Introduction
Glioma is the most aggressive type of adult brain
cancer (1,2). Despite research efforts, the average
lifespan for glioma patients postdiagnosis is ~15 months, with the
majority of patients experiencing tumor relapse and outgrowth
within seven months of initial radiation therapy (3,4).
Therefore, it is important to understanding the pathological
mechanisms for tumor initiation and progression.
MicroRNA (miR) is a small non-coding RNA molecule,
which functions in the transcriptional and posttranscriptional
regulation of gene expression. Several recent studies have
demonstrated that miRs have a critical role in cell proliferation,
apoptosis and metastasis. Accumulating evidence has demonstrated
the involvement of microRNAs in cancerous processes as either
oncogenes or tumor suppressor genes (5,6).
Early investigations demonstrated that miR-181 downregulated the
homeobox protein Hox-A11, a repressor of the differentiation
process, which revealed the existence of a functional correlation
between miR-181 and mammalian skeletal-muscle differentiation
(7). Subsequently, it was
identified that the expression levels of miR-181 were inversely
correlated with Tcl1 oncogene expression in B-cell chronic
lymphocytic leukemia samples (8).
Furthermore, miR-181 was reported to function as a tumor
suppressor, which triggered growth inhibition, induced apoptosis
and inhibited invasion in multiple tumor types, including breast,
colon and hepatocellular carcinoma (9–12).
However, whether miR-181 is involved in the development of glioma
remains largely unknown. Thus, the present study aimed to
investigate the role of miR-181 in glioma cell proliferation.
Materials and methods
Cell culture and tissue samples
Glioma cells (U251 and SHG-44) were obtained from
the American Type Culture Collection (Rockville, MD, USA). The
cells were cultured in Dulbecco’s modified Eagle’s medium (Sigma,
St. Louis, MO, USA) supplemented with 10% fetal bovine serum. The
cultures were maintained at 37°C in a humidified atmosphere with 5%
CO2. The tumor tissues and adjacent normal non-tumor
tissues were collected from routine therapeutic surgery at the
Department of Thoracic Surgery, Provincial Hospital Affiliated to
Shandong University (Jinan, China). All of the samples were
obtained with informed consent and the present study was approved
by the Ethics Committee of the Provincial Hospital Affiliated to
Shandong University.
Analysis of miRNA expression using TaqMan
reverse transcription polymerase chain reaction (RT-PCR)
Total RNA from tissue samples and the cell lines was
harvested using the miRNA Isolation kit (Ambion, Austin, TX, USA).
The expression of mature miRNAs was assayed using a Taqman MicroRNA
assay (Applied Biosystems, Shanghai, China) specific for
hsa-miR-181. Briefly, 10 ng of total RNA were reverse transcribed
to cDNA with the following specific stem-loop RT primers: Forward,
5′-UGGAAGGACGGGAAGUGGAA-3′ and reverse, 5′-CCAGUGCAGGGUCCGAGGUA-3′.
Quantitative (q)PCR was performed using an Applied Biosystems 7900
Real-time PCR system and a TaqMan Universal PCR Master mix (Applied
Biosystems). All of the primers were obtained from the TaqMan miRNA
assays. Small nuclear U6 snRNA (Applied Biosystems) was used as an
internal control.
Plasmid construction and
transfection
For the miR-181 expression plasmid, the human
miR-181 precursor was cloned into pSilencer 4.1 (Ambion). The
negative control plasmid consisted of a scrambled sequence
(Ambion). To inhibit miR-181 function, an Ambion miRNA inhibitor
for miR-181 was used, along with the negative control. For
transfection, a complex of Lipofectamine® 2000
(Invitrogen Life Technologies, Carlsbad, CA, USA) and 25 nM miRNA
mentioned above was prepared according to the manufacturer’s
instructions.
BrdU assays
A cell proliferation enzyme-linked immunosorbent
assay (BrdU kit; Beyotime, Nantong, China) was used to analyze the
incorporation of BrdU during DNA synthesis according to the
manufacturer’s instructions. All of the experiments were performed
in triplicate. Absorbance was measured at 450 nm in the Spectra Max
190 ELISA reader (Molecular Devices, Sunnyvale, CA, USA)
Western blotting
The cells or tissues were harvested and lysed with
ice-cold lysis buffer (50 mM Tris-HCl, pH 6.8; 100 mM 2-ME, 2% w/v
SDS and 10% glycerol). Following centrifugation at 20,000 × g for
10 min at 4°C, the proteins in the supernatants were quantified and
separated by 10% SDS PAGE, and transferred onto a nitrocellulose
membrane (Amersham Bioscience, Buckinghamshire, UK). Following
blocking with 10% non-fat milk in PBS, the membranes were
immunoblotted with cyclin B1 (Santa Cruz Biotechnology, Inc., Santa
Cruz, CA, USA) and GAPDH (Abcam, Cambridge, MA, USA) antibodies,
followed by anti-rabbit horseradish peroxidase-linked secondary
antibodies (Cell Signaling Technology, Inc., Beverly, MA, USA). The
signals were detected by a SuperSignal West Pico Chemiluminescent
Substrate kit (Pierce Biotechnology, Inc., Rockford, IL, USA)
according to manufacturer’s instructions. Anti-cyclin B1 antibodies
were purchased from Cell Signaling Technology, Inc.. The protein
levels were normalized to total GAPDH, using a mouse anti-GAPDH
antibody (Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA).
Luciferase reporter assay
Total cDNA from the U251 cells was used to amplify
the 3′ untranslated region (UTR) of cyclin B1 by PCR. The cyclin B1
3′UTR was cloned into pMir-Report (Ambion), yielding
pMir-Report-cyclin B1. Mutations were introduced in potential
miR-181 binding sites using the QuikChange site-directed
mutagenesis kit (Stratagene, La Jolla, CA, USA). The cells were
transfected with the 3′-UTR luciferase reporter and the miR-181
precursor plasmids for 36 h. The pRL-SV40 vector (Promega
Corporation, Madison, WI, USA) carrying the Renilla
luciferase gene was used as an internal control to normalize the
transfection efficiency. The luciferase values were determined
using the Dual-Luciferase Reporter Assay system (Promega
Corporation).
Statistical analysis
Data are expressed as the mean ± standard error of
the mean from at least three separate experiments. The differences
between the groups were analyzed using Student’s t-test. P<0.05
was considered to indicate a statistically significant
difference.
Results
miR-181 expression levels are
downregulated in patients with glioma
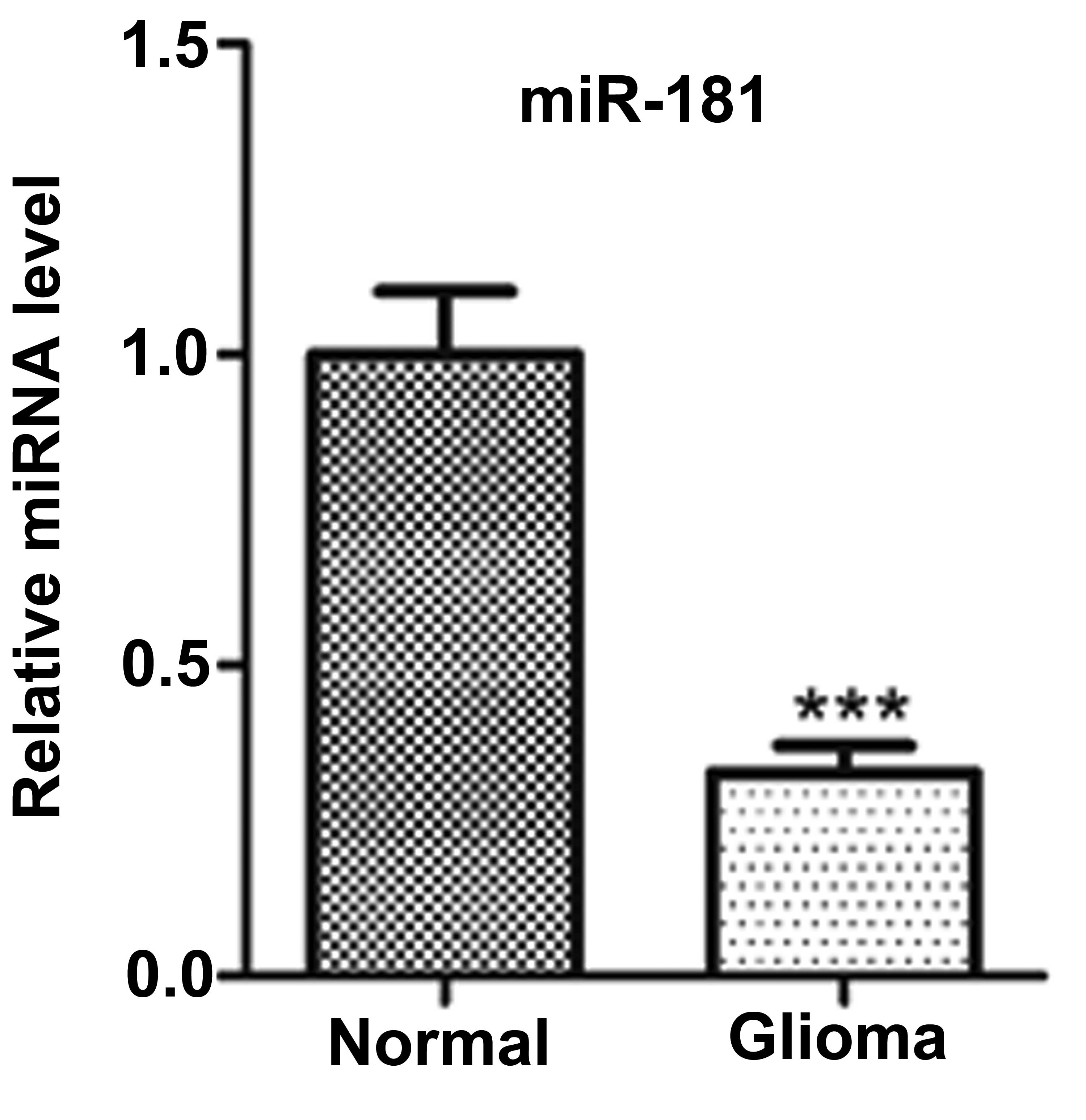
Firstly, to examine whether the miR-181 is
differentially expressed in human glioma, its expression level was
determined using TaqMan qPCR in 30 pairs of human glioma tissues
and pair-matched adjacent non-cancerous tissues. The results
demonstrated that the expression level of miR-181 was significantly
decreased in glioma tissues compared with the adjacent
non-cancerous tissues (Fig.
1).
miR-181 overexpression inhibits cell
proliferation
In order to assess the effects of miR-181 on glioma
cell growth, the miR-181 precursor was transfected into the U251
and SHG-44 cells, and cell growth post-transfection was examined.
The miR-181 precursor was found to upregulate miR-181 expression
(Fig. 2A and B), significantly
reduce the cell number and inhibit the proliferation of cells
post-transfection (Fig. 2C–F).
Inhibition of miR-181 promotes the
proliferation of glioma cells
As described above, miR-181 has a critical role in
the proliferation of glioma cells. However, whether inhibiting
miR-181 enhances cell proliferation is unclear. Therefore, the two
cell lines were transfected with antisense miR-181 and it was
revealed that the ectopic expression of antisense hsa-miR-181
promoted the growth of U251 and SHG-44 cells, compared with that of
the NC-transfected cells (Fig.
3A–D).
miR-181 directly targets cyclin B1 in
glioma cells
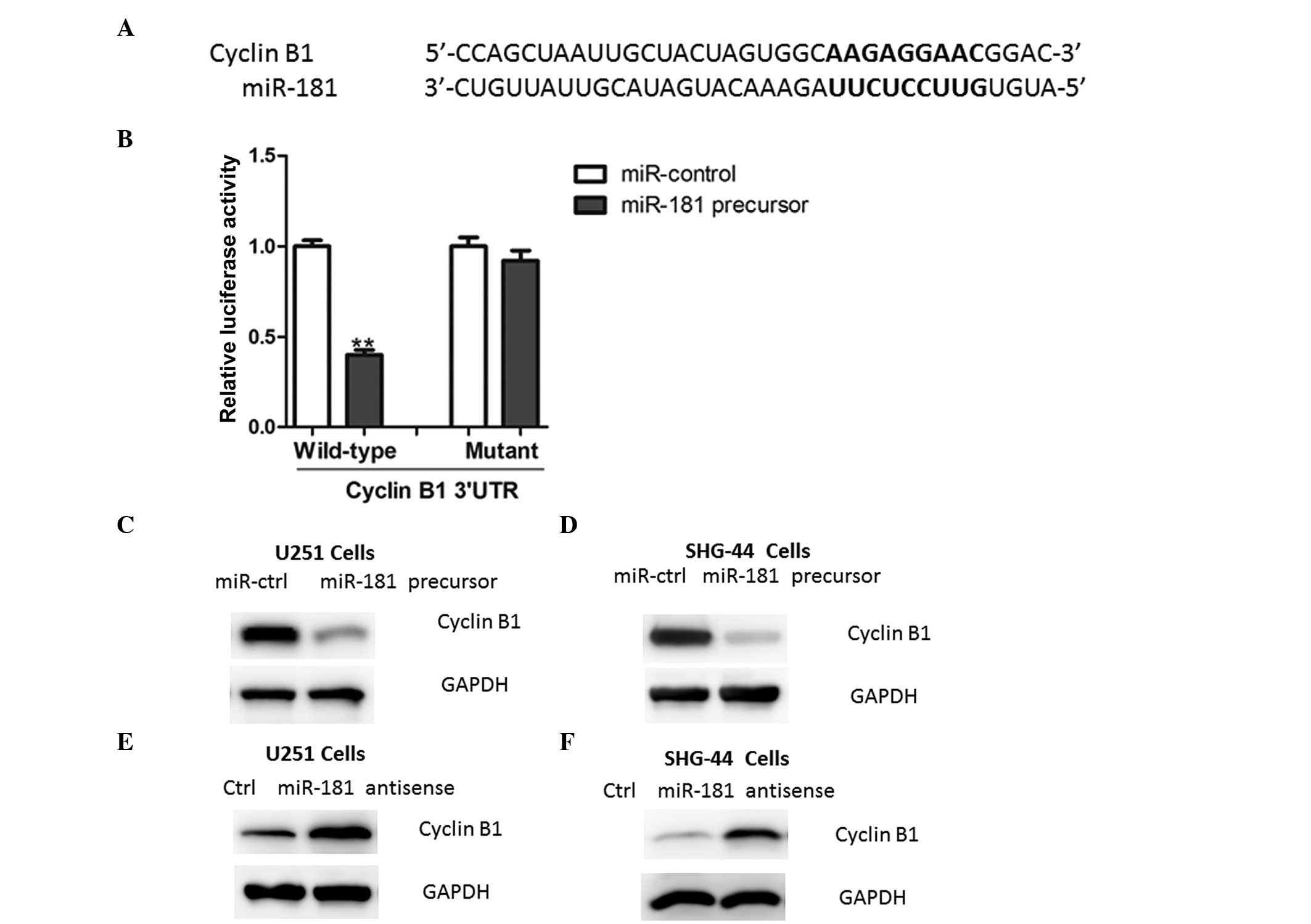
Using a stringent bioinformatics approach (miRWalk
software, Heidelberg, Germany; http://www.umm.uni-heidelberg.de/apps/zmf/mirwalk/),
24 putative human miR-181 target genes were identified (data not
shown), among which the gene encoding cyclin B1 harbored a
potential miR-181 binding site (Fig.
4A). The overexpression of miR-181 led to a reduction of
luciferase activity when the reporter construct contained the
cyclin B1 3′UTR (Fig. 4B). By
contrast, mutations in the conserved miR-181 binding motif
abrogated the reduced luciferase expression (Fig. 4B). Furthermore, the overexpression
of miR-181 in glioma cells led to reduced cyclin B1 protein
expression (Fig. 4C–D).
Consistently, the inhibition of miR-181 led to an increased
expression of cyclin B1 (Fig.
4E–F), further indicating that cyclin B1 is a target of miR-181
in glioma cells.
Discussion
In the present study, it was demonstrated that
miR-181 expression is downregulated in glioma tissues. To the best
of our knowledge, the present study was the first to identify at a
molecular level that miR-181 regulated cyclin B1 expression by
targeting its 3′UTR. Collectively, these findings suggest that the
downregulation of miR-181 may promote the initiation and
progression of glioma. Notably, a recent study demonstrated that
transiently overexpressed miR-181 significantly sensitized
malignant glioma cells to radiation treatment, which was concurrent
with the downregulation of B cell lymphoma/leukemia-2 (Bcl-2)
protein expression (13). This
indicates that miR-181 may modulate radiosensitivity by targeting
Bcl-2 in human malignant glioma cells (13), suggesting that miR-181 may be a
target for enhancing the effect of radiation treatment on malignant
glioma cells. Therefore, the precise roles of miR-181 may be
diverse in glioma cells.
It has been reported that several miRNAs were
misregulated in glioma tissues or cells (14,15).
For example, miR-92b controls glioma proliferation and invasion by
regulating Wnt/β-catenin signaling via Nemo-like kinase (16). In addition, miR-200b targets CREB1
and suppresses cell growth in human malignant glioma (17). Furthermore, the downregulation of
miR-383 promotes glioma cell invasion by targeting insulin-like
growth factor 1 receptor (18). By
contrast, miR-107 inhibits U87 glioma stem cell growth and invasion
by modulating Notch2 expression (19,20).
Therefore, miRNA expression appears to have a key role in
regulating cellular processes in glioma, which requires further
investigation in the future.
In conclusion, the key finding of the present study
is that miR-181 is able to promote the proliferation of glioma cell
lines by targeting cyclin B1. This data indicates that miR-181 has
an essential role in the regulation of glioma cell proliferation
and may function as a tumor suppressor.
References
|
1
|
Wang Y and Jiang T: Understanding high
grade glioma: molecular mechanism, therapy and comprehensive
management. Cancer Lett. 331:139–146. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chaudhry NS, Shah AH, Ferraro N, Snelling
BM, Bregy A, Madhavan K and Komotar RJ: Predictors of long-term
survival in patients with glioblastoma multiforme: advancements
from the last quarter century. Cancer Invest. 31:287–308. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hirst TC, Vesterinen HM, Sena ES, Egan KJ,
Macleod MR and Whittle IR: Systematic review and meta-analysis of
temozolomide in animal models of glioma: was clinical efficacy
predicted? Br J Cancer. 108:64–71. 2013. View Article : Google Scholar
|
|
4
|
Marsh JC, Goldfarb J, Shafman TD and Diaz
AZ: Current status of immunotherapy and gene therapy for high-grade
gliomas. Cancer Control. 20:43–48. 2013.PubMed/NCBI
|
|
5
|
Pritchard CC, Cheng HH and Tewari M:
MicroRNA profiling: approaches and considerations. Nat Rev Genet.
13:358–369. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kasinski AL and Slack FJ: Epigenetics and
genetics. MicroRNAs en route to the clinic: progress in validating
and targeting microRNAs for cancer therapy. Nat Rev Cancer.
11:849–864. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Naguibneva I, Ameyar-Zazoua M, Polesskaya
A, Ait-Si-Ali S, Groisman R, Souidi M, Cuvellier S and Harel-Bellan
A: The microRNA miR-181 targets the homeobox protein Hox-A11 during
mammalian myoblast differentiation. Nat Cell Biol. 8:278–284. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Pekarsky Y, Santanam U, Cimmino A,
Palamarchuk A, Efanov A, Maximov V, Volinia S, Alder H, Liu CG,
Rassenti L, Calin GA, Hagan JP, Kipps T and Croce CM: Tcl1
expression in chronic lymphocytic leukemia is regulated by miR-29
and miR-181. Cancer Res. 66:11590–11593. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Neel JC and Lebrun JJ: Activin and TGFβ
regulate expression of the microRNA-181 family to promote cell
migration and invasion in breast cancer cells. Cell Signal.
25:1556–1566. 2013.
|
|
10
|
Kim CH, Kim HK, Rettig RL, Kim J, Lee ET,
Aprelikova O, Choi IJ, Munroe DJ and Green JE: miRNA signature
associated with outcome of gastric cancer patients following
chemotherapy. BMC Med Genomics. 4:792011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ji J, Yamashita T, Budhu A, Forgues M, Jia
HL, Li C, Deng C, Wauthier E, Reid LM, Ye QH, Qin LX, Yang W, Wang
HY, Tang ZY, Croce CM and Wang XW: Identification of microRNA-181
by genome-wide screening as a critical player in EpCAM-positive
hepatic cancer stem cells. Hepatology. 50:472–480. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang B, Hsu SH, Majumder S, Kutay H, Huang
W, Jacob ST and Ghoshal K: TGFbeta-mediated upregulation of hepatic
miR-181b promotes hepatocarcinogenesis by targeting TIMP3.
Oncogene. 9:1787–1797. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen G, Zhu W, Shi D, Lv L, Zhang C, Liu P
and Hu W: MicroRNA-181a sensitizes human malignant glioma U87MG
cells to radiation by targeting Bcl-2. Oncol Rep. 23:997–1003.
2010.PubMed/NCBI
|
|
14
|
Goodenberger ML and Jenkins RB: Genetics
of adult glioma. Cancer Genet. 205:613–621. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Odjélé A, Charest D and Morin P Jr: miRNAs
as important drivers of glioblastomas: a no-brainer? Cancer
Biomark. 11:245–252. 2012.PubMed/NCBI
|
|
16
|
Wang K, Wang X, Zou J, Zhang A, Wan Y, Pu
P, Song Z, Qian C, Chen Y, Yang S and Wang Y: miR-92b controls
glioma proliferation and invasion through regulating
Wnt/beta-catenin signaling via Nemo-like kinase. Neuro Oncol.
15:578–588. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Peng B, Hu S, Jun Q, Luo D, Zhang X, Zhao
H and Li D: MicroRNA-200b targets CREB1 and suppresses cell growth
in human malignant glioma. Mol Cell Biochem. 379:51–58. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
He Z, Cen D, Luo X, Li D, Li P, Liang L
and Meng Z: Downregulation of miR-383 promotes glioma cell invasion
by targeting insulin-like growth factor 1 receptor. Med Oncol.
30:5572013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Chen L, Chen XR, Zhang R, Li P, Liu Y, Yan
K and Jiang XD: MicroRNA-107 inhibits glioma cell migration and
invasion by modulating Notch2 expression. J Neurooncol. 112:59–66.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen L, Chen XR, Chen FF, Liu Y, Li P,
Zhang R, Yan K, Yi YJ, Xu ZM and Jiang XD: MicroRNA-107 inhibits
U87 glioma stem cells growth and invasion. Cell Mol Neurobiol.
33:651–657. 2013. View Article : Google Scholar : PubMed/NCBI
|