Introduction
Congenital heart disease (CHD) is one of the most
common birth defects in humans worldwide, occurring in ~7–8% of
infants born annually (1).
Improved surgical treatments have decreased the mortality of
children with CHD, but not all infants with CHD survive to
adulthood (2,3). The mechanism of heart development is
complex, and involves multiple genetic and environmental factors in
its regulation (4). Therefore,
investigations of relevant target genes and microRNAs (miRNAs) are
important to fully comprehend the mechanism of heart development
and to provide therapeutic targets for treatment of CHD in
children.
miRNAs are endogenous non-coding RNAs of 20–22
nucleotides in length that have diverse functions in biological
processes at the transcriptional or post-transcriptional level, by
targeting the 3′-UTR of genes (5).
Previous studies have revealed that miRNAs are involved in the
development of the heart, including cardiomyocyte differentiation,
cell cycle and the conducting system of the heart (6,7). The
muscle-specific miR-1 has been reported to be important in heart
development. For example, miR-1 transcription is affected by the
regulation of myogenic differentiation 1 (MyoD), myocyte enhancer
factor (Mef), and serum response factor (SRF) (8). High levels of miR-1 expression in
mice lead to cardiomyocyte cell cycle arrest at an early stage and
attenuated cell proliferation (9,10).
The mouse teratoma-derived P19 cells are pluripotent, and thus can
be differentiated into cardiomyocytes, skeletal muscle cells and
neurons, allowing the application of P19 cells in cell replacement
therapy and myocardial tissue engineering (11,12).
Although several studies have reported significant roles of miR-1
in regulating heart development in mice and in human cells, few
have explored the possible role of miR-1 in regulating the heart
development in P19 cells.
The present study aimed to investigate the role of
miR-1 in heart development and to reveal a possible mechanism of
action. Endogenous expression of miR-1 was assessed in P19 cells,
as was the effect of miR-1 overexpression on the biological
processes of P19 differentiated cardiomyocytes. In addition, the
effect of miR-1 overexpression on cell viability and cell
apoptosis-related protein expression was examined.
Materials and methods
Cell culture and cell
differentiation
P19 cells (American Type Culture Collection,
Manassas, VA, USA) were cultured in Gibco α-modified Eagle's medium
(α-MEM; Thermo Fisher Scientific, Inc., Waltham, MA, USA)
supplemented with 10% Gibco fetal bovine serum (FBS; Thermo Fisher
Scientific, Inc.), 100 U/ml penicillin, and 100 µg/ml streptomycin
at 37°C in 5% CO2.
For the cardiac differentiation assay (13), 1×106 cells/ml P19 cells
were plated onto 10 cm bacterial dishes in 15 ml α-MEM containing
1% dimethyl sulphoxide (DMSO; Sigma-Aldrich; Merck Millipore,
Darmstadt, Germany), 10% FBS, 100 U/ml penicillin, and 100 µg/ml
streptomycin at 37°C in 5% CO2. Following 96 h of
incubation, cells were transferred onto 6 cm bacterial dishes and
cultured in α-MEM containing 10% FBS for another 6 days of
incubation.
Cell transfection
The miRNA mimic (Gene ID: 100314077; Sangon Biotech
Co., Ltd., Shanghai, China) was transfected into the differentiated
P19 cells using the Lipofectamine® 2000 protocol (Thermo
Fisher Scientific, Inc.). Cells transfected with the scrambled RNA
(catalog no. CS7005; Sangon Biotech, Co., Ltd.) were used as the
control.
Cell proliferation assay
Cell proliferation ability was assessed using the
3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyltetrazolium bromide (MTT)
assay, as previously described (14). Briefly, following transfection for
24 h, 5×103 cells were seeded into 96-well plates.
Following 24 h of incubation, cells cultured at 37°C were
centrifuged at 4,000 × g for 5 min, and the supernatant was
removed. MTT (20 µl) was added into the cells and then cultured for
another 4 h. Finally, 150 µl DMSO was mixed with the cells for 10
min to stop the reaction at room temperature. Absorbance of cells
in each well was observed at 570 nm with an absorption
spectrophotometer (Olympus Corporation, Tokyo, Japan).
Cell apoptosis assay
Apoptotic cells were measured using flow cytometry
following staining with the Annexin V-FITC apoptosis kit
(Invitrogen; Thermo Fisher Scientific, Inc.), according to the
manufacturer's protocol. Briefly, following transfection for 36 h,
cells were cultured in fresh serum-free α-MEM medium for 12 h. Then
total cells were harvested and washed 3 times with PBS buffer,
followed by resuspension in the kit staining buffer. Then, 5 µl of
Annexin V-FITC and 5 µl of propidium iodide (PI) were added into
the cells at room temperature for 10 min. Mixtures were analyzed
using FACS can flow cytometry (BD Biosciences, Franklin Lakes, NJ,
USA). The number of early-stage apoptotic cells (Annexin
V+ and PI− cells) was then analyzed.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Extraction of total RNA from cells was performed
using TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.),
according to the manufacturer's protocol. The extracted RNA was
treated with RNase-free Dnase I (Promega Corporation, Madison, WI,
USA) to remove the contaminating DNA, and concentration and purity
were measured using SMA 400 UV-VIS (Merinton, Shanghai, China).
Purified RNA dissolved in nuclease-free water at a concentration of
0.5 µg/µl was used for cDNA synthesis with the PrimerScript 1st
Strand cDNA Synthesis kit (Invitrogen; Thermo Fisher Scientific,
Inc.). Expression of targets was analyzed using an ABI 7900 (PE
Applied Biosystems; Thermo Fisher Scientific, Inc.) and the SYBR
ExScript RT-PCR kit (Takara Biotechnology Co., Ltd., Dalian,
China). GAPDH was selected as the internal control for target gene
expression, U6 small nuclear RNA (U6) was used as the internal
control for the miRNA expression. Primers used for target
amplification are listed in Table
I (15).
 | Table I.Primers used for target
amplification. |
Table I.
Primers used for target
amplification.
| Gene target | Sequence (5′-3′) |
|---|
| GAPDH |
F-GGGTGGAGCCAAACGGGTC |
|
|
R-GGAGTTGCTGTTGAAGTCGCA |
| GATA4 |
F-CCTGCGGCCTCTACATGA |
|
|
R-AGGGTCTCACCAGCAGGA |
| Nkx2-5 |
F-CCTGCGGCCTCTACATGA |
|
|
R-AGGGTCTCACCAGCAGGA |
| Hand2 |
F-TACCAGCTACATCGCCTACCT |
|
|
R-TCACTGCTTGAGCTCCAGGG |
| Caspase-3 |
F-TACCCTGAAATGGGCTTGTGT |
|
|
R-GTTAACACGAGTGAGGATGTG |
| miR-1 |
F-GTAGGCACCTGAAATGGAA |
|
|
R-TTGATGGTGCCTACAGTACAT |
| U6 |
F-CGCTTCACGAATTTGCGTGTCAT |
|
|
R-AACGCTTCACGAATTTGCGT |
Western blotting
Cells were lysed with radioimmunoprecipitation assay
buffer at 4°C for 5 min (RIPA; Sangon Biotech Co., Ltd.) containing
phenylmethylsulfonyl fluoride (PMSF; Sigma-Aldrich; Merck
Millipore), and then were centrifuged at 4,000 × g at 4°C for 10
min. Protein concentration was detected using a bicinchoninic acid
assay kit (Pierce; Thermo Fisher Scientific, Inc.). For western
blotting, total protein (30 µl) was subjected to 12% SDS-PAGE,
followed by transfer onto a polyvinylidenefluoride (PVDF) membrane.
The PVDF membranes were blocked with TBS/0.1% Tween-20 (TBST)
buffer containing 5% non-fat milk at room temperature for 1 h. Then
the membranes were incubated with rabbit primary antibodies against
heart and neural crest derivatives expressed 2 (Hand 2; 1:100;
catalog no. ab10131; Abcam, Cambridge, MA, USA), caspase-3 (1:100;
catalog no. ab2171), cleaved caspase-3 (1:100; catalog no. ab13585)
or GAPDH (1:100; catalog no. ab8245) obtained from Invitrogen;
Thermo Fisher Scientific, Inc., overnight at 4°C, then
horseradish-peroxidase labeled goat anti-rat secondary antibody
(catalog no. ab7097; 1:1,000; Abcam) at room temperature for 1 h.
Finally, the PVDF membranes were washed 3 times with TBST buffer
for 10 min each wash. Signals were detected following incubation
with a chromogenic substrate using an enhanced chemiluminescence
kit (Sigma-Aldrich; Merck Millipore). GAPDH served as the internal
control.
Statistical analysis
Data are presented as the mean ± standard deviation
of 3 independent replicates. Statistical analysis between two
groups was performed using a t-test, whereas the multiple
comparisons were analyzed by post-hoc tests that followed one-way
analysis of variance. All significant differences were analyzed
using SPSS 19.0 statistical software (IBM SPSS, Armonk, NY, USA).
P<0.05 was considered to indicate a statistically significant
difference.
Results
Cell differentiation of P19 cells
Firstly, successful differentiation of P19 cells
into cardiomyocytes was established. Following treatment with DMSO,
mRNA expression levels of the cardiomyocyte differentiation markers
GATA binding protein 4 (GATA4; Fig.
1A) and NK2 homeobox 5 (Nkx2-5; Fig. 1B) were analyzed. The results
demonstrated that the relative mRNA expression levels for GATA4 and
Nkx2-5 in the cells increased in a time-dependent manner for the
whole duration of the 10 days of differentiation treatment,
compared with that at day 0 (Fig.
1). However, their levels in the undifferentiated P19 cells
were not significantly different to t=0 at days 5 and 10
(P>0.05; Fig. 1). These results
demonstrated that the P19 cells were successfully differentiated
into cardiomyocytes in the present study.
miR-1 expression in differentiated P19
cells
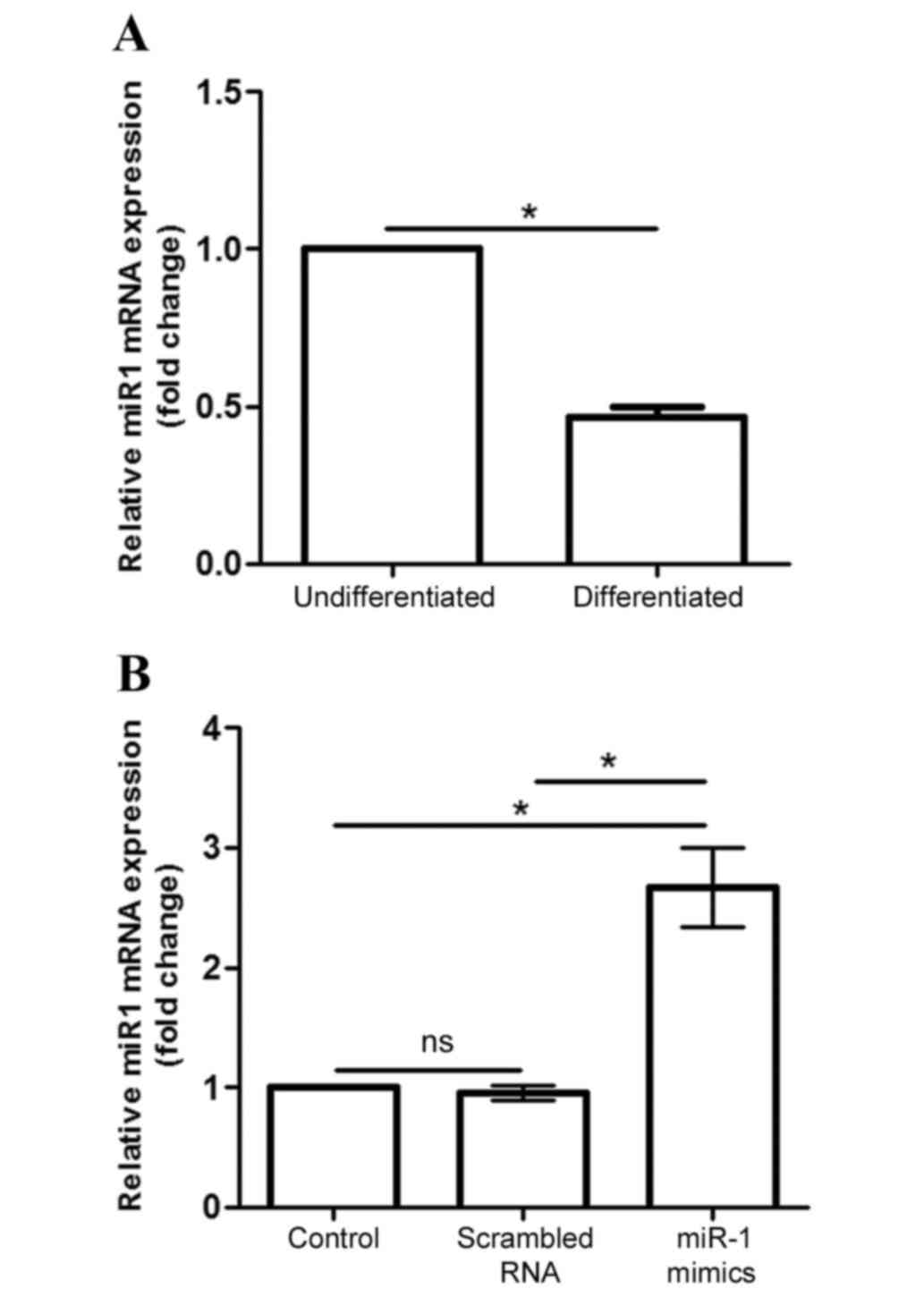
Endogenous miR-1 mRNA expression was significantly
decreased in differentiated P19 cells compared with
undifferentiated cells (P<0.05; Fig. 2A). Following transfection of
differentiated cells with siRNAs, miR-1 mimic and scramble, for 24
h, miR-1 mRNA expression was analyzed to verify whether the
transfection experiment was successful. The results demonstrated
that miR-1 mRNA levels were significantly increased following
transfection with the miR-1 mimic, compared with both transfected
cells and cells transfected with control scrambled RNA (P<0.05;
Fig. 2B).
miR-1 overexpression increases P19
cell viability
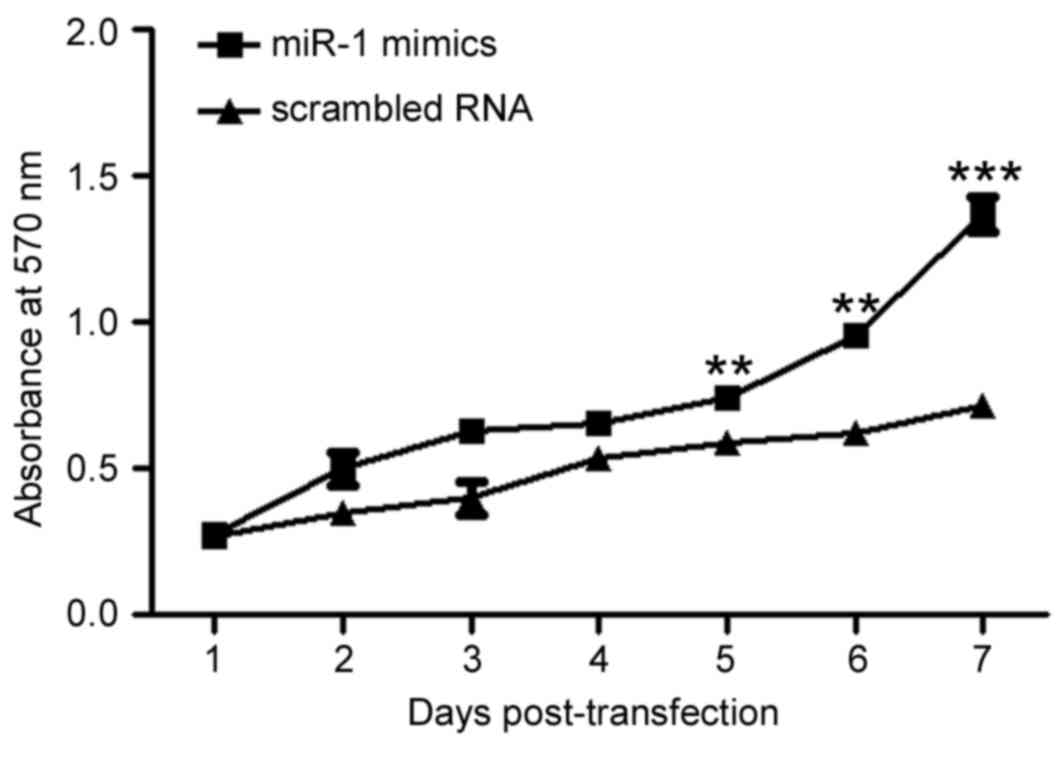
To assess the effect of miR-1 overexpression on cell
viability, viable cells were measured by MTT assay (Fig. 3). The results demonstrated that the
number of viable cells was increased in a time-dependent manner in
the P19 cells transfected with miR-1 mimic compared with the cells
transfected with scrambled RNA control (P<0.01; Fig. 3), suggesting that miR-1
overexpression may promote P19 cell proliferation.
miR-1 overexpression suppresses P19
cell apoptosis
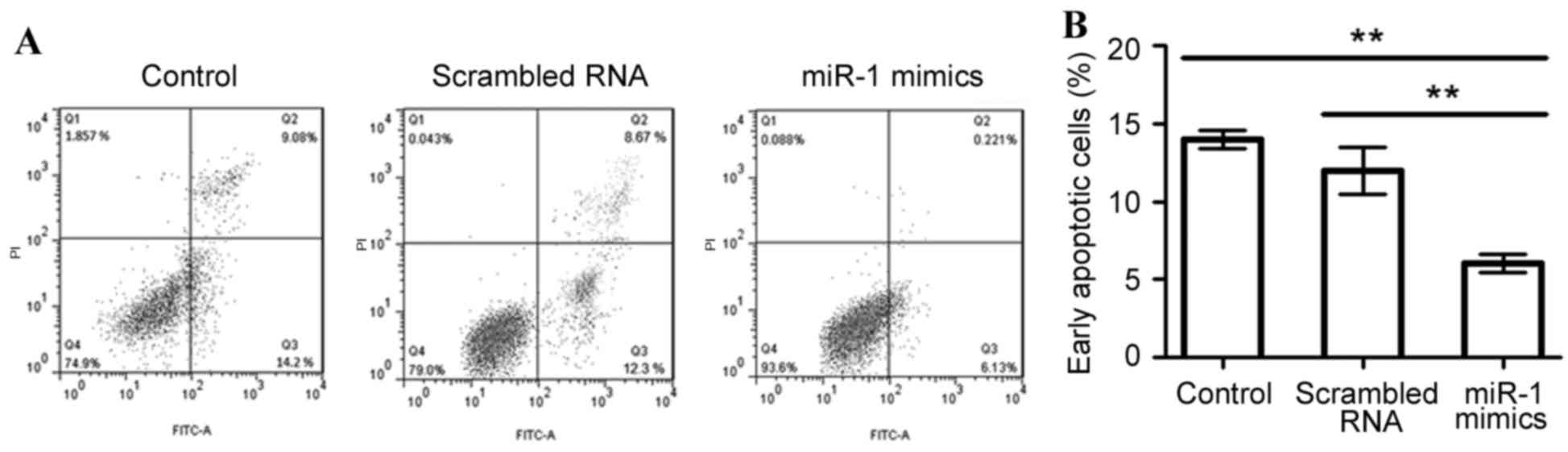
To assess the effect of miR-1 overexpression on cell
apoptosis in differentiated P19 cells, apoptotic cells were
measured using an Annexin V-FITC labeling assay (Fig. 4). The mean percentage of
early-stage apoptotic cells was significantly decreased when miR-1
was overexpressed (6.13%), compared with the untransfected or
scrambled RNA-transfected control cells (14.2 and 12.3%,
respectively; P<0.01; Fig.
4).
Effect of miR-1 overexpression on
Hand2 and caspase-3 expression
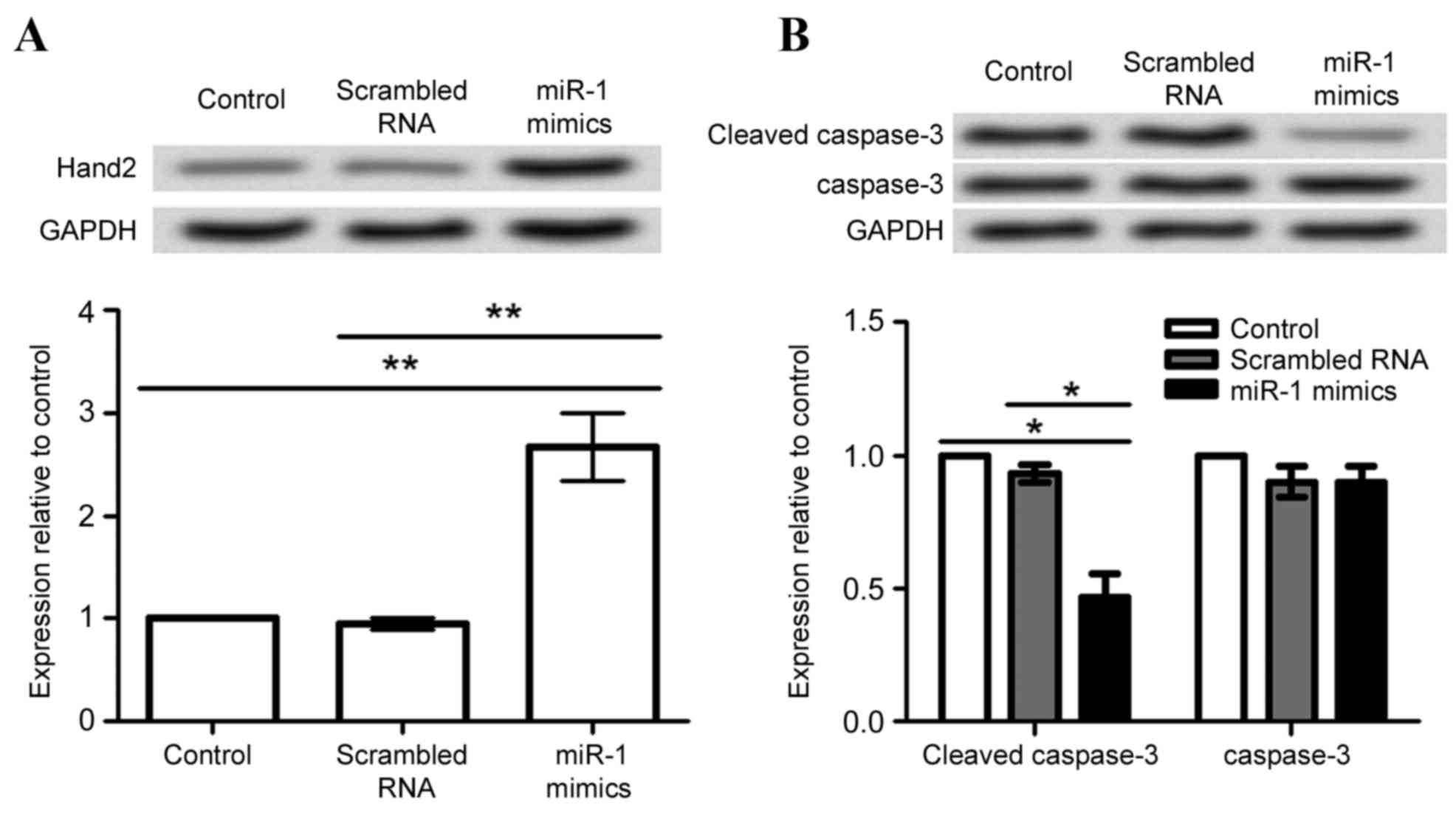
In order to investigate the possible mechanism of
miR-1 in cardiomyocytes, expression of Hand2 and caspase-3 proteins
was analyzed in differentiated P19 cells following transfection
with miR-1 mimic and scrambled RNA. Hand2 protein expression levels
were significantly increased by miR-1 overexpression, compared with
the untransfected cells and cells transfected with scrambled RNA
(P<0.01; Fig. 5A). By contrast,
caspase-3 cleavage was significantly decreased by miR-1
overexpression, compared with the untransfected cells and cells
transfected with scrambled RNA (P<0.05; Fig. 5B).
Discussion
Previous studies have demonstrated the importance of
miRNAs in the regulation of heart development, including miR-1 and
miR-133 (16), however, few
studies have explored the potential role of miR-1 in P19
differentiated cardiomyocytes. In the present study, expression of
miR-1 was evaluated in cardiomyocyte-differentiated and
undifferentiated P19 cells, and its effect on the viability and
apoptosis of cardiomyocyte-differentiated P19 cells was
examined.
In agreement with previous studies (17–20),
the present study confirmed that following treatment of P19 cells
with DMSO, mRNA expression levels of the GATA4 and Nkx2-5
differentiation markers were significantly increased compared with
untreated P19 cells (Fig. 1),
indicating that P19 cells were successfully induced towards
cardiomyocyte differentiation. When the differentiated P19 cells
were examined, a significant decrease in endogenous miR-1
expression was observed compared with the undifferentiated cells
(Fig. 2), suggesting an
association between miR-1 expression levels and P19 cell
differentiation state. The expression of miR-1 in P19 cells has not
been previously reported. However, Thomson et al (21) demonstrated that miR-1 and miR-133
were involved in the regulation of P19 embryonal teratocarcinoma at
the post-transcriptional level. The current study suggests that
abnormal expression of miR-1 may be associated with the
cardiomyocyte differentiation of P19 cells.
Consequently, the effect of miR-1 overexpression on
cell viability and apoptosis in cardiomyocyte-differentiated P19
cells was assessed. miR-1 upregulation in skeletal muscle has been
demonstrated to be positively correlated with muscle proliferation
and differentiation (22). By
contrast, miR-1 results in suppressed cardiomyocyte apoptosis by
targeting HSP60 and caspase-9 (9).
Similarly, lung cancer cell apoptosis is suppressed by upregulation
of miR-1 (23). In the present
study, cardiomyocyte-differentiated P19 cell viability was
increased while apoptosis was suppressed by miR-1 overexpression,
suggesting that miR-1 may be important in cardiac cell development
by regulating proliferation and apoptosis.
The current study demonstrated that miR-1
overexpression resulted in increased Hand2 expression but decreased
caspase-3 cleavage in cardiomyocyte-differentiated P19 cells
(Fig. 5). Hand2 is asymmetrically
expressed in the developing ventricular chambers and is important
in cardiac morphogenesis (24).
Yelon et al (25)
demonstrated that Hand2 served parallel roles to cell proliferation
and apoptosis in the heart development of zebrafish, and Olson
(26) demonstrated that Hand2 was
preferentially expressed in the derivative of the heart field.
Hence, it was hypothesized that miR-1 upregulation may contribute
to cardiac differentiation in P19 cells. Caspase-3 is a cell
apoptosis executor and its high expression indicates a high
percentage of apoptotic cells (27,28).
Izarra et al (16)
demonstrated that miR-1 overexpression results in reduced cell
apoptosis in pluripotent stem cells during cardiac differentiation.
In addition, Shan et al (29) demonstrated that caspase-3 levels
are decreased by miR-1 upregulation in a rat model of myocardial
infarction. The present study indicates that miR-1 suppresses cell
apoptosis in cardiomyocyte-differentiated P19 cells by decreasing
caspase-3 cleavage.
In conclusion, the present study revealed that miR-1
regulates heart development through the cell proliferation and
apoptosis processes, by increasing Hand2 expression and suppressing
caspase-3 cleavage in cardiomyocyte-differentiated P19 cells. The
present study may provide a theoretical basis for the role of miR-1
in regulating cardiomyocytes development and may indicate miR-1 as
a potential target in the therapeutic treatment of CHD in infants.
Further experimental studies are required to fully understand the
mechanism of miR-1 in the regulation of the P19 cells and the heart
development.
References
|
1
|
Al Mazrouei SK, Moore J, Ahmed F, Mikula
EB and Martin GR: Regional implementation of newborn screening for
critical congenital heart disease screening in Abu Dhabi. Pediatr
Cardiol. 34:1299–1306. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Dale MT, Solberg O, Holmstrøm H, Landolt
MA, Eskedal LT and Vollrath ME: Mothers of infants with congenital
heart defects: Well-being from pregnancy through the child's first
six months. Qual Life Res. 21:115–122. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ratanachu-Ek S and Pongdara A: Nutritional
status of pediatric patients with congenital heart disease: Pre-
and post cardiac surgery. J Med Assoc Thai. 94:(Suppl 3).
S133–S137. 2011.PubMed/NCBI
|
|
4
|
Moons P, Bovijn L, Budts W and Gewillig M:
Abstract 1866: Actual prospects to survive into adulthood in
patients with congenital heart disease. Circulation. 120:(Suppl).
S5612009.
|
|
5
|
Song R: Expression and function of small
non-coding RNAs in the mouse testis. PhD dissertation. University
of Nevada. ProQuest/UMI, Publication no. AAT 3472784. Reno, NV:
2011
|
|
6
|
Moll R, Sievers E, Hämmerling B, Schmidt
A, Barth M, Kuhn C, Grund C, Hofmann I and Franke WW: Endothelial
and virgultar cell formations in the mammalian lymph node sinus:
Endothelial differentiation morphotypes characterized by a special
kind of junction (complexus adhaerens). Cell Tissue Res.
335:109–141. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Kawashima K and Koshimizu U: Method for
proliferation cardiomyocytes using micro-rna. US Patent 20140213634
A1. Filed April 4, 2014; issued July 31. 2014.
|
|
8
|
L'Honore A, Rana V, Arsic N, Franckhauser
C, Lamb NJ and Fernandez A: Identification of a new hybrid serum
response factor and myocyte enhancer factor 2-binding element in
MyoD enhancer required for MyoD expression during myogenesis. Mol
Biol Cell. 18:1992–2001. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Xu C, Lu Y, Pan Z, Chu W, Luo X, Lin H,
Xiao J, Shan H, Wang Z and Yang B: The muscle-specific microRNAs
miR-1 and miR-133 produce opposing effects on apoptosis by
targeting HSP60, HSP70 and caspase-9 in cardiomyocytes. J Cell Sci.
120:3045–3052. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Grobe JL, Mecca AP, Lingis M, Shenoy V,
Bolton TA, Machado JM, Speth RC, Raizada MK and Katovich MJ:
Prevention of angiotensin II-induced cardiac remodeling by
angiotensin-(1–7). Am J Physiol Heart Circ Physiol. 292:H736–H742.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
McBurney MW, Jones-Villeneuve EM, Edwards
MK and Anderson PJ: Control of muscle and neuronal differentiation
in a cultured embryonal carcinoma cell line. Nature. 299:165–167.
1982. View
Article : Google Scholar : PubMed/NCBI
|
|
12
|
Choi SC, Choi JH, Shim WJ and Lim DS: P19
Embryonal carcinoma cells: A new model for the study of endothelial
cell differentiation. Biotechnol Lett. 30:1169–1175. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Paquin J, Danalache BA, Jankowski M,
McCann SM and Gutkowska J: Oxytocin induces differentiation of P19
embryonic stem cells to cardiomyocytes. Proc Natl Acad Sci USA.
99:9550–9555. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Arseculeratne SN, Atapattu DN, Kumarasiri
R, Perera D, Ekanayake D and Rajapakse J: The use of MTT [3-(4,
5-dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide]-
reduction as an indicator of the effects of strain-specific,
polyclonal rabbit antisera on Candida albicans and C. Indian J Med
Microbiol. 25:267–271. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Methods. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Izarra A, Moscoso I, Cañón S, Carreiro C,
Fondevila D, Martín-Caballero J, Blanca V, Valiente I, Díez-Juan A
and Bernad A: miRNA-1 and miRNA-133a are involved in early
commitment of pluripotent stem cells and demonstrate antagonistic
roles in the regulation of cardiac differentiation. J Tissue Eng
Regen Med. Dec 10–2014.(Epub ahead of print). PubMed/NCBI
|
|
17
|
Brown CO III, Chi X, Garcia-Gras E, Shirai
M, Feng XH and Schwartz RJ: The cardiac determination factor,
Nkx2-5, is activated by mutual cofactors GATA-4 and Smad1/4 via a
novel upstream enhancer. J Biol Chem. 279:10659–10669. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Haveri H, Ashorn M, Iltanen S, Wilson DB,
Andersson LC and Heikinheimo M: Enhanced expression of
transcription factor GATA4 in inflammatory bowel disease and its
possible regulation by TGF-beta1. J Clin Immunol. 29:444–453. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhou Y and Xiao-Yu HE: Expression of the
transcription factor GATA-4 in human heart development. J Med
Postg. 2008.
|
|
20
|
Zaglia T, Dedja A, Candiotto C, Cozzi E,
Schiaffino S and Ausoni S: Cardiac interstitial cells express GATA4
and control dedifferentiation and cell cycle re-entry of adult
cardiomyocytes. J Mol Cell Cardiol. 46:653–662. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Thomson JM, Newman M, Parker JS,
Morin-Kensicki EM, Wright T and Hammond SM: Extensive
post-transcriptional regulation of microRNAs and its implications
for cancer. Gene Dev. 20:2202–2207. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chen JF, Mandel EM, Thomson JM, Wu Q,
Callis TE, Hammond SM, Conlon FL and Wang DZ: The role of
microRNA-1 and microRNA-133 in skeletal muscle proliferation and
differentiation. Nat Genet. 38:228–233. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Nasser MW, Datta J, Nuovo G, Kutay H,
Motiwala T, Majumder S, Wang B, Suster S, Jacob ST and Ghoshal K:
Down-regulation of micro-RNA-1 (miR-1) in lung cancer. Suppression
of tumorigenic property of lung cancer cells and their
sensitization to doxorubicin-induced apoptosis by miR-1. J Biol
Chem. 283:33394–33405. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
VanDusen NJ, Casanovas J, Vincentz JW,
Firulli BA, Osterwalder M, Lopez-Rios J, Zeller R, Zhou B,
Grego-Bessa J, De La Pompa JL, et al: Hand2 is an essential
regulator for two Notch-dependent functions within the embryonic
endocardium. Cell Rep. 9:2071–2083. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Yelon D, Ticho B, Halpern ME, Ruvinsky I,
Ho RK, Silver LM and Stainier DY: The bHLH transcription factor
hand2 plays parallel roles in zebrafish heart and pectoral fin
development. Development. 127:2573–2582. 2000.PubMed/NCBI
|
|
26
|
Olson EN: Gene regulatory networks in the
evolution and development of the heart. Science. 313:1922–1927.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sun J, Shen X, Xie B, Zhong YS, Lu Q and
Sun Y: Effects of high glucose on apoptosis of human umbilical vein
endothelial cells and expression of Caspase-3. J Shang Jiaotong
Univ. 34:1709–1713. 2014.
|
|
28
|
Isa SA, Mainwaring LS, Webb R and Thomas
AW: The non-genomic effects of high doses of Rosiglitazone on cell
growth and apoptosis in cultured monocytic cells. Bayero J Pure
Appl Sci. 2:1–8. 2009.
|
|
29
|
Shan ZX, Lin QX, Fu YH, Deng CY, Zhou ZL,
Zhu JN, Liu XY, Zhang YY, Li Y, Lin SG and Yu XY: Upregulated
expression of miR-1/miR-206 in a rat model of myocardial
infarction. Biochem Biophys Res Commun. 381:597–601. 2009.
View Article : Google Scholar : PubMed/NCBI
|