Introduction
Osteosarcoma is the most common malignant primary
bone tumor and frequently undergoes distal metastasis, thus making
it a severe threat to both adolescents and young adults (1). The 5-year survival rate of localized
osteosarcoma has reached 80% in recent years due to major
developments in clinical treatment (2). However, patients with metastatic
osteosarcoma have a 5-year survival rate of only 20% (3). Although advanced therapies have been
developed for distal metastases from osteosarcoma, knowledge on
novel therapy targets that would prevent metastasis is still
lacking (4,5). Recent studies have demonstrated that
non-coding RNAs play a critical role in osteosarcoma pathology,
particularly during metastasis, which has prompted a search for
novel strategies for osteosarcoma treatment and metastasis
prevention (6–8). It is important to identify novel
molecular mechanisms involved in osteosarcoma metastasis.
MicroRNAs (miRNAs/miRs) are small, non-coding RNAs
that play critical roles in almost all cellular processes via
post-transcriptional regulation of mRNAs, including cancer cell
metastasis (9). miRNAs are involved
at each step of development of osteosarcoma (10,11). RNA
sequencing of osteosarcoma tissues and pulmonary metastatic
osteosarcoma tissues has led to the identification of miRNAs, long
non-coding RNAs and mRNAs with differential expression in primary
osteosarcoma, pulmonary metastatic osteosarcoma and normal
controls, and has uncovered the molecular mechanisms and signaling
networks that contribute to osteosarcoma progression (6). However, only a few differentially
expressed miRNAs have been identified, including miR-340,
miR-323-3p and miR-193a-3p/5p (6,12,13).
miR-545-5p has recently been reported to be upregulated in
metastatic colorectal cancer cells compared with its localized
strain (14), suggesting that
miR-545-5p plays a role in cancer cell metastasis. Currently, only
a single downstream target of miR-545-5p, delta-aminolevulinic acid
dehydratase (15), which is a
critical regulator of heme biosynthesis and pigmentation in
invertebrates (16,17), has been identified. Reportedly,
miR-545 serves as a tumor regulator in different types of cancer,
including hepatocellular carcinoma, cervical cancer and oral
squamous cell carcinoma (18–20).
Recently, Miao et al (21)
demonstrated that downregulation of miR-545-5p expression in colon
adenocarcinoma is associated with the survival and proliferation of
colon cancer cells. However, the role of miR-545-5p in osteosarcoma
remains unclear.
The present study aimed to further reveal the role
of miR-545-5p in the metastasis of osteosarcoma, by determining the
expression of miR-545-5p in clinical osteosarcoma samples and
Saos-2, U2OS and MG63 osteosarcoma cell lines, and by assessing its
roles in apoptosis and metastasis by transfection with miR-545-5p
mimics in osteosarcoma cell lines. This study may help to provide
new insights on the understanding and prevention of osteosarcoma
metastasis.
Materials and methods
Clinical samples and cell culture
A total of 40 patients with osteosarcoma (32 men and
8 women; mean age ± SD, 30.82±5.92; age range, 18–48 years) at
Honghui Hospital (Xi'an, China) were enrolled in the present study
between May 2018 and April 2019. The inclusion criteria were that
all patients were diagnosed by pathological and imaging examination
and all cases were newly diagnosed. The exclusion criteria were as
follows: i) Recurrent patients; ii) patients who had received
immunotherapy, radiotherapy or chemotherapy and iii) patients with
multiple newly diagnosed clinical disorders. Tumor tissues and
adjacent normal tissues (0.5 cm from the tumor boundary) were
collected via surgical resection. Both RNAs and proteins were
extracted from the samples and conserved in liquid nitrogen
(−196°C) for subsequent experimentation. The present study was
approved by the Ethics Committee of Honghui Hospital (Xi'an, China;
approval no. 202003059) and written informed consent was provided
by all patients prior to the study start.
The human osteoblast cell line, hFOB1.19, and
osteosarcoma cell lines, Saos-2, U2OS and MG63 were purchased from
The Cell Bank of Type Culture Collection of The Chinese Academy of
Sciences, and cultured according to the manufacturer's
instructions. Briefly, cryopreserved hFOB1.19 cells were recovered
from liquid nitrogen using the AccuVital cell recovery kit (AccuRef
Scientific) and cultured in 1:1 mixture of F12 medium (SH30023.01,
HyClone; Cytiva) and DMEM (SH30021.01; HyClone; Cytiva), with 2.5
mM L-glutamine (25030149; Gibco; Thermo Fisher Scientific, Inc.),
0.3 mg/ml G418 (A2513; APeXBIO Technology LLC) and 10% fetal bovine
serum (FBS, 10099141; Gibco; Thermo Fisher Scientific, Inc.).
Saos-2 and U2OS cells were maintained in McCoy's 5A medium
(16600082; Gibco; Thermo Fisher Scientific, Inc.) supplemented with
15% FBS. MG63 cells were maintained in MEM (MEL06-500ML; Caisson
Labs) supplemented with 10% FBS. All cells were cultured at 37°C
with 95% air and 5% CO2.
In situ hybridization of
miR-545-5p
In situ hybridization was performed as
previously described (18). The
miR-545-5p probe tagged with Digoxigenin (DIG) modifications and
modified with Locked Nucleic Acid (LNA) nucleotides was purchased
from Redlandbio Technology. On day 1, 4-µm sections of the TMA
blocks were placed in a heater at 60°C overnight to attach Super
Frost Plus Slides (Thermo Fisher Scientific, Inc.). Next, sections
were deparaffinized with xylene twice for 5 min, rehydrated with an
ethanol gradient, washed with DEPC water for 1 min and digested
with proteinase K (15 µg/ml; MilliporeSigma) in PK buffer [5 mM
Tris-HCl (pH 7.5) and 1 mM NaCl] at 37°C for 10 min. After this,
TMAs were rehydrated with ethanol and air-dried, followed by
denaturing of LNA-probes at 90°C for 5 min. Following this,
LNA-probes miR-545-5p (25 nM) against miR-545-5p were incubated
with the samples at 60°C for 5 min and then at 37°C overnight. On
day 2, samples were washed with saline sodium citrate and 2% bovine
serum albumin for 5 min at 4°C twice and incubated with
anti-DIG/alkaline phosphate conjugated antibody (dilution, 1:400;
cat. no. 11207733910; MilliporeSigma) at 37°C for 0.5 h. Next, the
blue color was generated by incubation with with Alexa Fluor
488-labeled antibody (dilution, 1:100; cat. no. A-11078;
Invitrogen; Thermo Fisher Scientific, Inc.) at 37°C for 1 h. The
slides were then washed with TBS twice at room temperature and
mounted using prolong gold anti-fade reagent with DAPI (cat. no.
P36935; Invitrogen; Thermo Fisher Scientific, Inc.). Slides were
air dried in a dark room overnight and covered with nail polish
before imaging in a Zeiss LSM 880 inverted confocal microscope
(Carl Zeiss AG).
Cell proliferation assay
The Cell Counting Kit-8 (CCK-8) assay (AccuRef
Scientific) was performed to detect cell proliferation, according
to the manufacturer's instructions. Briefly, 2×103 cells
were plated into a 96-well plate and technical repeats were
performed in triplicate. After 24 seeding, cells were transfected
with miR-545-5p mimic with or without co-transfection of DIMT1
overexpressing plasmid using Lipofactamine® 2000
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocols. The cells were incubated at 37°C for 48
h. Following 48 h of transfection, 10 µl of CCK-8 solution was
added to each well and incubated at 37°C for 2 h. The optical
density of each sample was measured at a wavelength of at 450 nm,
using a microplate reader (Thermo Fisher Scientific, Inc.).
In addition, the EdU assay was performed using an
EdU proliferation kit (AccuRef Scientific), according to the
manufacturer's instructions. A total of 2×103 cells were
seeded into a 96-well plate and incubated with 20 µM EdU for 2 h,
24 h post-seeding. Subsequently, cells were fixed with 4%
paraformaldehyde at room temperature for 30 min and stained with
EdU additive solution at room temperature for 2 h. Samples were
analyzed using a MoFlo Astrios cell sorter (Beckman Coulter,
Inc.).
Cell transfection
miR-545-5p inhibitor (50 nM,
5′-UCAUCUAAUAAACAUUUACUGA-3′), inhibitor negative control (NC, 50
nM, 5′-AACCUUUAGGGUUCUAGGGAGG-3′), miR-545-5p mimic (50 nM,
5′-UCAGUAAAUGUUUAUUAGAUGA-3′) and mimic NC (50 nM,
5′-UUGUACUACACAAAAGUACUG-3′) were purchased from Changzhou Ruibo
Bio-Technology Co., Ltd. Specific small interfering (si)RNAs
against DIMT1 (si-DIMT1, 50 nM, 5′-CCATAATATTGTCAGTGCT-3′) and
si-NC (50 nM. 5′-CACUGAUUUCAAAUGGUGCUAUU-3′) were synthesized by
Shanghai GenePharma Co., Ltd. The plasmid construction and the
corresponding NC were also synthesized by Shanghai GenePharma Co.,
Ltd. Plasmid transfection technology was applied to overexpress
DIMT1, with empty vector as its control. All transfections were
performed using Lipofectamine® 2000 (Invitrogen; Thermo
Fisher Scientific, Inc.), according to the manufacturer's protocol.
Briefly, miR-545-5p mimic, miR-545-5p inhibitor, DIMT1 siRNA or
plasmid (10 µg) were diluted in serum-free MEM (Caisson Labs) or
McCoy's 5A medium (Gibco; Thermo Fisher Scientific, Inc.), mixed
with Lipofectamine® 2000, and incubated for 20 min at
room temperature. Subsequently, cells were incubated with the
transfected mixture at 37°C with 5% CO2. Following
incubation for 6 h, the medium was replaced with MEM or McCoy's 5A
medium containing 10% FBS, and cells were harvested and subjected
to subsequent experimentation 48 h post-transfection. Transfection
efficiency was assessed via reverse transcription-quantitative
(RT-q)PCR analysis.
RT-qPCR
RNA was extracted from tissues or cells using
TRIzol® reagent (cat. no. 15596-018, Invitrogen; Thermo
Fisher Scientific, Inc.), according to the manufacturer's
instructions. RT was performed using 500 µg RNA and the
PrimeScript™ RT reagent kit (cat. no. RR047A; Takara Biotechnology
Co., Ltd.) according to the manufacturer's protocol. qPCR was
subsequently performed using SYBR Green (cat. no. RR420A; Takara
Biotechnology Co., Ltd.) in an ABI 7500 Real time PCR instrument
(Applied Biosystems) using the following conditions: 95°C for 5
min, followed by 40 cycles of 95°C for 15 sec, 58°C for 30 sec and
72°C for 10 sec. Fold-changes were calculated using the
2−ΔΔCq method (22). Each
sample was analyzed in triplicate. The following primer sequences
were used for qPCR: DIMT1 forward, 5′-GGCTGCCTTAAGACCAACTG-3′ and
reverse, 5′-CGTGCCCTGAACTCTTTTGT-3′; GAPDH forward,
5′-ACCAGGAAATGAGCTTGACA-3′ and reverse, 5′-GACCACAGTCCATGCCATC-3′;
miR-545-5p forward, 5′-TCAGTAAATGTTTATTAGATGA-3′ and reverse,
universal oligo(dT) reverse primer; and U6 forward,
5′-CTCGCTTCGGCAGCACA-3′ and reverse,
5′-AACGCTTCACGAATTTGCGT-3′.
Cell apoptosis analysis
Cell apoptosis was analyzed using the Annexin V-FITC
Apoptosis kit (cat. no. K101; Biovision, Inc.), according to the
manufacturer's instructions. Briefly, cells were seeded into 6-well
plates at a density of 3×105 cells/well and transfected
with miR-545-5p mimic with or without co-transfection of DIMT1
overexpressing plasmid for 48 h. Cells were collected and washed
with PBS. Following resuspension with Annexin binding buffer, cells
were labeled with Annexin V-FITC for 15 min at 4°C followed by
incubation with propidium iodide (PI) for 5 min at 4°C. Apoptotic
cells were subsequently analyzed using a flow cytometer (FACSCanto
II; BD Biosciences) and CellQuest 3.0 software (BD
Biosciences).
Wound healing assay
Following the indicated treatments, a monolayer of
Saos-2 or MG63 cells was scratched using a 10-µl pipette tip.
Scratched cells were removed by rinsing twice with PBS. Cells were
incubated at 37°C for an additional 48 h in serum-free DMEM. The
wound of each sample was pictured at 0 and 48 h using a light
microscope (Olympus Corporation) at ×200 magnification. The wound
distances were quantitatively evaluated using ImageJ software
(version 1.49; National Institutes of Health), as previously
described (23). Each wound healing
analysis was repeated in triplicate, using independent samples.
Transwell invasion assay
A Transwell invasion analysis was performed as
previously described (24). Briefly,
a 1:1 mix of matrigel and the indicated growth medium was used to
coat the Transwell (Costar). The coated Transwell was then
incubated at 37°C for 1 h before 1×105 cells/well were
seeded to the upper chamber, each with three replicates. The lower
chamber was flooded with culture medium containing 10% FBS. After
24 h of culture, the cells were fixed with 4% polyoxymethylene and
stained with crystal violet at room temperature for 15 min.
Following this, the stained results were captured and analyzed
under a light microscope (Olympus Corporation) at magnification of
×200.
Western blotting
Protein collection and western blot analysis were
performed as previously described (25). Cells and tissue samples were lysed on
ice for 30 min using RIPA lysis buffer [50 mM Tris (pH 7.4), 150 mM
NaCl, 1% NP-40, 0.5% sodium deoxycholate; Beijing Solarbio Science
& Technology Co., Ltd.] before centrifuging at 4°C, 17,000 × g
for 30 min. Proteins were quantified using the bicinchoninic acid
assay method (Thermo Fisher Scientific, Inc.), and equivalent
protein (25 µg) was loaded to each lane and separated on 10% gels
using SDS-PAGE. Following this, proteins were transferred to
polyvinylidene fluoride membrane and blocked with 5% skimmed milk
solution at room temperature for 1 h. Subsequently, membranes were
incubated with mouse anti-human primary antibodies against DIMT1
(cat. no. ab69434; Abcam) and rabbit anti-human GAPDH (cat. no.
ab97627; Abcam) at 4°C overnight. After this, membranes were
incubated with the rat anti-mouse (cat. no. ab6728; Abcam) or mouse
anti-rabbit (cat. no. ab99697; Abcam) secondary antibody at room
temperature for 1 h. Finally, protein bands were visualized using a
chemiluminescence kit (cat. no. AR21PN003; AccuRef Scientific) and
qualified using ImageJ software (version 1.49; National Institutes
of Health).
In vivo analysis of tumor growth
For the subcutaneous xenograft study, 8-week-old
male BALB/c nude mice (n=16, 20–25 g) were purchased from the
Laboratory Animal Center of Xi'an Jiaotong University, and housed
at 25°C with a 12 h light/dark cycle. The mice were reared under
specific pathogen-free conditions (25°C with 55% humidity), with
free access to food and water.
A total of 3×106 Saos-2 or MG63 cells
transfected with control miRNA (mimic NC) or miR-545-5p mimics for
24 h were trypsinized, resuspend in PBS and subcutaneously injected
into the left or right back of each mouse. The xenografts were
measured every 2 days and their volumes were calculated using the
following formula: axb2/2 [where (a) is the largest, and
(b) is the smallest]. The mice were euthanized by CO2
inhalation (CO2 flow rate, 30% of cage volume) (26) 14 days post-transplantation and
mortality was confirmed via cessation of heartbeat. The tumor
tissues were dissected, photographed and weighed. RNA was
subsequently extracted from the xenografts and RT-qPCR analysis.
All animal experiments were performed in accordance with the Guide
for the Care and Use of Laboratory Animals of the National
Institutes of Health (27) and
approved by the Animal Care and Use Committee of Hong Hui Hospital,
Xi'an Jiaotong University (Xi'an, China; approval no.
202003059).
Dual-luciferase reporter assay
DNA fragments containing wild-type (WT) or mutated
(MUT) DIMT1 DNA binding sites were synthesized by BGI. (https://en.genomics.cn/), and cloned into a pGL3
firefly reporter plasmid (Promega Corporation). All plasmids were
purchased from Guangzhou RiboBio Co., Ltd. Cells were cultured to a
confluence of 70–80% in 24-well plates, followed by transfection
with 100 ng pGL3-WT or pGL3-MUT, 20 ng of the transfection control
Renilla vector (pRL-TK; Promega Corporation) and 100 nM
miR-545-5p mimic or mimic NC (Guangzhou RiboBio Co., Ltd.), using 1
µl Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol. After 2
days, luciferase activities were detected via the dual-luciferase
reporter assay (Promega Corporation). The firefly luciferase
activity was normalized to Renilla luciferase activity, and
the relative luciferase activity in each group was calculated using
the ratio of firefly luciferase activity to Renilla
luciferase activity compared with that of the NC group.
Target prediction
miR-545-5p targeted candidate genes and the binding
sites of miR-545-5p were predicted using the online tools,
TargetScanV7.2 (http://www.targetscan.org/vert_72), miRDB (http://www.mirdb.org/mirdb/index.html)
and DIANA-microT (http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=microT_CDS/index).
The following parameters were set for each database: DIANA-microT
(miTG score ≥0.8); miRDB (Target score ≥80) and TargetScan
(Cumulative weighted context++ score ≤-0.5). The Venn diagram was
constructed using VENNY 2.1(https://bioinfogp.cnb.csic.es/tools/venny). The
miRNA-mRNA regulation network was constructed using Cytoscape
(version 3.6.1) (27).
Immunohistochemistry staining
The protein expression of DIMT1 in clinical samples
was determined using immunohistochemistry. Briefly, clinical tissue
samples were washed with PBS to remove blood on the surface and
fixed with 4% formaldehyde at 4°C for 48 h. The tissues were then
dehydrated with gradient ethanol and paraffin-embedded at 60°C
overnight. Tissues were cut into 5-µm thick sections, heated at
60°C for 1 h, de-waxed using xylene and rehydrated with gradient
alcohol. Following this, antigens on sections were retrieved using
a microwave oven and boiled 0.01 M citrate buffer (pH 6.0) for 3
min, three times at 5-min intervals. After this, sections were
incubated with 3% H2O2 for 10 min at room
temperature to block endogenous peroxidase. Next, sections were
blocked with 5% goat serum (Beyotime Institute of Biotechnology) at
room temperature for 1 h. The sections were then incubated with
mouse anti-human DIMT1 antibody (dilution, 1:500; cat. no.
HPA042944; MilliporeSigma) overnight at 4°C. The next day, sections
were incubated with donkey anti-rabbit secondary antibody
(dilution, 1:400; cat. no. ab207999; Abcam) at room temperature for
1 h and visualized using a DAB Substrate kit (cat. no. ab64238;
Abcam) according to the manufacturer's protocol. The nuclei on the
sections were counterstained with hematoxylin at room temperature
for 5–10 min and washed with running water. Following this,
sections were dehydrated with ethanol, hyalinized by xylene and
mounted with neutral resins (cat. no. E675007; Sangon Bioteck Co.
Ltd.). Subsequently, images of the sections were captured and
analyzed under a light microscope (Olympus Corporation).
Statistical analysis
Statistical analysis was performed using GraphPad
Prism 5.0 software (GraphPad Software, Inc.). All experiments were
performed in triplicate and data are presented as the mean ± SD.
Paired Student's t-test was used to compare differences between
tumor tissues and adjacent normal tissues. Unpaired Student's
t-test was used to compare differences between two groups, while
one-way ANOVA followed by Tukey's post hoc test were used to
compare differences between multiple groups. According to the
median value of miR-545-5p expression, patients were divided into
the high and low expression groups, and comparisons between them
were compared with χ2 analysis. P<0.05 was considered
to indicate a statistically significant difference.
Results
miR-545-5p expression is downregulated
in osteosarcoma
To understand the role that miR-545-5p plays in
osteosarcoma, RT-qPCR analysis was performed to detect miR-545-5p
expression in both clinical osteosarcoma tissues and adjacent
normal tissues. The results demonstrated that miR-545-5p expression
was significantly downregulated in osteosarcoma tissues compared
with adjacent normal tissues (P<0.05; Fig. 1A). miR-545-5p expression was also
detected in osteosarcoma cell lines. The results demonstrated that
miR-545-5p expression was downregulated in all three osteosarcoma
cell lines compared with the human normal osteoblast cell line,
hFOB1.19, and its expression was downregulated by >50% in Saos-2
and MG63 cells (P<0.05; Fig. 1B).
To confirm these results, in situ hybridization was
performed for miR-545-5p in clinical osteosarcoma tissues and
adjacent normal tissues. The results revealed a notable decrease in
the number of miR-545-5p positive cells in osteosarcoma tissues
compared with adjacent normal tissues (Fig. 1C).
To determine the clinical significance of miR-545-5p
in patients with osteosarcoma, the association between miR-545-5p
expression and the clinicopathological characteristics of 40
patients with osteosarcoma was assessed. Based on the median
miR-545-5p expression level in clinical samples, patient samples
were grouped into either a high or low expression groups (n=10 for
each group). The results demonstrated that miR-545-5p expression
was significantly associated with tumor size, metastasis and
differentiation (all P<0.05; Table
I).
 | Table I.Association between miR-545-5p
expression and the clinicopathological characteristics of patients
with osteosarcoma (n=40). |
Table I.
Association between miR-545-5p
expression and the clinicopathological characteristics of patients
with osteosarcoma (n=40).
|
| miR-545-5p
expression |
|
|---|
|
|
|
|
|---|
| Characteristic | Low (n=20) | High (n=20) | P-value |
|---|
| Age, years |
|
| 0.525 |
|
<55 | 10 | 12 |
|
|
≥55 | 10 | 8 |
|
| Sex |
|
| 0.236 |
|
Male | 18 | 14 |
|
|
Female | 2 | 6 |
|
| Tumor size, cm |
|
| 0.025 |
|
<3 | 15 | 8 |
|
|
>3 | 5 | 12 |
|
| Location |
|
| 0.185 |
|
Tibia/femur | 17 | 25 |
|
|
Elsewhere | 13 | 5 |
|
| Clinical stage |
|
| 0.507 |
| I | 14 | 12 |
|
| II | 6 | 8 |
|
| Distant
metastasis |
|
| 0.018 |
|
Absent | 16 | 12 |
|
|
Present | 4 | 8 |
|
| TNM stage |
|
| 0.011 |
|
I–II | 13 | 5 |
|
|
III–IV | 7 | 15 |
|
| Differentiated
degree |
|
| 0.044 |
|
High/middle | 15 | 6 |
|
|
Low/undifferentiated | 5 | 14 |
|
miR-545-5p inhibits the proliferation,
migration and invasion of osteosarcoma cells
To determine the role of miR-545-5p in osteosarcoma,
cells were transfected with miR-545-5p mimics to increase
miR-545-5p expression in osteosarcoma cells (Fig. S1A). Notably, the proliferation of
Saos-2 and MG63 cells decreased following overexpression of
miR-545-5p (both P<0.05; Fig.
2A). The EdU assay was performed to detect the proliferation of
osteosarcoma cells, and the results demonstrated that
overexpression of miR-545-5p inhibited the proliferation of Saos-2
and MG63 cells (both P<0.05; Fig.
2B). Notably, transfection with miR-545-5p mimics significantly
induced apoptosis of both Saos-2 and MG63 cells (both P<0.05;
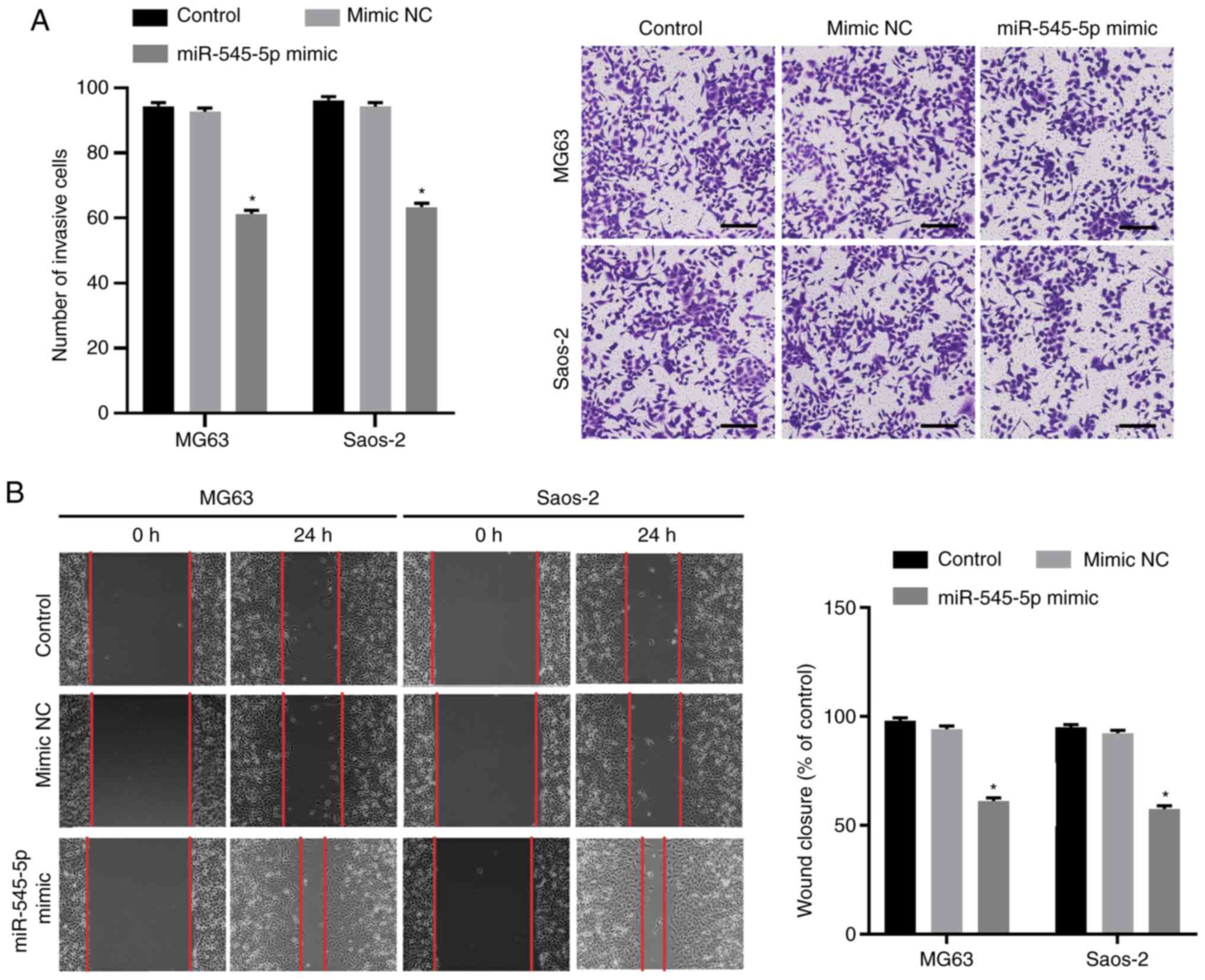
Fig. 2C). The Transwell and wound
healing assays were performed to evaluate the effect of miR-545-5p
overexpression on the invasion and migration of osteosarcoma cells,
respectively. The results demonstrated that the number of cells
infiltrating through the Matrigel significantly decreased following
transfection with miR-545-5p mimics (both P<0.05; Fig. 3A), suggesting that overexpression of
miR-545-5p decreases the invasive ability of Saos-2 and MG63 cells.
In addition, the wound healing speed decreased in both Saos-2 and
MG63 cells following transfection with miR-545-5p mimics (both
P<0.05; Fig. 3B). Taken together,
these results suggest that miR-545-5p facilitates apoptosis and
inhibits the migratory and invasive abilities of osteosarcoma
cells.
miR-545-5p restricts tumor growth
Given that the results of the present study
demonstrated that miR-545-5p inhibited proliferation and promoted
apoptosis of osteosarcoma cells, tumor growth in vivo was
assessed following overexpression of miR-545-5p. Xenografts
generated from MG63 (P<0.05; Fig.
4A) and Saos-2 (P<0.05; Fig.
4D) cells, with ectopic miR-545-5p expression, were
significantly smaller than those transfected with mimic NC. In
addition, both the weight and volume of the xenografts generated
from MG63 (P<0.05; Fig. 4B and C)
and Saos-2 (P<0.05; Fig. 4E and
F) cells overexpressing miR-545-5p significantly decreased
compared with the mimic NC group, suggesting that miR-545-5p exerts
an antitumor effect in vivo.
DIMT1 is a downstream target of
miR-545-5p
To determine the underlying molecular mechanism by
which miR-545-5p regulates osteosarcoma progression, downstream
miR-545-5p targets were predicted using the online databases,
DIANA-microT, miRDB and TargetScan. The following parameters were
set for each database: DIANA-microT (miTG score ≥0.8); miRDB
(Target score ≥80) and TargetScan (Cumulative weighted context++
score ≤-0.5). With these parameters, 14 potential downstream target
genes were identified (Fig. 5A).
RT-qPCR analysis was performed to detect the expression levels of
all 14 genes in Saos-2 cells transfected with miR-545-5p inhibitor.
The results demonstrated that transfection with miR-545-5p
inhibitor significantly upregulated DIMT1 expression compared with
cells transfected with inhibitor NC (P<0.001; Fig. 5B).
 | Figure 5.DIMT1 is a target of miR-545-5p. (A)
DIANA-microT, miRDB and TargetScan were used to predict the target
mRNAs of miR-545-5p. The Venn diagram depicts 14 potential targets.
The regulation network of the 14 potential target mRNAs is depicted
in the top right panel. The oval nodes represent
miR-545-5p-targeted mRNAs. The darker the color, the greater the
possibility. (B) Verification of potential targets in MG63 cells
transfected with 50 nM miR-545-5p inhibitor or inhibitor NC. (C)
Left panel: Sequence alignment depicting the potential interactions
between miR-545-5p and DIMT1-WT or DIMT1-MUT 3′-UTR of DIMT1. Right
panel: Luciferase reporter activities of Saos-2 and MG63 cells
co-transfected with plasmid containing WT or MUT DIMT1 3′-UTR and
miR-545-5p mimics or mimic NC. (D) RT-qPCR analysis was performed
to detect DIMT1 exp ression in MG63 cells transfected with 50 nM
mimic NC, miR-545-5p mimic, inhibitor NC or miR-545-5p inhibitor.
(E) Western blot analysis was performed to detect DIMT1 protein
expression in MG63 cells transfected with 50 nM mimic NC,
miR-545-5p mimic, inhibitor NC or miR-545-5p inhibitor. (F)
Representative immunohistochemistry staining of DIMT1 in
osteosarcoma tissues and adjacent normal tissues. Scale bar, 50 µm
and magnification, ×200. (G) RT-qPCR analysis was performed to
detect miR-545-5p expression in osteosarcoma tissues and adjacent
normal tissues. *P<0.05 and ***P <0.001 vs. inhibitor NC,
corresponding NC group or normal adjacent tissue. DIMT1,
dimethyladenosine transferase 1; miR, microRNA; NC, negative
control; WT, wild-type; MUT, mutant; UTR, untranslated region;
RT-qPCR, reverse transcription-quantitative PCR. |
To determine the direct regulatory association
between miR-545-5p and DIMT1, luciferase reporter plasmids carrying
WT DIMT1 3′-untranslated region (UTR) sequence (DIMT1-WT) or a
DIMT1 3′-UTR sequence with mutated binding sites (DIMT1-MUT) were
co-transfected with miR-545-5p mimics or mimic NC into osteosarcoma
cells (Fig. 5C). The results
demonstrated that in both Saos-2 and MG63 cells, luciferase
activities were significantly inhibited when miR-545-5p was
co-transfected with luciferase reporter plasmids carrying DIMT1-WT
(P<0.05; Fig. 5C). Notably,
miR-545-5p had no effect on the mutated DIMT1 3′-UTR, suggesting
that miR-545-5p binds to the mRNA of DIMT1 via the complementary
sequences predicted (Fig. 5C).
Changes in DIMT1 mRNA and protein expression levels
in MG63 cells were detected following transfection with mimic NC,
miR-545-5p mimic, inhibitor NC or miR-545-5p inhibitor,
respectively. The successful transfections of miR-545-5p mimic and
miR-545-5p inhibitor were confirmed using RT-qPCR (Fig. S1A and B). Notably, transfection with
miR-545-5p mimic suppressed DIMT1 expression (P<0.05), whereas
transfection with miR-545-5p inhibitor significantly increased
DIMT1 expression in Saos-2 cells (P<0.001; Fig. 5D and E), confirming that DIMT1 is a
downstream target of miR-545-5p. DIMT1 protein and mRNA expression
levels were also detected in clinical samples via
immunohistochemistry and RT-qPCR analyses. The results demonstrated
that both DIMT1 protein and mRNA expression levels were upregulated
in clinical osteosarcoma samples (P<0.05; Fig. 5F and G).
DIMT1 facilitates metastasis of
osteosarcoma
The present study investigated the role of DIMT1 in
osteosarcoma. DIMT1 protein expression was detected in five
clinical osteosarcoma samples and the results demonstrated that
DIMT1 expression was upregulated in all samples compared with
adjacent normal tissues (Fig. 6A).
In addition, DIMT1 expression was notably upregulated in Saos-2 and
MG63 cells compared with hFOB1.19 human osteoblast cells (Fig. 6B). DIMT1 expression was knocked down
at the mRNA level via transfection with siRNA, which decreased
DIMT1 expression in both Saos-2 and MG63 cells (all P<0.05;
Fig. 6C and D). Notably, DIMT1
knockdown significantly decreased the migratory and invasive
abilities of Saos-2 and MG63 cells (all P<0.05; Fig. 6E and F). These results are similar to
those observed following overexpression of miR-545-5p, suggesting
that miR-545-5p functions by regulating DIMT1 expression.
 | Figure 6.DIMT1 promotes migration and invasion
of osteosarcoma. (A) Western blot analysis was performed to detect
DIMT1 protein expression in osteosarcoma tissues and adjacent
normal tissues (n=5). Tissues were collected from five different
patients. GAPDH was used as the loading control. (B) Western blot
analysis was performed to detect DIMT1 protein expression in the
human osteoblast cell line, hFOB1.19, and osteosarcoma cell lines,
Saos-2 and MG63. GAPDH was used as the loading control. (C) Saos-2
and MG63 cells were transfected with 50 nM si-NC or si-DIMT1 for 48
h and western blot analysis was performed to detect DIMT1 protein
expression. GAPDH was used as the loading control. (D) Reverse
transcription-quantitative PCR analysis was performed to detect
DIMT1 expression in Saos-2 and MG63 cells transfected with si-NC or
si-DIMT1 for 48 h. (E) The Transwell assay was performed to detect
the invasive ability of Saos-2 and MG63 cells transfected with 50
nM si-NC or si-DIMT1 for 24 h. Quantification analysis is depicted
in the middle right panel. Scale bar, 50 µm and magnification,
×100. (F) Wound healing analysis of Saos-2 and MG63 cells
transfected with si-NC or si-DIMT1. Scale bar, 50 µm. *P<0.05
vs. si-NC or control. DIMT1, dimethyladenosine transferase 1; si,
small interfering; NC, negative control; C, osteosarcoma tissues;
N, adjacent normal tissues. |
miR-545-5p suppresses metastasis of
osteosarcoma by targeting DIMT1
To further validate the function of miR-545-5p via
DIMT1, miR-545-5p and DIMT1 were co-overexpressed in Saos-2 and
MG63 cells (P<0.01; Fig. S1C).
The results demonstrated that overexpression of DIMT1 attenuated
DIMT1 reduction induced by simultaneous overexpression of
miR-545-5p (Fig. 7A). Notably,
overexpression of DIMT1 in both Saos-2 and MG63 cells transfected
with mi-545-5p mimics restored cell viability to similar levels as
the mimic NC group (all P<0.05; Fig.
7B). Furthermore, the migratory ability of both Saos-2 and MG63
cells was also rescued following overexpression of DIMT1 in cells
transfected with miR-545-5p mimics (all P<0.05; Fig. 7C).
 | Figure 7.miR-545-5p functions by targeting
DIMT1. (A) Saos-2 and MG63 cells were transfected with 50 nM
miR-545-5p mimic or mimic NC, with or without co-transfection of 10
µg DIMT1 overexpression plasmid for 48 h and western blot analysis
was performed to detect DIMT1 protein expression. GAPDH was used as
the loading control. (B) Saos-2 and MG63 cells were transfected
with 50 nM miR-545-5p mimic or mimic NC, with or without
co-transfection of 10 µg DIMT1 overexpression plasmid for 48 h and
the Cell Counting Kit-8 assay was performed to detect cell
viability. (C) Saos-2 and MG63 cells were transfected with 50 nM
miR-545-5p mimic or mimic NC, with or without co-transfection of 10
µg DIMT1 overexpression plasmid for 24 h and the Transwell assay
was performed to detect cell invasion. Quantification analysis is
depicted in the lower panel. Scale bar, 50 µm and magnification,
×100. (D) Flow cytometric analysis of Saos-2 and MG63 cells
transfected with 50 nM miR-545-5p mimic or mimic NC, with or
without co-transfection of 10 µg DIMT1 overexpression plasmid for
48 h. Cells were harvested and labeled with Annexin V-FITC and PI.
The numbers in each quadrant indicate positive percentages of the
entire population. (E) Saos-2 and MG63 cells were transfected with
50 µm miR-545-5p mimic or mimic NC, with or without co-transfection
of 10 µg DIMT1 overexpression plasmid and the wound healing assay
was performed to detect cell migration. Quantification analysis is
depicted in the right panel. Scale bar, 50 µm. *P<0.05 vs. mimic
NC. #P<0.05 vs. miR-545-5p mimic. miR, microRNA;
DIMT1, dimethyladenosine transferase 1; NC, negative control; PI,
propidium iodide. |
To evaluate the effect of overexpressing DIMT1 on
the apoptosis of osteosarcoma cells, flow cytometric analysis of
cells labeled with Annexin V and PI was performed. The results
demonstrated that overexpression of DIMT1 in Saos-2 and MG63 cells
transfected with miR-545-5p mimics significantly decreased the
percentage of apoptotic cells compared with the control group
(P<0.05; Fig. 7D). In addition,
the migratory ability of Saos-2 and MG63 cells following
co-overexpression with miR-545-5p and DIMT1 was assessed. The
results demonstrated that overexpression of DIMT1 partially rescued
the reduced migratory ability of Saos-2 and MG63 cells following
overexpression of miR-545-5p (Fig.
7E), which suggests that miR-545-5p functions by inhibiting
DIMT1 expression. Taken together, the results of the present study
suggest that miR-545-5p functions as a tumor suppressor by
targeting DIMT1.
The results of the present study demonstrated that
miR-545-5p expression was downregulated in clinical osteosarcoma
patient samples and cell lines. In addition, overexpression of
miR-545-5p increased apoptosis, and inhibited migration and
invasion of osteosarcoma cells by targeting DIMT1. Furthermore,
overexpression of miR-545-5p successfully inhibited in vivo
xenograft growth. Collectively, these results suggest that
miR-545-5p is a novel miRNA that functions as a tumor suppressor in
osteosarcoma, thus providing novel therapeutic targets for
osteosarcoma.
Discussion
miRNA expression can be altered during the
development of certain diseases, and thus may act as a critical
biomarker for disease (28–30). The identification of novel miRNAs
with altered expression levels in osteosarcoma may assist current
therapeutic strategies since miRNAs are promising targets for drug
development (31). The results of
the present study demonstrated that miR-545-5p expression was
downregulated in osteosarcoma. However, the role of miR-545-5p in
cancer development remains unknown. To the best of our knowledge,
the present study is the first to provide experimental evidence
supporting the function of miR-545-5p in osteosarcoma development.
However, to further clarify the role of this potential tumor
suppressor, transgenic mice are required. miR-545-5p knockout mice
will be used to further determine the anticancer role of this miRNA
in an in vivo model. The results of the present study
demonstrated that miR-545-5p has antiproliferative and proapoptotic
roles in osteosarcoma. Notably, overexpression of miR-545-5p
decreased the migratory and invasive abilities of osteosarcoma
cells.
miR-545-5p target several downstream targets in
multiple signaling pathways that might serve critical roles in
tumorigenesis and development, such as murine double minute 2,
polo-like kinase 1 and δ-anubikevulinic acid dehydratase,
suggesting that miR-545-5p functions as a major switch for a series
of cellular processes (21,32,33). In
the present study, activating miR-545-5p alone inhibited cancer
growth by regulating the expression of DIMT1. The in vivo
xenograft experiments clearly demonstrated that overexpression of
miR-545-5p significantly suppressed xenograft growth. However, not
all the observed phenotypes can be explained by alterations in
DIMT1 levels due to modified miR-545-5p expression in osteosarcoma.
Prospective studies should include administering miR-545-5p to mice
with osteosarcoma to determine whether it induces tumor inhibition
and prevents a localized osteosarcoma from metastasizing.
A previous study suggested that miR-545-5p
expression was upregulated in colorectal cancer SW620 cells (a cell
line derived from a metastatic site) compared with SW480 cells (a
cell line derived from a local site in the same patient from which
the SW620 line was derived) (14).
However, the present study did not provide functional data to
support the hypothesis that miR-545-5p participates in the
transformation of a localized colon cancer cell to a metastatic
colon cancer cell. The conclusion that miR-545-5p inhibits
metastasis is supported by both the migration and invasion assays,
as overexpression of miR-545-5p inhibited cancer cell migration.
Notably, the results of the present study demonstrated that DIMT1,
a downstream target of miR-545-5p, facilitated the migration and
invasion of osteosarcoma cells. Thus, miR-545-5p is an inhibitor of
osteosarcoma metastasis. However, the role of miR-545-5p in
cancers, particularly in colon cancer, requires further
investigation.
The results of the present study revealed that
miR-545-5p regulates DIMT1 expression by binding to the 3′-UTR of
DIMT1 mRNA. DIMT1, also known as DIM1, was initially identified as
a homolog of 18S rRNA dimethylase in Saccharomyces
cerevisiae (gene ID, 27292). DIMT1 functions as a downstream
target of miR-210, which regulates gastric epithelial cell
proliferation (34). In addition,
DIMT1 has been demonstrated to be elevated in gastric cancer cells,
and it is considered a predictor of tumor progression and prognosis
in patients with gastric carcinoma (35). Furthermore, miR-210 regulates the
DIMT1-interferon regulatory factor 4 oncogenic axis in multiple
myeloma under hypoxic stress (36).
Although an oncogenic role for DIMT1 has been suggested, whether
DIMT1 functions as an oncogene in osteosarcoma remains unclear.
DIMT1 is regulated by multiple miRNAs (miR-210 and
miR-101) (36,37), and three different miRNAs target
DIMT1 simultaneously. Although the expression levels of miR-210 and
miR-101 in osteosarcoma cells remain unclear, it is possible that
the alteration of different miRNA expression levels creates
different phenotypes, rather than alterations in the expression
levels of DIMT1. However, to understand the regulatory network
controlling DIMT1, further expression analyses are required. The
results of the present study suggest that miR-545-5p controls
apoptosis, migration and invasion of osteosarcoma cells via DIMT1.
However, alterations in all pathways involved were not investigated
in the present study. It is important to perform high throughput
RNA sequencing for cells with or without high miR-545-5p levels to
determine the underlying molecular mechanisms of how miR-545-5p
regulates osteosarcoma development.
In conclusion, the results of the present study
suggest two potential targets for inhibiting osteosarcoma
metastasis, miR-545-5p and DIMT1. One can utilize miR-545-5p mimics
in situ or through the circulation to induce an anticancer
effect, while the other can antagonize DIMT1 via small molecule
cancer drugs, although the development of such drugs requires
further investigation. Taken together, these results provide novel
therapeutic targets for osteosarcoma and molecular insights into
their regulatory network.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
This study was supported by the Science and
Technology Planning Project of Xi'an [grant no.
2019115013YX005SF038(13)].
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author upon reasonable
request.
Authors' contributions
HZZ, BC and ZCT conceived the present study and
designed the experiments. JJD, NZ and YXS performed the
experiments. KZ and XJL contributed to data analysis. HZZ, BC and
ZCT drafted the initial manuscript. All authors have read and
approved the final manuscript. ZCT and KZ confirm the authenticity
of all the raw data.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Honghui Hospital of Xi'an Jiaotong University (Xi'an,
China) and written informed consent was provided by all patients
prior to the study start. All animal experiments were approved by
the Institutional Animal Care and Use Committee of Honghui Hospital
of Xi'an Jiaotong University (Xi'an, China). Study approval no.
202003059.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Gianferante DM, Mirabello L and Savage SA:
Germline and somatic genetics of osteosarcoma-connecting aetiology,
biology and therapy. Nat Rev Endocrinol. 13:480–491. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Colding-Rasmussen T, Thorn AP, Horstmann
P, Rechnitzer C, Hjalgrim LL, Krarup-Hansen A and Petersen MM:
Survival and prognostic factors at time of diagnosis in high-grade
appendicular osteosarcoma. Acta Oncol. 57:420–425. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lindsey BA, Markel JE and Kleinerman ES:
Osteosarcoma overview. Rheumatol Ther. 4:25–43. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kansara M, Teng MW, Smyth MJ and Thomas
DM: Translational biology of osteosarcoma. Nat Rev Cancer.
14:722–735. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
5
|
Vos HI, Coenen MJ, Guchelaar HJ and Te Loo
DM: The role of pharmacogenetics in the treatment of osteosarcoma.
Drug Discov Today. 21:1775–1786. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Xie L, Yao Z, Zhang Y, Li D, Hu F, Liao Y,
Zhou L, Zhou Y, Huang Z, He Z, et al: Deep RNA sequencing reveals
the dynamic regulation of miRNA, lncRNAs, and mRNAs in osteosarcoma
tumorigenesis and pulmonary metastasis. Cell Death Dis. 9:7722018.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ma J, Huang K, Ma Y, Zhou M and Fan S: The
TAZ-miR-224-SMAD4 axis promotes tumorigenesis in osteosarcoma. Cell
Death Dis. 8:e25392017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ji Q, Xu X, Li L, Goodman SB, Bi W, Xu M,
Xu Y, Fan Z, Maloney WJ, Ye Q and Wang Y: miR-216a inhibits
osteosarcoma cell proliferation, invasion and metastasis by
targeting CDK14. Cell Death Dis. 8:e31032017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Raza U, Zhang JD and Sahin O: MicroRNAs:
Master regulators of drug resistance, stemness, and metastasis. J
Mol Med (Berl). 92:321–336. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sampson VB, Yoo S, Kumar A, Vetter NS and
Kolb EA: MicroRNAs and potential targets in osteosarcoma: Review.
Front Pediatr. 3:692015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang C, Jing J and Cheng L: Emerging roles
of non-coding RNAs in the pathogenesis, diagnosis and prognosis of
osteosarcoma. Invest New Drugs. 36:1116–1132. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Su P, Mu S and Wang Z: Long noncoding RNA
SNHG16 promotes osteosarcoma cells migration and invasion via
sponging miR-340. DNA Cell Biol. 38:170–175. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Pu Y, Zhao F, Cai W, Meng X, Li Y and Cai
S: miR-193a-3p and miR-193a-5p suppress the metastasis of human
osteosarcoma cells by downregulating Rab27B and SRR, respectively.
Clin Exp Metastasis. 33:359–372. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yan W, Yang W, Liu Z and Wu G:
Characterization of microRNA expression in primary human colon
adenocarcinoma cells (SW480) and their lymph node metastatic
derivatives (SW620). OncoTargets Ther. 11:4701–4709. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Li C, Wang M, Wang Y, Zhang J and Sun N: A
new model of the mechanism underlying lead poisoning: SNPs in miRNA
target region influence the delta-aminolevulinic acid dehydratase
expression level. Epigenomics. 9:1353–1361. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bergdahl IA, Grubb A, Schutz A, Desnick
RJ, Wetmur JG, Sassa S and Skerfving S: Lead binding to
delta-aminolevulinic acid dehydratase (ALAD) in human erythrocytes.
Pharmacol Toxicol. 81:153–158. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang C, Han XS, Li FF, Huang S, Qin YW,
Zhao XX and Jing Q: Forkhead containing transcription factor Albino
controls tetrapyrrole-based body pigmentation in planarian. Cell
Discov. 2:160292016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Liu Z, Dou C, Yao B, Xu M, Ding L, Wang Y,
Jia Y, Li Q, Zhang H, Tu K, et al: Ftx non coding RNA-derived
miR-545 promotes cell proliferation by targeting RIG-I in
hepatocellular carcinoma. Oncotarget. 7:25350–25365. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Hu C, Wang Y, Li A, Zhang J, Xue F and Zhu
L: Overexpressed circ_0067934 acts as an oncogene to facilitate
cervical cancer progression via the miR-545/EIF3C axis. J Cell
Physiol. 234:9225–9232. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yuan G, Wu H, Du Y and He F: Tumor
suppressor role of microRNA-545 in oral squamous cell carcinoma.
Oncol Lett. 17:2063–2068. 2019.PubMed/NCBI
|
|
21
|
Miao Z, Liu S, Xiao X and Li D: LINC00342
regulates cell proliferation, apoptosis, migration and invasion in
colon adenocarcinoma via miR-545-5p/MDM2 axis. Gene.
743:1446042020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Lu YF, Yu JR, Yang Z, Zhu GX, Gao P, Wang
H, Chen SY, Zhang J, Liu MY, Niu Y, et al: Promoter hypomethylation
mediated upregulation of MicroRNA-10b-3p targets FOXO3 to promote
the progression of esophageal squamous cell carcinoma (ESCC). J Exp
Clin Cancer Res. 37:3012018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Nunes JPS and Dias AAM: ImageJ macros for
the user-friendly analysis of soft-agar and wound-healing assays.
Biotechniques. 62:175–179. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Marshall J: Transwell (®)
invasion assays. Methods Mol Biol. 769:97–110. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chen X, Cao X, Sun X, Lei R, Chen P, Zhao
Y, Jiang Y, Yin J, Chen R, Ye D, et al: Bcl-3 regulates TGFbeta
signaling by stabilizing Smad3 during breast cancer pulmonary
metastasis. Cell Death Dis. 7:e25082016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
National Research Council, . National
Institutes of Health Guide for the Care and Use of Laboratory
Animals: 8th Edition. The National Academies Press; Washington, DC:
2011
|
|
28
|
Laferriere CA and Pang DS: Review of
intraperitoneal injection of sodium pentobarbital as a method of
euthanasia in laboratory rodents. J Am Assoc Lab Anim Sci.
59:254–263. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yu AM, Jian C, Yu AH and Tu MJ: RNA
therapy: Are we using the right molecules? Pharmacol Ther.
196:91–104. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Nugent M: microRNA and bone cancer. Adv
Exp Med Biol. 889:201–230. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Xie M, Ma L, Xu T, Pan Y, Wang Q, Wei Y
and Shu Y: Potential regulatory roles of MicroRNAs and long
noncoding RNAs in anticancer therapies. Mol Ther Nucleic Acids.
13:233–243. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang H, Zhang K, Xu Z, Xu Z, Chen Z, Wang
Q, Wang C and Cui J: MicroRNA-545 suppresses progression of ovarian
cancer through mediating PLK1 expression by a direct binding and an
direct regulation involving KDM4B-mediated demethylation. BMC
Cancer. 21:1632021. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ikeda S, Kitadate A, Abe F, Saitoh H,
Michishita Y, Hatano Y, Kawabata Y, Kitabayashi A, Teshima K, Kume
M, et al: Hypoxia-inducible microRNA-210 regulates the DIMT1-IRF4
oncogenic axis in multiple myeloma. Cancer Sci. 108:641–652. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Liu G, Peng X, Cai Y, Cheng A, Zha L and
Wang Z: DIMT1 overexpression correlates with progression and
prognosis in gastric carcinoma. Hum Pathol. 70:35–42. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Kiga K, Mimuro H, Suzuki M,
Shinozaki-Ushiku A, Kobayashi T, Sanada T, Kim M, Ogawa M, Iwasaki
YW, Kayo H, et al: Epigenetic silencing of miR-210 increases the
proliferation of gastric epithelium during chronic Helicobacter
pylori infection. Nat Commun. 5:44972014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Guo ZX, Zhou FZ, Song W, Yu LL, Yan WJ,
Yin LH, Sang H and Zhang HY: Suppression of microRNA-101 attenuates
hypoxia-induced myocardial H9c2 cell injury by targeting
DIMT1-Sp1/survivin pathway. Eur Rev Med Pharmacol Sci.
22:6965–6976. 2018.PubMed/NCBI
|