|
1
|
Williams JC, Barratt-Boyes BG and Lowe JB:
Supravalvular aortic stenosis. Circulation. 24:1311–1318. 1961.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Beuren AJ, Apitz J and Harmjanz D:
Supravalvular aortic stenosis in association with mental
retardation and a certain facial appearance. Circulation.
26:1235–1240. 1962. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wuang YP and Tsai HY: Sensorimotor and
visual perceptual functioning in school-aged children with Williams
syndrome. J Intellect Disabil Res. Nov 30–2016.(Epub ahead of
print). PubMed/NCBI
|
|
4
|
Sammour ZM, de Bessa J Jr, Hisano M,
Bruschini H, Kim CA, Srougi M and Gomes CM: Lower urinary tract
symptoms in children and adolescents with Williams-Beuren syndrome.
J Pediatr Urol. Nov 2–2016.(Epub ahead of print). View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Hirai M, Muramatsu Y, Mizuno S, Kurahashi
N, Kurahashi H and Nakamura M: Typical visual search performance
and atypical gaze behaviors in response to faces in Williams
syndrome. J Neurodev Disord. 8:382016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Strømme P, Bjørnstad PG and Ramstad K:
Prevalence estimation of Williams syndrome. J Child Neurol.
17:269–271. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Abbas E, Cox DM, Smith T and Butler MG:
The 7q11.23 microduplication syndrome: A clinical report with
review of literature. J Pediatr Genet. 5:129–140. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Barak B and Feng G: Neurobiology of social
behavior abnormalities in autism and Williams syndrome. Nat
Neurosci. 19:647–655. 2016. View
Article : Google Scholar : PubMed/NCBI
|
|
9
|
Pinto D, Darvishi K, Shi X, Rajan D,
Rigler D, Fitzgerald T, Lionel AC, Thiruvahindrapuram B, Macdonald
JR, Mills R, et al: Comprehensive assessment of array-based
platforms and calling algorithms for detection of copy number
variants. Nat Biotechnol. 29:512–520. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Falkmer T, Anderson K, Falkmer M and
Horlin C: Diagnostic procedures in autism spectrum disorders: A
systematic literature review. Eur Child Adolesc Psychiatry.
22:329–340. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
de Bildt A, Sytema S, Zander E, Bölte S,
Sturm H, Yirmiya N, Yaari M, Charman T, Salomone E, LeCouteur A, et
al: Autism diagnostic interview-revised (adi-r) algorithms for
toddlers and young preschoolers: Application in a non-us sample of
1,104 children. J Autism Dev Disord. 45:2076–2091. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zander E, Willfors C, Berggren S,
Choque-Olsson N, Coco C, Elmund A, Moretti ÅH, Holm A, Jifält I,
Kosieradzki R, et al: The objectivity of the autism diagnostic
observation schedule (ADOS) in naturalistic clinical settings. Eur
Child Adolesc Psychiatry. 25:769–780. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yu TY, Chen KL, Chou W, Yang SH, Kung SC,
Lee YC and Tung LC: Intelligence quotient discrepancy indicates
levels of motor competence in preschool children at risk for
developmental delays. Neuropsychiatr Dis Treat. 12:501–510.
2016.PubMed/NCBI
|
|
14
|
Trumpff C, De Schepper J, Vanderfaeillie
J, Vercruysse N, Van Oyen H, Moreno-Reyes R, Tafforeau J, Vanderpas
J and Vandevijvere S: Thyroid-stimulating hormone (TSH)
concentration at birth in belgian neonates and cognitive
development at preschool age. Nutrients. 7:9018–9032. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Goud TM, Al Salmani KK, Al Harasi SM, Al
Musalhi M, Wasifuddin SM and Rajab A: Importance of FISH combined
with morphology, immunophenotype and cytogenetic analysis of
childhood/adult acute lymphoblastic leukemia in Omani patients.
Asian Pac J Cancer Prev. 16:7343–7350. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Peterson JF, Aggarwal N, Smith CA, Gollin
SM, Surti U, Rajkovic A, Swerdlow SH and Yatsenko SA: Integration
of microarray analysis into the clinical diagnosis of hematological
malignancies: How much can we improve cytogenetic testing.
Oncotarget. 6:18845–18862. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sun G, Tan Z, Fan L, Wang J, Yang Y and
Zhang W: 1q21.1 microduplication in a patient with mental
impairment and congenital heart defect. Mol Med Rep. 12:5655–5658.
2015.PubMed/NCBI
|
|
18
|
MacDonald JR, Ziman R, Yuen RK, Feuk L and
Scherer SW: The database of genomic variants: A curated collection
of structural variation in the human genome. Nucleic Acids Res.
42:D986–D992. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Reiss AL, Feinstein C, Rosenbaum KN,
Borengasser-Caruso MA and Goldsmith BM: Autism associated with
Williams syndrome. J Pediatr. 106:247–249. 1985. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Gillberg C and Rasmussen P: Brief report:
Four case histories and a literature review of Williams syndrome
and autistic behavior. J Autism Dev Disord. 24:381–393. 1994.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Gosch A and Pankau R: Personality
characteristics and behaviour problems in individuals of different
ages with Williams syndrome. Dev Med Child Neurol. 39:527–533.
1997. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Herguner S and Mukaddes NM: Autism and
Williams syndrome: A case report. World J Biol Psychiatry.
7:186–188. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Leyfer OT, Woodruff-Borden J, Klein-Tasman
BP, Fricke JS and Mervis CB: Prevalence of psychiatric disorders in
4 to 16-year-olds with Williams syndrome. Am J Med Genet B
Neuropsychiatr Genet. 141B:615–622. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
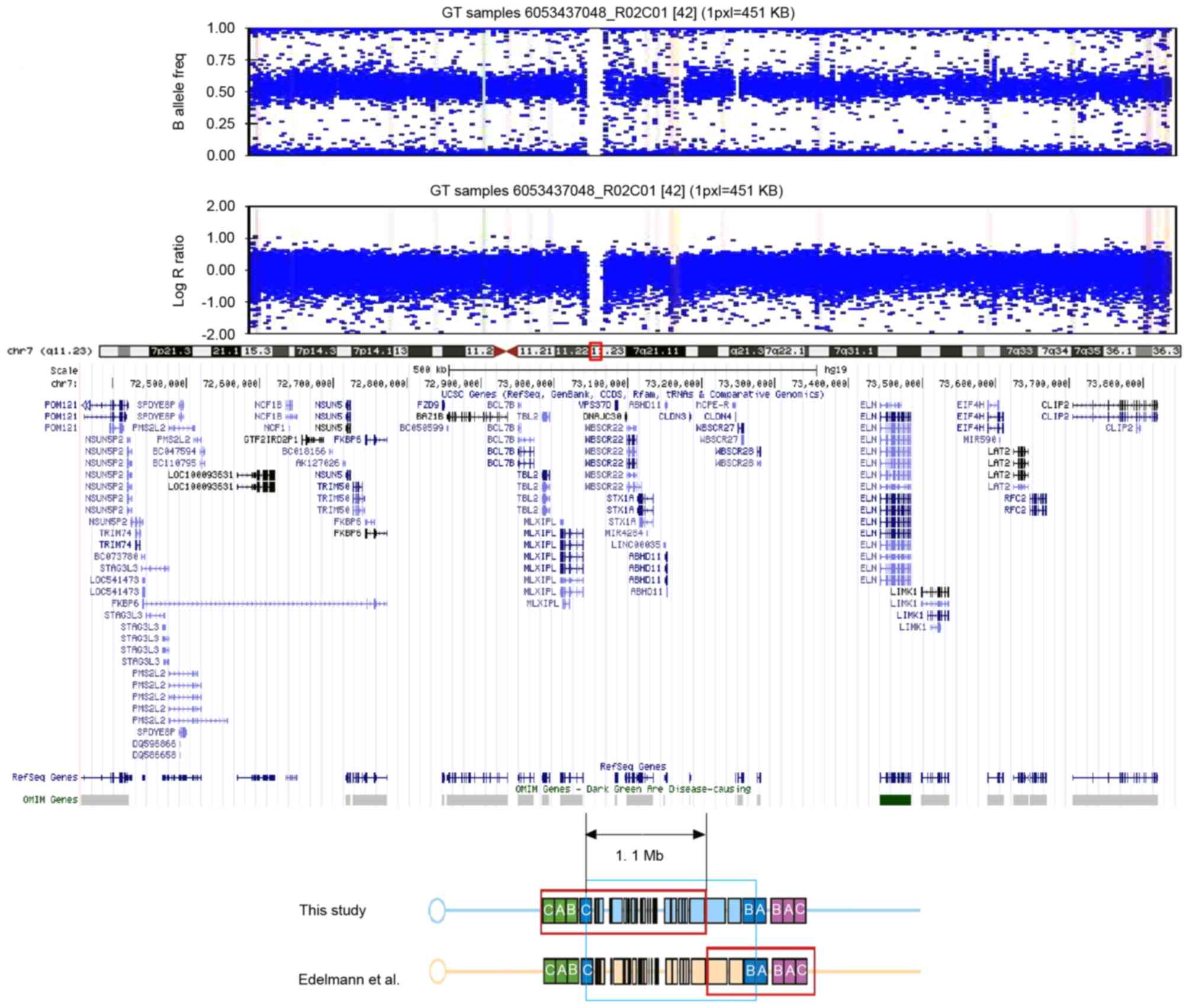
Edelmann L, Prosnitz A, Pardo S, Bhatt J,
Cohen N, Lauriat T, Ouchanov L, González PJ, Manghi ER, Bondy P, et
al: An atypical deletion of the Williams-Beuren syndrome interval
implicates genes associated with defective visuospatial processing
and autism. J Med Genet. 44:136–143. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Lincoln AJ, Searcy YM, Jones W and Lord C:
Social interaction behaviors discriminate young children with
autism and Williams syndrome. J Am Acad Child Adolesc Psychiatry.
46:323–331. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Klein-Tasman BP, Mervis CB, Lord C and
Phillips KD: Socio-communicative deficits in young children with
Williams syndrome: Performance on the autism diagnostic observation
Schedule. Child Neuropsychol. 13:444–467. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tordjman S, Anderson GM, Botbol M, Toutain
A, Sarda P, Carlier M, Saugier-Veber P, Baumann C, Cohen D,
Lagneaux C, et al: Autistic disorder in patients with
Williams-Beuren syndrome: A reconsideration of the Williams-Beuren
syndrome phenotype. PLoS One. 7:e307782012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Tordjman S, Anderson GM, Cohen D,
Kermarrec S, Carlier M, Touitou Y, Saugier-Veber P, Lagneaux C,
Chevreuil C and Verloes A: Presence of autism, hyperserotonemia,
and severe expressive language impairment in Williams-Beuren
syndrome. Mol Autism. 4:292013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Sanders SJ, Ercan-Sencicek AG, Hus V, Luo
R, Murtha MT, Moreno-De-Luca D, Chu SH, Moreau MP, Gupta AR,
Thomson SA, et al: Multiple recurrent de novo CNVs, including
duplications of the 7q11.23 Williams syndrome region, are strongly
associated with autism. Neuron. 70:863–885. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Curran ME, Atkinson DL, Ewart AK, Morris
CA, Leppert MF and Keating MT: The elastin gene is disrupted by a
translocation associated with supravalvular aortic stenosis. Cell.
73:159–168. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Pober BR: Williams-Beuren syndrome. N Engl
J Med. 362:239–252. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Gagliardi C, Bonaglia MC, Selicorni A,
Borgatti R and Giorda R: Unusual cognitive and behavioural profile
in a Williams syndrome patient with atypical 7q11.23 deletion. J
Med Genet. 40:526–530. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Hirota H, Matsuoka R, Chen XN, Salandanan
LS, Lincoln A, Rose FE, Sunahara M, Osawa M, Bellugi U and
Korenberg JR: Williams syndrome deficits in visual spatial
processing linked to GTF2IRD1 and GTF2I on chromosome 7q11.23.
Genet Med. 5:311–321. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Morris CA, Mervis CB, Hobart HH, Gregg RG,
Bertrand J, Ensing GJ, Sommer A, Moore CA, Hopkin RJ, Spallone PA,
et al: GTF2I hemizygosity implicated in mental retardation in
Williams syndrome: Genotype-phenotype analysis of five families
with deletions in the Williams syndrome region. Am J Med Genet A.
123A:45–59. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Pankau R, Gosch A, Simeoni E and Wessel A:
Williams-Beuren syndrome in monozygotic twins with variable
expression. Am J Med Genet. 47:475–477. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Blanco-Dávila F and Olveda-Rodriguez JA:
Cleft palate in a patient with Williams' syndrome. J Craniofac
Surg. 12:145–147. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Vincent C, Mercier JM and David A: Cleft
palate and Williams syndrome. Rev Stomatol Chir Maxillofac.
109:44–47. 2008.(In French). View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Domenico S, Orlando C, Graziana FF, Papi P
and Giulia A: Cleft palate in Williams syndrome. Ann Maxillofac
Surg. 3:84–86. 2013. View Article : Google Scholar : PubMed/NCBI
|