Introduction
Colorectal cancer (CRC) is one of the most prevalent
cancers in the world (1). It is
the second leading cause of cancer-related mortality worldwide. In
Japan, the incidence of CRC has doubled over the past 20 years such
that CRC is now the second most deadly neoplastic disease (2,3).
Surgery is still the most effective treatment for
CRC. Among the patients that undergo curative surgery, some develop
local recurrence or distant metastases that lead to shorter
survival times (4). Distant
metastasis has a critical influence on the prognosis of CRC.
Clinicopathological indicators such as the TNM classification
proposed by the International Union Against Cancer (UICC) remain
the indicator of prognosis and provide the basis for therapeutic
decision making. However, the current TNM classification system is
limited in that it cannot predict prognosis for individual patients
(5). In order to develop
personalized therapeutic regimens, it is therefore critical that
novel genes involved in distant metastasis are identified that can
serve as prognostic biomarkers (6). Microarray is a particularly powerful
tool for identifying potential biomarker genes for use in cancer
prognosis (7–9). Using microarray analysis it is now
possible to investigate several thousand cancer-related or
cancer-specific genes at once.
Chromosomal structural alterations play an important
role in cancer development. In CRC, copy number aberrations (CNAs),
including gains on chromosomes 7, 8, 13 and 20, and losses on
chromosomes 1p, 8p, 17p and 18, are frequently observed (10–13).
Some of these CNAs are related to metastasis of CRC and can thus be
used in prognosis. The single nucleotide polymorphism microarray
(SNP array) analysis has become a useful tool for examining CNAs,
permitting highly accurate exploration of thousands of genetic
markers in a single study (14).
Studies of the relationship between chromosomal
aberrations and gene expression in cancers, including CRC, have
shown that CNAs directly influence gene expression (15–19).
Several groups have therefore suggested that integrating gene
expression analysis with genomic profiling represents an efficient
approach for the discovery of cancer-related genes (20–22).
Genes that show a strong positive correlation between expression
and copy number may play an important role in cancer progression.
In this study, we therefore integrated gene expression and copy
number analyses to identify novel genes associated with the distant
metastasis of CRC. We focused on the genes that are overexpressed
and have an amplified copy number in cases of distant metastasis
because such characteristics indicate that these genes have the
potential to serve as useful therapeutic targets or clinical
biomarkers.
Using the aforementioned comprehensive analysis, we
identified S100 calcium-binding protein A2 (S100A2) as a gene
involved in the distant metastasis of stage II and III CRC. It has
been suggested that S100A2 plays an important role in cell cycle
progression, and overexpression of S100A2 has been reported in
several cancers (23–28). This is the first study to
demonstrate the prognostic significance of S100A2 expression in
CRC, using integrated copy number and gene expression analyses of
clinical tissue samples.
Patients and methods
Patients
Primary tumors from 278 patients who underwent
curative surgery for stage II and III CRC between 2002 and 2009 at
the Tokyo Medical and Dental University Hospital (Tokyo, Japan)
were studied. Written informed consent was obtained from all the
patients, and the study was approved by the ethics committee of
Tokyo Medical and Dental University, and all the following
procedures were performed strictly in accordance with the ethical
standards established by this committee. Clinical data were
obtained from the medical records of each patient, and
histopathological evaluations were assessed by reference to the
criteria of the TNM-system of the UICC, 7th edition. A total of 125
patients were assigned to the comprehensive analyses for extraction
of candidate genes. All of the patients were assigned to the gene
expression and the CNA study, including 66 patients with stage II
and 59 patients with stage III disease. The median follow-up time
for these patients was 62 months (range, 1–76 months). Quantitative
reverse transcription polymerase chain reaction (RT-PCR) assays
were performed for validation using samples from 50 stage II and
III CRC patients, including 24 patients with stage II and 26
patients with stage III disease. The median follow-up time for
these patients was 61 months (range, 7–96 months). Furthermore, 161
patients, including patients subjected to RT-PCR validation, were
analyzed using immunohistochemistry (IHC). The IHC study included
80 patients with stage II and 81 patients with stage III disease.
The median follow-up time for these patients was 86 months (range,
1–96 months). The patients enrolled in the comprehensive analyses
were excluded from these validation studies.
DNA extraction
After resection, cancer tissues were immediately
embedded in Tissue-Tek OCT compound medium (Sakura Finetek Japan,
Tokyo, Japan). Serial frozen sections of 9-mm in thickness were
mounted onto a 90 FOIL-SL25 foil-coated glass slide (Leica
Microsystems, Wetzlar, Germany). Laser capture microdissection
(LCM) was performed using an Application Solutions LCM System
(Leica Microsystems). Tumor DNA was extracted and purified using a
QIAamp DNA micro kit (Qiagen, Hilden, Germany) according to the
manufacturer's instructions. Non-neoplastic tissues were
homogenized in microtubes, and DNA was extracted and purified from
these tissues using a QIAamp DNA mini kit (Qiagen) according to the
manufacturer's instructions.
CNA analysis
Copy number analysis was performed using a
GeneChip® Human Mapping 250K Sty array (Affymetrix,
Santa Clara, CA, USA) in strict adherence to the assay manual.
Genomic DNA was digested using the enzyme StyI, and a
Sty1 adaptor was used prior to the PCR reaction. Amplicons
were fragmented after purification and then labeled. After
hybridization, the microarrays were transferred to a totally
automated GeneChip® Fluidics Station 450 (Affymetrix)
for the washing and staining steps. After fluorescence staining,
microarray images were scanned using a GeneChip® Scanner
3000 7G (Affymetrix). The microarray data from the scanner were
used for copy number analysis with the Chromosome Copy Number
Analysis Tool (Affymetrix). Data were analyzed using R statistical
software (version 2.12.1; http://www.r-project.org/).
RNA extraction
Cancer cells were microdissected using LCM. Total
RNA was extracted from cancer cells and purified using an RNeasy
micro kit (Qiagen) with on-column DNase digestion, according to the
manufacturer's instructions. Total RNA collected from bulk samples
of cancer tissues and adjacent non-neoplastic tissues was extracted
and purified using an RNeasy mini kit (Qiagen) with on-column DNase
digestion, according to the manufacturer's instructions. The
integrity of the total RNA was assessed using an Agilent 2100
BioAnalyzer (Agilent Technologies, Palo Alto, CA, USA). Samples
with an RNA integrity number >5.0 were used for the rest of the
experiments.
Gene expression analysis
Complementary RNA was prepared from total RNA using
two-cycle target labeling and a control reagents kit (Affymetrix).
The experiment was performed using the GeneChip® Human
Genome U133 Plus 2.0 Array (Affymetrix), according to the
manufacturer's instructions. Statistical analyses of microarray
data were normalized using the robust multi-array average method
with R statistical software (version 2.12.1; http://www.r-project.org/) together with the
BioConductor package (http://www.bioconductor.org/).
Extraction of candidate genes
We defined patients with metastatic recurrence of
stage II and III CRC as the recurrence group, and patients without
any recurrence as the non-recurrence group. Local recurrence and
lymph node recurrence were excluded from the recurrence group. The
CNA data and the gene expression data were analyzed and compared
between the 2 groups to identify genes involved in the metastatic
recurrence of stage II and III CRC.
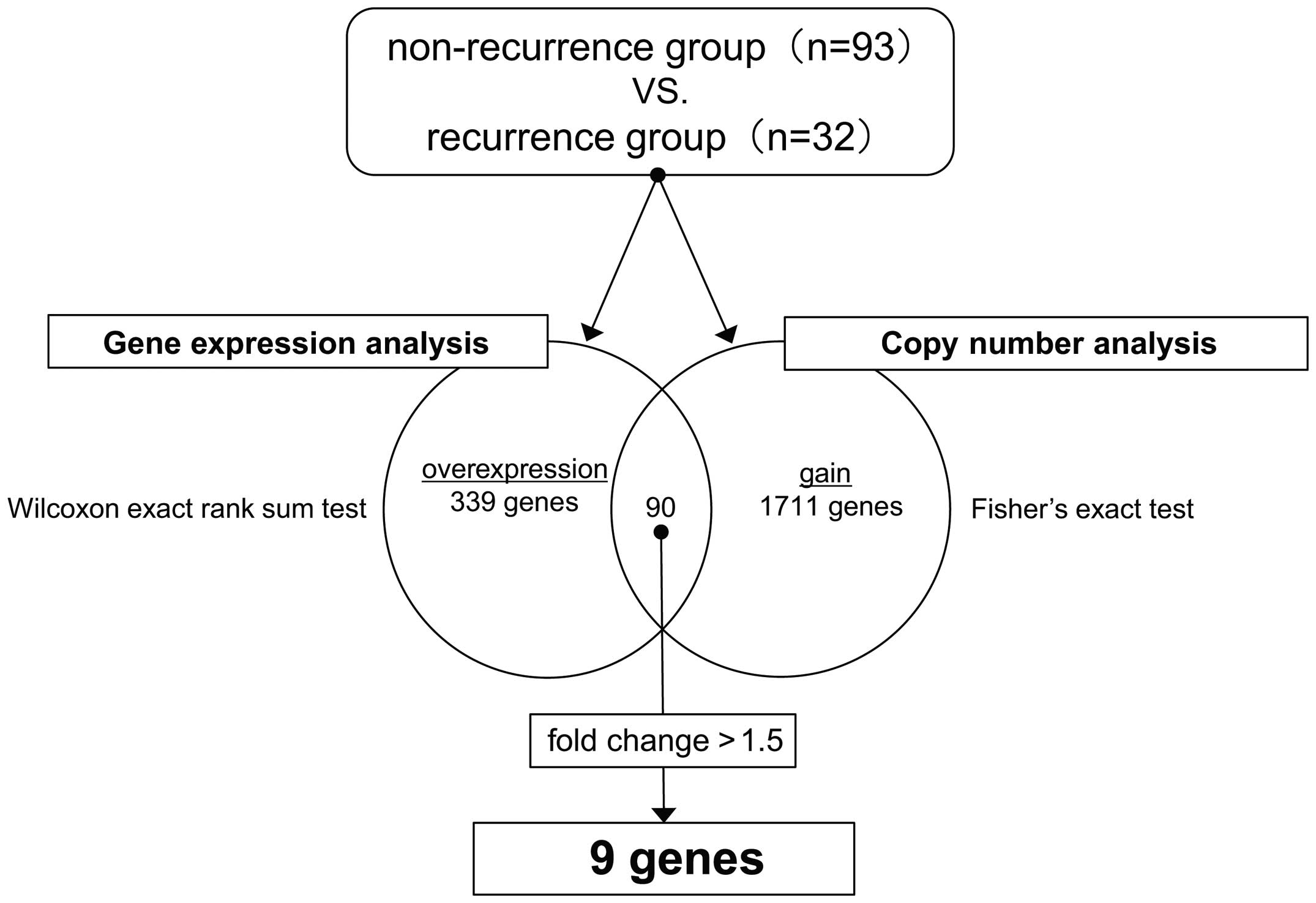
Data regarding genes which showed a copy number gain
in the recurrence group were extracted using Fisher's exact test
(P<0.01). Data regarding genes that were significantly
upregulated in the recurrence group were extracted using the
Wilcoxon exact rank sum test (P<0.01). Among the genes that were
common to both groups, those that were overexpressed (fold change
>1.5) in the recurrence group were selected as candidates for
further analysis.
The gene expression data and DNA copy number data
were deposited in the Gene Expression Omnibus (GEO) under accession
IDs GSE64256, GSE64257 and GSE64258.
Quantitative RT-PCR
Total RNA collected from bulk samples of cancer
tissues and adjacent non-neoplastic tissues was reverse-transcribed
into cDNA using a High Capacity cDNA Reverse Transcription kit
(Applied Biosytems, Foster City, CA, USA) according to the
manufacturer's instructions. A TaqMan® gene expression
assay (Applied Biosystems; S100A2; Hs00195582_m1, β-actin;
Hs99999903_m1) was used to investigate the expression of S100A2,
and β-actin was used as an internal control. The PCR reaction was
carried out using TaqMan® Universal PCR Master Mix
(Applied Biosystems). The thermal cycling conditions were as
follows: 50ºC for 2 min, 95ºC for 10 min, and 40 cycles of
denaturation at 95ºC for 15 sec and annealing at 60ºC for 1 min.
All calculated concentrations of target genes were normalized by
the amount of the endogenous reference using the comparative Ct
method for relative quantification with Relative Quantification
Study Software (7300 Sequence Detection System version 1.2.1,
Applied Biosystems).
Immunohistochemistry
IHC analyses of S100A2 were conducted on
formalin-fixed paraffin-embedded tissue blocks from each patient.
For S100A2 staining, antigen retrieval by autoclave treatment was
carried out for 15 min in 1X TE (1X Tris-EDTA, pH 8.0) at 121ºC
after deparaffinization in xylene and rehydration through a series
of incubations in graded concentrations of ethanol. The slides were
then incubated in a solution of 3% hydrogen peroxide in 100%
methanol for 15 min at room temperature in order to quench
endogenous peroxidase activity. Subsequently, the slides were
incubated with mouse monoclonal antibody against S100A2
(Sigma-Aldrich), at a 1:50 dilution, for 30 min at room
temperature. The slides were then incubated with peroxidase-labeled
antibody [Histofine Simple Stain Max PO (MULTI; Nichirei
Bioscience)] for 30 min at room temperature. Peroxidase activity
was detected with DAB Solution (Histofine Simple Stain DAB
Solution; Nichirei BioScience). Finally, the slides were
counterstained with 1% Mayer's hematoxylin.
All the sections were divided into four stages
(negative, weak, moderate, or strong) by staining intensity.
Expression was graded by two independent observers who were blinded
to the patient information.
Statistical analysis of S100A2
expression
Statistical analyses of S100A2 expression were
carried out using SPSS (version 17.0, SPSS Inc, Chicago, IL, USA)
software for Windows. To estimate the significance of differences
between groups, Wilcoxon signed-rank, Mann-Whitney U, and
C2 tests were used where appropriate. Survival curves
were estimated using the Kaplan-Meier method, and curves were
compared using the log-rank test. Survival times were determined
from the date of surgery. Prognostic factors were examined with
univariate and multivariate analyses using the Cox proportional
hazards model. A P-value <0.05 was considered statistically
significant.
Results
Gene expression and copy number
analyses
In the copy number and gene expression analyses, 9
genes (S100A2, PROX1, TCN1, PROM1, CHRM3, ZNF678, CREB5, PPARGC1A,
and ATF6) were identified that fulfilled the specified criteria. Of
the 9 genes with both elevated copy number and expression, the
expression was upregulated with a fold change >1.5 (Fig. 1). Only S100A2 and TCN1 of these
genes have been shown to be associated with cancer. Overexpression
of S100A2 has been reported to occur in several cancers, although
it has not been reported in relation to prognosis in CRC (23–29).
We therefore focused on S100A2 in the subsequent analyses.
S100A2 mRNA and protein expression
Quantitative RT-PCR analysis of stage II and III CRC
tissue from 50 patients showed that expression of S100A2 mRNA is
significantly higher in cancerous tissue than in neighboring
non-neoplastic tissue (data not shown). The cellular localization
of the S100A2 protein was investigated using cancer tissue from the
same 50 CRC patients that were analyzed using RT-PCR. IHC indicated
that the S100A2 protein is localized in the cytoplasm of CRC cells,
whereas staining for the S100A2 protein in normal epithelial cells
adjacent to the cancer cells was negative or weak. There was no
difference in staining intensity of the invasive tumor front and
the marginal tissue of the tumor. As a result of these data, we
then estimated the extent and intensity of S100A2 staining in
sections containing the area of infiltration in stage II and III
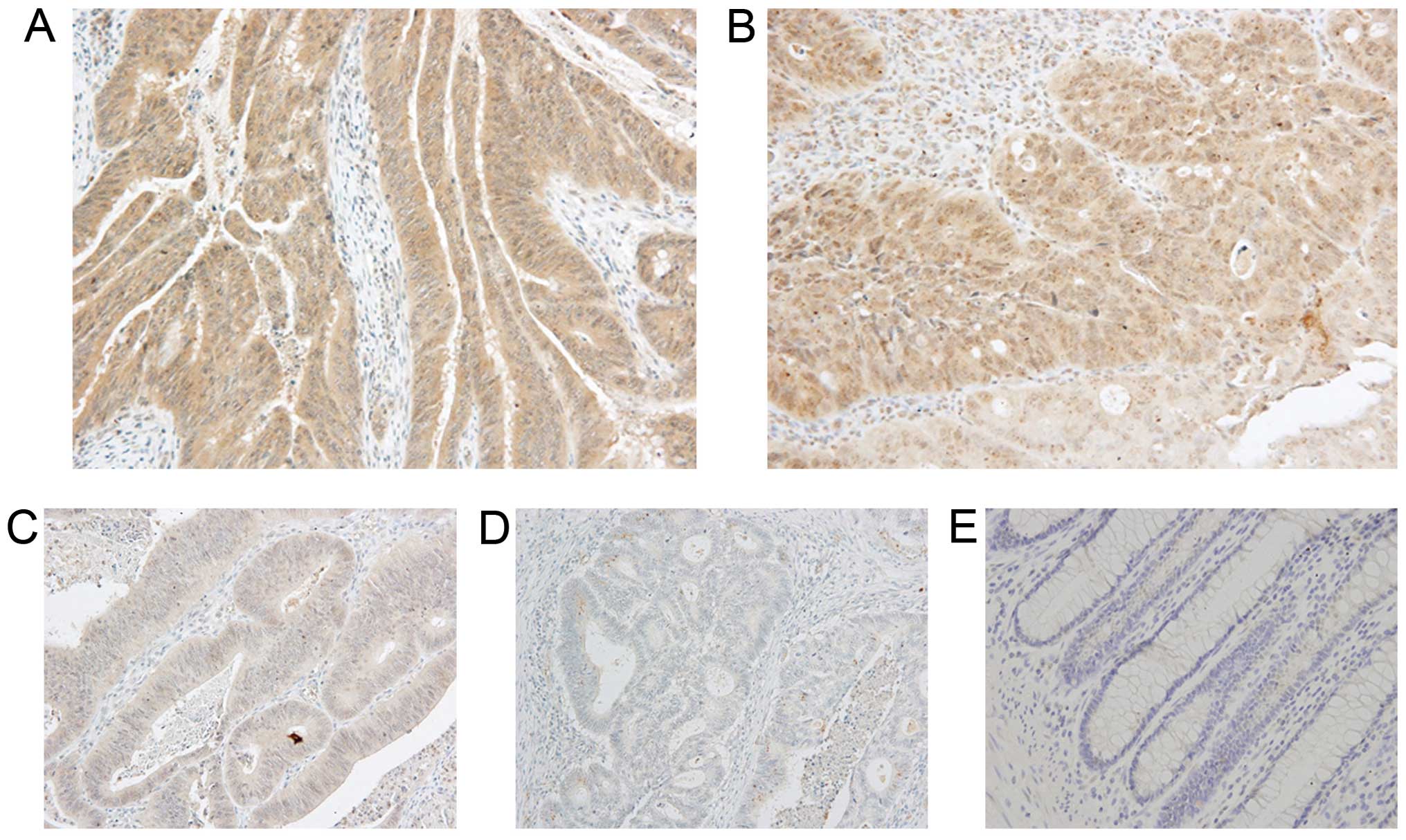
CRC tissue samples from 161 patients. As shown in Fig. 2, in the cytoplasm of cancer cells,
staining was observed in three patterns, strong (Fig. 2A), moderate (Fig. 2B) and weak (Fig. 2C).
Relationship between the expression of
S100A2 and patient characteristics
For statistical evaluation purposes, the 161 samples
that underwent IHC were divided into 2 groups: a high-expression
group (strong staining spread of >10% or moderate staining
spread of >70%, n=91) and a low-expression group (the others,
n=70).
The correlation between S100A2 expression and
various clinicopathological factors in 161 patients with stage II
or III CRC is shown in Table I.
Location (rectum; P=0.001), venous invasion (positive; P=0.011),
and the presence or absence of recurrence (recurrence group;
P<0.001) were significantly associated with overexpression of
S100A2.
 | Table IRelationship between clinicopathologic
variables and S100A2 expression in patients with stage II–III
colorectal cancer. |
Table I
Relationship between clinicopathologic
variables and S100A2 expression in patients with stage II–III
colorectal cancer.
| S100A2
expression | |
|---|
|
| |
|---|
| Variables | Low (n=70) | High (n=91) | P-value |
|---|
| Age (median),
years | 31–92 (67) | 20–86 (68) | 0.63 |
| Gender | | | 0.555 |
| Male | 43 | 60 | |
| Female | 27 | 31 | |
| Histology | | | 0.301 |
| Well | 27 | 28 | |
| Moderate, poor and
others | 43 | 63 | |
| Location | | | 0.001 |
| Colon | 53 | 46 | |
| Rectum | 17 | 45 | |
| Depth | | | 0.933 |
| T1/T2/T3 | 42 | 54 | |
| T4 | 28 | 37 | |
| Lymphatic
invasion | | | 0.139 |
| Negative | 20 | 17 | |
| Positive | 50 | 74 | |
| Venous
invasion | | | 0.011 |
| Negative | 10 | 3 | |
| Positive | 60 | 88 | |
| Lymph node
metastasis | | | 0.306 |
| Negative | 38 | 42 | |
| Positive | 32 | 49 | |
| CEA (ng/ml) | | | 0.201 |
| <5 | 46 | 50 | |
| ≥5 | 20 | 38 | |
| Recurrence | | | <0.001 |
| Non-rec | 61 | 56 | |
| Rec | 9 | 35 | |
S100A2 expression and prognosis of stage
II and III patients
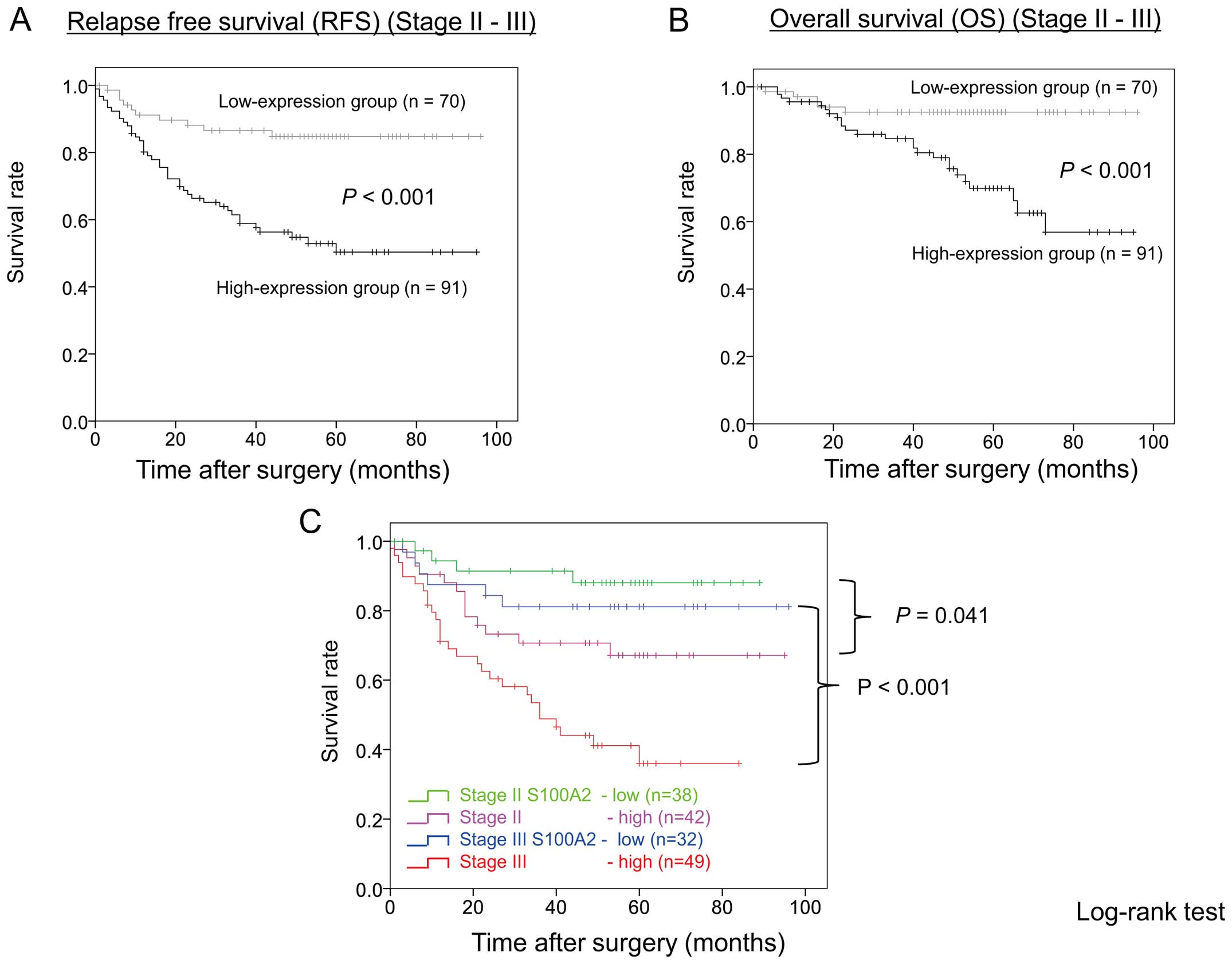
The relapse-free survival (RFS) rate was
significantly lower (P<0.001) in the S100A2 high-expression
group than in the low-expression group (Fig. 3A). Univariate analysis indicated
that gender (P=0.024), histology (P=0.022), location (P=0.003),
lymphatic invasion (P=0.024), lymph node metastasis (P=0.007), CEA
level (5.0 ng/ml or higher; P=0.003), and S100A2 expression
(P<0.001) were significantly associated with RFS. Multivariate
analysis indicated that overexpression of S100A2 is a significant
prognostic factor of RFS for Stage II or III CRC patients (P=0.007;
relative risk (RR) = 2.726; 95% confidence interval, 1318–5.638)
(Table II).
 | Table IIUnivariate and multivariate analysis
of clinicopathologic factors affecting relapse-free survival in
patients with stage II–III colorectal cancer. |
Table II
Univariate and multivariate analysis
of clinicopathologic factors affecting relapse-free survival in
patients with stage II–III colorectal cancer.
| | | Multivariate
analysis |
|---|
| | |
|
|---|
| Variables | No. of
patients | Univariate analysis
P-value | Relative risk (95%
confidence interval) | P-value |
|---|
| Age | | 0.705 | | |
| ≤65 | 81 | | | |
| >65 | 80 | | | |
| Gender | | 0.024 | | |
| Male | 103 | | | |
| Female | 58 | | | |
| Histology | | 0.022 | | 0.149 |
| Well | 55 | | | |
| Moderate, poor and
others | 106 | | | |
| Location | | 0.003 | | 0.078 |
| Colon | 99 | | | |
| Rectum | 62 | | | |
| Depth | | 0.091 | | |
| T1/T2/T3 | 96 | | | |
| T4 | 65 | | | |
| Lymphatic
invasion | | 0.024 | | 0.332 |
| Negative | 37 | | | |
| Positive | 124 | | | |
| Venous
invasion | | 0.446 | | |
| Negative | 13 | | | |
| Positive | 148 | | | |
| Lymph node
metastasis | | 0.007 | | 0.195 |
| Negative | 80 | | | |
| Positive | 81 | | | |
| CEA (ng/ml) | | 0.003 | | 0.335 |
| <5 | 96 | | | |
| ≥5 | 58 | | | |
| S100A2
expression | | <0.001 | 2.726 | 0.007 |
| Low | 70 | | (1.318–5.683) | |
| High | 91 | | | |
Likewise, the overall survival (OS) rate was
significantly lower (P<0.001) in the S100A2 high-expression
group than in the low-expression group (Fig. 3B). Univariate analysis indicated
that gender (P=0.007), histology (P=0.005), location (P=0.009),
tumor depth (P=0.007), lymphatic invasion (P=0.013), lymph node
metastasis (P=0.024), CEA level (P=0.004), and S100A2 expression
(P<0.001) were significantly associated with OS. Multivariate
analysis indicated that S100A2 overexpression is an independent and
significant prognostic factor of OS for patients with stage II or
III CRC (P=0.008; RR=3.941; 95% confidence interval, 1.434–10.830)
(Table III).
 | Table IIIUnivariate and multivariate analysis
of clinicopathologic factors affecting overall survival in patients
with stage II–III colorectal cancer. |
Table III
Univariate and multivariate analysis
of clinicopathologic factors affecting overall survival in patients
with stage II–III colorectal cancer.
| | | Multivariate
analysis |
|---|
| | |
|
|---|
| Variables | No. of
patients | Univariate analysis
P-value | Relative risk (95%
confidence interval) | P-value |
|---|
| Age | | 0.061 | | |
| ≤65 | 81 | | | |
| >65 | 80 | | | |
| Gender | | 0.007 | | 0.058 |
| Male | 103 | | | |
| Female | 58 | | | |
| Histology | | 0.005 | | 0.115 |
| Well | 55 | | | |
| Moderate, poor and
others | 106 | | | |
| Location | | 0.009 | | 0.111 |
| Colon | 99 | | | |
| Rectum | 62 | | | |
| Depth | | 0.007 | | 0.07 |
| T1/T2/T3 | 96 | | | |
| T4 | 65 | | | |
| Lymphatic
invasion | | 0.013 | | 0.363 |
| Negative | 37 | | | |
| Positive | 124 | | | |
| Venous
invasion | | 0.257 | | |
| Negative | 13 | | | |
| Positive | 148 | | | |
| Lymph node
metastasis | | 0.024 | | 0.613 |
| Negative | 80 | | | |
| Positive | 81 | | | |
| CEA (ng/ml) | | 0.004 | | 0.533 |
| <5 | 96 | | | |
| ≥5 | 58 | | | |
| S100A2
expression | | <0.001 | 3.941 | 0.008 |
| Low | 70 | | (1.434–10.830) | |
| High | 91 | | | |
Fig. 3C shows RFS
curves stratified by TNM-7th stage and S100A2 expression level
group. RFS curves were significantly separated by S100A2 expression
level group in both stage II and III patients. RFS was
significantly worse in the S100A2 high expression group than in the
S100A2 low expression group in stage II (P=0.041) as well as in
stage III (P<0.001) patients.
Discussion
This study is the first to demonstrate the
prognostic significance of intratumoral S100A2 expression in
clinical tissue samples of CRC. S100A2 was identified as a
recurrence-related gene in the combined analysis of gene expression
and copy number, and high S100A2 expression was an independent and
significant prognostic factor of distant recurrence after curative
surgery for stage II or III CRC.
The S100A2 gene is located on chromosome 1q.21. The
human S100A2 is a member of the S100 family. It has been suggested
that this family promotes tumor progression and metastasis by
regulating the cell cycle, motility, and invasion in many human
neoplasms (30–34). S100A2 is an EF-hand calcium-binding
protein that regulates protein phosphorylation, cytoskeletal
components, and calcium homeostasis both inside and outside of
cells (30,35). The S100A2 protein, which is found
in the cytoplasm and nucleus of epithelial cells including those in
the esophagus and colon, is involved in TGF-β signaling (36,37).
It has also been suggested to play a role in the cell cycle
regulation of p53. S100A2 protein overexpression has been reported
in lung cancer, brain cancer, and several types of
gastroenterological cancers such as pancreatic and esophageal
cancers (35). The S100A2 protein
is reported to be involved in the chemotactic activity of tumor
cells in colon cancer (37,38).
In addition, S100A2 knockdown has been reported to reduce
TGF-β-induced cellular chemotaxis (37). In the present study, high
expression of S100A2 was significantly related with recurrence with
distant metastasis. Our results support the idea that S100A2 might
play an important role in distant metastasis of CRC. Further
studies are warranted to investigate the roles and functions of
S100A2 in CRC.
At present, recurrence risk and prognosis are
predicted largely based on pathological tumor staging (TNM
classification). The usefulness of postoperative adjuvant
chemotherapy for stage II CRC has not been established yet, and it
is recommended that determination of whether or not to use adjuvant
chemotherapy should be based on the recurrence risk predicted for
each patient. Major western guidelines, such as the National
Comprehensive Cancer Network of Clinical Practice Guidelines in
Oncology, recommend adjuvant chemotherapy when patients have risk
factors including T4 lesions, less than 12 lymph nodes examined,
perforation, poorly differentiated histology, and lymphovascular
involvement. Our results suggested that stage II CRC patients with
high S100A2 expression might be candidates for adjuvant
chemotherapy as high-risk patients of distant recurrence. For stage
III CRC, postoperative chemotherapy is recommended without
exception. However, when stage III is further divided into IIIA,
IIIB, and IIIC according to the TNM-7th classification and each
sub-stage is considered separately, it has been reported that an
additive effect of oxaliplatin cannot be anticipated in stage IIIA
patients, and that stratification of the recurrence risk is
important for stage III colon cancer as is the case for stage II
CRC (39,40). Our study suggests that stage III
CRC patients with high S100A2 expression may require strong
adjuvant chemotherapy. Validation of the usefulness of risk-guided
treatment is important in postoperative chemotherapy for stage II
and III CRC and further clinical studies need to be conducted.
In conclusion, this study demonstrated that S100A
was expressed at a significantly increased level in the CRC
recurrence group. In our screen, we focused on highly expressed
genes, because we intended to use the identified gene, S100A2, as a
blood biomarker in actual clinical practice. The results of our
study suggest the potential of the S100A protein as a
recurrence-predicting factor and a target for molecular-targeted
drugs for CRC.
Acknowledgements
The authors thank Y. Takagi and M. Itoda for
excellent technical assistance.
References
|
1
|
Ricchi P, Zarrilli R, Di Palma A and
Acquaviva AM: Nonsteroidal anti-inflammatory drugs in colorectal
cancer: From prevention to therapy. Br J Cancer. 88:803–807. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Tsukuma H, Ajiki W and Oshima A: Cancer
incidence in Japan. Gan To Kagaku Ryoho. 31:840–846. 2004.In
Japanese. PubMed/NCBI
|
|
3
|
Matsuda T, Marugame T, Kamo K, Katanoda K,
Ajiki W and Sobue T; Japan Cancer Surveillance Research Group.
Cancer incidence and incidence rates in Japan in 2004: Based on
data from 14 population-based cancer registries in the Monitoring
of Cancer Incidence in Japan (MCIJ) Project. Jpn J Clin Oncol.
40:1192–1200. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kobayashi H, Mochizuki H, Sugihara K,
Morita T, Kotake K, Teramoto T, Kameoka S, Saito Y, Takahashi K,
Hase K, et al: Characteristics of recurrence and surveillance tools
after curative resection for colorectal cancer: A multicenter
study. Surgery. 141:67–75. 2007. View Article : Google Scholar
|
|
5
|
Weitz J, Koch M, Debus J, Höhler T, Galle
PR and Büchler MW: Colorectal cancer. Lancet. 365:153–165. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ross JS, Torres-Mora J, Wagle N, Jennings
TA and Jones DM: Biomarker-based prediction of response to therapy
for colorectal cancer: Current perspective. Am J Clin Pathol.
134:478–490. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wang Y, Jatkoe T, Zhang Y, Mutch MG,
Talantov D, Jiang J, McLeod HL and Atkins D: Gene expression
profiles and molecular markers to predict recurrence of Dukes' B
colon cancer. J Clin Oncol. 22:1564–1571. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Shih W, Chetty R and Tsao MS: Expression
profiling by microarrays in colorectal cancer (Review). Oncol Rep.
13:517–524. 2005.PubMed/NCBI
|
|
9
|
Nannini M, Pantaleo MA, Maleddu A, Astolfi
A, Formica S and Biasco G: Gene expression profiling in colorectal
cancer using microarray technologies: Results and perspectives.
Cancer Treat Rev. 35:201–209. 2009. View Article : Google Scholar
|
|
10
|
Aragane H, Sakakura C, Nakanishi M,
Yasuoka R, Fujita Y, Taniguchi H, Hagiwara A, Yamaguchi T, Abe T,
Inazawa J, et al: Chromosomal aberrations in colorectal cancers and
liver metastases analyzed by comparative genomic hybridization. Int
J Cancer. 94:623–629. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kurashina K, Yamashita Y, Ueno T, Koinuma
K, Ohashi J, Horie H, Miyakura Y, Hamada T, Haruta H, Hatanaka H,
et al: Chromosome copy number analysis in screening for
prognosis-related genomic regions in colorectal carcinoma. Cancer
Sci. 99:1835–1840. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Nakao M, Kawauchi S, Furuya T, Uchiyama T,
Adachi J, Okada T, Ikemoto K, Oga A and Sasaki K: Identification of
DNA copy number aberrations associated with metastases of
colorectal cancer using array CGH profiles. Cancer Genet Cytogenet.
188:70–76. 2009. View Article : Google Scholar
|
|
13
|
Yamamoto S, Midorikawa Y, Morikawa T,
Nishimura Y, Sakamoto H, Ishikawa S, Akagi K and Aburatani H:
Identification of chromosomal aberrations of metastatic potential
in colorectal carcinoma. Genes Chromosomes Cancer. 49:487–496.
2010.PubMed/NCBI
|
|
14
|
Yau C and Holmes CC: CNV discovery using
SNP genotyping arrays. Cytogenet Genome Res. 123:307–312. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Phillips JL, Hayward SW, Wang Y, Vasselli
J, Pavlovich C, Padilla-Nash H, Pezullo JR, Ghadimi BM, Grossfeld
GD, Rivera A, et al: The consequences of chromosomal aneuploidy on
gene expression profiles in a cell line model for prostate
carcinogenesis. Cancer Res. 61:8143–8149. 2001.PubMed/NCBI
|
|
16
|
Hyman E, Kauraniemi P, Hautaniemi S, Wolf
M, Mousses S, Rozenblum E, Ringnér M, Sauter G, Monni O, Elkahloun
A, et al: Impact of DNA amplification on gene expression patterns
in breast cancer. Cancer Res. 62:6240–6245. 2002.PubMed/NCBI
|
|
17
|
Pollack JR, Sørlie T, Perou CM, Rees CA,
Jeffrey SS, Lonning PE, Tibshirani R, Botstein D, Børresen-Dale AL
and Brown PO: Microarray analysis reveals a major direct role of
DNA copy number alteration in the transcriptional program of human
breast tumors. Proc Natl Acad Sci USA. 99:12963–12968. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Tsafrir D, Bacolod M, Selvanayagam Z,
Tsafrir I, Shia J, Zeng Z, Liu H, Krier C, Stengel RF, Barany F, et
al: Relationship of gene expression and chromosomal abnormalities
in colorectal cancer. Cancer Res. 66:2129–2137. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ramakrishna M, Williams LH, Boyle SE,
Bearfoot JL, Sridhar A, Speed TP, Gorringe KL and Campbell IG:
Identification of candidate growth promoting genes in ovarian
cancer through integrated copy number and expression analysis. PLoS
One. 5:e99832010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Nigro JM, Misra A, Zhang L, Smirnov I,
Colman H, Griffin C, Ozburn N, Chen M, Pan E, Koul D, et al:
Integrated array-comparative genomic hybridization and expression
array profiles identify clinically relevant molecular subtypes of
glioblastoma. Cancer Res. 65:1678–1686. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Cardoso J, Boer J, Morreau H and Fodde R:
Expression and genomic profiling of colorectal cancer. Biochim
Biophys Acta. 1775:103–137. 2007.
|
|
22
|
Yoshida T, Kobayashi T, Itoda M, Muto T,
Miyaguchi K, Mogushi K, Shoji S, Shimokawa K, Iida S, Uetake H, et
al: Clinical omics analysis of colorectal cancer incorporating copy
number aberrations and gene expression data. Cancer Inform.
9:147–161. 2010.PubMed/NCBI
|
|
23
|
Pedrocchi M, Schäfer BW, Mueller H,
Eppenberger U and Heizmann CW: Expression of Ca(2+)-binding
proteins of the S100 family in malignant human breast-cancer cell
lines and biopsy samples. Int J Cancer. 57:684–690. 1994.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Böni R, Heizmann CW, Doguoglu A, Ilg EC,
Schäfer BW, Dummer R and Burg G: Ca(2+)-binding proteins S100A6 and
S100B in primary cutaneous melanoma. J Cutan Pathol. 24:76–80.
1997. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Lauriola L, Michetti F, Maggiano N, Galli
J, Cadoni G, Schäfer BW, Heizmann CW and Ranelletti FO: Prognostic
significance of the Ca(2+) binding protein S100A2 in laryngeal
squamous-cell carcinoma. Int J Cancer. 89:345–349. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Feng G, Xu X, Youssef EM and Lotan R:
Diminished expression of S100A2, a putative tumor suppressor, at
early stage of human lung carcinogenesis. Cancer Res. 61:7999–8004.
2001.PubMed/NCBI
|
|
27
|
Gupta S, Hussain T, MacLennan GT, Fu P,
Patel J and Mukhtar H: Differential expression of S100A2 and S100A4
during progression of human prostate adenocarcinoma. J Clin Oncol.
21:106–112. 2003. View Article : Google Scholar
|
|
28
|
Tsai ST, Jin YT, Tsai WC, Wang ST, Lin YC,
Chang MT and Wu LW: S100A2, a potential marker for early recurrence
in early-stage oral cancer. Oral Oncol. 41:349–357. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Simonsen K, Rode A, Nicoll A, Villadsen G,
Espelund U, Lim L, Angus P, Arachchi N, Vilstrup H, Nexo E, et al:
Vitamin B12 and its binding proteins in hepatocellular carcinoma
and chronic liver diseases. Scand J Gastroenterol. 49:1096–1102.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Salama I, Malone PS, Mihaimeed F and Jones
JL: A review of the S100 proteins in cancer. Eur J Surg Oncol.
34:357–364. 2008. View Article : Google Scholar
|
|
31
|
Haase-Kohn C, Wolf S, Lenk J and Pietzsch
J: Copper-mediated cross-linking of S100A4, but not of S100A2,
results in proinflammatory effects in melanoma cells. Biochem
Biophys Res Commun. 413:494–498. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Jamieson NB, Carter CR, McKay CJ and Oien
KA: Tissue biomarkers for prognosis in pancreatic ductal
adenocarcinoma: a systematic review and meta-analysis. Clin Cancer
Res. 17:3316–3331. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
McKiernan E, McDermott EW, Evoy D, Crown J
and Duffy MJ: The role of S100 genes in breast cancer progression.
Tumour Biol. 32:441–450. 2011. View Article : Google Scholar
|
|
34
|
Jin L, Shen Q, Ding S, Jiang W, Jiang L
and Zhu X: Immunohistochemical expression of Annexin A2 and S100A
proteins in patients with bulky stage IB-IIA cervical cancer
treated with neoadjuvant chemotherapy. Gynecol Oncol. 126:140–146.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wolf S, Haase-Kohn C and Pietzsch J:
S100A2 in cancero-genesis: A friend or a foe? Amino Acids.
41:849–861. 2011. View Article : Google Scholar
|
|
36
|
Ranganathan P, Agrawal A, Bhushan R,
Chavalmane AK, Kalathur RK, Takahashi T and Kondaiah P: Expression
profiling of genes regulated by TGF-beta: Differential regulation
in normal and tumour cells. BMC Genomics. 8:982007. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Naz S, Ranganathan P, Bodapati P, Shastry
AH, Mishra LN and Kondaiah P: Regulation of S100A2 expression by
TGF-β-induced MEK/ERK signalling and its role in cell
migration/invasion. Biochem J. 447:81–91. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Giráldez MD, Lozano JJ, Cuatrecasas M,
Alonso-Espinaco V, Maurel J, Mármol M, Hörndler C, Ortego J, Alonso
V, Escudero P, et al: Gene-expression signature of tumor recurrence
in patients with stage II and III colon cancer treated with
5-fluoruracil-based adjuvant chemotherapy. Int J Cancer.
132:1090–1097. 2013. View Article : Google Scholar
|
|
39
|
Haller DG, Tabernero J, Maroun J, de Braud
F, Price T, Van Cutsem E, Hill M, Gilberg F, Rittweger K and
Schmoll HJ: Capecitabine plus oxaliplatin compared with
fluorouracil and folinic acid as adjuvant therapy for stage III
colon cancer. J Clin Oncol. 29:1465–1471. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Gao P, Song YX, Wang ZN, Xu YY, Tong LL,
Sun JX, Yu M and Xu HM: Is the prediction of prognosis not improved
by the seventh edition of the TNM classification for colorectal
cancer? Analysis of the surveillance, epidemiology, and end results
(SEER) database. BMC Cancer. 13:1232013. View Article : Google Scholar : PubMed/NCBI
|