor in nonsmoking lung cancer. Clin Cancer Res. 11:5410–5416.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Yurgelun MB, Allen B, Kaldate RR, Bowles
KR, Judkins T, Kaushik P, Roa BB, Wenstrup RJ, Hartman AR and
Syngal S: Identification of a variety of mutations in cancer
predisposition genes in patients with suspected lynch syndrome.
Gastroenterology. 149:604–613.e20. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Pearlman R, Frankel WL, Swanson B, Zhao W,
Yilmaz A, Miller K, Bacher J, Bigley C, Nelsen L, Goodfellow PJ, et
al: Prevalence and spectrum of germline cancer susceptibility gene
mutations among patients with early-onset colorectal cancer. JAMA
Oncol. 3:464–471. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Antoniou AC, Casadei S, Heikkinen T,
Barrowdale D, Pylkäs K, Roberts J, Lee A, Subramanian D, De Leeneer
K, Fostira F, et al: Breast-cancer risk in families with mutations
in PALB2. N Engl J Med. 371:497–506. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Thompson D, Duedal S, Kirner J, McGuffog
L, Last J, Reiman A, Byrd P, Taylor M and Easton DF: Cancer risks
and mortality in heterozygous ATM mutation carriers. J Natl Cancer
Inst. 97:813–822. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Konstantinopoulos PA, Ceccaldi R, Shapiro
GI and D'Andrea AD: Homologous recombination deficiency: Exploiting
the fundamental vulnerability of ovarian cancer. Cancer Discov.
5:1137–1154. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
da Cunha Colombo Bonadio RR, Fogace RN,
Miranda VC and Diz MDPE: Homologous recombination deficiency in
ovarian cancer: A review of its epidemiology and management.
Clinics (Sao Paulo). 73 (Suppl 1):e450s2018.PubMed/NCBI
|
|
29
|
Chao EC and Lipkin SM: Molecular models
for the tissue specificity of DNA mismatch repair-deficient
carcinogenesis. Nucleic Acids Res. 34:840–852. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Gentles L, Goranov B, Matheson E, Herriott
A, Kaufmann A, Hall S, Mukhopadhyay A, Drew Y, Curtin NJ and
O'Donnell RL: Exploring the frequency of homologous recombination
DNA repair dysfunction in multiple cancer types. Cancers (Basel).
11(pii): E3542019. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Cerbinskaite A, Mukhopadhyay A, Plummer
ER, Curtin NJ and Edmondson RJ: Defective homologous recombination
in human cancers. Cancer Treat Rev. 38:89–100. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Miranda E, Destro A, Malesci A, Balladore
E, Bianchi P, Baryshnikova E, Franchi G, Morenghi E, Laghi L,
Gennari L and Roncalli M: Genetic and epigenetic changes in primary
metastatic and nonmetastatic colorectal cancer. Br J Cancer.
95:1101–1107. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Malapelle U, Mayo de-Las-Casas C, Rocco D,
Garzon M, Pisapia P, Jordana-Ariza N, Russo M, Sgariglia R, De Luca
C, Pepe F, et al: Development of a gene panel for next-generation
sequencing of clinically relevant mutations in cell-free DNA from
cancer patients. Br J Cancer. 116:802–810. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Li H and Durbin R: Fast and accurate short
read alignment with Burrows-Wheeler transform. Bioinformatics.
25:1754–1760. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
McKenna A, Hanna M, Banks E, Sivachenko A,
Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly
M and DePristo MA: The genome analysis toolkit: A MapReduce
framework for analyzing next-generation DNA sequencing data. Genome
Res. 20:1297–1303. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Cibulskis K, Lawrence MS, Carter SL,
Sivachenko A, Jaffe D, Sougnez C, Gabriel S, Meyerson M, Lander ES
and Getz G: Sensitive detection of somatic point mutations in
impure and heterogeneous cancer samples. Nat Biotechnol.
31:213–219. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Sherry ST, Ward MH, Kholodov M, Baker J,
Phan L, Smigielski EM and Sirotkin K: dbSNP: The NCBI database of
genetic variation. Nucleic Acids Res. 29:308–311. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Vucic EA, Thu KL, Robison K, Rybaczyk LA,
Chari R, Alvarez CE and Lam WL: Translating cancer ‘omics’ to
improved outcomes. Genome Res. 22:188–195. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Le DT, Durham JN, Smith KN, Wang H,
Bartlett BR, Aulakh LK, Lu S, Kemberling H, Wilt C, Luber BS, et
al: Mismatch-repair deficiency predicts response of solid tumors to
PD-1 blockade. Science. 357:409–413. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Copija A, Waniczek D, Witkos A, Walkiewicz
K and Nowakowska-Zajdel E: Clinical significance and prognostic
relevance of microsatellite instability in sporadic colorectal
cancer patients. Int J Mol Sci. 18(pii): E1072017. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Kaplan KB, Burds AA, Swedlow JR, Bekir SS,
Sorger PK and Näthke IS: A role for the adenomatous polyposis Coli
protein in chromosome segregation. Nat Cell Biol. 3:429–432. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Leoz ML, Carballal S, Moreira L, Ocana T
and Balaguer F: The genetic basis of familial adenomatous polyposis
and its implications for clinical practice and risk management.
Appl Clin Genet. 8:95–107. 2015.PubMed/NCBI
|
|
44
|
Krausova M and Korinek V: Wnt signaling in
adult intestinal stem cells and cancer. Cell Signal. 26:570–579.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Witkiewicz AK, McMillan EA, Balaji U, Baek
G, Lin WC, Mansour J, Mollaee M, Wagner KU, Koduru P, Yopp A, et
al: Whole-exome sequencing of pancreatic cancer defines genetic
diversity and therapeutic targets. Nat Commun. 6:67442015.
View Article : Google Scholar : PubMed/NCBI
|