Introduction
Prostate cancer (PCa) is the second most common
cancer and one of the leading causes of mortality among males
worldwide by 2012 (1). The incidence
of PCa in China is considered reduced compared with that in Western
countries; however, it has been progressively increasing over the
past 30 years (2).
Genetic susceptibility to PCa has been well
established and almost 100 common risk loci have been identified by
genome wide association studies (GWAS) among European,
African-American, Japanese and Chinese populations (3,4). However,
only 10 of these loci were initially identified from GWAS in
Japanese and Chinese populations. Among previous evaluation and
validation studies, a part of the loci was revealed to be
associated with PCa risk in Chinese population (5). Since these risk-associated single
nucleotide polymorphisms (SNPs) exhibited a cumulative effect on
PCa risk, the genetic risk scores (GRS) derived from PCa
risk-associated SNPs were able to evaluate an individual's risk of
PCa. The GRS based on the Chinese population is established and
demonstrated to be a significant predictor of biopsy outcome in
previous studies (5–9).
With an increasing sample size used in GWAS through
combined data, a meta-analysis of a multi-ethnic population, which
included 87,040 individuals, identified 23 new susceptibility loci
for PCa (including 15 in European, 7 in multiethnic and 1 in the
early onset analysis) (10). These
PCa risk-associated alleles exhibited decreased effects with odds
ratios (ORs) ranging between 1.06 and 1.14 (10). However, since the Chinese population
was not included in the study, the effects of these 23 novel risk
variants in individuals of Chinese descent remains unknown.
The objective of the present study was to evaluate
the 23 newly identified susceptibility loci for PCa in a Chinese
population and assess whether any validated loci contributed to the
GRS in predicting the risk of PCa in a Chinese population.
Materials and methods
Population
The baseline characteristics of the present study
subjects were summarized (Table I). A
total of 2,425 subjects including 1,417 patients with PCa and 1,008
controls were recruited in the present study. All patients were
part of the China PCa consortium from the southeast of China
(11–13) recruited during January 2010 and
December 2011, from which data were obtained. All cases were
pathologically diagnosed with primary PCa and all the controls were
recruited from the community or selected from subjects who had
undergone routine physical examination in local hospitals. Written
informed consent was obtained from subjects for their participation
in the present study and a blood sample was taken from each subject
at the time of recruitment for DNA extraction. The present study
was reviewed and approved by the Institutional Review Board of
every participating institution.
 | Table I.Characteristics of study
population. |
Table I.
Characteristics of study
population.
| Variables | PCa cases
(n=1,417) | Controls
(n=1,008) |
|---|
| Age,
yearsa,b | 71.3±8.1 | 62.1±10.0 |
| PSA,
ng/mla,c |
|
|
|
0–3.99 | 54 (4.0) | 965 (95.9) |
|
4–9.99 | 187 (14.0) | 32 (3.2) |
|
10–19.99 | 305 (22.8) | 6 (0.6) |
| ≥20 | 791 (59.2) | 3 (0.3) |
|
Missing | 80 (5.6) | 2 (0.2) |
| Gleason
scorec |
|
|
| ≤7 | 809 (60.1) | N/A |
| ≥8 | 537 (39.9) | N/A |
|
Missing | 71 (5) | N/A |
Genotyping and quality control
DNA samples were genotyped in the Center for Cancer
Genomics at Wake Forest University (Winston-Salem, NC, USA) using
the Illumina HumanOmniExpress BeadChips (Illumina, Inc., San Diego,
CA, USA), which included 731,458 SNPs. For PCa risk-associated SNPs
that were not included in the GWAS array, imputation was performed
using IMPUTE 2.2.2 based on the combined data of the 1,000 Genomes
project and HapMap3 data (14). A
posterior probability of >0.9 was applied to call imputed
genotypes. Imputed SNPs were excluded if they exhibited: i) A call
rate <95%; ii) a minor allele frequency <0.05; or iii)
P<1×10−3 in a Hardy-Weinberg equilibrium test in
controls, as previously described (13).
Assessment of genetic risk
A GRS was calculated for each subject based on
genotypes of the SNPs and weighted by their ORs and risk allele
frequency, as described previously (15). GRS was calculated as
GRS=∏i=1nORigiWi
where gi is the genotype of SNP
i for an individual (0, homozygous of non-risk allele; 1,
heterozygous; 2 homozygous of risk allele). ORi
is the OR of SNP i estimated from external study (16), Wi is the average
population risk of SNP i, calculated as Wi
= fi2ORi2
+ 2fi(1-fi) ORi +
(1-fi)2, where fi is
the risk allele frequency of SNP i based on the 1,000 Genome
Project of the CHB (Han Chinese in Bejing, China) population
(17). Therefore, a GRS value of 1.0
represents a population average risk.
Statistical analysis
A logistic regression model was used to analyze the
association of each SNP with PCa risk, assuming an additive genetic
model, which was implemented in PLINK version 1.07 (18). ORs and 95% confident intervals (CIs)
were estimated from logistic regression analysis with adjustment
for age and the highest eigen value. Student's t-tests were used to
analyze the differences in means of normally distributed variables
between 2 groups. For variables that were not normally distributed,
2 tests were performed: i) A nonparametric method using the
Wilcoxon rank sum test and ii) Student's t-tests for different
means between 2 groups following log-transformation. Differences in
binary variables were investigated using χ2 tests. Area
under the receiver operating characteristic curve (AUC) was used to
evaluate the performance of GRS in discriminating between 2 groups
of subjects. The difference between two AUCs was determined using
Delong's test (19). P<0.05 was
considered to indicate a statistically significant difference. All
statistical analyses were performed using SPSS 19.0 (SPSS; IBM
Corporation, Armonk, NY, USA).
Results
SNPs and PCa risk
The present study evaluated 19 newly reported SNPs.
Among the 19 SNPs, only 1 (rs7153648 at 14q23) was associated with
PCa risk in the China PCa cohort (OR=1.206, P<0.05). The
direction of the effect was consistent with the previous
multiethnic meta-analysis (10). The
other 18 SNPs that had previously demonstrated genome-wide
significance in European ancestry meta-analysis and multiethnic
meta-analysis (Table II) either were
not associated with PCa risk or did not demonstrate the same
magnitude of effect in the Chinese population investigated in the
present study.
 | Table II.Association results for 19 novel risk
variants for PCa in Chinese males. |
Table II.
Association results for 19 novel risk
variants for PCa in Chinese males.
|
|
|
|
|
|
|
| Risk allele
frequency |
|
|
|---|
|
|
|
|
|
|
|
|
|
|
|
|---|
| Origin of GWAS | SNP ID | Chromosome
positiona | Region | Gene | Alleles | Risk allele | PCA cases | Controls | Odds ratio | P-value |
|---|
| European | rs636291 | 1p35 | Intron | PEX14 | A/G | G | 0.254 | 0.241 | 1.12 | 0.148 |
| European | rs17599629 | 1q21 | Intron | GOLPH3L | G/A | G | 0.103 | 0.098 | 1.03 | 0.780 |
| Multi-ethnic | rs1775148 | 1q32 | Intergenic | SLC41A1 | C/T | C | 0.508 | 0.486 | 1.13 | 0.109 |
| European | rs9287719 | 2p25 | Intergenic | NOL10 | C/T | C | 0.372 | 0.350 | 1.14 | 0.075 |
| European | rs10009409 | 4q13 | Intergenic | COX18 | T/C | C | 0.484 | 0.468 | 1.07 | 0.310 |
| European | rs4713266 | 6p24 | Intron | NEDD9 | C/T | C | 0.187 | 0.177 | 1.11 | 0.263 |
| European | rs115457135 | 6p22 | Intron | TRIM31 | A/G | A | 0.144 | 0.161 | 1.03 | 0.903 |
| European | rs115306967 | 6p21 | Intergenic | HLA-DRB6 | G/C | C | 0.135 | 0.126 | 1.12 | 0.570 |
| Multi-ethnic | rs9443189 | 6q14 | Intron | MYO6 | G/A | A | 0.633 | 0.632 | 1.03 | 0.728 |
| European | rs56232506 | 7p12 | Intron | TNS3 | A/G | A | 0.383 | 0.383 | 1.02 | 0.764 |
| European | rs17694493 | 9p21 | Intron | CDKN2B-AS1 | G/C | G | 0.030 | 0.029 | 1.13 | 0.575 |
| European | rs76934034 | 10q11 | Intron | 41706 | T/C | T | 1.000 | 0.999 | 1.00 | 0.999 |
| European | rs11214775 | 11q23 | Intron | HTR3B | G/A | G | 0.798 | 0.796 | 0.97 | 0.756 |
| Multi-ethnic | rs7153648 | 14q23 | Intergenic | SIX1 | C/G | C | 0.178 | 0.153 | 1.21 | 0.045 |
| European | rs8014671 | 14q24 | Intergenic | TTC9 | G/A | G | 0.300 | 0.298 | 0.99 | 0.904 |
| Multi-ethnic | rs12051443 | 16q22 | Intron | PHLPP2 | A/G | A | 0.762 | 0.758 | 0.91 | 0.243 |
| Multi-ethnic | rs12480328 | 20q13 | Intron | ADNP | T/C | T | 0.924 | 0.919 | 0.94 | 0.616 |
| Multi-ethnic | rs1041449 | 21q22 | Intergenic | TMPRSS2 | G/A | G | 0.164 | 0.161 | 1.05 | 0.674 |
| Multi-ethnic | rs2238776 | 22q11 | Intron | TBX1 | G/A | G | 0.515 | 0.515 | 1.07 | 0.376 |
SNPs and PCa aggressiveness
The association between the 19 SNPs and PCa
aggressiveness was also investigated (cases with a Gleason score
≥7; Table III). The results did not
demonstrate a significant association between rs7153648 and PCa
aggressiveness, whereas rs636291 at 1p23 was significantly
associated with PCa aggressiveness (OR=1.123, P<0.05).
 | Table III.Association results for 19 novel risk
variants for aggressive PCa in Chinese males. |
Table III.
Association results for 19 novel risk
variants for aggressive PCa in Chinese males.
|
|
|
|
|
|
|
| Risk allele
frequency |
|
|
|---|
|
|
|
|
|
|
|
|
|
|
|
|---|
| Origin of GWAS | SNP ID | Chromosome
positiona | Region | Gene | Alleles | Risk allele | PCa cases | Controls | Odds ratio | P-value |
|---|
| European | rs636291 | 1p35 | Intron | PEX14 | A/G | G | 0.262 | 0.239 | 1.16 | 0.049 |
| European | rs17599629 | 1q21 | Intron | GOLPH3L | G/A | G | 0.102 | 0.100 | 1.00 | 0.984 |
| Multi-ethnic | rs1775148 | 1q32 | Intergenic | SLC41A1 | C/T | C | 0.508 | 0.490 | 1.11 | 0.148 |
| European | rs9287719 | 2p25 | Intergenic | NOL10 | C/T | C | 0.373 | 0.356 | 1.07 | 0.303 |
| European | rs10009409 | 4q13 | Intergenic | COX18 | T/C | C | 0.477 | 0.475 | 0.98 | 0.746 |
| European | rs4713266 | 6p24 | Intron | NEDD9 | C/T | T | 0.820 | 0.818 | 0.99 | 0.862 |
| European | rs115457135 | 6p22 | Intron | TRIM31 | A/G | A | 0.139 | 0.132 | 0.95 | 0.790 |
| European | rs115306967 | 6p21 | Intergenic | HLA-DRB6 | G/C | C | 0.142 | 0.124 | 1.21 | 0.308 |
| Multi-ethnic | rs9443189 | 6q14 | Intron | MYO6 | G/A | G | 0.378 | 0.362 | 1.10 | 0.141 |
| European | rs56232506 | 7p12 | Intron | TNS3 | A/G | A | 0.388 | 0.379 | 1.07 | 0.374 |
| European | rs17694493 | 9p21 | Intron | CDKN2B-AS1 | G/C | G | 0.031 | 0.029 | 1.12 | 0.565 |
| European | rs76934034 | 10q11 | Intron | 41706 | T/C | T | 1.000 | 1.000 | 1.00 | 0.999 |
| European | rs11214775 | 11q23 | Intron | HTR3B | G/A | G | 0.800 | 0.796 | 0.97 | 0.661 |
| Multi-ethnic | rs7153648 | 14q23 | Intergenic | SIX1 | C/G | C | 0.171 | 0.161 | 1.05 | 0.571 |
| European | rs8014671 | 14q24 | Intergenic | TTC9 | G/A | G | 0.305 | 0.294 | 1.06 | 0.407 |
| Multi-ethnic | rs12051443 | 16q22 | Intron | PHLPP2 | A/G | A | 0.767 | 0.760 | 0.92 | 0.286 |
| Multi-ethnic | rs12480328 | 20q13 | Intron | ADNP | T/C | T | 0.929 | 0.919 | 0.89 | 0.352 |
| Multi-ethnic | rs1041449 | 21q22 | Intergenic | TMPRSS2 | G/A | A | 0.840 | 0.834 | 0.96 | 0.647 |
| Multi-ethnic | rs2238776 | 22q11 | Intron | TBX1 | G/A | G | 0.517 | 0.512 | 1.01 | 0.875 |
SNPs, GRS and PCa
GRS was calculated using rs7153648 and 29 previously
implicated SNPs (10). The mean GRS
based on the 30 SNPs (GRS30) was significantly increased in
patients with PCa compared with that among non-PCa individuals
(1.439 vs. 0.961, P=7.44×10−41; Table IV). As reported in a previous study,
it would be more efficient and reliable to calculate GRS using
race-specific disease-associated SNPs that demonstrated genome-wide
significance (20). Therefore, in the
present study, GRS was also calculated based on the 9 strongest
SNPs previously reported in individuals of Asian descent (GRS9;
Table V) (16). The mean GRS based on 9 SNPs was 1.26
in patients with PCa and 0.99 in non-PCa controls
(P=3.71×10−28).
 | Table IV.Genetic score and prostate biopsy
outcomes. |
Table IV.
Genetic score and prostate biopsy
outcomes.
| Parameter | 9 SNPs | 30 SNPs |
|---|
| Genetic
scorea |
|
|
|
PCa | 1.26±0.72 | 1.44±1.18 |
|
Non-PCa | 0.99±0.53 | 0.96±0.73 |
|
P-value |
3.71×10−28 |
7.44×10−41 |
| Association with
PCab |
|
|
| Genetic
score ≤1.0 | 1 | 1 |
| Genetic
score >1.0 | 2.47
(2.05–2.97) | 2.25
(1.96–2.58) |
|
P-value |
6.90×10−22 |
2.97×10−31 |
| Discrimination of
PCa |
|
|
|
AUC | 0.792 | 0.799 |
|
P-value | 0.138 |
| (AUC
comparison) |
|
|
 | Table V.Most significant SNPs previously
reported in Asian individualsa. |
Table V.
Most significant SNPs previously
reported in Asian individualsa.
|
|
|
|
| Reported SNPs in
meta-analysis |
|
|
|
|
|
|
|---|
|
|
|
|
|
|
|
|
|
|
|
|
|---|
| SNP | Locus |
Positionb | Nearby gene | Allelec | OR | P-value |
| Frequency in 1,000
Genome-CHB |
|
|---|
| rs12653946 | 5p15.33 | 1895829 | IRX4 | G/A | 0.78 |
1.94×10−12 | TT | 0.103 | TC | 0.474 | CC | 0.422 |
| rs339331 | 6q22.2 |
1.17×108 | RFX6 | C/T | 0.78 |
4.27×10−11 | CC | 0.103 | CT | 0.485 | TT | 0.412 |
| rs1512268 | 8p21.2 | 23526463 | NKX3-1 | C/T | 0.76 |
1.40×10−13 | TT | 0.113 | TC | 0.526 | CC | 0.361 |
| rs16901979 | 8q24.21 |
1.28×108 | POU5F1B | C/A | 0.72 |
1.00×10−16 | AA | 0.052 | AC | 0.392 | CC | 0.557 |
| rs6983267 | 8q24.21 |
1.28×108 | POU5F1B | G/T | 1.23 |
8.36×10−9 | GG | 0.113 | GT | 0.526 | TT | 0.361 |
| rs1447295 | 8q24.21 |
1.28×108 | LOC727677 | C/A | 0.65 |
3.92×10−23 | AA | 0.010 | AC | 0.237 | CC | 0.753 |
| rs10993994 | 10q11 | 51549496 | MSMB | C/T | 0.83 |
3.58×10−8 | TT | 0.165 | TC | 0.505 | CC | 0.330 |
| rs12791447 | 11p15.4 | 7556577 | PPFIBP2 | C/T | 1.23 |
3.59×10−8 | GG | 0.010 | GA | 0.155 | AA | 0.835 |
| rs58262369 | 14q23.2 | 64693912 | ESR2 | C/T | 0.78 |
6.05×10−10 | TT | 0.031 | TC | 0.216 | CC | 0.753 |
Following adjustment for age (Table IV), GRS9 and GRS30 remained
significantly associated with PCa (all P<0.01). The OR of the
GRS30 for the prediction of PCa risk was 2.25 (95% CI, 1.976–2.598;
P=2.97×10−31), decreased compared with that of GRS9
(OR=2.468; 95% CI, 2.053–2.967; P=6.9×10−22), although
no significant differences were identified. When comparing the
predictive ability of the GRS9 and GRS30, the AUCs were similar
(0.792 for GRS9 vs. 0.7994 for GRS30, P=0.138).
Discussion
Genetic susceptibility is a major risk factor for
PCa and is estimated to account for 42% of variation in the disease
(21). In the past few years, GWAS
and meta-analysis of combined data have identified 99 genomic
variants associated with PCa in multiple populations of European,
African-American, Japanese, Latino and Chinese ancestry (10). In the present study, 23 novel
susceptibility loci detected in European ancestry or multi-ethnic
analysis were investigated and their association in a Chinese
population was evaluated.
Of the 19 SNPs evaluated in the present study, only
1 was identified to be associated with PCa. The estimate of risk of
this SNP in the Chinese population was similar to that in European
and multi-ethnic populations (10).
Despite reaching genome-wide significance in European or
multi-ethnic populations, the other 18 loci were not identified to
be significant in the population of the present study. The
discrepancy may be explained in multiple ways. First, since the
effects of 18 SNPs (not including rs636291 at 1p36) were relatively
low, with ORs ranging between 1.06 and 1.13, the present study may
not possess the power to identify the small effects of these SNPs.
This was also one of the reasons why rs7153648 did not reach a
significant level following Bonferroni correction (P=0.05/19).
Second, the risk allele frequencies in European and Chinese
ancestry differed between SNPs evaluated (Table II); this difference may also
influence the detection of significant effects of these SNPs in
populations of Chinese ancestry. Finally, besides the different
genetic backgrounds between European ancestry (or other
populations) and Chinese ancestry, environmental factors,
dietary-habit and other non-genetic factors may also affect the
penetrance of these alleles, which may result in the difference of
risk profiles.
When evaluating the association between the 19 SNPs
and aggressive PCa (Gleason score, ≥7), the results demonstrated
that rs636291 at 1p36 reached a significant level (P<0.05). This
SNP reached genome-wide significance in early onset disease in
European ancestry (10); however, a
similar analysis could not be performed in the present study due to
the lack of cases (only 34 patients with PCa were diagnosed <55
years of age). Nevertheless, this result may indicate that this
risk variant was associated with more advanced PCa and should be
further validated in an independent study.
In the comparison of the two GRS-based risk models,
the results revealed that the performance was approximately the
same between the two models. This may be attributed to the fact
that certain risk variants were not strongly associated with PCa
and others conferred a decreased effect to the risk of PCa in
Chinese population compared with that in European whites. In a
previous study, the plateau effect of PCa risk-associated SNPs was
evaluated in predicting PCa in a Chinese population and it was
identified that the predictive performance increased when the top
13 highest impact PCa risk-associated SNPs were included in the GRS
(9). The results were similar in the
present study; therefore, this may indicate that further SNPs
weakly associated with PCa may not improve the predictive
performance of GRS for PCa. Therefore, GRS only including the
strongest SNPs may be appropriate while balancing the predictive
performance and economic benefit.
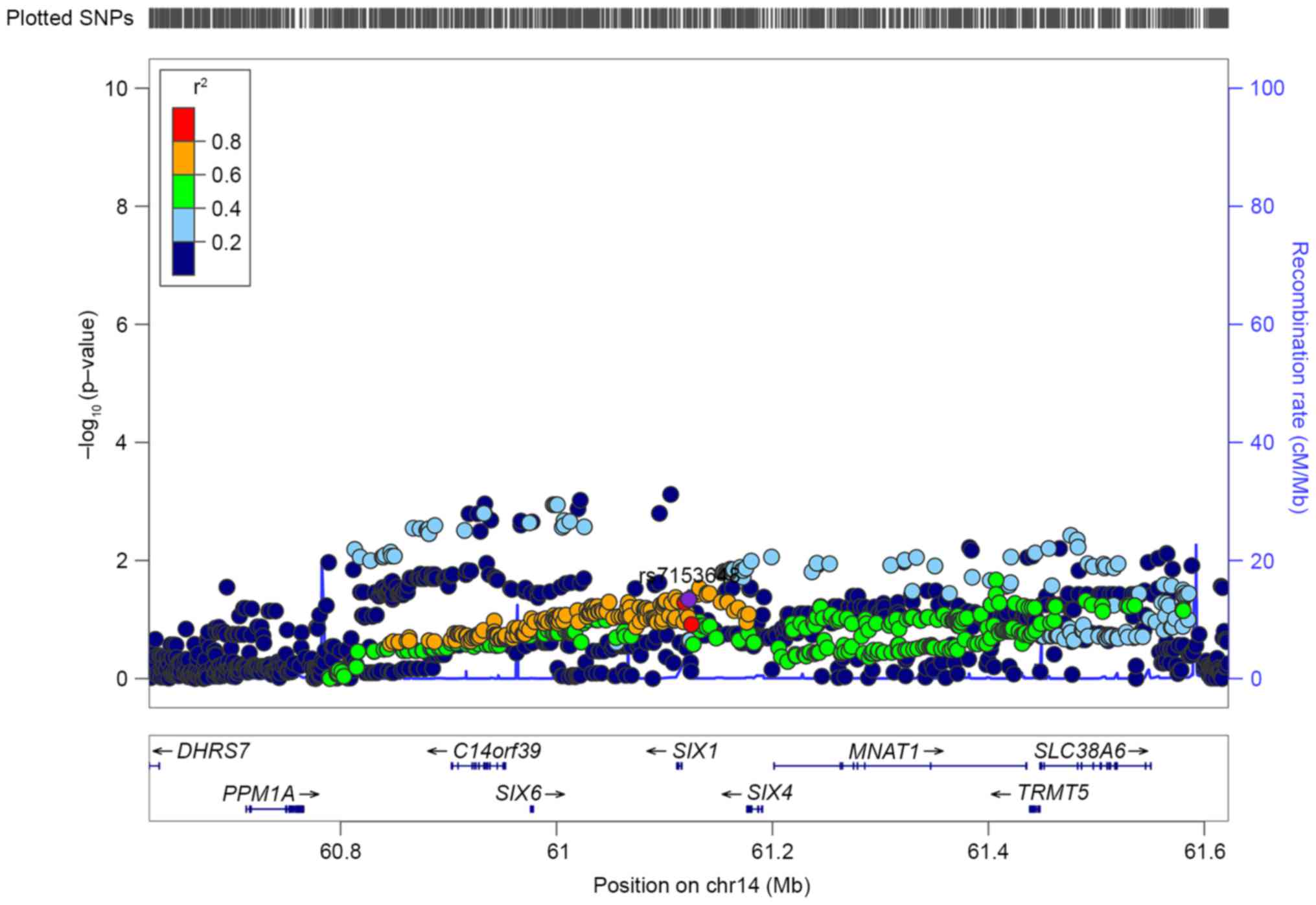
In the present study, the variant rs7153648 at 14q23
that we demonstrated to be associated with PCa is located in the
intergenic region of SIX homeobox 1. The regional information of
the confirmed SNP (rs7153648) was presented (Fig. 1). In the LocusZoom plots of this loci,
multiple SNPs located upstream of rs7153648 demonstrated marked
association (P<0.01) but a weak correlation (dark blue circles),
which may suggest the presence of multiple potential independent
association signals. Variant rs636291 at 1p36, which was associated
with early-onset PCa in European ancestry and was identified to be
associated with aggressive PCa in Chinese ancestry in the current
study, is located in intron 2 of peroxisomal biogenesis factor 14
and is associated with a variant (rs616488) reported in a GWAS of
breast cancer (22).
There were multiple limitations to the present
study. First, only 19 SNPs, rather than 23 of the novel identified
loci, were genotyped or imputed due to 4 SNPs not being included in
the GWAS panel and failing to impute using the CHB population of
the 1,000 Genome project. Among the 4 SNPs, rs80130819 at 12q13 was
not polymorphous in the CHB population, while the remaining 3 were
polymorphous in the CHB population. Second, due to the open nature
of the China PCa cohort, the clinical characterization of the cases
was not consistent between distinct hospitals (e.g., Gleason score
diagnosis in the present study), which limited further analysis of
clinical phenotypes.
To conclude, by evaluating 19 PCa risk-associated
SNPs identified in a large meta-analysis of GWAS from a European
and multiethnic population, the results of the present study
identified 1 SNP that was associated with PCa risk and 1 that was
associated with aggressive PCa in a Chinese population. However,
the validated small-effect SNP and other SNPs that weakly
associated with PCa are not likely to improve the predictive
ability of existing GRS in Chinese populations.
Acknowledgements
The present study was supported by the National
Natural Science Foundation of China (grant nos. 81202269 and
81402339), the Clinical Science and Technology Innovation Project
of Shanghai Shen Kang Hospital Development Center (grant no.
SHDC12015105) and Scientific Research Project supported by Huashan
Hospital, Fudan University (grant no. 2016QD079).
References
|
1
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Liu M, Wang JY, Zhang YG, Zhu SC, Lu ZH
and Wan B: Detection of urological and male genital tumors
diagnosed in Beijing Hospital 1995–2004. Zhonghua Yi Xue Za Zhi.
87:2423–2425. 2007.(In Chinese). PubMed/NCBI
|
|
3
|
Xu J, Sun J and Zheng SL: Prostate cancer
risk-associated genetic markers and their potential clinical
utility. Asian J Androl. 15:314–322. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Eeles R, Goh C, Castro E, Bancroft E, Guy
M, Al Olama AA, Easton D and Kote-Jarai Z: The genetic epidemiology
of prostate cancer and its clinical implications. Nat Rev Urol.
11:18–31. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Na R, Liu F, Zhang P, Ye D, Xu C, Shao Q,
Qi J, Wang X, Chen Z, Wang M, et al: Evaluation of reported
prostate cancer risk-associated SNPs from genome-wide association
studies of various racial populations in Chinese men. Prostate.
73:1623–1635. 2013.PubMed/NCBI
|
|
6
|
Aly M, Wiklund F, Xu J, Isaacs WB, Eklund
M, D'Amato M, Adolfsson J and Grönberg H: Polygenic risk score
improves prostate cancer risk prediction: Results from the
Stockholm-1 cohort study. Eur Urol. 60:21–28. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Kader AK, Sun J, Reck BH, Newcombe PJ, Kim
ST, Hsu FC, D'Agostino RB Jr, Tao S, Zhang Z, Turner AR, et al:
Potential impact of adding genetic markers to clinical parameters
in predicting prostate biopsy outcomes in men following an initial
negative biopsy: Findings from the REDUCE trial. Eur Urol.
62:953–961. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Jiang H, Liu F, Wang Z, Na R, Zhang L, Wu
Y, Zheng J, Lin X, Jiang D, Sun J, et al: Prediction of prostate
cancer from prostate biopsy in Chinese men using a genetic score
derived from 24 prostate cancer risk-associated SNPs. Prostate.
73:1651–1659. 2013.PubMed/NCBI
|
|
9
|
Ren S, Xu J, Zhou T, Jiang H, Chen H, Liu
F, Na R, Zhang L, Wu Y, Sun J, et al: Plateau effect of prostate
cancer risk-associated SNPs in discriminating prostate biopsy
outcomes. Prostate. 73:1824–1835. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Al Olama AA, Kote-Jarai Z, Berndt SI,
Conti DV, Schumacher F, Han Y, Benlloch S, Hazelett DJ, Wang Z,
Saunders E, et al: A meta-analysis of 87,040 individuals identifies
23 new susceptibility loci for prostate cancer. Nature Genetics.
46:1103–1109. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Liu F, Hsing AW, Wang X, Shao Q, Qi J, Ye
Y, Wang Z, Chen H, Gao X, Wang G, et al: Systematic confirmation
study of reported prostate cancer risk-associated single nucleotide
polymorphisms in Chinese men. Cancer Sci. 102:1916–1920. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang M, Liu F, Hsing AW, Wang X, Shao Q,
Qi J, Ye Y, Wang Z, Chen H, Gao X, et al: Replication and
cumulative effects of GWAS-identified genetic variations for
prostate cancer in Asians: A case-control study in the ChinaPCa
consortium. Carcinogenesis. 33:356–360. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Xu J, Mo Z, Ye D, Wang M, Liu F, Jin G, Xu
C, Wang X, Shao Q, Chen Z, et al: Genome-wide association study in
Chinese men identifies two new prostate cancer risk loci at 9q31.2
and 19q13.4. Nat Genet. 44:1231–1235. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Marchini J, Howie B, Myers S, McVean G and
Donnelly P: A new multipoint method for genome-wide association
studies by imputation of genotypes. Nat Genet. 39:906–913. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wu Y, Zhang N, Li K, Chen H, Lin X, Yu Y,
Gou Y, Hou J, Jiang D, Na R, et al: Genetic scores based on
risk-associated single nucleotide polymorphisms (SNPs) can reveal
inherited risk of renal cell carcinoma. Oncotarget. 7:18631–18637.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang M, Takahashi A, Liu F, Ye D, Ding Q,
Qin C, Yin C, Zhang Z, Matsuda K, Kubo M, et al: Large-scale
association analysis in Asians identifies new susceptibility loci
for prostate cancer. Nat Commun. 6:84692015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
1000 Genomes Project Consortium, . Auton
A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, Marchini
JL, McCarthy S, McVean GA and Abecasis GR: A global reference for
human genetic variation. Nature. 526:68–74. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Purcell S, Neale B, Todd-Brown K, Thomas
L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ
and Sham PC: PLINK: A tool set for whole-genome association and
population-based linkage analyses. Am J Hum Genet. 81:559–575.
2007. View
Article : Google Scholar : PubMed/NCBI
|
|
19
|
DeLong ER, DeLong DM and Clarke-Pearson
DL: Comparing the areas under two or more correlated receiver
operating characteristic curves: A nonparametric approach.
Biometrics. 44:837–845. 1988. View
Article : Google Scholar : PubMed/NCBI
|
|
20
|
Na R, Ye D, Qi J, Liu F, Lin X, Helfand
BT, Brendler CB, Conran C, Gong J, Wu Y, et al: Race-specific
genetic risk score is more accurate than nonrace-specific genetic
risk score for predicting prostate cancer and high-grade diseases.
Asian J Androl. 18:525–529. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lichtenstein P, Holm NV, Verkasalo PK,
Iliadou A, Kaprio J, Koskenvuo M, Pukkala E, Skytthe A and Hemminki
K: Environmental and heritable factors in the causation of
cancer-analyses of cohorts of twins from Sweden, Denmark, and
Finland. N Engl J Med. 343:78–85. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Michailidou K, Hall P, Gonzalez-Neira A,
Ghoussaini M, Dennis J, Milne RL, Schmidt MK, Chang-Claude J,
Bojesen SE, Bolla MK, et al: Large-scale genotyping identifies 41
new loci associated with breast cancer risk. Nat Genet. 45:353–361.
2013. View
Article : Google Scholar : PubMed/NCBI
|